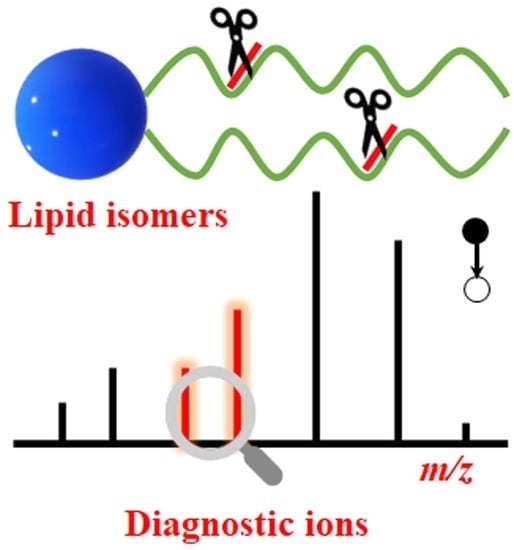
Review of Recent Advances in Lipid Analysis of Biological Samples via Ambient Ionization Mass Spectrometry
Abstract
:1. Introduction
2. Mechanism and Instrumental Setup of Various Strategies for Lipid Structural Elucidation
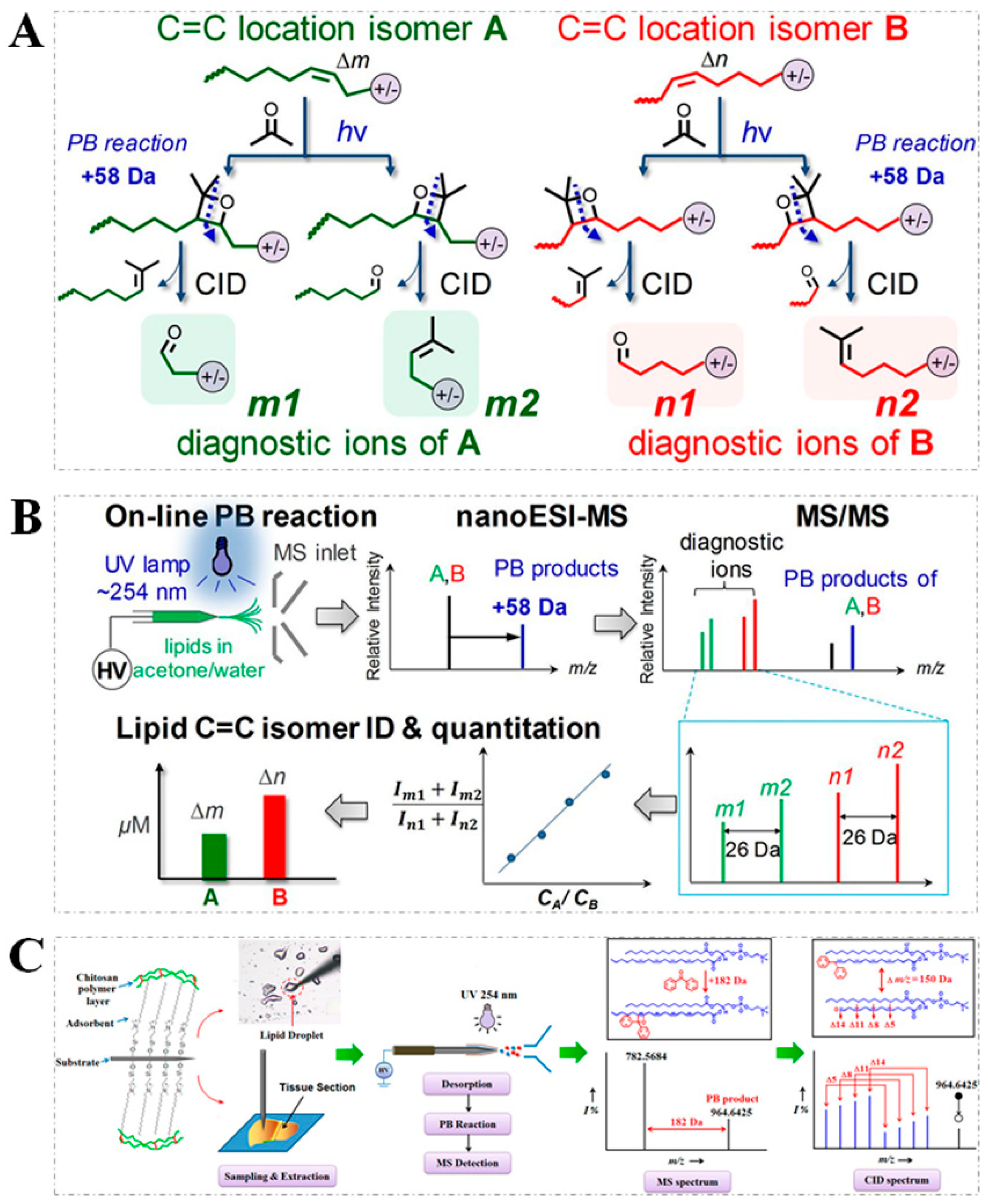
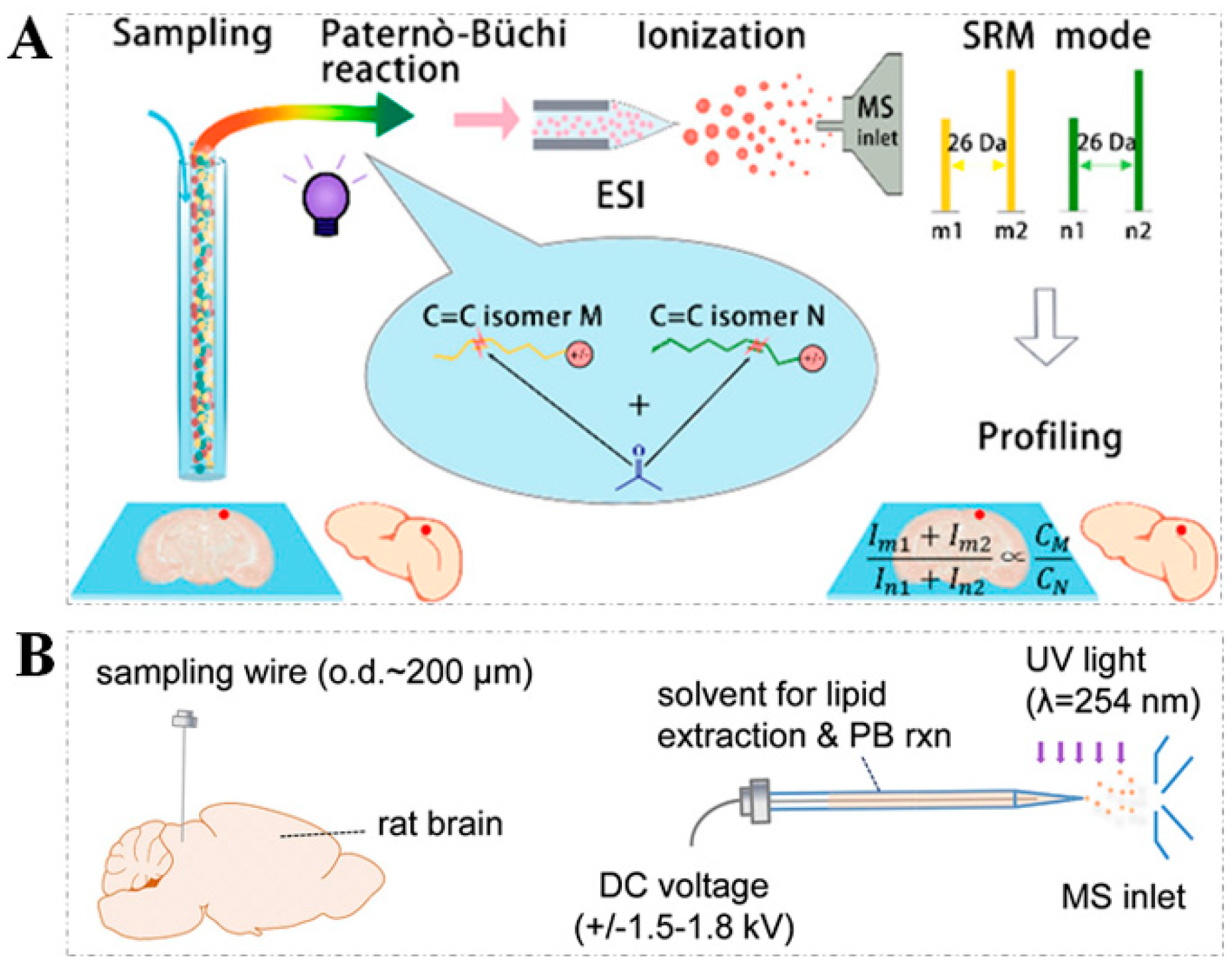
2.1. PB Reaction
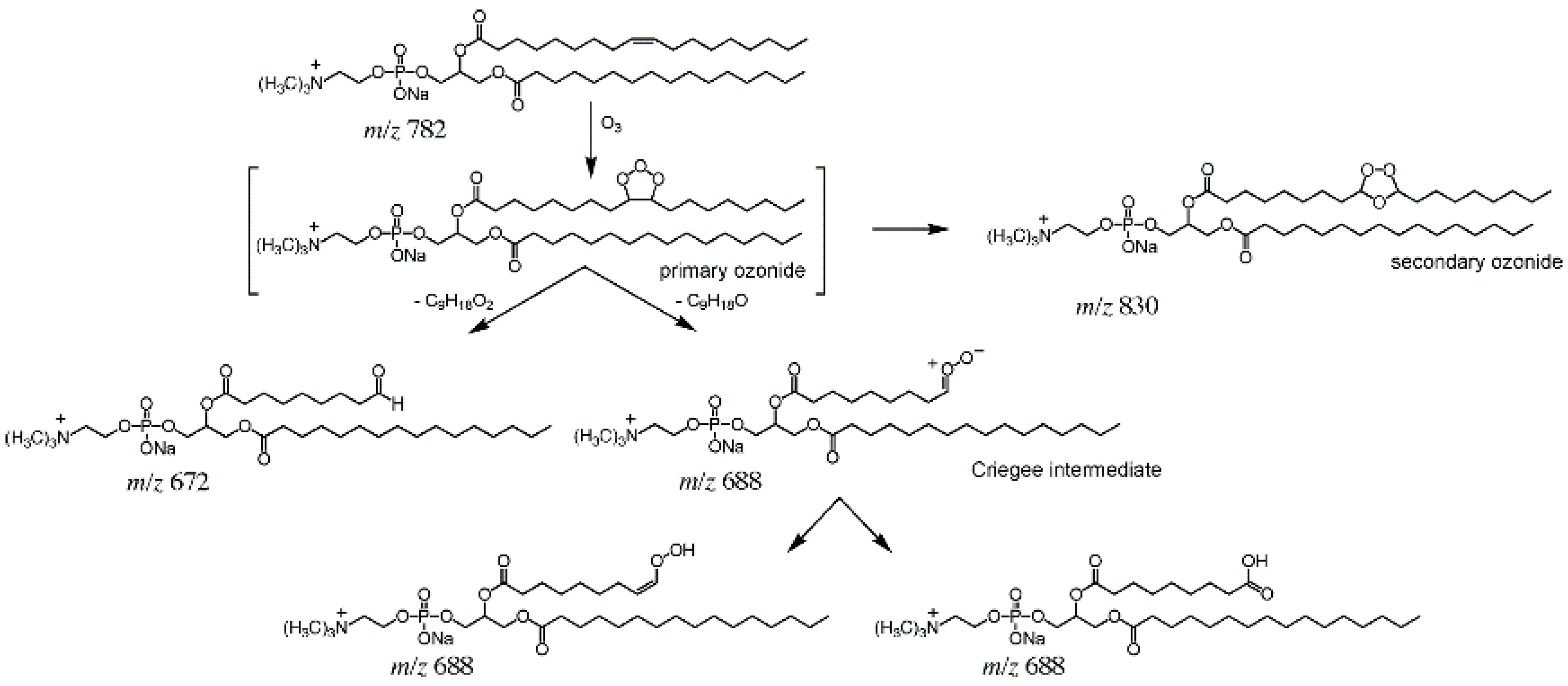
2.2. OzID and UVPD
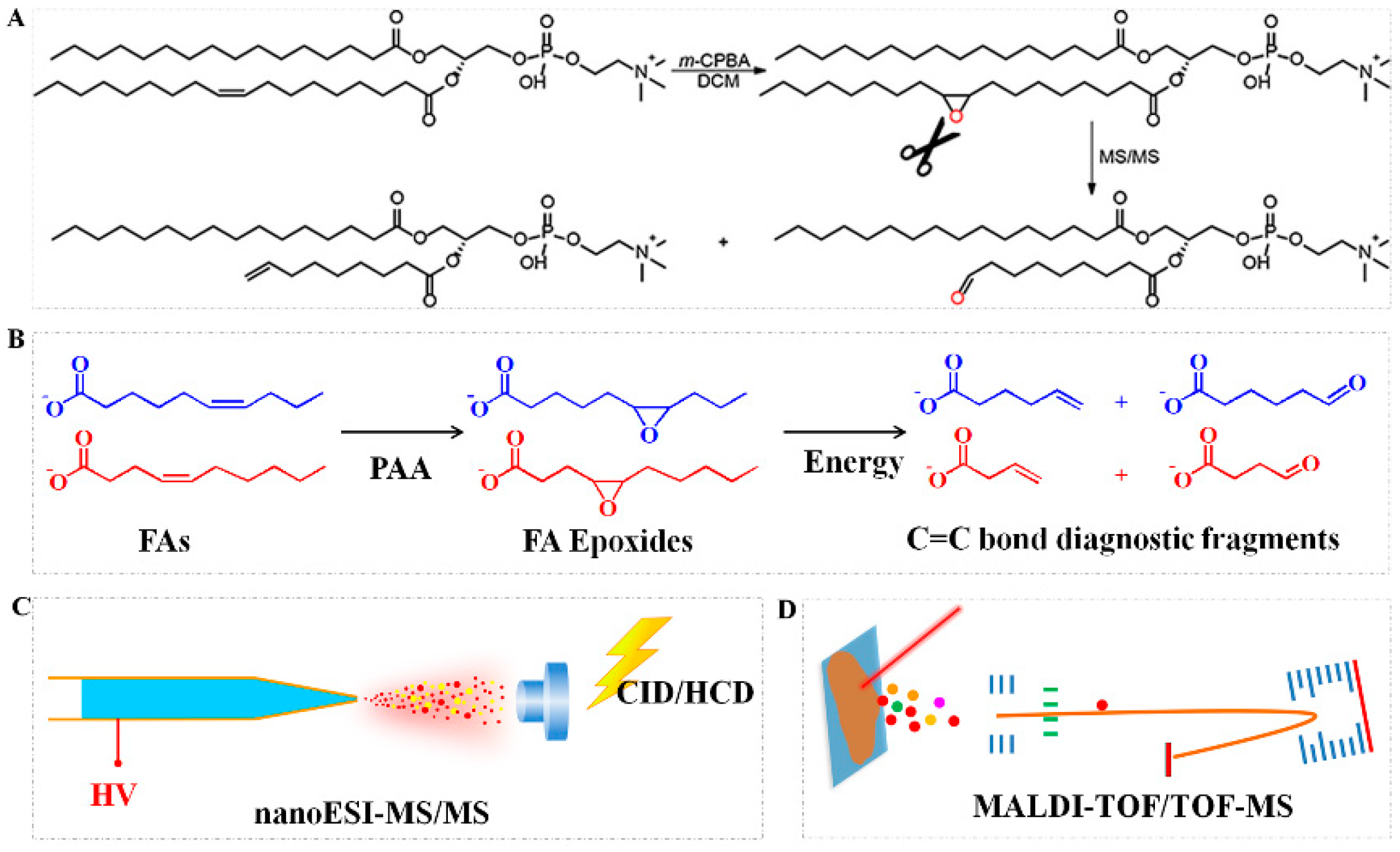
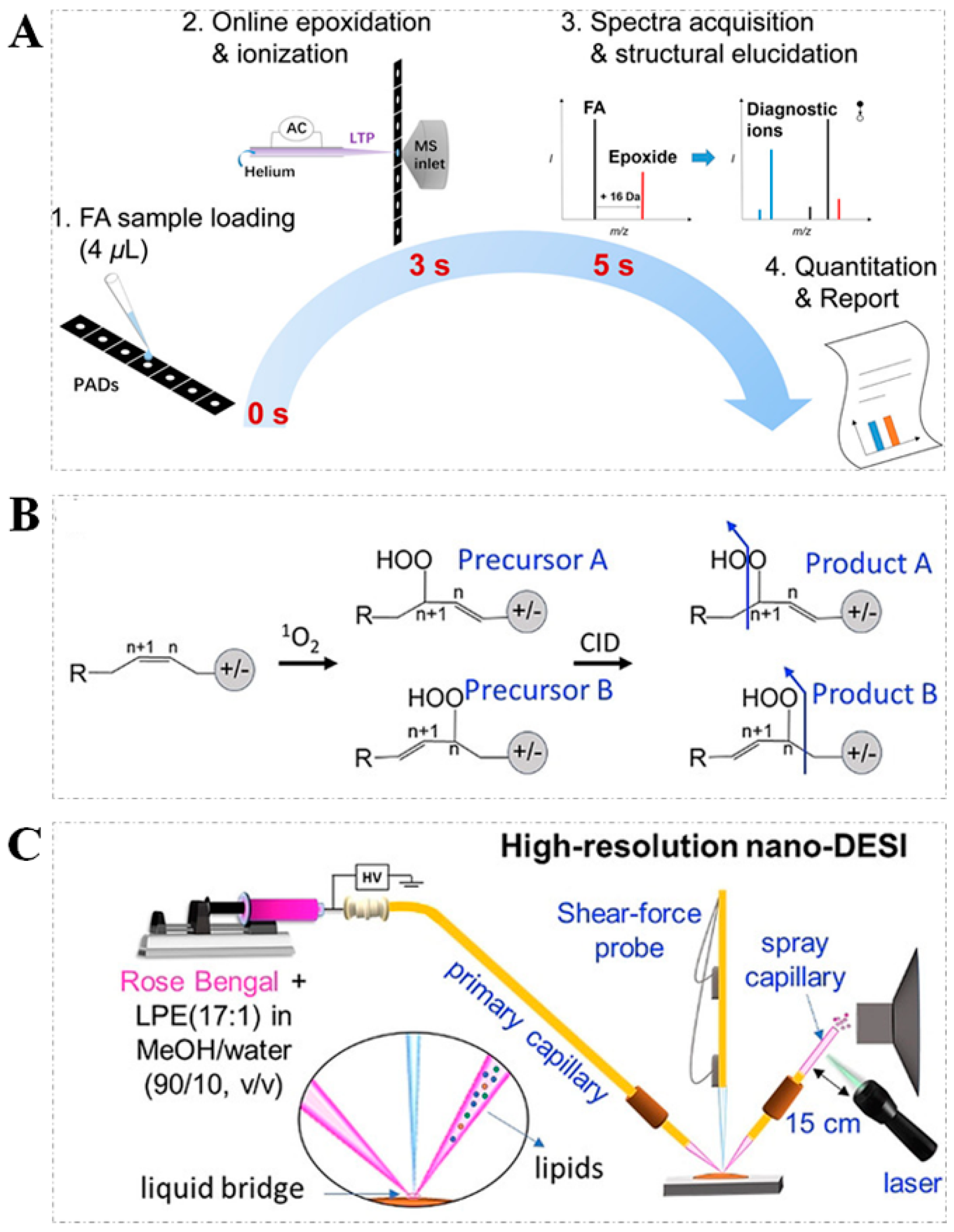
2.3. Epoxidation Reaction
2.4. Other Strategies
3. The Applications of AIMS in Lipid Structural Elucidation
3.1. Direct Characterization of Unsaturated Lipid Isomers in Tissue Samples
3.2. Direct Characterization of Unsaturated Lipid Isomers in Biofluid Samples
3.3. Direct Characterization of Unsaturated Lipid Isomers in Cell Samples
3.4. Spatial Visualization of Unsaturated Lipid Isomers within Tissue Sections
3.5. Direct Characterization of Unsaturated Lipid Isomers in Other Samples
4. Conclusions and Future Directions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
References
- Klein, D.R.; Blevins, M.S.; Macias, L.A.; Douglass, M.V.; Trent, M.S.; Brodbelt, J.S. Localization of Double Bonds in Bacterial Glycerophospholipids Using 193 nm Ultraviolet Photodissociation in the Negative Mode. Anal. Chem. 2020, 92, 5986–5993. [Google Scholar] [CrossRef] [PubMed]
- Unsihuay, D.; Su, P.; Hu, H.; Qiu, J.; Kuang, S.; Li, Y.; Sun, X.; Dey, S.K.; Laskin, J. Imaging and Analysis of Isomeric Unsaturated Lipids through Online Photochemical Derivatization of Carbon-Carbon Double Bonds *. Angew Chem. Int. Ed. Engl. 2021, 60, 7559–7563. [Google Scholar] [CrossRef] [PubMed]
- Feng, G.F.; Hao, Y.H.; Wu, L.; Chen, S.M. A visible-light activated [2 + 2] cycloaddition reaction enables pinpointing carbon-carbon double bonds in lipids. Chem. Sci. 2020, 11, 7244–7251. [Google Scholar] [CrossRef] [PubMed]
- Xu, T.; Hu, C.; Xuan, Q.; Xu, G. Recent advances in analytical strategies for mass spectrometry-based lipidomics. Anal Chim Acta 2020, 1137, 156–169. [Google Scholar] [CrossRef]
- Zhang, H.; Xu, M.; Shi, X.; Liu, Y.; Li, Z.; Jagodinsky, J.C.; Ma, M.; Welham, N.V.; Morris, Z.S.; Li, L. Quantification and molecular imaging of fatty acid isomers from complex biological samples by mass spectrometry. Chem. Sci. 2021, 12, 8115–8122. [Google Scholar] [CrossRef]
- Zhu, Y.; Wang, W.; Yang, Z. Combining Mass Spectrometry with Paternò–Büchi Reaction to Determine Double-Bond Positions in Lipids at the Single-Cell Level. Anal. Chem. 2020, 92, 11380–11387. [Google Scholar] [CrossRef]
- Zhang, W.P.; Shang, B.; Ouyang, Z.; Xia, Y. Enhanced Phospholipid Isomer Analysis by Online Photochemical Derivatization and RPLC-MS. Anal. Chem. 2020, 92, 6719–6726. [Google Scholar] [CrossRef]
- Zhang, W.P.; Zhang, D.H.; Chen, Q.H.; Wu, J.H.; Ouyang, Z.; Xia, Y. Online photochemical derivatization enables comprehensive mass spectrometric analysis of unsaturated phospholipid isomers. Nat. Commun. 2019, 10, 79. [Google Scholar] [CrossRef]
- Rohrig, F.; Schulze, A. The multifaceted roles of fatty acid synthesis in cancer. Nat. Rev. Cancer 2016, 16, 732–749. [Google Scholar] [CrossRef]
- Bazinet, R.P.; Laye, S. Polyunsaturated fatty acids and their metabolites in brain function and disease. Nat. Rev. Neurosci. 2014, 15, 771–785. [Google Scholar] [CrossRef]
- Cao, W.B.; Cheng, S.M.; Yang, J.; Feng, J.X.; Zhang, W.P.; Li, Z.S.; Chen, Q.H.; Xia, Y.; Ouyang, Z.; Ma, X.X. Large-scale lipid analysis with C=C location and sn-position isomer resolving power. Nat. Commun. 2020, 11, 375. [Google Scholar] [CrossRef] [Green Version]
- Stinson, C.A.; Xia, Y. A method of coupling the Paterno-Buchi reaction with direct infusion ESI-MS/MS for locating the C=C bond in glycerophospholipids. Analyst 2016, 141, 3696–3704. [Google Scholar] [CrossRef]
- Zhang, H.; Chingin, K.; Li, J.; Lu, H.; Huang, K.; Chen, H. Selective Enrichment of Phosphopeptides and Phospholipids from Biological Matrixes on TiO2 Nanowire Arrays for Direct Molecular Characterization by Internal Extractive Electrospray Ionization Mass Spectrometry. Anal. Chem. 2018, 90, 12101–12107. [Google Scholar] [CrossRef]
- Zhang, H.; Lu, H.; Huang, K.; Li, J.; Wei, F.; Liu, A.; Chingin, K.; Chen, H. Selective detection of phospholipids in human blood plasma and single cells for cancer differentiation using dispersed solid-phase microextraction combined with extractive electrospray ionization mass spectrometry. Analyst 2020, 145, 7330–7339. [Google Scholar] [CrossRef]
- Wang, J.; Wang, C.; Han, X. Tutorial on lipidomics. Anal Chim Acta 2019, 1061, 28–41. [Google Scholar] [CrossRef]
- Ma, X.X.; Chong, L.; Tian, R.; Shi, R.Y.; Hu, T.Y.; Ouyang, Z.; Xia, Y. Identification and quantitation of lipid C=C location isomers: A shotgun lipidomics approach enabled by photochemical reaction. Proc. Natl. Acad. Sci. USA 2016, 113, 2573–2578. [Google Scholar] [CrossRef] [Green Version]
- Porta Siegel, T.; Ekroos, K.; Ellis, S.R. Reshaping Lipid Biochemistry by Pushing Barriers in Structural Lipidomics. Angew. Chem. Int. Ed. Engl. 2019, 58, 6492–6501. [Google Scholar] [CrossRef] [Green Version]
- Paine, M.R.L.; Poad, B.L.J.; Eijkel, G.B.; Marshall, D.L.; Blanksby, S.J.; Heeren, R.M.A.; Ellis, S.R. Mass Spectrometry Imaging with Isomeric Resolution Enabled by Ozone-Induced Dissociation. Angew. Chem. Int. Ed. 2018, 57, 10530–10534. [Google Scholar] [CrossRef] [Green Version]
- Knowles, S.L.; Vu, N.; Todd, D.A.; Raja, H.A.; Rokas, A.; Zhang, Q.B.; Oberlies, N.H. Orthogonal Method for Double-Bond Placement via Ozone-Induced Dissociation Mass Spectrometry (OzID-MS). J. Nat. Prod. 2019, 82, 3421–3431. [Google Scholar] [CrossRef]
- Jones, J.W.; Thompson, C.J.; Carter, C.L.; Kane, M.A. Electron-induced dissociation (EID) for structure characterization of glycerophosphatidylcholine: Determination of double-bond positions and localization of acyl chains. J. Mass Spectrom. 2015, 50, 1327–1339. [Google Scholar] [CrossRef] [Green Version]
- Feider, C.L.; Macias, L.A.; Brodbelt, J.S.; Eberlin, L.S. Double Bond Characterization of Free Fatty Acids Directly from Biological Tissues by Ultraviolet Photodissociation. Anal. Chem. 2020, 92, 8386–8395. [Google Scholar] [CrossRef]
- Xie, X.B.; Xia, Y. Analysis of Conjugated Fatty Acid Isomers by the Paterno-Buchi Reaction and Trapped Ion Mobility Mass Spectrometry. Anal. Chem. 2019, 91, 7173–7180. [Google Scholar] [CrossRef]
- Ma, X.X.; Xia, Y. Pinpointing Double Bonds in Lipids by Paterno-Buchi Reactions and Mass Spectrometry. Angew. Chem. Int. Ed. 2014, 53, 2592–2596. [Google Scholar] [CrossRef]
- Feng, Y.; Chen, B.; Yu, Q.; Li, L. Identification of Double Bond Position Isomers in Unsaturated Lipids by m-CPBA Epoxidation and Mass Spectrometry Fragmentation. Anal. Chem. 2019, 91, 1791–1795. [Google Scholar] [CrossRef]
- Kuo, T.-H.; Chung, H.-H.; Chang, H.-Y.; Lin, C.-W.; Wang, M.-Y.; Shen, T.-L.; Hsu, C.-C. Deep Lipidomics and Molecular Imaging of Unsaturated Lipid Isomers: A Universal Strategy Initiated by mCPBA Epoxidation. Anal. Chem. 2019, 91, 11905–11915. [Google Scholar] [CrossRef]
- Xie, X.B.; Zhao, J.; Lin, M.; Zhang, J.L.; Xia, Y. Profiling of Cholesteryl Esters by Coupling Charge-Tagging Paterno-Buchi Reaction and Liquid Chromatography-Mass Spectrometry. Anal. Chem. 2020, 92, 8487–8496. [Google Scholar] [CrossRef]
- Jeck, V.; Korf, A.; Vosse, C.; Hayen, H. Localization of double-bond positions in lipids by tandem mass spectrometry succeeding high-performance liquid chromatography with post-column derivatization. Rapid Commun. Mass Spectrom. 2019, 33, 86–94. [Google Scholar] [CrossRef] [Green Version]
- Hakova, E.; Vrkoslav, V.; Mikova, R.; Schwarzova-Peckova, K.; Bosakova, Z.; Cvacka, J. Localization of double bonds in triacylglycerols using high-performance liquid chromatography/atmospheric pressure chemical ionization ion-trap mass spectrometry. Anal. Bioanal. Chem. 2015, 407, 5175–5188. [Google Scholar] [CrossRef]
- Lu, H.; Zhang, H.; Chingin, K.; Xiong, J.; Fang, X.; Chen, H. Ambient mass spectrometry for food science and industry. Trac-Trend Anal. Chem. 2018, 107, 99–115. [Google Scholar] [CrossRef]
- Cooks, R.G.; Ouyang, Z.; Takats, Z.; Wiseman, J.M. Ambient mass spectrometry. Science 2006, 311, 1566–1570. [Google Scholar] [CrossRef]
- Balog, J.; Sasi-Szabo, L.; Kinross, J.; Lewis, M.R.; Muirhead, L.J.; Veselkov, K.; Mirnezami, R.; Dezso, B.; Damjanovich, L.; Darzi, A.; et al. Intraoperative Tissue Identification Using Rapid Evaporative Ionization Mass Spectrometry. Sci. Transl. Med. 2013, 5, 194ra93. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.L.; Rector, J.; Lin, J.Q.; Young, J.H.; Sans, M.; Katta, N.; Giese, N.; Yu, W.D.; Nagi, C.; Suliburk, J.; et al. Nondestructive tissue analysis for ex vivo and in vivo cancer diagnosis using a handheld mass spectrometry system. Sci. Transl. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lu, H.; Zhang, H.; Chingin, K.; Wei, Y.; Xu, J.; Ke, M.; Huang, K.; Feng, S.; Chen, H. Sequential Detection of Lipids, Metabolites, and Proteins in One Tissue for Improved Cancer Differentiation Accuracy. Anal. Chem. 2019, 91, 10532–10540. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Zhang, H.; Xiao, Y.; Chingin, K.; Dai, C.; Wei, F.; Wang, N.; Frankevich, V.; Chagovets, V.; Zhou, F.; et al. Comparative study of alterations in phospholipid profiles upon liver cancer in humans and mice. Analyst 2020, 145, 6470–6477. [Google Scholar] [CrossRef]
- Jackson, S.N.; Muller, L.; Roux, A.; Oktem, B.; Moskovets, E.; Doroshenko, V.M.; Woods, A.S. AP-MALDI Mass Spectrometry Imaging of Gangliosides Using 2,6-Dihydroxyacetophenone. J. Am. Soc. Mass Spectrom. 2018, 29, 1463–1472. [Google Scholar] [CrossRef]
- Abdelhamid, H.N.; Bhaisare, M.L.; Wu, H.F. Ceria nanocubic-ultrasonication assisted dispersive liquid-liquid microextraction coupled with matrix assisted laser desorption/ionization mass spectrometry for pathogenic bacteria analysis. Talanta 2014, 120, 208–217. [Google Scholar] [CrossRef]
- Hua, P.Y.; Manikandan, M.; Abdelhamid, H.N.; Wu, H.F. Graphene nanoflakes as an efficient ionizing matrix for MALDI-MS based lipidomics of cancer cells and cancer stem cells. J. Mater Chem. B 2014, 2, 7334–7343. [Google Scholar] [CrossRef]
- Keller, C.; Maeda, J.; Jayaraman, D.; Chakraborty, S.; Sussman, M.R.; Harris, J.M.; Ane, J.M.; Li, L. Comparison of Vacuum MALDI and AP-MALDI Platforms for the Mass Spectrometry Imaging of Metabolites Involved in Salt Stress in Medicago truncatula. Front. Plant Sci. 2018, 9, 1238. [Google Scholar] [CrossRef]
- Chen, B.; OuYang, C.; Tian, Z.; Xu, M.; Li, L. A high resolution atmospheric pressure matrix-assisted laser desorption/ionization-quadrupole-orbitrap MS platform enables in situ analysis of biomolecules by multi-mode ionization and acquisition. Anal. Chim. Acta 2018, 1007, 16–25. [Google Scholar] [CrossRef]
- Caleb Bagley, M.; Garrard, K.P.; Muddiman, D.C. The development and application of matrix assisted laser desorption electrospray ionization: The teenage years. Mass Spectrom. Rev. 2021. [Google Scholar] [CrossRef]
- Adam, W.; Peters, K.; Peters, E.M.; Stegmann, V.R. Hydroxy-Directed Regio- and Diastereoselective [2 + 2] Photocycloaddition (Paternò−Büchi Reaction) of Benzophenone to Chiral Allylic Alcohols. J. Am. Chem. Soc. 2000, 122, 2958–2959. [Google Scholar] [CrossRef]
- Waldchen, F.; Spengler, B.; Heiles, S. Reactive Matrix-Assisted Laser Desorption/Ionization Mass Spectrometry Imaging Using an Intrinsically Photoreactive Paterno-Buchi Matrix for Double-Bond Localization in Isomeric Phospholipids. J. Am. Chem. Soc. 2019, 141, 11816–11820. [Google Scholar] [CrossRef]
- Chonglo, L.; Tian, R.; Shi, R.Y.; Ouyang, Z.; Xia, Y. Coupling the Paterno-Buchi (PB) Reaction With Mass Spectrometry to Study Unsaturated Fatty Acids in Mouse Model of Multiple Sclerosis. Front. Chem. 2019, 7. [Google Scholar] [CrossRef]
- Xu, T.F.; Pi, Z.F.; Song, F.R.; Liu, S.; Liu, Z.Q. Benzophenone used as the photochemical reagent for pinpointing C=C locations in unsaturated lipids through shotgun and liquid chromatography-mass spectrometry approaches. Anal. Chim. Acta 2018, 1028, 32–44. [Google Scholar] [CrossRef]
- Zhao, J.; Xie, X.; Lin, Q.; Ma, X.; Su, P.; Xia, Y. Next-Generation Paterno-Buchi Reagents for Lipid Analysis by Mass Spectrometry. Anal. Chem. 2020, 92, 13470–13477. [Google Scholar] [CrossRef]
- Xu, S.L.; Wu, B.F.; Oresic, M.; Xie, Y.; Yao, P.; Wu, Z.Y.; Lv, X.; Chen, H.; Wei, F. Double Derivatization Strategy for High-Sensitivity and High- Coverage Localization of Double Bonds in Free Fatty Acids by Mass Spectrometry. Anal. Chem. 2020, 92, 6446–6455. [Google Scholar] [CrossRef]
- Deng, J.; Yang, Y.; Liu, Y.; Fang, L.; Lin, L.; Luan, T. Coupling Paterno-Buchi Reaction with Surface-Coated Probe Nanoelectrospray Ionization Mass Spectrometry for In Vivo and Microscale Profiling of Lipid C horizontal lineC Location Isomers in Complex Biological Tissues. Anal. Chem. 2019, 91, 4592–4599. [Google Scholar] [CrossRef]
- Zhang, W.P.; Chiang, S.; Li, Z.S.; Chen, Q.H.; Xia, Y.; Ouyang, Z. A Polymer Coating Transfer Enrichment Method for Direct Mass Spectrometry Analysis of Lipids in Biofluid Samples. Angew. Chem. Int. Ed. 2019, 58, 6064–6069. [Google Scholar] [CrossRef]
- Thomas, M.C.; Mitchell, T.W.; Harman, D.G.; Deeley, J.M.; Nealon, J.R.; Blanksby, S.J. Ozone-Induced Dissociation: Elucidation of Double Bond Position within Mass-Selected Lipid Ions. Anal. Chem. 2008, 80, 303–311. [Google Scholar] [CrossRef] [Green Version]
- Klein, D.R.; Brodbelt, J.S. Structural Characterization of Phosphatidylcholines Using 193 nm Ultraviolet Photodissociation Mass Spectrometry. Anal. Chem. 2017, 89, 1516–1522. [Google Scholar] [CrossRef] [Green Version]
- Williams, P.E.; Klein, D.R.; Greer, S.M.; Brodbelt, J.S. Pinpointing Double Bond and sn-Positions in Glycerophospholipids via Hybrid 193 nm Ultraviolet Photodissociation (UVPD) Mass Spectrometry. J. Am. Chem. Soc. 2017, 139, 15681–15690. [Google Scholar] [CrossRef]
- Brodbelt, J.S.; Morrison, L.J.; Santos, I. Ultraviolet Photodissociation Mass Spectrometry for Analysis of Biological Molecules. Chem. Rev. 2020, 120, 3328–3380. [Google Scholar] [CrossRef]
- Luo, K.; Chen, H.; Zare, R.N. Location of carbon-carbon double bonds in unsaturated lipids using microdroplet mass spectrometry. Analyst 2021, 146, 2550–2558. [Google Scholar] [CrossRef]
- Zhang, J.I.; Tao, W.A.; Cooks, R.G. Facile determination of double bond position in unsaturated fatty acids and esters by low temperature plasma ionization mass spectrometry. Anal. Chem. 2011, 83, 4738–4744. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Zhao, Y.; Zhang, L.; Ma, X.; Zhang, S.; Zhang, X. Rapid Analysis of Unsaturated Fatty Acids on Paper-Based Analytical Devices via Online Epoxidation and Ambient Mass Spectrometry. Anal. Chem. 2018, 90, 2070–2078. [Google Scholar] [CrossRef] [PubMed]
- Steyer, D.J.; Kennedy, R.T. High-Throughput Nanoelectrospray Ionization-Mass Spectrometry Analysis of Microfluidic Droplet Samples. Anal. Chem. 2019, 91, 6645–6651. [Google Scholar] [CrossRef] [PubMed]
- Domon, B.; Aebersold, R. Review—Mass spectrometry and protein analysis. Science 2006, 312, 212–217. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tang, S.L.; Cheng, H.Y.; Yan, X. On-Demand Electrochemical Epoxidation in Nano-Electrospray Ionization Mass Spectrometry to Locate Carbon-Carbon Double Bonds. Angew. Chem. Int. Ed. 2020, 59, 209–214. [Google Scholar] [CrossRef] [Green Version]
- Zhang, X.; Wang, W.; Zare, R.N.; Min, Q. Peptide and protein assays using customizable bio-affinity arrays combined with ambient ionization mass spectrometry. Chem. Sci. 2021, 12, 10810–10816. [Google Scholar] [CrossRef]
- Bergman, H.M.; Lanekoff, I. Profiling and quantifying endogenous molecules in single cells using nano-DESI MS. Analyst 2017, 142, 3639–3647. [Google Scholar] [CrossRef]
- Kirschbaum, C.; Greis, K.; Lettow, M.; Gewinner, S.; Schollkopf, W.; Meijer, G.; von Helden, G.; Pagel, K. Non-covalent double bond sensors for gas-phase infrared spectroscopy of unsaturated fatty acids. Ana. Bioanal. Chem. 2021, 413, 3643–3653. [Google Scholar] [CrossRef]
- Barrientos, R.C.; Vu, N.; Zhang, Q.B. Structural Analysis of Unsaturated Glycosphingolipids Using Shotgun Ozone-Induced Dissociation Mass Spectrometry. J. Am. Soc. Mass Spectrom. 2017, 28, 2330–2343. [Google Scholar] [CrossRef]
- Vu, N.; Brown, J.; Giles, K.; Zhang, Q.B. Ozone-induced dissociation on a traveling wave high-resolution mass spectrometer for determination of double-bond position in lipids. Rapid Commun. Mass Spectrom. 2017, 31, 1415–1423. [Google Scholar] [CrossRef]
- Cetraro, N.; Cody, R.B.; Yew, J.Y. Carbon-carbon double bond position elucidation in fatty acids using ozone-coupled direct analysis in real time mass spectrometry. Analyst 2019, 144, 5848–5855. [Google Scholar] [CrossRef]
- Tu, A.; Garrard, K.P.; Said, N.; Muddiman, D.C. In situ Detection of Fatty Acid C=C Positional Isomers by Coupling On-tissue mCPBA Epoxidation with IR-MALDESI Mass Spectrometry. Rapid Commun. Mass Spectrom. 2021, e9119. [Google Scholar] [CrossRef]
- Lu, H.; Zhang, H.; Wei, Y.; Chen, H. Ambient mass spectrometry for the molecular diagnosis of lung cancer. Analyst 2020, 145, 313–320. [Google Scholar] [CrossRef]
- Tang, F.; Guo, C.G.; Ma, X.X.; Zhang, J.; Su, Y.; Tian, R.; Shi, R.Y.; Xia, Y.; Wang, X.H.; Ouyang, Z. Rapid In Situ Profiling of Lipid C=C Location Isomers in Tissue Using Ambient Mass Spectrometry with Photochemical Reactions. Anal. Chem. 2018, 90, 5612–5619. [Google Scholar] [CrossRef]
- Su, Y.; Ma, X.X.; Page, J.; Shi, R.Y.; Xia, Y.; Ouyang, Z. Mapping lipid C=C location isomers in organ tissues by Coupling photochemical derivatization and rapid extractive mass spectrometry. Int. J. Mass Spectrom. 2019, 445, 116206. [Google Scholar] [CrossRef]
- Zhao, X.; Chen, J.; Zhang, W.Y.; Yang, C.D.; Ma, X.X.; Zhang, S.C.; Zhang, X.R. Lipid Alterations during Zebrafish Embryogenesis Revealed by Dynamic Mass Spectrometry Profiling with C=C Specificity. J. Am. Soc. Mass Spectrom. 2019, 30, 2646–2654. [Google Scholar] [CrossRef]
- Deng, J.; Yang, Y.; Zeng, Z.; Xiao, X.; Li, J.; Luan, T. Discovery of Potential Lipid Biomarkers for Human Colorectal Cancer by In-Capillary Extraction Nanoelectrospray Ionization Mass Spectrometry. Anal. Chem. 2021, 93, 13089–13098. [Google Scholar] [CrossRef]
- Ma, X.X.; Zhao, X.; Li, J.J.; Zhang, W.P.; Cheng, J.X.; Ouyang, Z.; Xia, Y. Photochemical Tagging for Quantitation of Unsaturated Fatty Acids by Mass Spectrometry. Anal. Chem. 2016, 88, 8931–8935. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Y.Y.; Zhao, H.S.; Zhao, X.; Jia, J.; Ma, Q.; Zhang, S.C.; Zhang, X.R.; Chiba, H.; Hui, S.P.; Ma, X.X. Identification and Quantitation of C=C Location Isomers of Unsaturated Fatty Acids by Epoxidation Reaction and Tandem Mass Spectrometry. Anal. Chem. 2017, 89, 10270–10278. [Google Scholar] [CrossRef]
- Cao, W.B.; Ma, X.X.; Li, Z.S.; Zhou, X.Y.; Ouyang, Z. Locating Carbon-Carbon Double Bonds in Unsaturated Phospholipids by Epoxidation Reaction and Tandem Mass Spectrometry. Anal. Chem. 2018, 90, 10286–10292. [Google Scholar] [CrossRef]
- Chintalapudi, K.; Badu-Tawiah, A.K. An integrated electrocatalytic nESI-MS platform for quantification of fatty acid isomers directly from untreated biofluids. Chem. Sci. 2020, 11, 9891–9897. [Google Scholar] [CrossRef]
- Gaston, R.; Maria Eugenia, P.; Das, U.N.; Eynard, A.R. Polyunsaturated Fatty Acids Differentially Modulate Cell Proliferation and Endocannabinoid System in Two Human Cancer Lines. Arch. Med. Res. 2017, 48, 46–54. [Google Scholar] [CrossRef]
- Nakkarach, A.; Foo, H.L.; Song, A.A.; Mutalib, N.E.A.; Nitisinprasert, S.; Withayagiat, U. Anti-cancer and anti-inflammatory effects elicited by short chain fatty acids produced by Escherichia coli isolated from healthy human gut microbiota. Microb. Cell Fact. 2021, 20, 36. [Google Scholar] [CrossRef]
- Hu, R.; Li, Y.; Yang, Y.; Liu, M. Mass spectrometry-based strategies for single-cell metabolomics. Mass Spectrom. Rev. 2021. [Google Scholar] [CrossRef]
- Wu, C.; Dill, A.L.; Eberlin, L.S.; Cooks, R.G.; Ifa, D.R. Mass spectrometry imaging under ambient conditions. Mass Spectrom. Rev. 2013, 32, 218–243. [Google Scholar] [CrossRef] [Green Version]
- Handberg, E.; Chingin, K.; Wang, N.; Dai, X.; Chen, H. Mass spectrometry imaging for visualizing organic analytes in food. Mass Spectrom. Rev. 2015, 34, 641–658. [Google Scholar] [CrossRef]
- Gemperline, E.; Chen, B.; Li, L. Challenges and recent advances in mass spectrometric imaging of neurotransmitters. Bioanalysis 2014, 6, 525–540. [Google Scholar] [CrossRef] [Green Version]
- Johnson, J.; Sharick, J.T.; Skala, M.C.; Li, L. Sample preparation strategies for high-throughput mass spectrometry imaging of primary tumor organoids. J. Mass Spectrom. 2020, 55, e4452. [Google Scholar] [CrossRef] [PubMed]
- Marshall, D.L.; Criscuolo, A.; Young, R.S.E.; Poad, B.L.J.; Zeller, M.; Reid, G.E.; Mitchell, T.W.; Blanksby, S.J. Mapping Unsaturation in Human Plasma Lipids by Data-Independent Ozone-Induced Dissociation. J. Am. Soc. Mass Spectrom. 2019, 30, 1621–1630. [Google Scholar] [CrossRef] [PubMed]
- Bednarik, A.; Bolsker, S.; Soltwisch, J.; Dreisewerd, K. An On-Tissue Paterno-Buchi Reaction for Localization of Carbon-Carbon Double Bonds in Phospholipids and Glycolipids by Matrix-Assisted Laser-Desorption-Ionization Mass-Spectrometry Imaging. Angew. Chem. Int. Ed. 2018, 57, 12092–12096. [Google Scholar] [CrossRef] [PubMed]
- Claes, B.S.R.; Bowman, A.P.; Poad, B.L.J.; Young, R.S.E.; Heeren, R.M.A.; Blanksby, S.J.; Ellis, S.R. Mass Spectrometry Imaging of Lipids with Isomer Resolution Using High-Pressure Ozone-Induced Dissociation. Anal. Chem. 2021, 93, 9826–9834. [Google Scholar] [CrossRef]
- Bednarik, A.; Preisler, J.; Bezdekova, D.; Machalkova, M.; Hendrych, M.; Navratilova, J.; Knopfova, L.; Moskovets, E.; Soltwisch, J.; Dreisewerd, K. Ozonization of Tissue Sections for MALDI MS Imaging of Carbon-Carbon Double Bond Positional Isomers of Phospholipids. Anal. Chem. 2020, 92, 6245–6250. [Google Scholar] [CrossRef]
- Sun, C.; Liu, W.; Geng, Y.; Wang, X. On-Tissue Derivatization Strategy for Mass Spectrometry Imaging of Carboxyl-Containing Metabolites in Biological Tissues. Anal. Chem. 2020, 92, 12126–12131. [Google Scholar] [CrossRef]
- Klein, D.R.; Feider, C.L.; Garza, K.Y.; Lin, J.Q.; Eberlin, L.S.; Brodbelt, J.S. Desorption Electrospray Ionization Coupled with Ultraviolet Photodissociation for Characterization of Phospholipid Isomers in Tissue Sections. Anal. Chem. 2018, 90, 10100–10104. [Google Scholar] [CrossRef] [Green Version]
- Xia, F.; Wan, J.B. Chemical derivatization strategy for mass spectrometry-based lipidomics. Mass Spectrom. Rev. 2021, e21729. [Google Scholar] [CrossRef]
| Derivatization Strategies | Reagents | AIMS Techniques | Sample | Analytes | LOD | Ref. |
|---|---|---|---|---|---|---|
| PB | Acetone or acetonitrile solution containing benzophenone | nanoESI-MS | Single cells | FAs, SMs, and PCs | 0.1 pM | [6] |
| Acetone | LMJ-SSP-MS | The sections of rat brain, lung, liver, spleen, and kidney, as well as in normal and diseased rat tissues | FAs and PCs | N/A | [67] | |
| nanoESI-MS | Rat brain tissue and rat organ tissues (including kidney, liver, and muscle), normal and cancerous mouse breast tissues | FAs, PEs, LPEs, LPSs, PIs, LPIs, PCs, LPCs, and GPs | N/A | [16] | ||
| Rat brain, kidney, and liver tissues | FAs, PAs, LPAs, PGs, PSs, LPGs, PEs, and PIs | N/A | [68] | |||
| Human plasma, whole blood, and cell lines | FAs | 15 nM | [71] | |||
| Zebrafish embryos | PCs | N/A | [69] | |||
| Bovine blood, human blood, rat blood, and homogenized mouse brain samples | FAs, PEs, PSs, and PCs | N/A | [48] | |||
| Benzophenone | SCP-nanoESI-MS | Perilla seed, Zebrafish muscle, and human intestinal tissue | FAs, PCs, and TAGs | N/A | [47] | |
| CH3OH/CH2Cl2/H2O (v/v/v = 8:1:1) containing 0.4 mg/mL benzophenone and 0.4 mg/mL NH4HCO3 | ICE-nanoESI-MS | Human colorectal cancer tumors and paired distal noncancerous tissues | PCs, LPs, SMs, FAs, PAs, LPAs, PSs, LPSs, PGs, LPGs, PIs, LPIs, PEs, LPEs, CLs, and Cers | N/A | [70] | |
| OzID | Ozone | DART-MS | Hawaiian Drosophila species | FAs | N/A | [64] |
| Epoxidation | peracetic acid | nanoESI-MS | (HPDE/E6E7) cells and (PANC-1) cells | FAs | 4.2 nM | [5] |
| m-CPBA | IR-MALDESI | Tissue sections of rat liver and human bladder | FAs | 2–4 fmol per laser spot | [65] | |
| DESI-MS | Mouse kidney and metastatic lung tissue | FAs and PGs | 10~100 pmole | [25] | ||
| HAuCl4 | microdroplet-MS | Standards | LA, RA, IA, NA, DOPC, and lysoPC | N/A | [53] | |
| Electro-epoxidation | Hydrochloric acid and an acetonitrile/water | nanoESI-MS | Chicken egg yolk | PGs, PSs, and FAs | 10 nM | [58] |
| Electro-oxidation | Ir and Ru | Serum | FAs | 1.18–8.00 μM | [74] | |
| Oxidation | 1O2 | nano-DESI-MS | Rat brain, mouse uterine, and gastrocnemius muscle tissue | FAs, LPEs, PEs, and PCs | N/A | [2] |
| UVPD | 193 nm | DESI-MS | Mouse brain and kidney tissues, pancreas, kidney, lung, fallopian, ovarian tissue, and endometrial tissue | PCs | ∼500 nM | [87] |
| Mouse brain tissue, human ovarian tumor tissue, and breast cancer tissue | FAs | N/A | [21] | |||
| LTP | Pure helium | nanoESI-MS | Human plasma | FAs | 0.07 μM | [72] |
| Oxygen | Bovine liver polar extract | PCs, PAs, PEs, PGs, and PIs | N/A | [73] | ||
| Reactive oxygen species in the plasma | LTP-MS | Human, Equine, and Fetal Bovine Serums | FAs | 0.1 μM | [55] | |
| Ozone | Salmonella enterica typhimurium INSP24 (SARA1) | FAs and FAEE | N/A | [54] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lu, H.; Zhang, H.; Xu, S.; Li, L. Review of Recent Advances in Lipid Analysis of Biological Samples via Ambient Ionization Mass Spectrometry. Metabolites 2021, 11, 781. https://doi.org/10.3390/metabo11110781
Lu H, Zhang H, Xu S, Li L. Review of Recent Advances in Lipid Analysis of Biological Samples via Ambient Ionization Mass Spectrometry. Metabolites. 2021; 11(11):781. https://doi.org/10.3390/metabo11110781
Chicago/Turabian StyleLu, Haiyan, Hua Zhang, Shuling Xu, and Lingjun Li. 2021. "Review of Recent Advances in Lipid Analysis of Biological Samples via Ambient Ionization Mass Spectrometry" Metabolites 11, no. 11: 781. https://doi.org/10.3390/metabo11110781
APA StyleLu, H., Zhang, H., Xu, S., & Li, L. (2021). Review of Recent Advances in Lipid Analysis of Biological Samples via Ambient Ionization Mass Spectrometry. Metabolites, 11(11), 781. https://doi.org/10.3390/metabo11110781