1. Introduction
Foodborne diseases include several illnesses resulting from ingestion of foods contaminated with microorganisms or chemicals. Food can be contaminated at any stage from production to consumption, because of several factors, such as environmental contamination (pollution of water, soil or air) or incorrect conservation and preparation procedures. Although great security awareness and the improvements in food safety and quality control worldwide, foodborne diseases still represent a significant public health problem with a significant rate of morbidity and mortality, in both developed and developing countries [
1]. According to the WHO studies about 30 hazards (including viruses, bacteria, protozoa, helminths and chemicals) caused 600 million foodborne illnesses globally and 420,000 deaths every year [
2]. Foodborne diseases represent not only a health problem, but has a strong socioeconomic impact, since it affects the health care systems, and national economies, tourism and trade.
Listeria monocytogenes is a gram-positive bacterium and is one of the most common foodborne pathogens affecting millions of people annually.
L. monocytogenes is a ubiquitous, intracellular pathogen and can cause both invasive or non-invasive listeriosis [
3,
4]. Outbreaks of listeriosis have been caused by a variety of foods, including milk, soft cheeses, meat, fermented sausages, poultry, seafood and vegetable products [
5,
6]. This pathogen represents a potential risk to human health, since it is able to overcome food preservation and safety barriers, due to the adaptation and survival mechanisms allowing the persistence and growth in different environmental conditions (cold temperatures, low pH and high salt concentration) [
7].
FDA regulation for
L. monocytogenes in ready-to-eat foods imposes strict limits and corrective solutions for contaminated food. In particular the limits correspond to 0.04 CFU of
L. monocytogenes per gram of food that supports the growth of
L. monocytogenes, while in foods that do not support the growth of the microorganism the limits is extended to 100 CFU/g [
8,
9,
10,
11].
The high incidence of foodborne pathogen contamination require the development of new, sensitive, portable, high-throughput, and automated platforms to detect pathogens and their toxins in foods [
12,
13]. In particular several technological strategies have been proposed for the development of biosensors for pathogens identification coupled with different analytical strategies ranging from nucleic acids analysis [
14] to whole cells detection [
15]. Among the wide range of proposed technology, Due to its high sensitivity and easy setup, electrochemical impedance spectroscopy (EIS) has been widely used for the development of biosensor and in particular for pathogens detection. Electrochemical biosensors, in particular impedance-based systems, present great advantages in terms of ease of miniaturization, lack of reagents, sensitivity, and low cost. Impedance methods have been often used for bacterial identification by monitoring changes in the impedance of the solution induced by the release of ionic metabolites from the living cells (i.e., carbon dioxide and organic acids) as a result of bacterial metabolism and cell growth. The magnitude of these changes in the impedance of the solution and can be correlated to the number of living cells allowing a quantification of bacterial cells. Such an approach has been used by Yang and coworkers, who developed an impedimetric detection of viable
Salmonella spp. through a three-electrodes configuration set-up [
16]. The improvements in microfabrication, and the trend towards miniaturization, allows the translation of the traditional electrochemical set-up in compact and portable chip—offering new opportunities in the field of bacterial detection. thanks to a great improvement in sensitivity. Such chips, being cheap thanks to the mass production, can be used as disposable devices reducing possible cross-contamination errors.
In this respect, Yang et al., in 2004, achieves a quantitative detection of
Salmonella spp. in culture and milk samples by using interdigitated microelectrodes (IME) for impedance measurements [
17].
In addition, microfabricated systems allow the functionalization of the electrode surface with capture molecules specific for one or more pathogens of interests. In this case impedance spectroscopy allows the characterization of solution/electrode interface and can be used to monitor the attachment of molecules [
18,
19] and cells [
20,
21]. In this last reference, a gold interdigitated array microelectrode (IDAM) allowed the simultaneous detection of two different food pathogens [
19]. Thanks to specific biorecognition reactions, EIS methods allow the detection of various pathogen species simultaneously, even in complex matrices, resulting from a reduction of the unspecific signals. Moreover, compared with growth-based impedance methods that require at least 24 h, the analysis is much faster, requiring less than 2 h.
Ruan et al. developed an impedimetric biosensor for detection of
Escherichia coli O157:H7 with a detection limit of 6 × 10
3 cells/mL. The method exploited the immobilization of species-specific antibodies onto the electrode surface in order to capture
E. coli cells from a solution. A secondary antibody conjugated with alkaline phosphatase was used to amplify the signals [
22].
A step forward was done by Yang and colleagues, that avoided the amplification step by the development of a label-free immunosensor for detection of the same bacterial strain,
E. coli O157:H7, using indium tin oxide (ITO) interdigitated electrodes with a size comparable to that of bacterial cells [
23]. Bacterial cells attached to the specific antibodies immobilized on the electrodes induced an increase in the electron transfer resistance [
13]. A similar approach was used by Radhakrishnan and colleagues: Their electrochemical impedance spectroscopy system based on a gold electrode functionalized with an anti-
Listeria monoclonal antibody was able to detect
L. monocytogenes. The biochip achieved high sensitivity with a limit of detection (LoD) of 4–5 CFU/mL [
12].
The integration of sensors and microfluidics structures led to the development of lab on a chip technology (LOC), opening new scenarios in many fields and in particular in microbiology [
18] and food security monitoring [
21,
24]. LOC platforms may allow both the isolation and detection of bacterial cells, thanks to the integration of biosensors with additional modules, and microfluidic components—in order to perform all the analytical and pre-analytical procedures on the same chip.
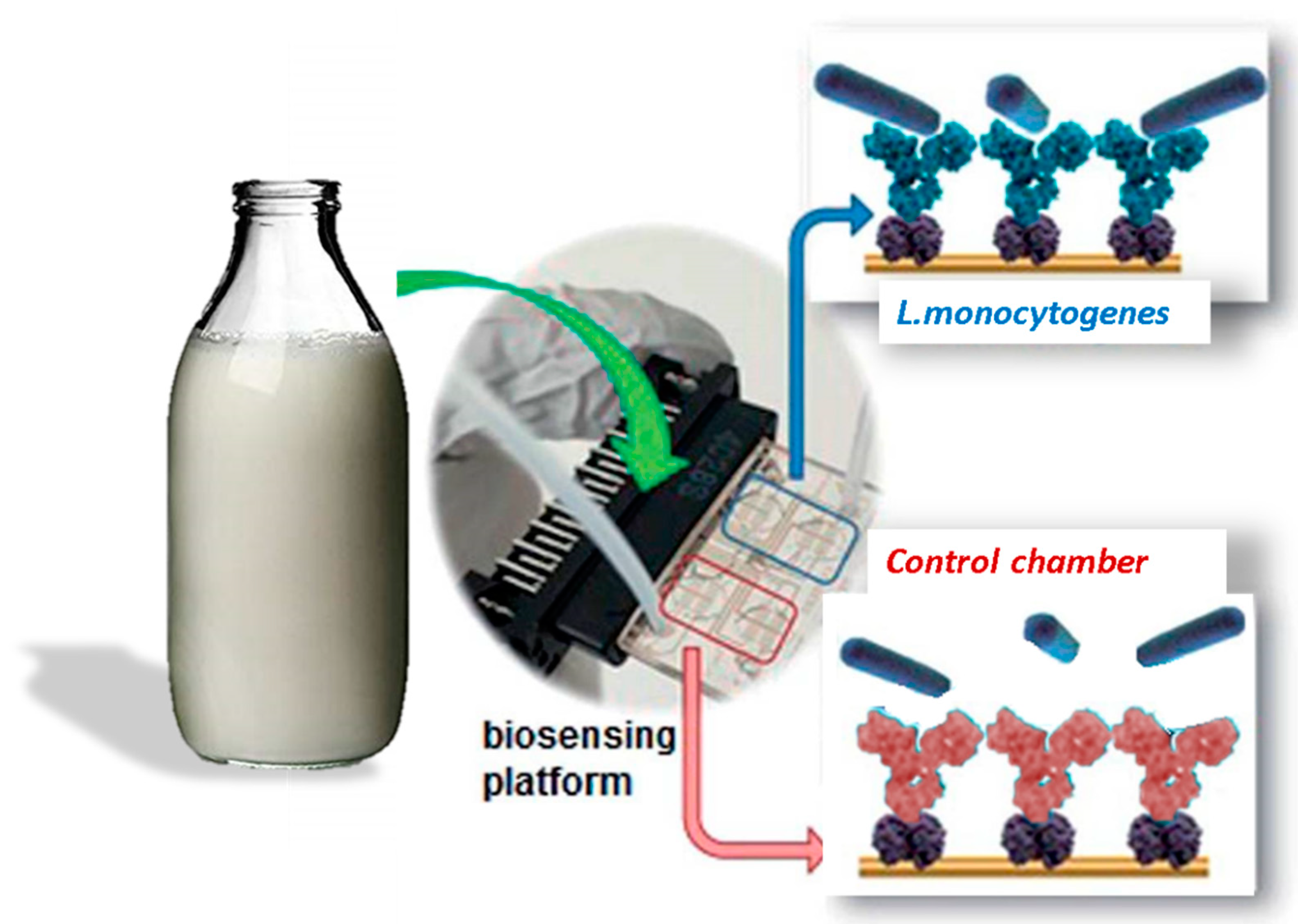
Here we describe a miniaturized, portable EIS biochip consisting of a microfluidic platform with EIS sensors, validated through detection of L. monocytogenes in artificially contaminated milk samples. The developed tool consists of an array of interdigited electrodes aligned at the bottom of a network of four microchambers and related microchannels. Microelectrodes in this case were functionalized with antibodies specific for L. monocytogenes. The network provides the possibility to separately fill the different chambers (to allow multiple functions, or to inject different samples from peripheral holes) or in turn to use a central inlet common to the four chambers, in order to detect different analytes from a single sample. In our experiment a chamber was used for the detection of L. monocytogenes and the other one as a negative control. The biochip was connected to a portable potentiostat and to a notebook, which makes the platform very suitable for on-field applications and in particular it exploits the advantages of flow immunoassays as quantitative results, speed, reduced sample handling and costs. In the developed protocol of analysis there is no need for fluorescent tagging of reagents, no PCR amplification and no complex instrumentation or skilled operators. Our biochips for detection of milk contamination down to 5 CFU/mL represents a robust, portable platform for a fast and low-cost analysis, which is very suitable for use in food factories, since it can allow testing, at the same time, different batches of products for their contamination levels. Due to the high flexibility, predisposition to further miniaturization, and the possibility to improve the number of analytes and the ease of use, the developed device is competitive with currently used methods and can be used as the point-of-care device for clinical and food diagnostics, as well as for biosecurity purposes.
2. Materials and Methods
2.1. Microfabrication and Functionalization of the Biochip
The device (
Figure 1) consists of an array of gold interdigitated electrodes on glass substrates arranged in four different sensing areas (
Figure 1B) aligned with a microfluidic network with microchambers and microchannels (
Figure 1A).
The interdigitated electrodes have a line-space period of 10 µm and cover a 2 mm × 1.5 mm area, they were fabricated by optical lithography (using a Karl Suss MJB3 mask aligner and AZ5214e resist) and thermal evaporation of Cr/Au (3 nm/15 nm). The microfluidic module includes channels and microchambers in order to handle and confine all the solution and reagents necessary for functionalization of the electrodes. The microfluidic module is obtained from a SU8 hard master by PDMS replica molding. The volume of each microchamber is of around 15 µL (3 mm diameter and 0.5 mm height) and the architecture of the network provides a common central inlet and four peripheral holes that can be used in turn to independently functionalize each chamber or to inject a sample to be tested in quadruplicate. In order to completely fill the chambers considering microchannels, a volume of 80 µL is required. The presence of the microfluidic module allows a high degree of reproducibility of experiments and to save a large quantity of reagents and samples.
The microelectrodes were functionalized with antibodies using reagents purchased from Sigma Aldrich (Saint Louis, MO, USA) at the maximum degree of purity.
The electrodes were incubated overnight with a solution of 11-mercaptoundecanoic acid (MUA) (0.2 mM) and 2-mercaptoethanol (2-ME) (1 mM) for the deposition of a mixed self-assembling monolayer, then COOH groups of MUA are activated by incubation with a solution of N-hydroxysuccinimide (NHS) and N-ethyl-N-(3-dimethylaminopropyl carbodiimide hydrochloride (EDC) at a ratio of 1:4 in milliQ water for 30 min.
Then a solution of protein A (50 µg/mL) in PBS was incubated for 2 h to allow a covalent binding to SAM. Protein A naturally binds the Fc region of antibodies allowing an oriented immobilization of antibodies. Ethanolamine (1 M) was used to block reactive groups while BSA (Bovine Serum Albumin) (1 mg/mL) to passivate and saturate the unbound surface. As final step each sensing area was functionalized with antibodies specific for L. monocytogenes.
2.2. EIS Measurements on Cell Chip
A pocketSTAT, a portable potentiostat and galvanostat with integrated impedance analyzer from IVIUM was used for measurements. Impedance spectra were acquired between 0.1 and 106 Hz using a sinusoidal ac voltage with a 15 mV RMS amplitude and no dc voltage applied (VDC = 0 V). The equivalent circuit consists of the ohmic resistance of the electrolytic solution consisting of PBS added with K3[Fe(CN)6]/K4[Fe(CN)6] (1:1) 10 mM (Rs), a double layer capacitance (Cdl), an interfacial electron-transfer resistance (Ret) and a Warburg impedance (Zw) associated to the diffusion of the redox probe. In particular, Cdl and Ret depend on the electrical properties of the solution/electrode interface and can be used to monitor the attachment of molecules and bacterial cells.
2.3. Samples Preparation
Pathogens growth conditions and measurement of concentration.
L. monocytogenes strains used in this study were isolated from food samples, grown in Half Frazer broth followed by a Listeria Enrichment broth (24 LEB) with selective polymixin and quinolone supplements (Oxoid, ThermoFisher, Milano, Italy) and identified by Real Time PCR. Subsequently, an aliquot of exponentially grown bacteria was serially diluted with an isotonic solution, and the serial dilutions were plated onto agar plates with solidified Agar Listeria Ottaviani and Agosti (ALOA). The number of CFU/mL was determined by counting the bacteria in plates containing spots well isolated and colonies well-spaced from each other. The samples were prepared in two replicates, so that each dilution could be stored frozen for further analyses. For EIS analysis, samples denatured by heating for 10 min were used.
Listeria monocytogenes suspensions in growth medium and in half-skimmed milk were prepared. Spiked milk samples were made by adding to half-skimmed milk a known concentration of pathogen suspension, in a ratio of 4:1.
2.4. Application of EIS Biochip to L. monocytogenes Detection
Pathogens’ suspensions were incubated in the device chamber. For dilution of the samples, an uncontaminated growth medium and half-skimmed milk were used. In order to calibrate the device for specific detection of the bacteria, different concentrations of
L. monocytogenes in culture medium and in the milk suspension were incubated above the sensing layer by filling the microfluidic chamber with a volume of 20 μL. Using serially diluted samples of
L. monocytogenes, (i.e., 2.2 × 10
1, 2.2 × 10
2, 2.2 × 10
3), the increase in impedance values was recorded. To obtain the reference EIS value for the negative control, a suspension containing
Salmonella enterica was used. After 1 h incubation, to allow the interaction between bacteria and antibodies immobilized onto the electrodes, the devices are washed with PBS, and a solution with the redox probes K
3[Fe(CN)
6]/K
4[Fe(CN)
6] (1:1) 10 mM is added. After the redox solution (20 μL) is injected in the chamber, EIS measurements are collected (see scheme in
Figure 2).
3. Results
Once the fabrication of the sensing platform was optimized, impedance spectroscopy measurements were performed and plotted as Nyquist curves, in which impedance values were collected in a frequency range from 10
6 to 0.1 Hz, and the diameter of the semicircle-shaped curves corresponds to the electric transfer resistance at the interface between the electrodes and the solution, in a complex system (Z
re vs. -Z
im). At first, the sensing layer given by immobilized antibodies on the electrode surface was characterized (functionalization steps, described in Materials and methods). Anti-
L. monocytogenes antibodies layer resulted in impedance values of around 27 kΩ (black line, inset in
Figure 2). In order to calibrate the device for the detection of the pathogen, different concentrations of
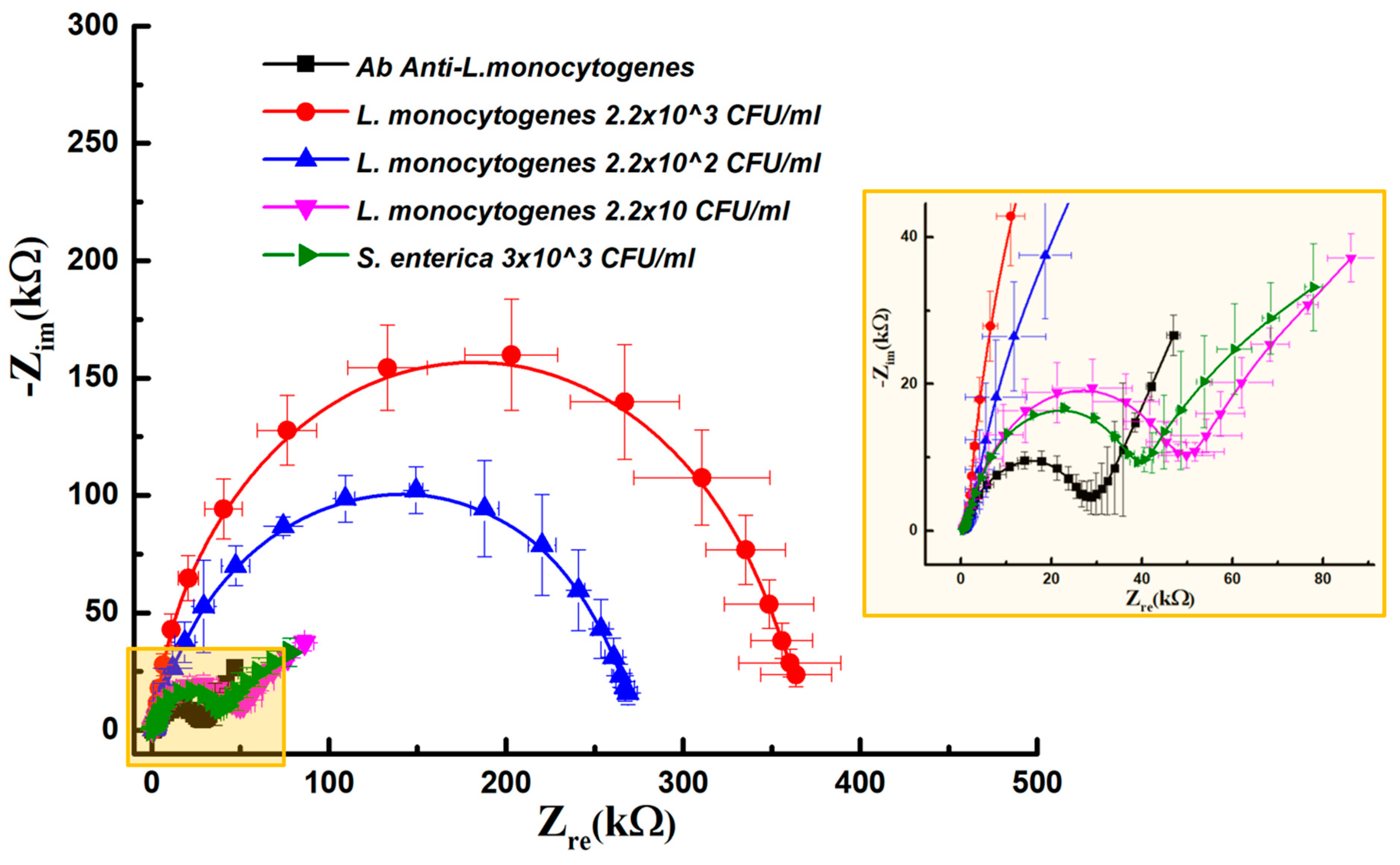
L. monocytogenes in culture medium suspension were incubated above the sensing layer. After the incubation time and a washing step, redox solution was replaced in the chamber and EIS measurements were collected. As shown in
Figure 3, a considerable increase in impedance values with respect to the antibody baseline can be associated to the incubation of the stock solution (2.2 × 10
3 CFU/mL) on the sensing layer (red line). As the concentration of
L. monocytogenes in suspension decreases, this results in a decrease in impedance values. As can be seen in the diagram, the presence of 2.2 × 10
2 CFU/mL produced EIS values of around 269 kΩ (blue line).
Using serially diluted samples of L. monocytogenes, (i.e., 2.2 × 103, 2.2 × 102, 2.2 × 101), a decrease in impedance values was recorded, from 269 kΩ to 51 kΩ. The suspension containing 22 CFU/mL was associated with an increment of impedance values of around 51 kΩ (pink line, inset).
As a negative control, we incubated a suspension containing 3 × 103 CFU/mL of Salmonella enterica. Impedance values measured in this case gave values of around 39 kΩ (green line, inset). Curves related to Ab baseline, the lowest concentration of L. monocytogenes and negative controls are very close, thus revealing a limit of the detection of the sensing platform realized. The impedance increase was 7 kΩ in the case of negative control, while the value obtained at the highest concentration tested (3 × 103 CFU/mL) produced values of about 350 kΩ, showing the high specificity of the optimized sensing system.
In order to test the possibility to use our optimized device for the detection of
L. monocytogenes in real samples, we prepared half-skimmed milk spiked samples with different amount of bacteria. As it was not possible to exactly replicate concentrations of
L. monocytogenes in a culture medium, we fixed a milk concentration of 75% and the remaining 25% of the sample was represented by the bacteria suspension, both in the case of
L. monocytogenes and of negative control (
S. enterica) (
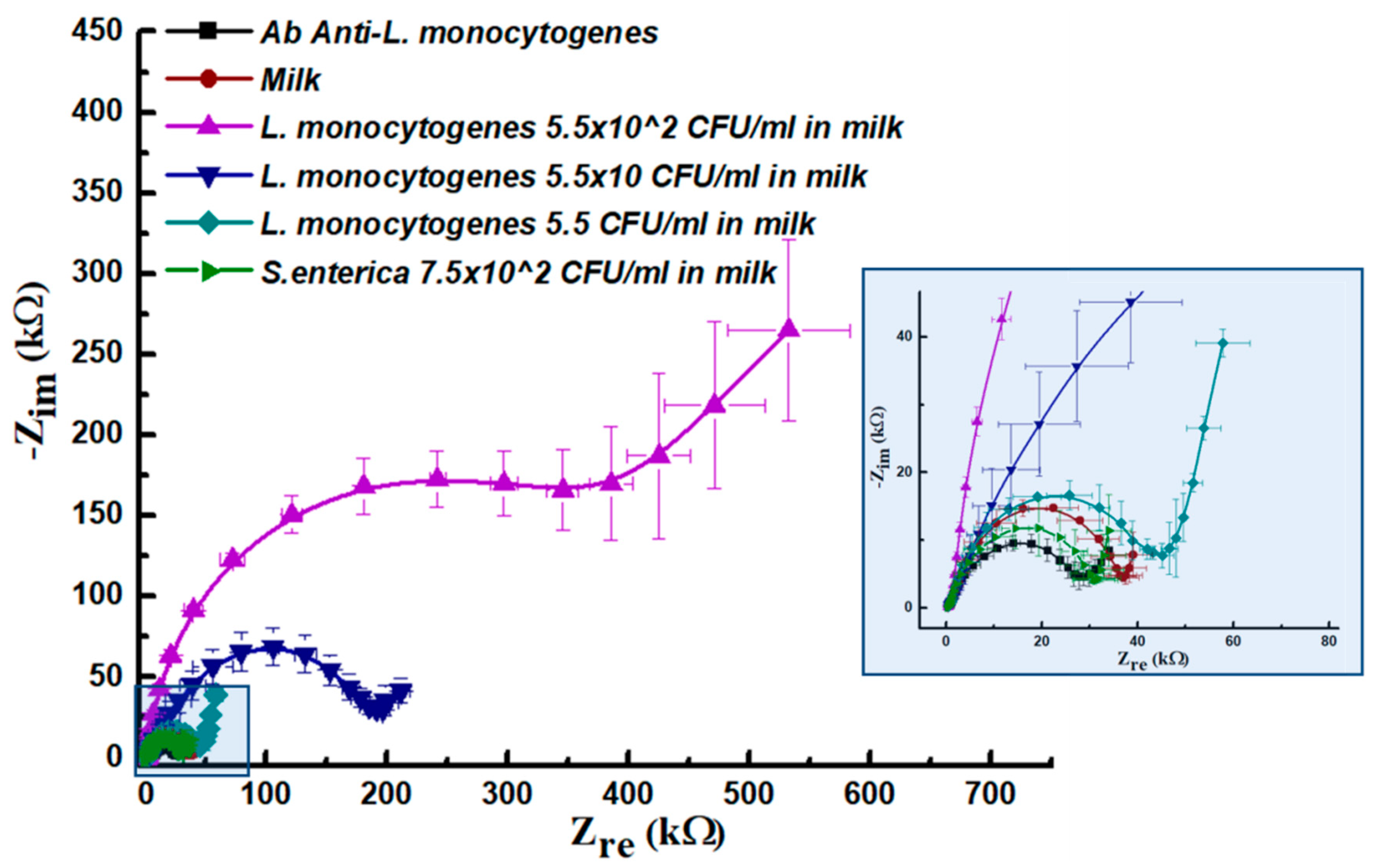
Figure 4).
In this case, we first tested the matrix contribution due to milk (100%) unspecific absorption (red line, inset in
Figure 4). As a result, a low impedance value (37 kΩ) was detected over the Ab baseline fixed at 27 kΩ, thus confirming once again the specificity of the sensing layer. In this experiment we incubated the first prepared spiked sample (75% spiked milk) containing 5.5 × 10
2 CFU/mL, collecting high impedance values of around 386 kΩ. At a concentration of 5.5 × 10 CFU/mL EIS values resulted in around 196 kΩ, while in the case of 5.5 CFU in milk we found an increased impedance response (45 kΩ) over the baseline related to antibodies and also over the milk matrix contribution, thus revealing a better performance of the assay in spiked milk samples with respect to the same assay carried on culture medium. In order to set up negative control by considering spiked
S. enteric samples made up of 75% of half-skimmed milk and 25% of culture medium containing 7.5 × 10
2 CFU/mL. In this case the related Nyquist plot had a diameter of 31 kΩ, thus very close to the antibody baseline (green curve in
Figure 4).
The aim of these experiments was to compare performances of the optimized platform to detect
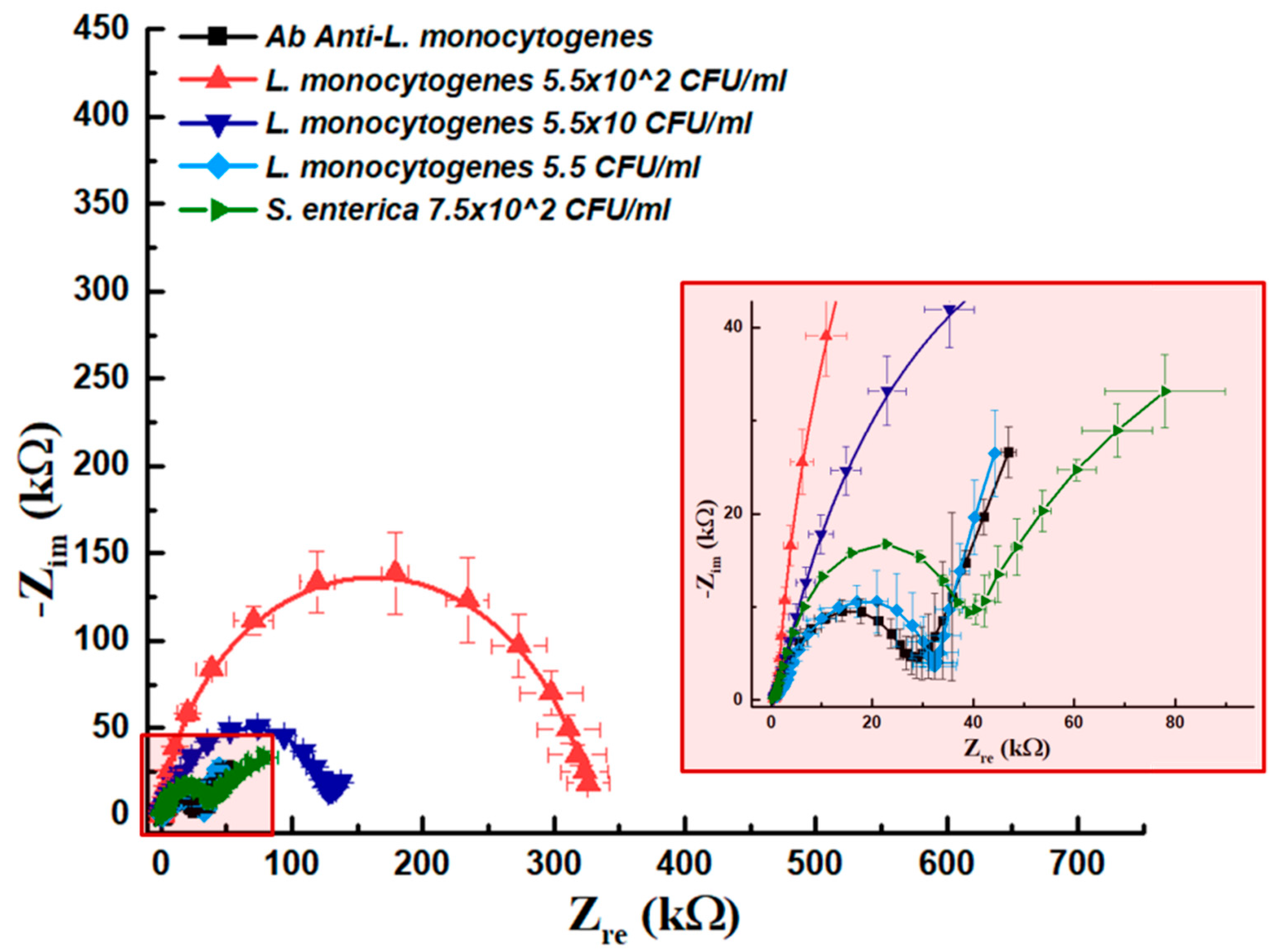
L. monocytogenes in culture media and in real samples, in order to verify the possibility to use it as a tool for on-field analysis. To allow this, a direct comparison of impedance values considering concentrations, was not possible, as skimmed milk spiked samples were prepared to contain 75% of the matrix, and 25% of suspended Listeria cells. Therefore, in order to bypass this differences and to compare the response of the system in presence of a different matrix, we prepared culture medium samples containing the same amount of
L. monocytogenes cells. As can be seen from
Figure 5, the two most concentrated suspensions gave a remarkable increase in impedance values over the sensing area. The
L. monocytogenes suspension containing 5.5 CFU/mL was instead overlapping with the antibody baseline, below or near the value associated with that obtained using
S. enterica as a negative control. These results, combined with the first calibration curves, allowed to set the limit of detection (LoD) for
L. monocytogenes dissolved in the medium between 22 and 5.5 CFU/mL.
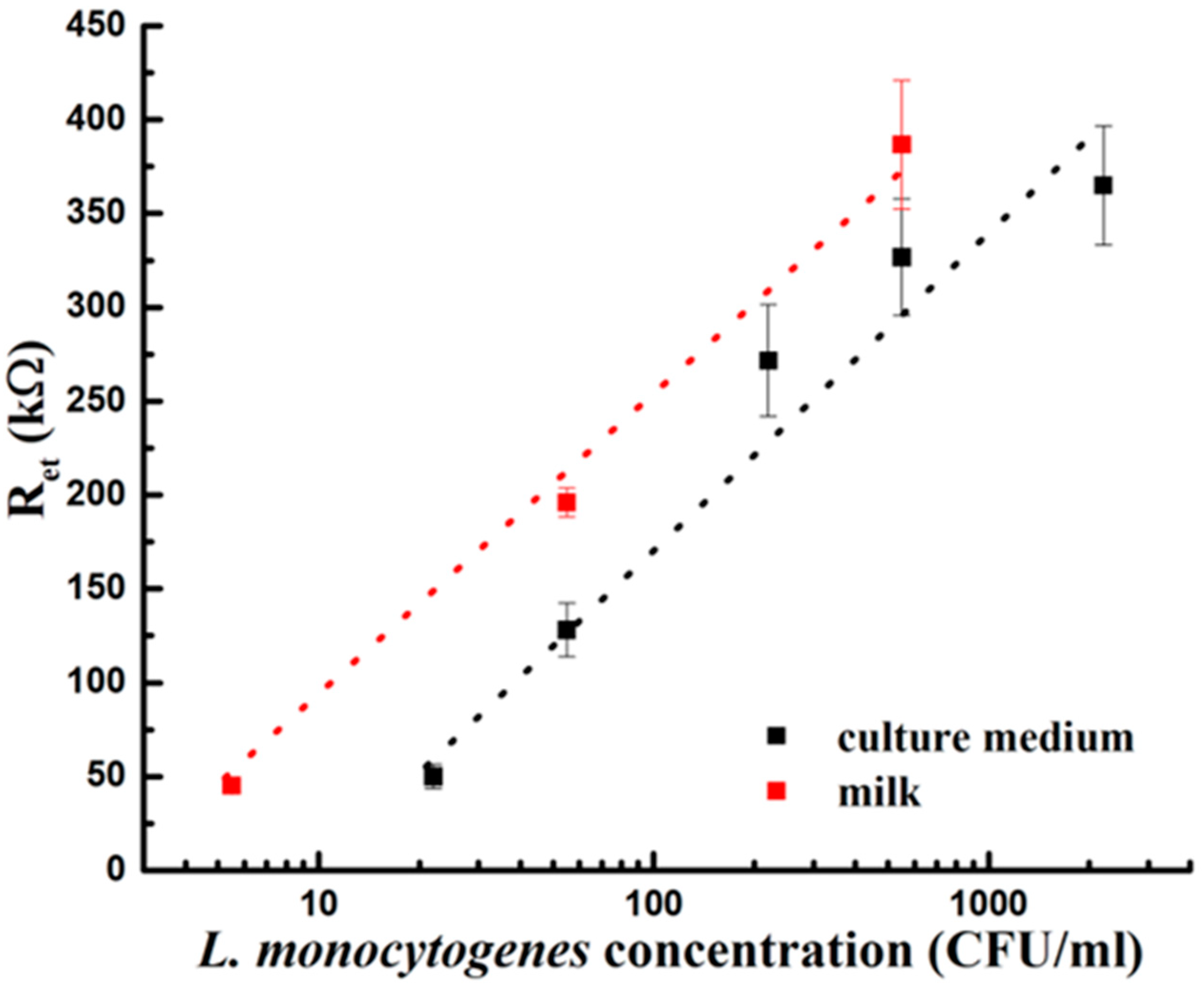
Once our platform was calibrated for
L. monocytogenes detection in medium culture and in milk spiked samples we obtained two calibration curves (
Figure 6), using the collected Ret values plotted in function of the different
L. monocytogenes concentrations.
4. Discussion
Food contamination is an eventuality that may happen in critical points of the food supply chain, and Listeria monocytogenes is one of the most threatening factors of contamination. In order to promptly identify such events, tools for the on-field analysis of dairy products, with the features of being low-costs portable, and easy to use, are mandatory. In this study, a biosensing platform was realized for the sensitive and specific on-chip detection of L. monocytogenes both in medium culture (reaching a limit of detection of 22 CFU/mL) and in milk matrix, reaching in this case a lower limit of detection, established as 5.5 CFU/mL, with negligible effects from matrix and unspecific pathogen detection (using S. enterica as negative control).
The presence of a matrix introduces a new variable, that in antibody-based biosensors consist in the unspecific binding of proteins and other solutes to the electrodes. In this study, the good performance of the platform tested with the food matrix could be attributed to the intrinsic features of the sample. In the case of milk, indeed, casein acts as a natural passivation agent [
25], and in laboratory protocols of sensing methods it is often added to improve the sensibility of the assay [
26]. These characteristics in this optimized assay allowed to match the legally-imposed limits of presence for
L. monocytogenes in foods, fixed at 100 CFU/g of food. However, ready-to-eat (RTE) foods and meats pose serious problems even with a minimum presence of the pathogen. For this reason, US safety agencies adopt different methods than EU and zero tolerance for
L. monocytogenes in RTE products [
10,
11].
Other types of substrates used for electrodes fabrication have been based on degenerate (highly doped) Si: The doped Si has very weak electrical conductivity and prevents the formation of a space charge layer during AC interrogation of the sensor interface. Radhakrishnan and Suni presented results demonstrating the possibility to regenerate the Si electrode during experiments during a 30-days storage using KSCN based solution [
27,
28]. The result showed the possible application and reusability of antibody-coated degenerate Si electrodes for EIS measurements, just performing the calibration on the day of use of electrodes.
Further advancements are envisaged through the application of superior surface modification methods, such as the use of bidendate thiol 16-[3,5-bis(mercaptomethyl)phenoxy]-hexadecanoic acid (BMPHA), the application of proper solutions to stabilize the capture ligands, and the application of AC electrokinetic (ACEK) microfluidics platforms and ACEK microflow mechanisms, such as AC electroosmosis (ACEO) and AC electrothermal (ACET) effects. Furthermore, an increase in the limit of detection may be achieved by combining the biosensor methods with immunoseparation of bacteria from larger volumes [
29,
30].
Several challenges and obstacles remain to be solved before their use in on-site detection. EIS sensor analysis needs operator skills in data processing. Sample preparation must be simple and performable on site. EIS methods have been often applied successfully to detects pathogens in real samples, such as plant extracts and sera [
31,
32].
EIS sensing platforms are an appealing device for the food industry and health officers: The immediate detection of pathogens [
33,
34] may allow a quick set up of countermeasures for protection of humans, plants and environment. In a recent study, we used the same anti-
Listeria antibodies, and obtained reproducible results on the specificity of antibodies [
21]. In addition, a protein chip was set up to discriminate five different species of pathogens on the same surface, i.e.,
L. monocytogenes,
S. aureus,
E. coli,
Salmonella spp. and
Campylobacter spp. [
33]. In that setting, the five pathogen-specific antibodies showed no binding to any of the other pathogens, so that specificity was high. Using the arrayed antibodies, Limit of Detection (LOD) of protein chips for
L. monocytogenes was 2.55 CFU/mL, even in presence of other bacterial species. In that study, the bacteria were from a 24 h enrichment culture, after immunoseparation from 50 mL or higher volumes using a Pathatrix instrument (Life Technologies, Carlsbad, CA, USA). Finally, validation assays remain to be performed, to show the robustness of detection, reproducibility and variability in multi-site installations.