Competitive Promoter-Associated Matrix Attachment Region Binding of the Arid3a and Cux1 Transcription Factors
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cell Culture
2.2. Transfections and Retroviral Transduction
2.3. Nuclear Protein Extraction
2.4. Electrophoretic Mobility Shift Assays (EMSA)
2.5. Cell Cycle Synchronization and Cell Cycle Analysis
2.6. In Vitro Transcription/Translation
2.7. Luciferase Assays
3. Results
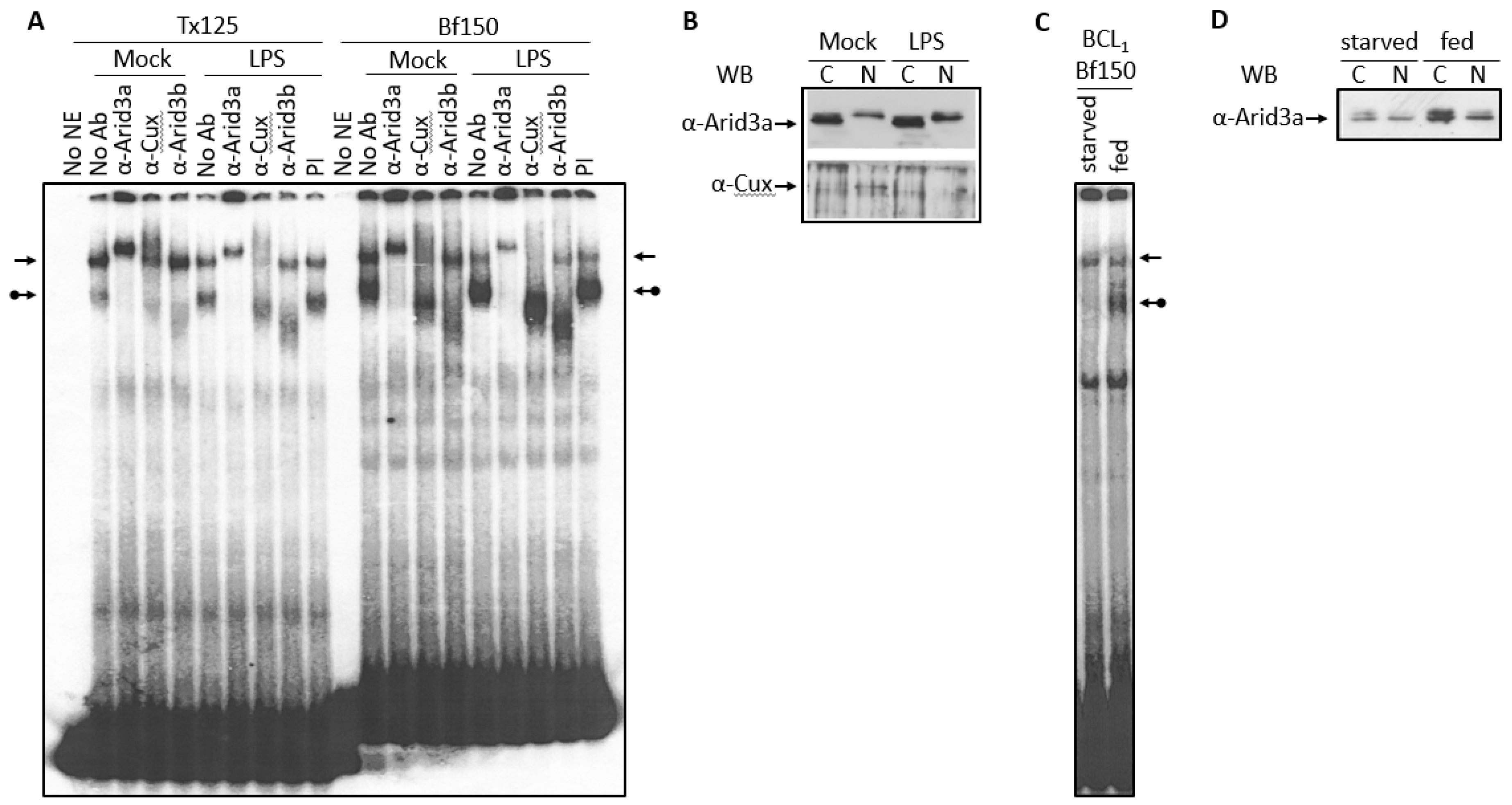
3.1. Components of the Arid3a and NF-μNR Complexes in B Cells
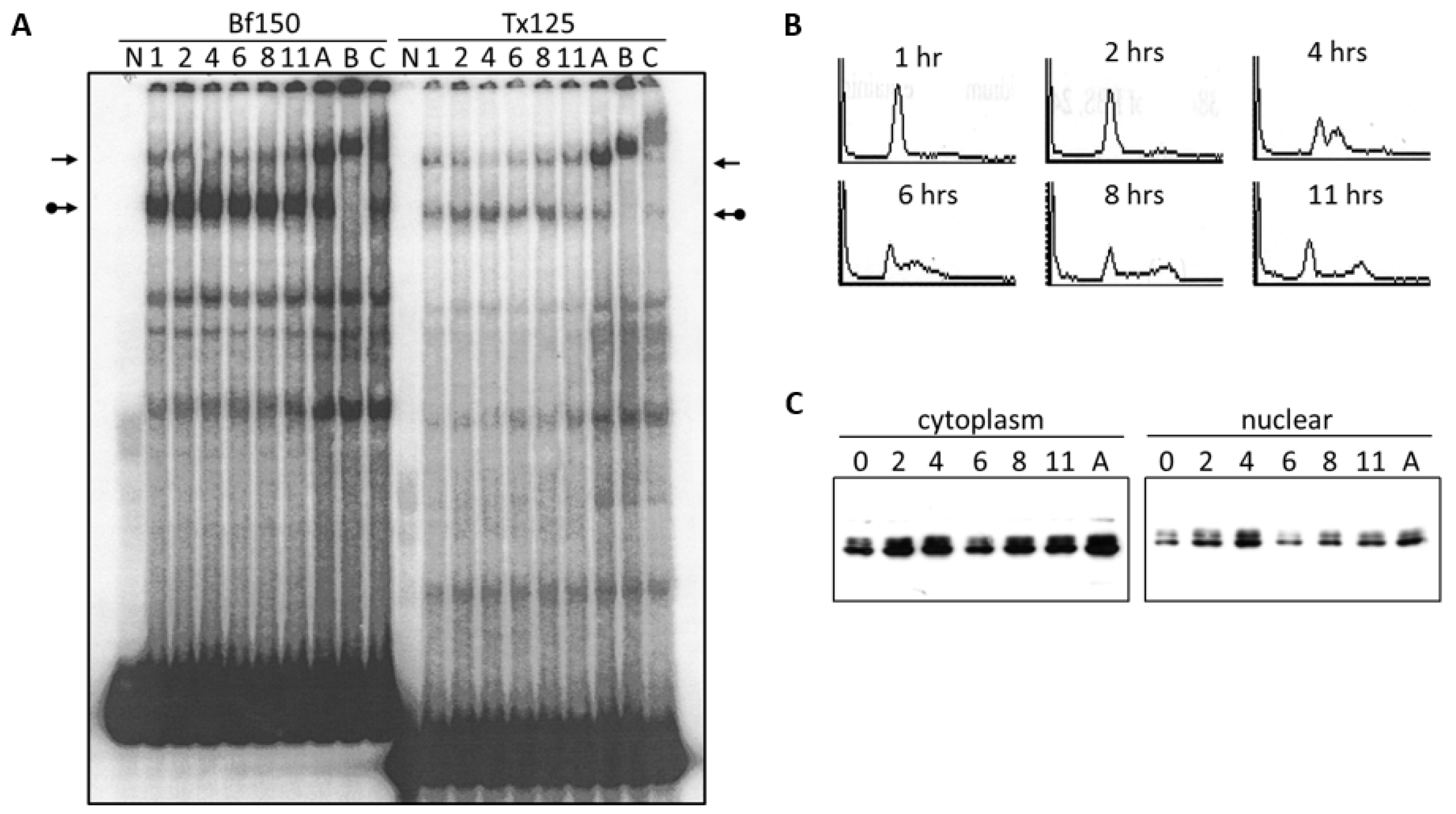
3.2. DNA Binding Activity of Arid3a Is Sensitive to Cell Cycle and Nuclear Localization
3.3. LPS Stimulates Formation of the NF-μNR Complex
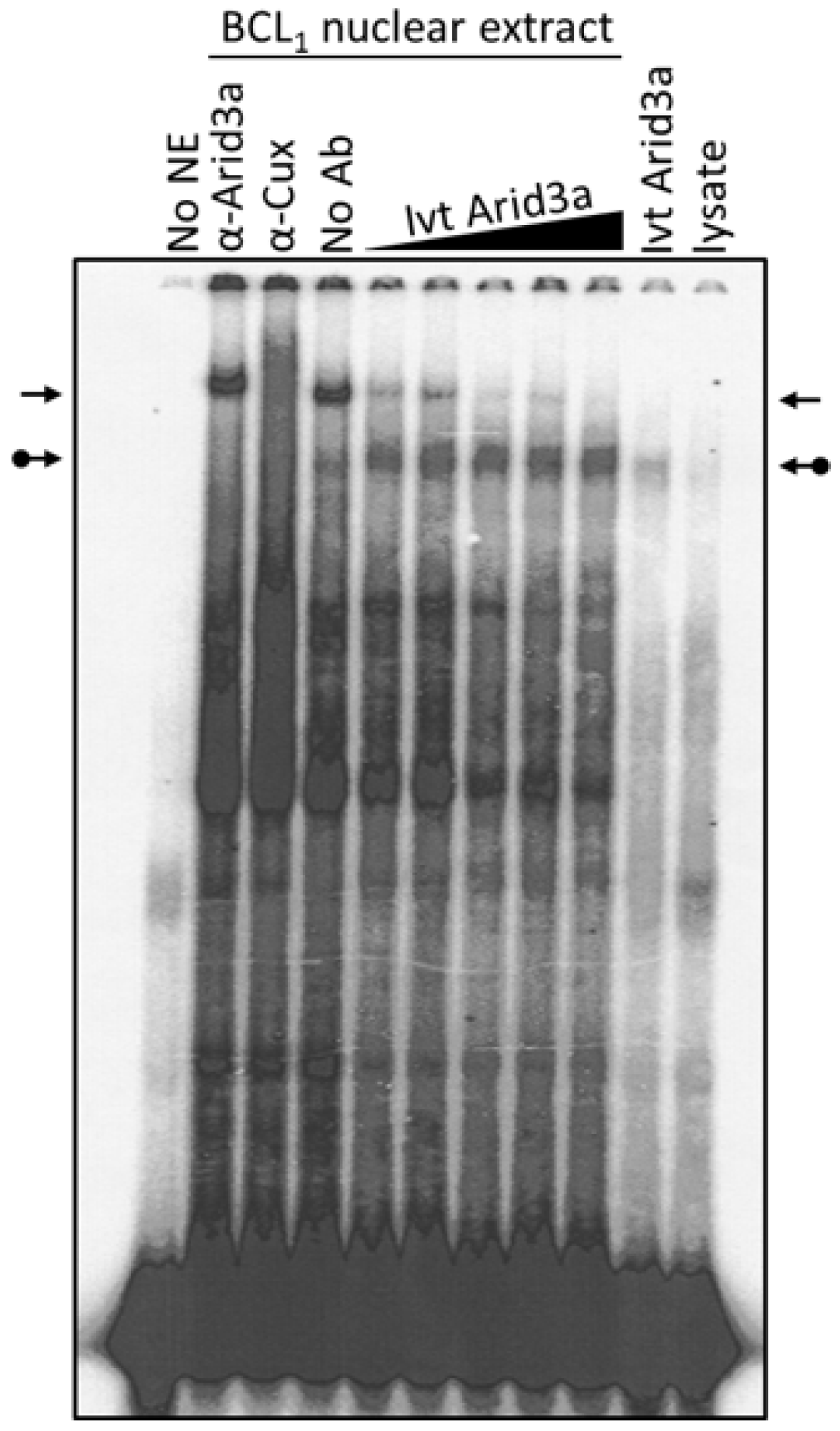
3.4. Increased Nuclear Levels of Arid3a Is Sufficient for High Affinity Arid3a-MAR Complex Formation
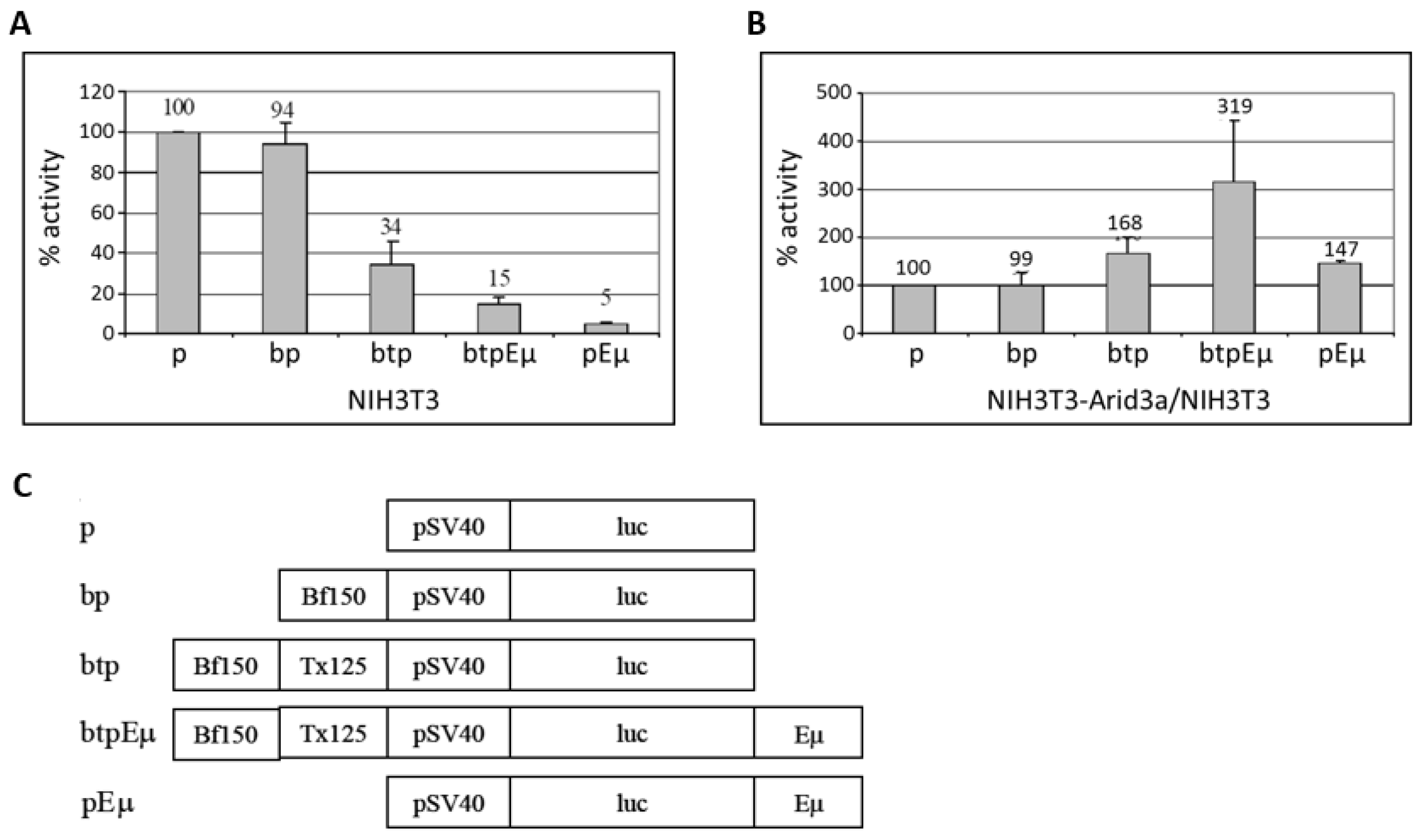
3.5. Transactivation Activity of Arid3a in Non-B Cells Is Modulated by Promoter-Associated MARs
4. Discussion
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Wasylyk, C.; Wasylyk, B. The immunoglobulin heavy-chain B-lymphocyte enhancer efficiently stimulates transcription in non-lymphoid cells. EMBO J. 1986, 5, 553–560. [Google Scholar] [PubMed]
- Matthias, P.; Baltimore, D. The immunoglobulin heavy chain locus contains another B-cell-specific 3’ enhancer close to the alpha constant region. Mol. Cell. Biol. 1993, 13, 1547–1553. [Google Scholar] [CrossRef] [PubMed]
- Chauveau, C.; Jansson, E.A.; Muller, S.; Cogne, M.; Pettersson, S. Cutting edge: Ig heavy chain 3’ HS1–4 directs correct spatial position-independent expression of a linked transgene to B lineage cells. J. Immunol. 1997, 163, 4637–4641. [Google Scholar]
- Le Noir, S.; Boyer, F.; Lecardeur, S.; Brousse, M.; Oruc, Z.; Cook-Moreau, J.; Denizot, Y.; Cogné, M. Functional anatomy of the immunoglobulin heavy chain 3’ super-enhancer needs not only core enhancer elements but also their unique DNA context. Nucleic Acids Res. 2017, 45, 5829–5837. [Google Scholar] [CrossRef] [PubMed]
- Cockerill, P.N.; Yuen, M.H.; Garrard, W.T. The enhancer of the immunoglobulin heavy chain locus is flanked by presumptive chromosomal loop anchorage elements. J. Biol. Chem. 1987, 262, 5394–5397. [Google Scholar] [PubMed]
- Banerji, J.; Olson, L.; Schaffner, W. A lymphocyte-specific cellular enhancer is located downstream of the joinings region in immunoglobulin heavy chain genes. Cell 1983, 33, 729–740. [Google Scholar] [CrossRef]
- Staudt, L.M. Immunoglobulin gene transcription. Annu. Rev. Immunol. 1991, 9, 373–398. [Google Scholar] [CrossRef] [PubMed]
- Perez-Mutul, J.; Macchi, M.; Wasylyk, B. Mutational analysis of the contribution of sequence motifs within the IgH enhancer to tissue specific transcriptional activation. Nucleic Acids Res. 1998, 16, 6085–6096. [Google Scholar] [CrossRef]
- Webb, C.F.; Zong, R.T.; Lin, D.; Wang, Z.; Kaplan, M.; Paulin, Y.; Smith, E.; Probst, L.; Bryant, J.; Goldstein, A.; et al. Differential regulation of immunoglobulin gene transcription via nuclear matrix-associated regions. Cold Spring Harb. Symp. Quant. Biol. 1999, 64, 109–118. [Google Scholar] [CrossRef] [PubMed]
- Kaplan, M.H.; Zong, R.T.; Herrscher, R.F.; Scheuermann, R.H.; Tucker, P.W. Transcriptional activation by a matrix associating region-binding protein. Contextual requirements for the function of bright. J. Biol. Chem. 2001, 276, 21325–21330. [Google Scholar] [CrossRef] [PubMed]
- Herrscher, R.F.; Kaplan, M.H.; Lelsz, D.L.; Das, C.; Scheuermann, R.; Tucker, P.W. The immunoglobulin heavy-chain matrix-associating regions are bound by Bright: A B cell-specific trans-activator that describes a new DNA-binding protein family. Genes Dev. 1995, 9, 3067–3082. [Google Scholar] [CrossRef] [PubMed]
- Avitahl, N.; Calame, K.A. 125 bp region of the Ig VH1 promoter is sufficient to confer lymphocyte-specific expression in transgenic mice. Int. Immunol. 1996, 8, 1359–1366. [Google Scholar] [CrossRef] [PubMed]
- Grosschedl, R.; Weaver, D.; Baltimore, D.; Costantini, F. Introduction of a mu immunoglobulin gene into the mouse germ line: Specific expression in lymphoid cells and synthesis of functional antibody. Cell 1988, 38, 647–658. [Google Scholar] [CrossRef]
- Aguilera, R.J.; Hope, T.J.; Sakano, H. Characterization of immunoglobulin enhancer deletions in murine plasmacytomas. EMBO J. 1985, 4, 3689–3693. [Google Scholar] [PubMed]
- Wang, Z.; Goldstein, A.; Zong, R.T.; Lin, D.; Neufeld, E.J.; Scheuermann, R.H.; Tucker, P.W. Cux/CDP homeoprotein is a component of NF-μNR and represses the immunoglobulin heavy chain intronic enhancer by antagonizing the Bright transcription activator. Mol. Cell. Biol. 1999, 19, 284–295. [Google Scholar] [CrossRef] [PubMed]
- Neufeld, E.J.; Skalnik, D.G.; Lievens, P.M.; Orkin, S.H. Human CCAAT displacement protein is homologous to the Drosophila homeoprotein, cut. Nat. Genet. 1992, 1, 50–55. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Moy, L.; Pittman, N.; Shue, G.; Aufiero, B.; Neufeld, E.J.; LeLeiko, N.S.; Walsh, M.J. Transcriptiona l repression of the cystic fibrosis transmembrane conductance regulator gene, mediated by CCAAT displacement protein/cut homolog, is associated with histone deacetylation. J. Biol. Chem. 1999, 274, 7803–7815. [Google Scholar] [CrossRef] [PubMed]
- Coqueret, O.; Berube, G.; Nepveu, A. The mammalian Cut homeodomain protein functions as a cell-cycle-dependent transcriptional repressor which downmodulates p21WAF1/CIP1/SDI1 in S phase. EMBO J. 1998, 17, 4680–4694. [Google Scholar] [CrossRef] [PubMed]
- Van Gurp, M.F.; Pratap, J.; Luong, M.; Javed, A.; Hoffmann, H.; Giordano, A.; Stein, J.L.; Neufeld, E.J.; Lian, J.B.; Stein, G.S.; et al. The CCAAT displacement protein/cut homeodomain protein represses osteocalcin gene transcription and forms complexes with the retinoblastoma protein-related protein p107 and cyclin A. Cancer Res. 1999, 59, 5980–5988. [Google Scholar] [PubMed]
- Numata, S.; Claudio, P.P.; Dean, C.; Giordano, A.; Croce, C.M. Bdp, a new member of a family of DNA-binding proteins, associates with the retinoblastoma gene product. Cancer Res. 1999, 59, 3741–3747. [Google Scholar] [PubMed]
- Kim, D.; Tucker, P.W. A regulated nucleocytoplasmic shuttle contributes to Bright’s function as a transcriptional activator of immunoglobulin genes. Mol. Cell. Biol. 2006, 26, 2187–2201. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Probst, L.; Das, C.; Tucker, P.W. REKLES is an ARID3-restricted multifunctional domain. J. Biol. Chem. 2007, 25, 15768–15777. [Google Scholar] [CrossRef] [PubMed]
- Laskov, R.; Kim, J.K.; Woods, V.L.; McKeever, P.E.; Asofsky, R. Membrane immunoglobulins of spontaneous B-lymphomas of aged BALB/c mice. Eur. J. Immunol. 1981, 11, 462–471. [Google Scholar] [CrossRef] [PubMed]
- Gronowicz, E.S.; Doss, C.A.; Howard, F.D.; Morrison, D.C.; Strober, S. An in vitro line of the B cell tumor BCL1 can be activated by LPS to secrete IgM1. J. Immunol. 1980, 125, 976–980. [Google Scholar] [PubMed]
- Johnson, D.R.; Levanat, S.; Bale, A.E. Isolation of intact nuclei for nuclear extract preparation from a fragile B-lymphocyte cell line. Bio-Techniques 1995, 19, 192–195. [Google Scholar]
- Webb, C.F.; Das, C.; Eneff, K.L.; Tucker, P.W. Identification of a matrix associated region 5’ of an immunoglobulin heavy chain variable region gene. Mol. Cell. Biol. 1991, 11, 5206–5211. [Google Scholar] [CrossRef] [PubMed]
- Webb, C.F.; Yamashita, Y.; Ayers, N.; Evetts, S.; Paulin, Y.; Conley, M.E.; Smith, E.A. The transcription factor Bright associates with Bruton’s tyrosine kinase, the defective protein in immunodeficiency disease. J. Immunol. 2000, 165, 6956–6965. [Google Scholar] [CrossRef] [PubMed]
- Zong, R.T.; Das, C.; Tucker, P.W. Regulation of matrix attachment region dependent, lymphocyte-restricted transcription through differential localization within promyelocytic leukemia nuclear bodies. EMBO J. 2000, 19, 4123–4133. [Google Scholar] [CrossRef] [PubMed]
- Van Wijnen, A.J.; van Gurp, M.F.; de Ridder, M.C.; Tufarelli, C.; Last, T.J.; Birnbaum, M.; Vaughan, P.S.; Giordano, A.; Krek, W.; Neufeld, E.J.; et al. CDP/cut is the DNA-binding subunit of histone gene transcription factor HiNF-D: A mechanism for gene regulation at the G1/S phase cell cycle transition point independent of transcription factor E2F. Proc. Natl. Acad. Sci. USA 1996, 93, 11516–11521. [Google Scholar] [CrossRef] [PubMed]
- Ho, A.; Dowdy, S.F. Regulation of G1 cell-cycle progression by oncogenes and tumor suppressor genes. Curr. Opin. Genet. Dev. 2002, 12, 47–52. [Google Scholar] [CrossRef]
- Ishov, A.M.; Sotnikov, A.G.; Negorev, D.; Vladimirova, O.V.; Neff, N.; Kamitani, T.; Yeh, E.T.; Strauss, J.F., III; Maul, G.G. PML is critical for ND10 formation and recruits the PML- interacting protein daxx to this nuclear structure when modified by SUMO-1. J. Cell Biol. 1999, 147, 221–234. [Google Scholar] [CrossRef] [PubMed]
- Schneider, R.; Grosschedl, R. Dynamics and interplay of nuclear architecture, genome organization, and gene expression. Gene Dev. 2007, 21, 3027–3043. [Google Scholar] [CrossRef] [PubMed]
- Osborne, C.S.; Chakalova, L.; Brown, K.E.; Carter, D.; Horton, A.; Debrand, E.; Goyenechea, B.; Mitchell, J.A.; Lopes, S.; Reik, W.; et al. Active genes dynamically colocalize to shared sites of ongoing transcription. Nat. Genet. 2004, 36, 1065–1071. [Google Scholar] [CrossRef] [PubMed]
- Peric-Hupkes, D.; Meuleman, W.; Pagie, L.; Bruggeman, S.W.; Solovei, I.; Brugman, W.; Graf, S.; Flicek, P.; Kerkhoven, R.M.; van Lohuizen, M.; et al. Molecular maps of the reorganization of genome-nuclear lamina interactions during differentiation. Mol. Cell 2010, 38, 603–613. [Google Scholar] [CrossRef] [PubMed]
- Reddy, K.L.; Zullo, J.M.; Bertolino, E.; Singh, H. Transcriptional repression mediated by repositioning of genes to the nuclear lamina. Nature 2008, 452, 243–247. [Google Scholar] [CrossRef] [PubMed]
- Kosak, S.T.; Skok, J.A.; Medina, K.L.; Riblet, R.; Le Beau, M.M.; Fisher, A.G.; Singh, H. Subnuclear compartmentalization of immunoglobulin loci during lymphocyte development. Science 2002, 296, 158–162. [Google Scholar] [CrossRef] [PubMed]
- Skok, J.A.; Brown, K.E.; Azuara, V.; Caparros, M.L.; Baxter, J.; Takacs, K.; Dillon, N.; Gray, D.; Perry, R.P.; Merkenschlager, M.; et al. Nonequivalent nuclear location of immunoglobulin alleles in B lymphocytes. Nat. Immunol. 2001, 2, 848–854. [Google Scholar] [CrossRef] [PubMed]
- Lafrenz, D.; Koretz, S.; Stratte, P.T.; Ward, R.B.; Strober, S. LPS-induced differentiation of a murine B cell leukemia (BCL1): Changes in surface and secreted IgM. J. Immunol. 1982, 129, 1329–1335. [Google Scholar] [PubMed]
- Brooks, K.; Yuan, D.; Uhr, J.W.; Krammer, P.H.; Vitetta, E.S. Lymphokine-induced IgM secretion by clones of neoplastic B cells. Nature 1983, 302, 825–826. [Google Scholar] [CrossRef] [PubMed]
- Webb, C.F.; Das, C.; Coffman, R.L.; Tucker, P.W. Induction of immunoglobulin µ mRNA in a B cell transfectant stimulated with interleukin-5 and T-dependent antigen. J. Immunol. 1989, 143, 3934–3939. [Google Scholar] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, D.; Schmidt, C.; Brown, M.A.; Tucker, H. Competitive Promoter-Associated Matrix Attachment Region Binding of the Arid3a and Cux1 Transcription Factors. Diseases 2017, 5, 34. https://doi.org/10.3390/diseases5040034
Kim D, Schmidt C, Brown MA, Tucker H. Competitive Promoter-Associated Matrix Attachment Region Binding of the Arid3a and Cux1 Transcription Factors. Diseases. 2017; 5(4):34. https://doi.org/10.3390/diseases5040034
Chicago/Turabian StyleKim, Dongkyoon, Christian Schmidt, Mark A. Brown, and Haley Tucker. 2017. "Competitive Promoter-Associated Matrix Attachment Region Binding of the Arid3a and Cux1 Transcription Factors" Diseases 5, no. 4: 34. https://doi.org/10.3390/diseases5040034
APA StyleKim, D., Schmidt, C., Brown, M. A., & Tucker, H. (2017). Competitive Promoter-Associated Matrix Attachment Region Binding of the Arid3a and Cux1 Transcription Factors. Diseases, 5(4), 34. https://doi.org/10.3390/diseases5040034




