5-CQA and Mangiferin, Two Leaf Biomarkers of Adaptation to Full Sun or Shade Conditions in Coffea arabica L.
Abstract
1. Introduction
2. Results
2.1. Identification of Phenolic Compounds in Mature Leaves of C. arabica Using LC-MS2
2.2. Influence of Light Intensity on Leaf Phenolic Content in One C. arabica Cultivar at Low or High Elevation
2.3. Influence of Light Intensity on the Leaf Phenolic Content of Numerous C. arabica Genotypes Grown in Different Environments
2.4. Influence of C. arabica Genetic Groups on Leaf Phenolic Contents in Two Different Environments
3. Discussion
3.1. LC-MS2 Analysis Enabled Identification of Two Novel Mangiferin Derivatives in C. arabica
3.2. Identification of Leaf Phenolic Contents as Biomarkers of Adaptation of C. arabica to Full-Sun or Shade Conditions
3.2.1. Preselection of Candidate Biomarkers of Adaptation of C. arabica to Full-Sun Conditions
3.2.2. Selection of Biomarkers for Adaptation of C. arabica to Full Sunlight Using Genetic Diversity and Contrasting Environments
3.2.3. Influence of C. arabica Genotype on the Phenolic Leaf Contents and Selection of Biomarkers of Adaptation to Full Sunlight or Shade in Breeding Programs
4. Materials and Methods
4.1. Locations and Plant Material
4.2. Leaf Sampling for Secondary Metabolite Analysis
4.3. Secondary Metabolite Analysis
4.4. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| Amu | atomic mass unit |
| APL | American pure lines |
| CGAs | chlorogenic acid group (CQA, diCQA, Coum-QA and FQA) |
| Coum-QA | coumaroyl quinic acid |
| 3-CQA | 3-O-caffeoylquinic acid |
| 4-CQA | 4-O-caffeoylquinic acid |
| 5-CQA | 5-O-caffeoylquinic acid |
| 3,4-diCQA | 3,4-O-dicaffeoylquinic acid |
| 3,5-diCQA | 3,5-O-dicaffeoylquinic acid |
| 4,5-diCQA | 4,5-O-dicaffeoylquinic acid |
| EWA | Ethiopian wild accessions |
| F-dihex | flavone-di-C-hexose |
| FQA | feruloylquinic acid |
| HF1 | hybrid F1 clone |
| HPLC-DAD | High performance liquid chromatograph/ photodiode array detection |
| K-dihex-dhex | kaempferol-3-O-di-hexose-deoxyhexose |
| K-hex-dhex | kaempferol-3-O-hexose-deoxyhexose |
| LC-DAD-MS2 | High performance liquid chromatograph/ photodiode array detection /mass spectrometry2 |
| pOH Mang | mangiferin parahydroxybenzoate |
| Q-dihex-dhex | quercetin-3-O-dihexose-deoxyhexose |
| Q-diGlu | quercetin-3-O-diglucoside |
| SNP | single-nucleotide polymorphism |
References
- Sylvain, P.G. Some observations on Coffea arabica L. in Ethiopia. Turrialba 1955, 5, 37–53. [Google Scholar]
- Fernie, L.M.; Greathead, D.J.; Meyer, F.G.; Monaco, L.C.; Narasimhaswamy, R.L. FAO Coffee Mission to Ethiopia, 1964–1965; FAO: Tome, Italy, 1968; p. 214. [Google Scholar]
- Scalabrin, S.; Toniutti, L.; Di Gaspero, G.; Scaglione, D.; Magris, G.; Vidotto, M.; Pinosio, S.; Cattonaro, F.; Magni, F.; Jurman, I.; et al. A single polyploidization event at the origin of the tetraploid genome of Coffea arabica is responsible for the extremely low genetic variation in wild and cultivated germplasm. Sci. Rep. 2020. [Google Scholar] [CrossRef]
- Anthony, F.; Combes, M.; Astorga, C.; Bertrand, B.; Graziosi, G.; Lashermes, P. The origin of cultivated Coffea arabica L. varieties revealed by AFLP and SSR markers. Theor. Appl. Genet. 2002, 104, 894–900. [Google Scholar] [PubMed]
- Silva, M.C.; Várzea, V.; Guerra-Guimarães, L.; Azinheira, H.G.; Fernandez, D.; Petitot, A.-S.; Bertrand, B.; Lashermes, P.; Nicole, M. Coffee resistance to the main diseases: Leaf rust and coffee berry disease. Braz. J. Plant Physiol. 2006, 18, 119–147. [Google Scholar]
- Van der Vossen, H.; Bertrand, B.; Charrier, A. Next generation variety development for sustainable production of Arabica coffee (Coffea arabica L.): A review. Euphytica 2015, 204, 243–256. [Google Scholar]
- Setotaw, T.A.; Texeira Caixeta, E.; Fereire Pena, G.; Zambolim, E.M.; Pereira, A.A.; Sakiyama, N.S. Breeding potential and genetic diversity of ‘‘Hibrido do Timor’’ coffee evaluated by molecular markers. Crop Breed. Appl. Biotechnol. 2010, 10, 298–304. [Google Scholar]
- Jha, S.; Bacon, C.M.; Philpott, S.M.; Rice, R.A.; Méndez, V.E.; Läderach, P. A review of ecosystem services, farmer livelihoods, and value chains in shade coffee agroecosystems. In Integrating Agriculture, Conservation and Ecotourism: Examples from the Field; Campbell, W.B., López-Ortiz, S., Eds.; Springer: Dordrecht, The Netherlands, 2011; pp. 141–208. [Google Scholar]
- Bertrand, B.; Marraccini, P.; Villain, L.; Breitler, J.C.; Etienne, H. Healthy tropical plants to mitigate the impact of climate change—As exemplified in coffee. In Climate Change and Agriculture Worldwide; Torquebiau, E., Ed.; Springer: Berlin/Heidelberg, Germany, 2016; pp. 83–95. [Google Scholar]
- Bennett, R.N.; Wallsgrove, R.M. Secondary metabolites in plant defence mechanisms. New Phytol. 1994, 127, 617–633. [Google Scholar] [CrossRef]
- Sampaio, B.L.; Edrada-Ebel, R.; Da Costa, F.B. Effect of the environment on the secondary metabolic profile of Tithonia diversifolia: A model for environmental metabolomics of plants. Sci. Rep. 2016, 6. [Google Scholar] [CrossRef]
- Winkel-Shirley, B. Biosynthesis of flavonoids and effects of stress. Curr. Opin. Plant Biol. 2002, 5, 218–223. [Google Scholar] [CrossRef]
- Tattini, M.; Galardi, C.; Pinelli, P.; Massai, R.; Remorini, D.; Agati, G. Differential accumulation of flavonoids and hydroxycinnamates in leaves of Ligustrum vulgare under excess light and drought stress. New Phytol. 2004, 163, 547–561. [Google Scholar] [CrossRef]
- Pavarini, D.P.; Pavarini, S.P.; Niehues, M.; Lopes, N.P. Exogenous influences on plant secondary metabolite levels. Anim. Feed Sci. Technol. 2012, 176, 5–16. [Google Scholar] [CrossRef]
- Clé, C.; Hill, L.M.; Niggeweg, R.; Martin, C.R.; Guisez, Y.; Prinsen, E.; Jansen, M.A.K. Modulation of chlorogenic acid biosynthesis in Solanum lycopersicum; consequences for phenolic accumulation and UV-tolerance. Phytochem. 2008, 69, 2149–2156. [Google Scholar] [CrossRef] [PubMed]
- Hectors, K.; Van Oevelen, S.; Geuns, J.; Guisez, Y.; Jansen, M.A.K.K.; Prinsen, E. Dynamic changes in plant secondary metabolites during UV acclimation in Arabidopsis thaliana. Physiol. Plant. 2014, 152, 219–230. [Google Scholar] [CrossRef] [PubMed]
- Grace, S.C.; Logan, B.A. Energy dissipation and radical scavenging by the plant phenylpropanoid pathway. Philos. Trans R. Soc. B Biol. Sci. 2000, 355, 1499–1510. [Google Scholar] [CrossRef] [PubMed]
- Yao, K.; Luca, V.D.; Brisson, N. Creation of a metabolic sink for tryptophan alters the phenylpropanoid pathway and the susceptibility of potato to phytophthora infestans. Plant Cell 1995, 7, 1787–1799. [Google Scholar] [CrossRef]
- Díaz, J.; Barceló, A.R.; De Cáceres, F.M. Changes in shikimate dehydrogenase and the end products of the shikimate pathway, chlorogenic acid and lignins, during the early development of seedlings of Capsicum annuum. New Phytol. 1997, 136, 183–188. [Google Scholar] [CrossRef]
- Ranheim, T.; Halvorsen, B. Coffee consumption and human health–beneficial or detrimental? Mechanisms for effects of coffee consumption on different risk factors for cardiovascular disease and type 2 diabetes mellitus. Mol. Nutr. Food Res. 2005, 49, 274–284. [Google Scholar] [CrossRef]
- Tajik, N.; Tajik, M.; Mack, I.; Enck, P. The potential effects of chlorogenic acid, the main phenolic components in coffee, on health: A comprehensive review of the literature. Eur. J. Nutr. 2017, 56, 2215–2244. [Google Scholar] [CrossRef]
- Gamboa-Becerra, R.; Hernández-Hernández, M.C.; González-Ríos, Ó.; Suárez-Quiroz, M.L.; Gálvez-Ponce, E.; Ordaz-Ortiz, J.J.; Winkler, R. Metabolomic Markers for the Early Selection of Coffea Canephora Plants with Desirable Cup Quality Traits. Metabolites 2019, 9, 214. [Google Scholar] [CrossRef]
- Anthony, F.; Clifford, M.N.; Noirot, M. Biochemical diversity in the genus Coffea L.: Chlorogenic acids, caffeine and mozambioside contents. Genet. Resour. Crop Evol. 1993, 40, 61–70. [Google Scholar] [CrossRef]
- Campa, C.; Rakotomalala, J.J.; de Kochko, A.; Hamon, S. Chlorogenic acids: Diversity in green beans of wild coffee species. Adv. Plant Physiol. 2008, 10, 421–437. [Google Scholar]
- Dussert, S.; Laffargue, A.; de Kochko, A.; Joët, T. Effectiveness of the fatty acid and sterol composition of seeds for the chemotaxonomy of Coffea subgenus Coffea. Phytochemistry 2008, 69, 2950–2960. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, B.; Villarreal, D.; Laffargue, A.; Posada, H.; Lashermes, P.; Dussert, S. Comparison of the effectiveness of fatty acids, chlorogenic acids and elements for the chemometric discrimination of coffee (Coffea arabica L.) varieties and growing origins. J. Agric. Food Chem. 2008, 56, 2273–2280. [Google Scholar] [CrossRef] [PubMed]
- Bertrand, B.; Boulanger, R.; Dussert, S.; Joët, T. Climatic factors directly impact the volatile organic compound fingerprint in green Arabica coffee bean as well as coffee beverage quality. Food Chem. 2012, 135, 2575–2583. [Google Scholar] [CrossRef] [PubMed]
- Ashihara, H.; Sano, H.; Crozier, A. Caffeine and related purine alkaloids: Biosynthesis, catabolism, function and genetic engineering. Phytochemistry 2008, 69, 841–856. [Google Scholar] [CrossRef] [PubMed]
- Magalhães, S.T.V.; Fernandes, F.L.; Demuner, A.J.; Picanço, M.C.; Guedes, R.N.C. Leaf alkaloids, phenolics, and coffee resistance to the leaf miner Leucoptera coffeella (Lepidoptera: Lyonetiidae). J. Econ. Entomol. 2010, 103, 1438–1443. [Google Scholar] [CrossRef] [PubMed]
- Martins, S.C.V.; Araujo, W.L.; Tohge, T.; Fernie, A.R.; Da Matta, F.M. In high-light-acclimated coffee plants the metabolic machinery is adjusted to avoid oxidative stress rather than to benefit from extra light enhancement in photosynthetic yield. PLoS ONE 2014, 9, e94862. [Google Scholar] [CrossRef]
- Campa, C.; Mondolot, L.; Rakotondravao, A.; Bidel, L.P.R.; Gargadennec, A.; Couturon, E.; La Fisca, P.; Rakotomalala, J.J.; Jay-Allemand, C.; Davis, A.P. A survey of mangiferin and hydroxycinnamic acid ester accumulation in coffee (Coffea) leaves: Biological implications and uses. Ann. Bot. 2012, 110, 595–613. [Google Scholar] [CrossRef]
- Trevisan, M.T.S.; Farias de Almeida, R.; Soto, G.; De Melo, V.F.E.; Ulrich, C.M.; Owen, R.W. Quantitation by HPLC-UV of mangiferin and isomangiferin in Coffee (Coffea arabica) leaves from brazil and costa rica after solvent extraction and infusion. Food Anal. Methods 2016, 9, 2649–2655. [Google Scholar] [CrossRef]
- Chen, X.-M.; Ma, Z.; Kitts, D.D. Effects of processing method and age of leaves on phytochemical profiles and bioactivity of coffee leaves. Food Chem. 2018, 249, 143–153. [Google Scholar] [CrossRef]
- De Almeida, R.F.; Salles Trevisan, M.T.; Thomaziello, R.A.; Breuer, A.; Klika, K.D.; Ulrich, C.M.; Owen, R.W. Nutraceutical compounds: Echinoids, flavonoids, xanthones and caffeine identified and quantitated in the leaves of coffea arabica trees from three regions of Brazil. Food Res. Int. 2019, 115, 493–503. [Google Scholar] [CrossRef]
- Fortunato, A.S.; Lidon, F.C.; Batista-Santos, P.; Leitão, A.E.; Pais, I.P.; Ribeiro, A.I.; Ramalho, J.C. Biochemical and molecular characterization of the antioxidative system of Coffea sp. under cold conditions in genotypes with contrasting tolerance. J. Plant Physiol. 2010, 167, 333–342. [Google Scholar] [CrossRef] [PubMed]
- Schütz, K.; Kammerer, D.R.; Carle, R.; Schieber, A. Characterization of phenolic acids and flavonoids in dandelion (Taraxacum officinale WEB. ex WIGG.) root and herb by high-performance liquid chromatography/electrospray ionization mass spectrometry. Rapid Commun. Mass Spectrom. 2005, 19, 179–186. [Google Scholar] [CrossRef] [PubMed]
- Willems, J.L.; Khamis, M.M.; Saeid, W.M.; Purves, R.W.; Katselis, G.; Low, N.H.; El-Aneed, A. Analysis of a series of chlorogenic acid isomers using differential ion mobility and tandem mass spectrometry. Anal. Chim. Acta 2016, 933, 164–174. [Google Scholar] [CrossRef] [PubMed]
- Ito, T.; Kakino, M.; Tazawa, S.; Oyama, M.; Maruyama, H.; Araki, Y.; Hara, H.; Iinuma, M. Identification of phenolic compounds in aquilaria crassna leaves via liquid chromatography-electrospray ionization mass spectroscopy. Food Sci. Technol. Res. 2012, 18, 259–262. [Google Scholar] [CrossRef]
- Mabry, T.; Markham, K.R.; Thomas, M.B. The ultraviolet spectra of flavones and flavonols. In The Systematic Identification of Flavonoids; Mabry, T., Markham, K.R., Thomas, M.B., Eds.; Springer: Berlin/Heidelberg, Germany, 1970; pp. 41–164. [Google Scholar]
- Zhong, R.-F.; Xu, G.-B.; Wang, Z.; Wang, A.-M.; Guan, H.-Y.; Li, J.; He, X.; Liu, J.-H.; Zhou, M.; Li, Y.-J.; et al. Identification of anti-inflammatory constituents from Kalimeris indica with UHPLC-ESI-Q-TOF-MS/MS and GC–M. J. Ethnopharmacol. 2015, 165, 39–45. [Google Scholar] [CrossRef]
- Schmidt, W.; Beerhues, L. Alternative pathways of xanthone biosynthesis in cell cultures of Hypericum and rosaenum L. FEBS Lett. 1997, 420, 143–146. [Google Scholar] [CrossRef]
- Fujita, M.; Takao, I. Further studies on the biosynthesis of mangiferin in anemarrhena asphodeloides: Hydroxylation of the shikimate-derived ring. Phytochemistry 1981, 20, 2183–2185. [Google Scholar] [CrossRef]
- Joubert, E.; de Beer, D.; Hernandez, I.; Munné-Bosch, S. Accumulation of mangiferin, isomangiferin, iriflophenone-3-C-b-glucoside and hesperidin in honeybush leaves (Cyclopia genistoides Vent.) in response to harvest time, harvest interval and seed source. Ind. Crops Prod. 2014, 56, 74–82. [Google Scholar] [CrossRef]
- Ratanamarno, S.; Surbkar, S. Caffeine and catechine in fresh coffee leaf (Coffea arabica) and coffee leaf tea. Maejo Int. J. Sci. Technol. 2017, 11, 211–218. [Google Scholar]
- Bernal, M.; Llorens, L.; Julkunen-Tiitto, R.; Badosa, J.; Verdaguer, D. Altitudinal and seasonal changes of phenolic compounds in Buxus sempervirens and cuticles. Plant Physiol. Biochem. 2013, 70, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Petrussa, E.; Braidot, E.; Zancani, M.; Peresson, C.; Bertolini, A.; Patui, S.; Vianello, A. Plant flavonoids—Biosynthesis, transport and involvement in stress responses. Int. J. Mol. Sci. 2013, 14, 14950–14973. [Google Scholar] [CrossRef] [PubMed]
- Sharma, A.; Shahzad, B.; Rehman, A.; Bhardwaj, R.; Landi, M.; Zheng, B. Response of phenylpropanoid pathway and the role of polyphenols in plants under abiotic stress. Molecules 2019, 24, 2452. [Google Scholar] [CrossRef] [PubMed]
- Franklin, G.; Conceição, L.F.R.; Kombrink, E.; Dias, A.C.P. Xanthone biosynthesis in hypericum perforatum cells provides antioxidant and antimicrobial protection upon biotic stress. Phytochemistry 2009, 70, 60–68. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Sadhukhan, P.; Sil, P.C. Mangiferin: A xanthonoid with multipotent anti-inflammatory potential. BioFactors 2016, 42, 459–474. [Google Scholar] [CrossRef]
- Imran, M.; Arshad, M.S.; Butt, M.S.; Kwon, J.-H.; Arshad, M.U.; Sultan, M.T. Mangiferin: A natural miracle bioactive compound against lifestyle related disorders. Lipids Health Dis. 2017, 16. [Google Scholar] [CrossRef]
- Danthu, P.; Lubrano, C.; Flavet, L.; Rahajanirina, V.; Behra, O.; Fromageot, C.; Rabevohitra, R.; Roger, E. Biological factors influencing production of xanthones in aphloia theiformis. Chem. Biodivers. 2010, 7, 140–150. [Google Scholar] [CrossRef]
- Downey, G.; Boussion, J. Authentication of coffee bean variety by near-infrared reflectance spectroscopy of dried extract. J. Sci. Food Agric. 1996, 71, 41–49. [Google Scholar] [CrossRef]
- Vaast, P.; Bertrand, B.; Perriot, J.-J.; Guyot, B.; Génard, M. Fruit thinning and shade improve bean characteristics and beverage quality of coffee (Coffea arabica L.) under optimal conditions. J. Sci. Food Agric. 2006, 86, 197–204. [Google Scholar] [CrossRef]
- Bertrand, B.; Vaast, P.; Alpizar, E.; Etienne, H.; Davrieux, F.; Charmetant, P. Comparison of bean biochemical composition and beverage quality of Arabica hybrids involving Sudanese-Ethiopian origins with traditional varieties at various elevations in Central America. Tree Physiol. 2006, 26, 1239–1248. [Google Scholar] [CrossRef]
- Okubo, N.; Kurata, Y. Nondestructive classification analysis of green coffee beans by using near-infrared spectroscopy. Foods 2019, 8, 82. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, M.; Morina, F.; Jovanovic, S.V. 2-Stimulation of various phenolics in plants under ambient UV-B radiation. In UV-B Radiation: From Environmental Stressor to Regulator of Plant Growth; Singh, V.P., Singh, S., Prasad, S.M., Parihar, P., Eds.; John Wiley & Sons Ltd.: Hoboken, NJ, USA, 2017; pp. 9–56. [Google Scholar]
- Ryan, K.G.; Swinny, E.E.; Winefielf, C.; Markham, K.R. Flavonoids and UV photoprotection in Arabidopsis mutants. Z. Für Nat. C 2001, 56, 745–754. [Google Scholar] [CrossRef] [PubMed]
- Campa, C.; Urban, L.; Mondolot, L.; Fabre, D.; Roques, S.; Lizzi, Y.; Aarouf, J.; Doulbeau, S.; Breitler, J.-C.; Letrez, C.; et al. Juvenile coffee leaves acclimated to low light are unable to cope with a moderate light increase. Front Plant Sci. 2017, 8, 1126. [Google Scholar] [CrossRef] [PubMed]
- Agati, G.; Cerovic, Z.G.; Pinelli, P.; Tattini, M. Light-induced accumulation of ortho-dihydroxylated flavonoids as non-destructively monitored by chlorophyll fluorescence excitation techniques. Environ. Exp. Bot. 2011, 73, 3–9. [Google Scholar] [CrossRef]
- Agati, G.; Brunetti, C.; Di Fernando, M.; Ferrini, F.; Pollastri, S.; Tattini, M. Functional roles of flavonoids in photoprotection: New evidence, lessons from the past. Plant. Physiol. Biochem. 2013, 72, 35–45. [Google Scholar] [CrossRef]
- Zhang, Q.; Liu, M.; Ruan, J. Metabolomics analysis reveals the metabolic and functional roles of flavonoids in light-sensitive tea leaves. BMC Plant. Biol. 2017, 17, 64. [Google Scholar] [CrossRef]
- R Development Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2005; ISBN 3-900051-07-0. Available online: http://www.R-project.org.
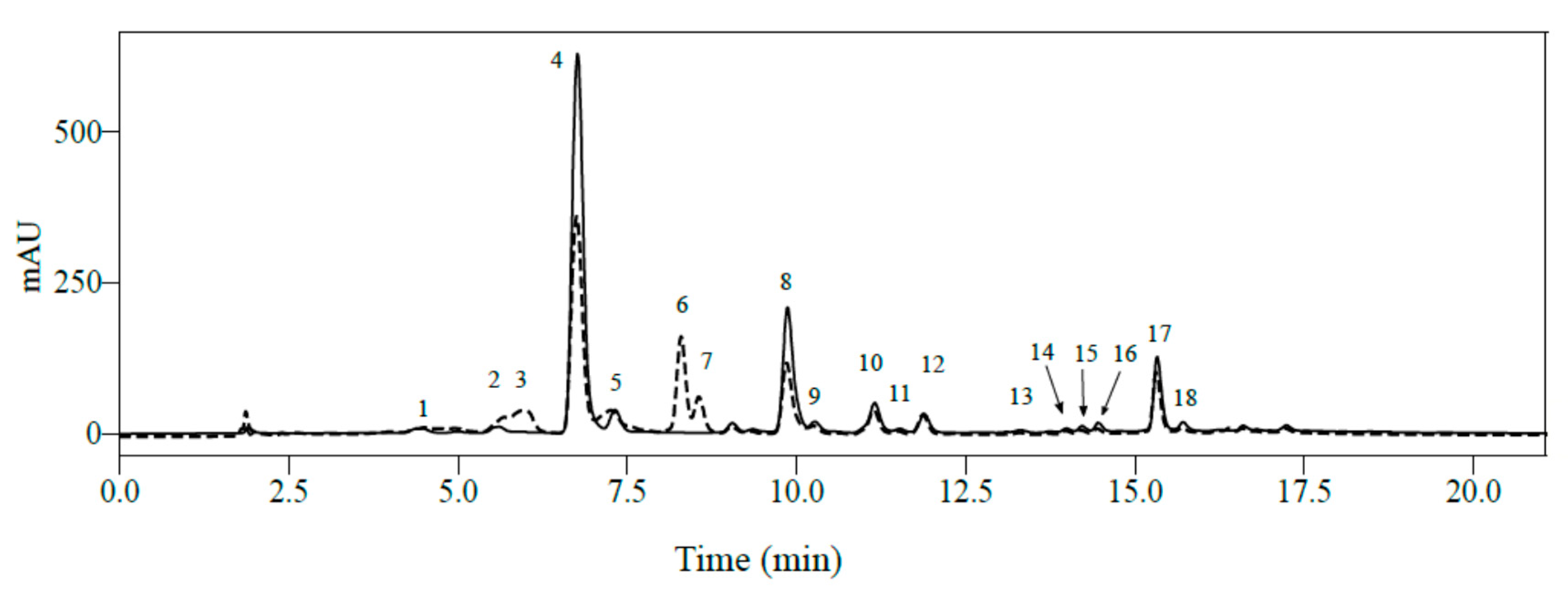
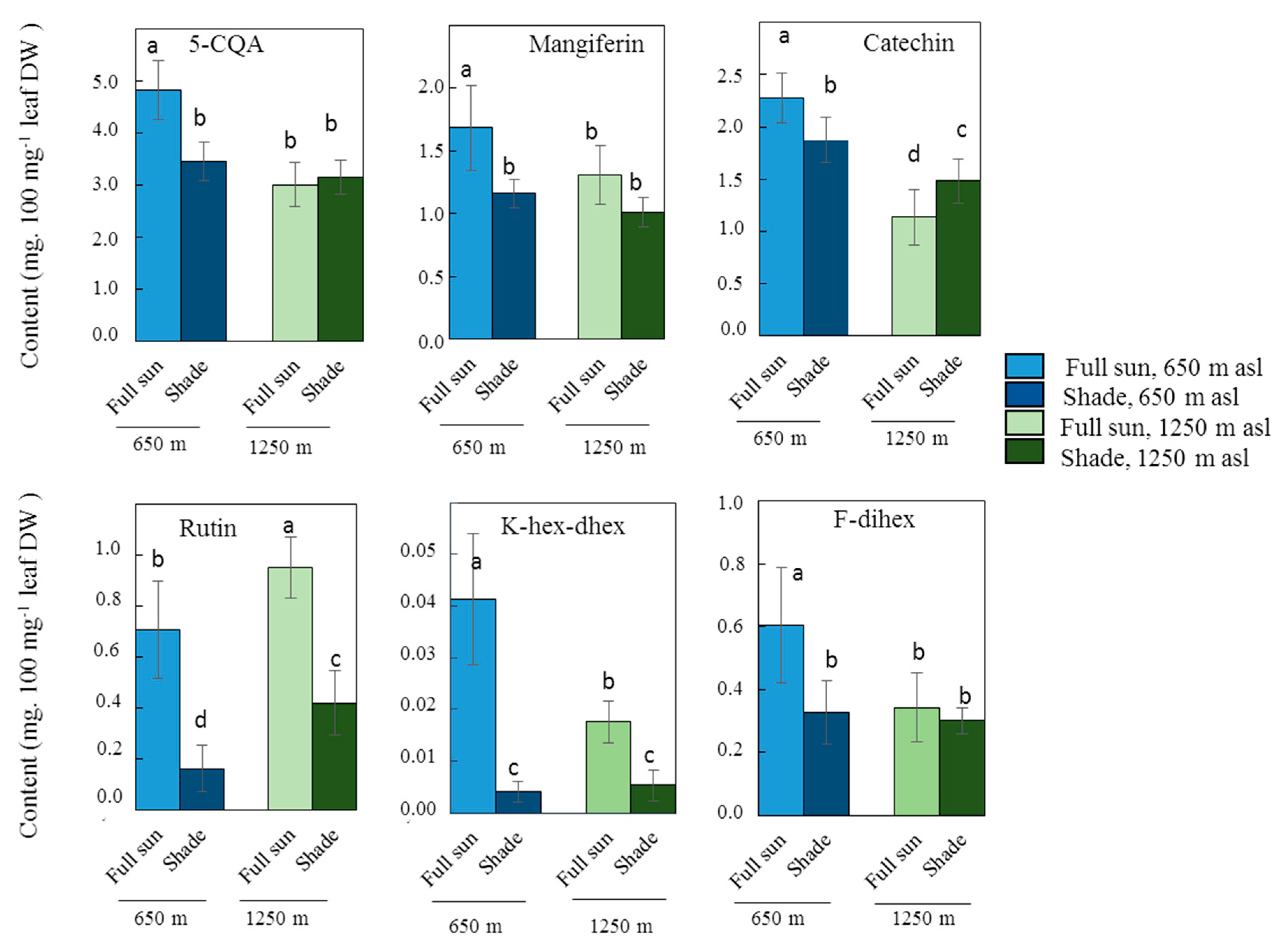

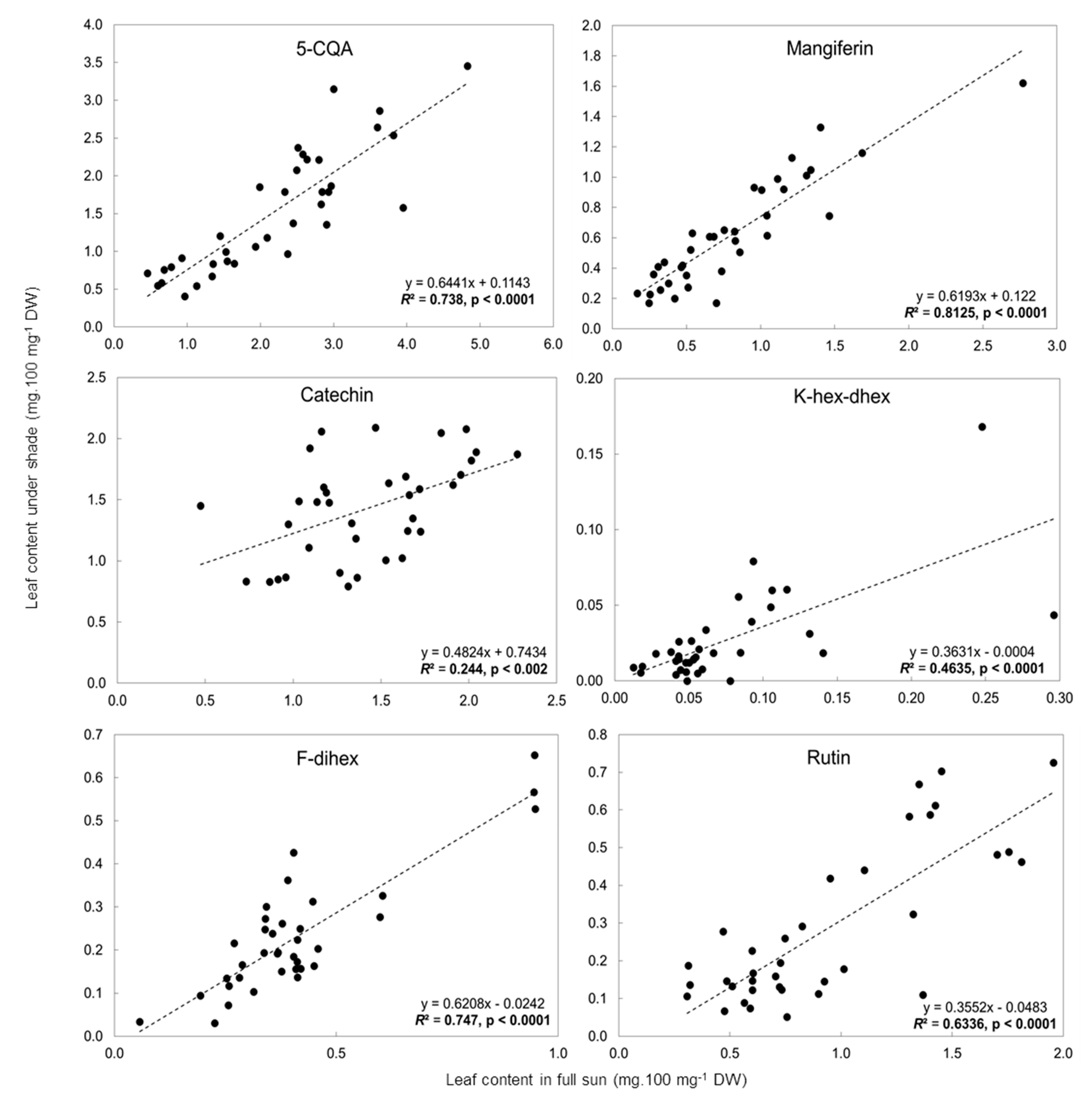

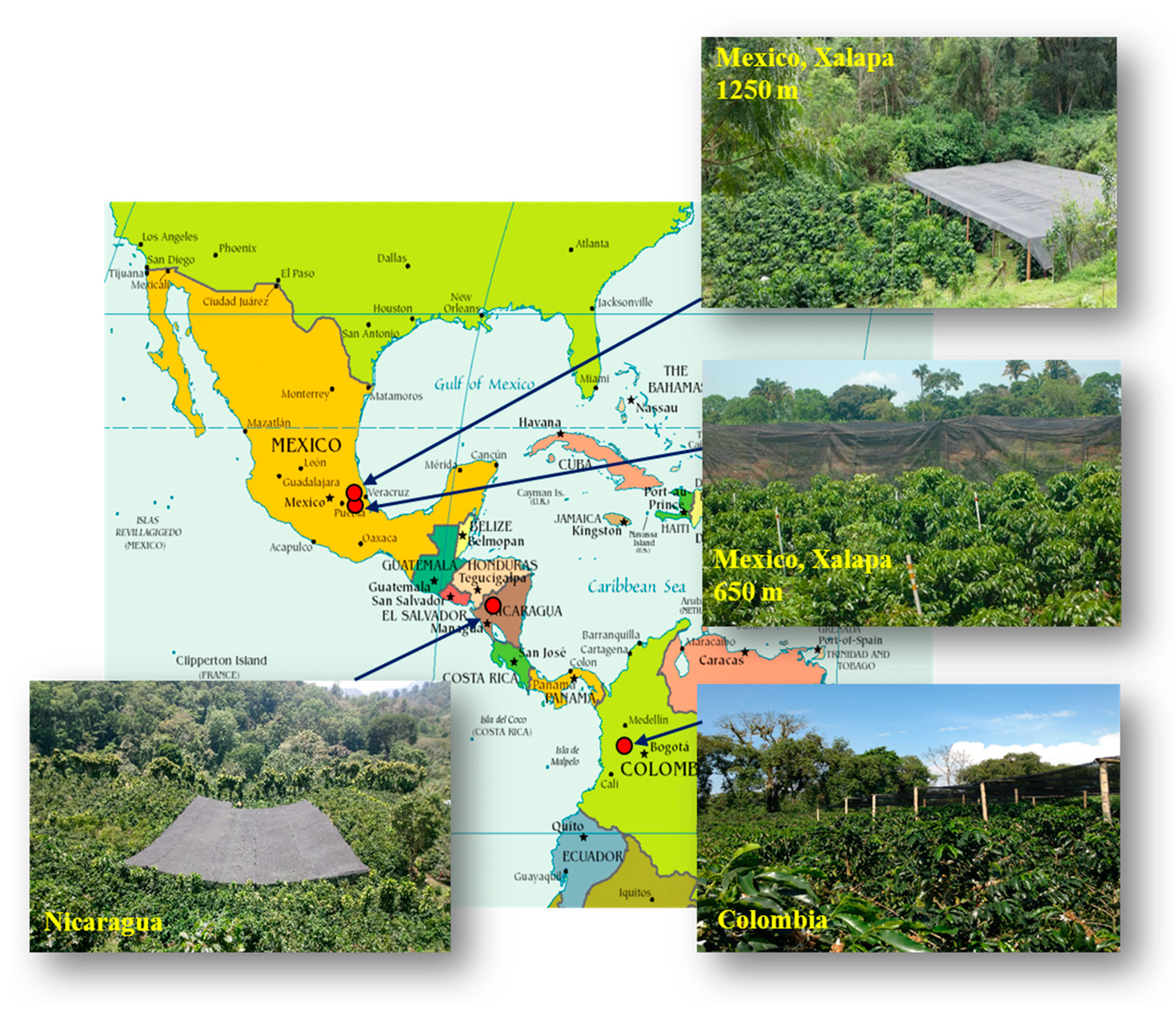
| Peak N° | RT (min) | Mass (Da) | Molecular Formula | Wavelength (nm) | Compound Name | [M − H]− | MS2 | [M+H]+ | MS2 |
|---|---|---|---|---|---|---|---|---|---|
| 1 | 2.61 | 180.1640 | C7H8N4O2 | 272 | Theobromine * | - | 181.0718 | 67 (100) 85 (78) 181 (74) 138 (65) 69 (61) 108 (61) 56 (43) 163 (43) 122 (28) 117 (13) 156 (9) | |
| 2 | 3.00 | 354.3087 | C16H18O10 | 300sh-325 | 3-O-Caffeoylquinic acid | 353.0901 | 191 (100) 179 (61) 135 (47) 173 (6) | 355.1002 | 163 (100) 145 (24) 89 (9) |
| 3 | 4.02 | 408.3561 | C19H20O10 | 286 | Iriflophenone-C-hexose | 407.1012 | 287 (100) 317 (23) 245 (61) 193 (47) 125 (8) | 409.1107 | 195 (100) 231 (79) 13 (61) 325 (46) 177 (43) 121 (39) 271 (39) 355 (14) 85 (7) |
| 4 | 4.20 | 290.2700 | C15H14O6 | 278 | (+)-Catechin * | 289.0769 | 109 (100) 123 (69) 125 (64) 203 (56) 151 (44) 245 (40) 137 (36) 205 (26) 97 (25) 121 (23) 289 (21) 187 (21) 188 (20) 149 (20) 221 (20) 161 (20) | 291.0858 | 139 (100) 123 (59) 147 (23) 161 (12) 207 (6) |
| 5 | 4.42 | 354.3087 | C16H18O10 | 300sh-325 | 5-O-Caffeoylquinic acid * | 353.0901 | 191 (100) 161 (2) 179 (2) 173 (1) | 355.1002 | 163 (100) 145 (11) 164 (11) 135 (4) 117 (3) 89 (2) |
| 6 | 4.74 | 354.3087 | C16H18O10 | 300sh-325 | 4-O-Caffeoylquinic acid | 353.0901 | 173 (100) 191 (85) 179 (73) 135 (51) 93 (20) | 355.1002 | 163 (100) 145 (14) 164 (9) 117 (6) 135 (5) 89 (4) |
| 7 | 4.90 | 194.1900 | C8H10N4O2 | 272 | Caffeine * | - | 195.089 | 138 (100) 110 (36) 195 (29) 69 (21) | |
| 8 | 5.42 | 290.2700 | C15H14O6 | 278 | (-)-Epicatechin * | 289.0709 | 109 (100) 125 (74) 203 (69) 123 (60) 151 (43) 245 (40) 137 (37) 205 (34) 161 (26) 97 (23) 188 (23) 221 (22) 187 (20) 121 (20) 179 (20) 289 (19) | 191.0844 | 139 (100) 123 (63) 147 (24) 161 (12) 207 (6) |
| 9 | 5.62 | 338.3093 | C16H18O8 | 310 | 5-O-Coumaroylquinic acid | 337.0953 | 191 (100) 133 (17) 53 (12) 93 (12) 253 (12) 145 (10) 75 (2) 302 (2) | 339.1085 | 147 (100) 75 (23) 97 (23) 236 (23) 127 (19) 258 (16) 341 (16) 65 (10) 189 (10) 217 (10) 306 (10) |
| 10 | 6.08 | 422.3400 | C19H18O11 | 257-317-365 | Mangiferin * | 421.0799 | 301 (100) 331 (85) 259 (12) 421 (15) 273 (10) | 423.0906 | 273 (100) 303 (74) 327 (51) 369 (22) 351 (18) 299 (15) 357 (9) |
| 11 | 6.79 | 368.3353 | C17H20O9 | 258-267sh-300sh-327 | 5-O-Feruloylquinic acid | 367.1058 | 191 (100) 93 (21) 173 (11) 111 (7) 134 (4) | 369.1157 | 177 (100) 145 (42) 117 (5) |
| 12 | 6.81 | 594.5181 | C27H30O15 | 256(sh)-265-330 | Flavone di-C-hexose | 593.1555 | 593 (100) 473 (11) 353 (10) 406 (10) 503 (8) | 595.1643 | 457 (100) 427 (67) 379 (35) 325 (29) 295 (12) |
| 13 | 6.86 | 772.6581 | C33H40O21 | 257-300sh-353 | Quercetin-3-O-dihexose-deoxyhexose | 771.2036 | 771 (100) 300 (53) 179 (5) 273 (5) 271 (5) | 773.2129 | 303 (100) 125 (53) 84 (26) 194 (13) |
| 14 | 6.97 | 626.5169 | C27H30O17 | 257-300sh-353 | Quercetin-3,4-di-O-glucoside * | 625.1449 | 625 (100) 300 (72) 271 (10) 179 (3) 445 (3) 463 (3) 505 (3) | 627.1514 | 303 (100) |
| 15 | 7.34 | 756.659 | C33H40O20 | 265-346 | Kaempferol-3-O-dihexose-deoxyhexose | 755.2128 | ND | 757.2131 | 287 (100) 449 (6) 129 (6) 757 (6) 612 (1) |
| 16 | 7.74 | 610.5175 | C27H30O16 | 257-353 | Quercetin-3-O-rutinoside * | 609.1539 | 609 (100) 300 (20) | 611.1595 | 303 (100) |
| 17 | 7.90 | 578.5202 | C30H26O12 | 276 | Procyanidin dimer | 577.1396 | 289 (100) 577 (83) 407 (72) 125 (61) 161 (17) 451 (11) | 579.1472 | 287 (100) 409 (94) 127 (84) 580 (64) 163 (31) 247 (19) 579 (13) |
| 18 | 7.96 | 464.3800 | C21H20O12 | 257-353 | Quercetin-3-O-glucoside * | 463.0908 | 300 (100) 463 (48) 191 (19) 271 (10) | 465.1018 | 303 (100) |
| 19 | 8.20 | 594.5181 | C27H30O15 | 265-346 | Kaempferol-3-O-hexose-deoxyhexose | 593.1540 | ND | 595.1629 | 287 (100) 596 (1) |
| 20 | 8.39 | 516.451 | C25H24O12 | 246-300sh-324 | 3,4-O-Dicaffeoylquinic acid * | 515.1221 | 173 (100) 353 (95) 179 (70) 191 (34) 354 (21) 515 (21) 335 (14) | 517.1323 | 163 (100) 63 (27) 140 (18) 191 (18) 370 (11) |
| 21 | 8.48 | 516.451 | C25H24O12 | 243-297sh-327 | 3,5-O-Dicaffeoylquinic acid * | 515.1229 | 191 (100) 353 (91) 179 (63) 135 (13) 173 (6) 515 (3) | 517.1323 | 140 (100) 163 (77) 519 (64) 213 (59) 117 (41) 303 (41) 189 (27) 238 (27) 367 (27) 442 (27) 492 (23) |
| 22 | 8.93 | 516.4510 | C25H24O12 | 243-297sh-327 | 4,5-O-Dicaffeoylquinic acid * | 515.1229 | 353 (100) 173 (92) 179 (53) 354 26) 191 (16) 135 (11) 137 (7) 203 (6) | 517.1323 | 163 (100) 111 (18) 383 (14) 435 (14) 224 (5) |
| 23 | 9.49 | 542.4451 | C26H22O13 | 257-317-365 | Mangiferin parahydroxybenzoate | 541.1017 | 301 (100) 331 (79) 541 (53) 259 (6) 385 (6) | 543.1118 | 303 (100) 121 (96) 405 (24) 327 (23) 543 (21) |
| Phenolic Compounds | Light | Elevation | Elevation x Light | |||
|---|---|---|---|---|---|---|
| F | p | F | p | F | p | |
| Chlorogenic Acids | ||||||
| 3-CQA | 1.72 | 0.198936 | 7.72 | 0.008836 | 0.00 | 0.954559 |
| 4-CQA | 0.05 | 0.823017 | 29.29 | 0.000005 | 5.81 | 0.021526 |
| 5-CQA | 18.69 | 0.000127 | 56.51 | 0.000000 | 28.72 | 0.000006 |
| 3,4-diCQA | 1.97 | 0.169890 | 35.48 | 0.000001 | 11.11 | 0.002085 |
| 3,5-diCQA | 6.88 | 0.012958 | 4.77 | 0.035947 | 11.71 | 0.001633 |
| 4,5-diCQA | 8.20 | 0.007112 | 49.98 | 0.000000 | 35.19 | 0.000001 |
| FQA | 0.03 | 0.865203 | 0.23 | 0.633213 | 4.21 | 0.048007 |
| Flavonoids | ||||||
| Flavanols | ||||||
| Epicatechin | 6.00 | 0.019604 | 63.86 | 0.000000 | 10.29 | 0.002912 |
| Catechin | 0.13 | 0.717829 | 100.22 | 0.000000 | 23.80 | 0.000025 |
| Glycosylated flavonoids | ||||||
| K-dihex-dhex | 26.33 | 0.000012 | 29.04 | 0.000005 | 15.76 | 0.000353 |
| K-hex-dhex | 117.43 | 0.000000 | 23.92 | 0.000024 | 29.81 | 0.000004 |
| F-dihex | 16.15 | 0.000307 | 13.02 | 0.000980 | 8.79 | 0.005506 |
| Q-dihex-dhex | 54.52 | 0.000000 | 11.64 | 0.001685 | 0.30 | 0.589041 |
| Q-diGlu | 100.65 | 0.000000 | 23.18 | 0.000030 | 0.01 | 0.903435 |
| Rutin | 142.72 | 0.000000 | 31.26 | 0.000003 | 0.02 | 0.881375 |
| Xanthones | ||||||
| Mangiferin | 31.19 | 0.000003 | 12.69 | 0.001112 | 2.39 | 0.131611 |
| Country | Geographic Coordinates | Elevation (m asl) | Plant Density (m) | Plants for Each Half Plot: Full Sun or under Shade (N) | Annual Average Temperature Min/Max (°C) | Average Surface Temperature (°C) | Rain (mm by Year) | Global Horizontal Irradiation (kWh.m−2) |
|---|---|---|---|---|---|---|---|---|
| Mexico | 18°51′46″ N 96°51′35″ W | 650 | 2 × 1.5 | 192 | 18.6/31.4 | 19.1 | 2650 | 1890 |
| Mexico | 19°23′43″ N 96°59′60″ W | 1250 | 2 × 1.5 | 192 | 14.2/28.2 | 17.0 | 2100 | 1848 |
| Nicaragua | 13°2′45″ N 85°53′30″ W | 1200 | 2 × 1 | 950 | 14.1/24.0 | 18.7 | 1760 | 1706 |
| Colombia | 4°58′17″ N 75°39′09″ W | 1380 | 1.3 × 1.5 for APL 1.5 × 1.5 for WEA | 490 | 17.4/26.8 | 22.1 | 3570 | 1851 |
| Genotype | Genealogy | Origin | Genetic Group |
|---|---|---|---|
| Mexican experimental plot | |||
| Marsellesa® | Timor hybrid CIFC 832/2 x cv. Villa Sarchi (Costa Rica) | CIRAD Nicaragua | APL of Sarchimor group |
| Nicaraguan experimental plot | |||
| ET08 A8 | EWA | Kaffa province, Ethiopia (ORSTOM prospecting, 1966) | “Jimma Bonga” (G1A) * |
| ET47 A4 | Kaffa province, Ethiopia (FAO prospecting, 1964–1965) | ||
| ET26 A1 | “Sheka” (G1B) * | ||
| ET25 A4 | G1G2 * | ||
| ET06 | Not yet characterized | ||
| T5175 | Timor hybrid CIFC 832/1 x cv. Caturra | Instituto del Café of Costa Rica (ICAFE) | APL of Catimor group |
| T8667 | Timor hybrid CIFC 832/1 x cv. Caturra | CATIE, Turrialba, Costa Rica | |
| T5296 | Timor hybrid CIFC 832/2 x cv. Villa Sarchi | APL of Sarchimor group | |
| T17931 | Timor hybrid CIFC1343 x cv. Caturra | APL Catimor line of the multiline var. Colombia | |
| Catuaí | cv. Mundo Novo x cv. Caturra | Instituto Agronômico de Campinas (IAC), Brazil | APL non-introgressed dwarf cultivar |
| T5175 x ET08 A8 | APL mother x EWA father F1 hybrid | CIRAD, Nicaragua | F1 hybrid clone |
| T5175 x ET26 A1 | |||
| T5175 x ET25 A4 | |||
| T5175 x T17931 | APL mother x APL father F1 hybrid | ||
| T8667 x ET47 A4 | APL mother x EWA father F1 hybrid | ||
| T8667 x ET26 A1 | |||
| T8667 x T5296 | APL mother x APL father F1 hybrid | ||
| T5296 x T17931 | |||
| T17931 x ET47 A4 | APL mother x EWA father F1 hybrid | ||
| T17931 x ET26 A1 | |||
| T17931 x ET25 A4 | |||
| Catuaí x ET47 A4 | |||
| Catuaí x ET26 A1 | |||
| Colombian experimental plot | |||
| E554 | EWA | Kaffa province, Ethiopia (FAO 1964-1965) | “Jimma Bonga” (G1A) * |
| E286 | Kaffa province, Ethiopia (FAO 1964-1965) | “Jimma Bonga” (G1A) * | |
| E057 | Kaffa province, Ethiopia (FAO 1964-1965) | “Jimma Bonga” (G1A) * | |
| CX2385 | Timor hybrid CIFC1343 x cv. Caturra | CENICAFE, Colombia | APL of Catimor group |
| CU1842 | |||
| CX2385 x E554 | APL mother x EWA father F1 hybrid | F1 hybrid clone | |
| CX2385 x E286 | |||
| CX2385 x E057 | |||
| CU 1842 x E554 | |||
| CU 1842 x E286 | |||
| CU 1842 x E057 | |||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duangsodsri, T.; Villain, L.; Vestalys, I.R.; Michalet, S.; Abdallah, C.; Breitler, J.-C.; Bordeaux, M.; Villegas, A.M.; Raherimandimby, M.; Legendre, L.; et al. 5-CQA and Mangiferin, Two Leaf Biomarkers of Adaptation to Full Sun or Shade Conditions in Coffea arabica L. Metabolites 2020, 10, 383. https://doi.org/10.3390/metabo10100383
Duangsodsri T, Villain L, Vestalys IR, Michalet S, Abdallah C, Breitler J-C, Bordeaux M, Villegas AM, Raherimandimby M, Legendre L, et al. 5-CQA and Mangiferin, Two Leaf Biomarkers of Adaptation to Full Sun or Shade Conditions in Coffea arabica L. Metabolites. 2020; 10(10):383. https://doi.org/10.3390/metabo10100383
Chicago/Turabian StyleDuangsodsri, Teerarat, Luc Villain, Ialy Rojo Vestalys, Serge Michalet, Cécile Abdallah, Jean-Christophe Breitler, Mélanie Bordeaux, Andres Mauricio Villegas, Marson Raherimandimby, Laurent Legendre, and et al. 2020. "5-CQA and Mangiferin, Two Leaf Biomarkers of Adaptation to Full Sun or Shade Conditions in Coffea arabica L." Metabolites 10, no. 10: 383. https://doi.org/10.3390/metabo10100383
APA StyleDuangsodsri, T., Villain, L., Vestalys, I. R., Michalet, S., Abdallah, C., Breitler, J.-C., Bordeaux, M., Villegas, A. M., Raherimandimby, M., Legendre, L., Etienne, H., Bertrand, B., & Campa, C. (2020). 5-CQA and Mangiferin, Two Leaf Biomarkers of Adaptation to Full Sun or Shade Conditions in Coffea arabica L. Metabolites, 10(10), 383. https://doi.org/10.3390/metabo10100383







