Ensemble Clustering Method via Robust Consensus Learning
Abstract
1. Introduction
- (1)
- A symmetric error matrix is introduced for each connective matrix to identify noise components effectively. Additionally, a reliable consensus matrix is recovered by designing a set of mapping models that address structural differences among the connective matrices and enable robust consensus learning.
- (2)
- Multi-order graph structures are designed to fully exploit the association relationships between samples in the feature space, thereby enhancing the structural quality of the consensus matrix.
- (3)
- A theoretical rank constraint is incorporated to reinforce the block-diagonal property of the consensus matrix, ensuring a clear cluster structure.
- (4)
- The experimental results on all adopted datasets demonstrate that ECM-RCL is effective compared to the state-of-the-art methods.
2. Preliminaries
2.1. Co-Association Matrix
2.2. Multi-Order Graph Structures
3. Ensemble Clustering Method via Robust Consensus Learning
3.1. The Objective Function of ECM-RCL
3.2. Optimization and Analysis
3.2.1. Optimization
| Algorithm 1: Optimization of Objective Function |
| Input:, multi-order graph structures . |
|
| Output: Consensus Matrix |
| Algorithm 2: ECM-RCL |
| Input:, the number of base clustering results , the number of ground-truth clusters , the number of clusters of base clustering results . |
|
| Output: Final ensemble clustering result. |
3.2.2. Complexity Analysis
4. Experimental Section
4.1. Experimental Organization
4.1.1. Datasets
4.1.2. Comparative Methods
4.1.3. Experimental Setup and Evaluation Indices
4.2. Discussions on Datasets
4.2.1. Clustering Performance
4.2.2. Statistical Analysis
4.2.3. Ablation Study
4.2.4. Parameter Sensitivity Analysis
4.2.5. Robustness Analysis of Base Clustering Results
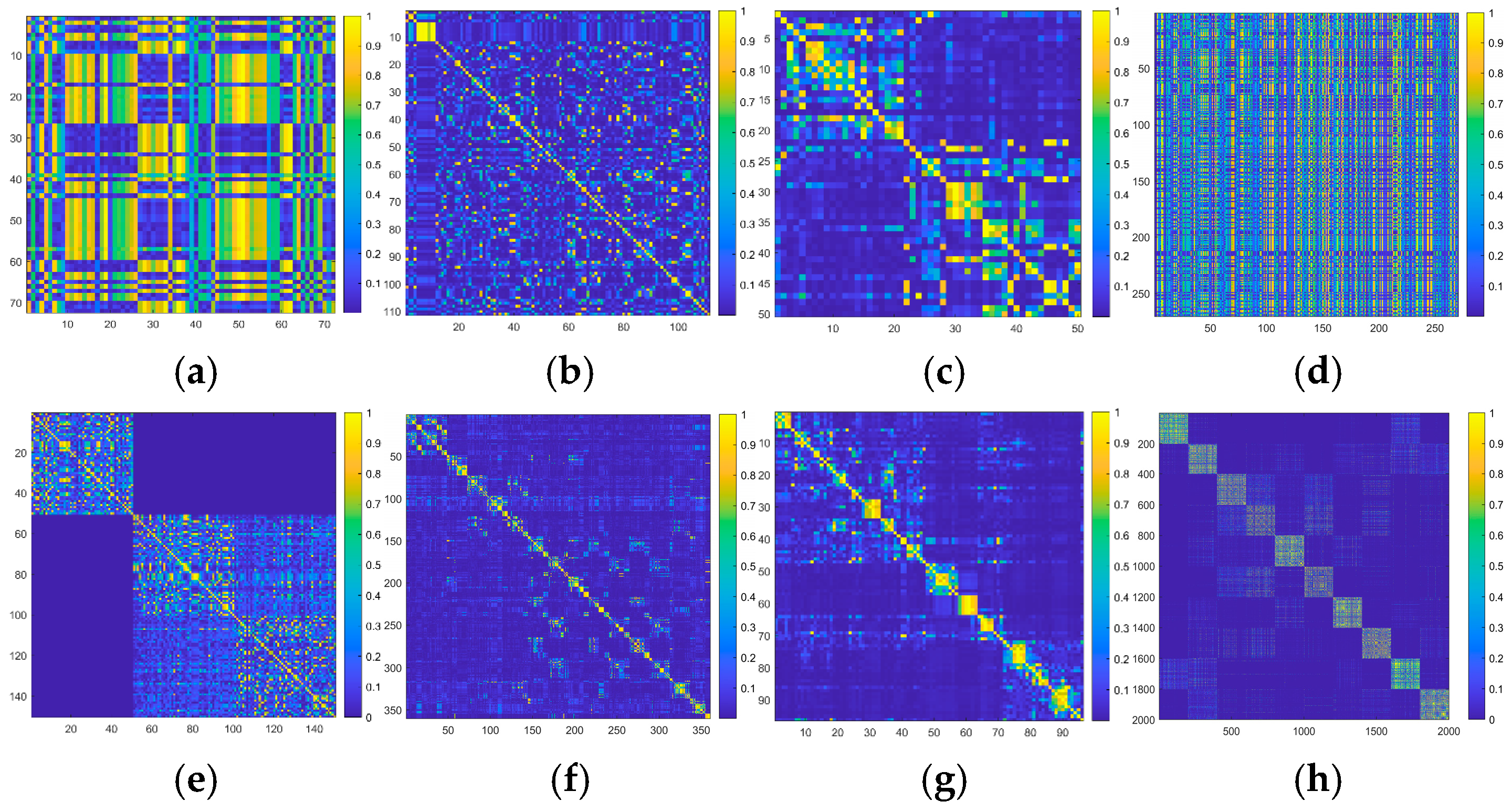
4.2.6. Visual Analysis
4.2.7. Execution Time and Convergence Analysis
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Appendix A
Appendix B

Appendix C







References
- Popescu, M.; Keller, J.; Bezdek, J.; Zare, A. Random projections fuzzy c-means (RPFCM) for big data clustering. In Proceedings of the 2015 IEEE International Conference on Fuzzy Systems, Istanbul, Turkey, 2–5 August 2015. [Google Scholar]
- Sun, H.; Liu, L.; Li, F. A lie group semi-supervised FCM clustering method for image segmentation. Pattern Recognit. 2024, 155, 110681. [Google Scholar] [CrossRef]
- Huang, D.; Wang, C.D.; Lai, J.H. Locally Weighted Ensemble Clustering. IEEE Trans. Cybern. 2018, 48, 1460–1473. [Google Scholar] [CrossRef]
- Huang, D.; Wang, C.-D.; Peng, H.; Lai, J.; Kwoh, C.-K. Enhanced Ensemble Clustering via Fast Propagation of Cluster-Wise Similarities. IEEE Trans. Syst. Man Cybern. Syst. 2021, 51, 508–520. [Google Scholar] [CrossRef]
- Ji, X.; Sun, J.; Peng, J.; Pang, Y.; Zhou, P. Clustering Ensemble Based on Fuzzy Matrix Self-Enhancement. IEEE Trans. Knowl. Data Eng. 2024, 37, 148–161. [Google Scholar] [CrossRef]
- Jia, Y.; Liu, H.; Hou, J.; Zhang, Q. Clustering Ensemble Meets Low-rank Tensor Approximation. In Proceedings of the Thirty-Fifth AAAI Conference on Artificial Intelligence, Online, 2–9 February 2021; Volume 35, pp. 7970–7978. [Google Scholar]
- Jia, Y.; Tao, S.; Wang, R.; Wang, Y. Ensemble Clustering via Co-Association Matrix Self-Enhancement. IEEE Trans. Neural Netw. Learn. Syst. 2023, 35, 11168–11179. [Google Scholar] [CrossRef]
- Shi, Y.; Yu, Z.; Chen, C.L.P.; Zeng, H. Consensus Clustering with Co-Association Matrix Optimization. IEEE Trans. Neural Netw. Learn. Syst. 2024, 35, 4192–4205. [Google Scholar] [CrossRef]
- Chen, M.S.; Lin, J.Q.; Wang, C.D.; Huang, D.; Lai, J.H. Contrastive Ensemble Clustering. IEEE Trans. Neural Netw. Learn. Syst. 2025, 36, 14678–14690. [Google Scholar] [CrossRef]
- Li, T.; Shu, X.; Wu, J.; Zheng, Q.; Lv, X.; Xu, J. Adaptive weighted ensemble clustering via kernel learning and local information preservation. Knowl.-Based Syst. 2024, 294, 111793. [Google Scholar] [CrossRef]
- Ay, M.; Özbakır, L.; Kulluk, S.; Gülmez, B.; Öztürk, G.; Özer, S. FC-Kmeans: Fixed-centered K-means algorithm. Expert Syst. Appl. 2023, 211, 118656. [Google Scholar] [CrossRef]
- Bagherinia, A.; Minaei-Bidgoli, B.; Hossinzadeh, M.; Parvin, H. Elite fuzzy clustering ensemble based on clustering diversity and quality measures. Appl. Intell. 2018, 49, 1724–1747. [Google Scholar] [CrossRef]
- Tao, Z.; Liu, H.; Li, S.; Ding, Z.; Fu, Y. Robust Spectral Ensemble Clustering via Rank Minimization. ACM Trans. Knowl. Discov. Data 2019, 13, 1–25. [Google Scholar] [CrossRef]
- Zhou, P.; Sun, B.; Liu, X.; Du, L.; Li, X. Active Clustering Ensemble with Self-Paced Learning. IEEE Trans. Neural Netw. Learn. Syst. 2023, 35, 12186–12200. [Google Scholar] [CrossRef] [PubMed]
- Zhou, P.; Du, L.; Li, X. Adaptive Consensus Clustering for Multiple K-Means Via Base Results Refining. IEEE Trans. Knowl. Data Eng. 2023, 35, 10251–10264. [Google Scholar] [CrossRef]
- Gu, Q.; Wang, Y.; Wang, P.; Li, X.; Chen, L.; Xiong, N.N.; Liu, D. An improved weighted ensemble clustering based on two-tier uncertainty measurement. Expert Syst. Appl. 2024, 238, 121672. [Google Scholar] [CrossRef]
- Xu, J.; Li, T.; Zhang, D.; Wu, J. Ensemble clustering via fusing global and local structure information. Expert Syst. Appl. 2024, 237, 121557. [Google Scholar] [CrossRef]
- Xu, J.; Li, T.; Wu, J.; Zhang, D. Ensemble clustering via dual self-enhancement by alternating denoising and topological consistency propagation. Appl. Soft Comput. J. 2024, 167, 112299. [Google Scholar] [CrossRef]
- Yang, X.; Zheng, Z.; Xie, J.; Zhao, W.; Xue, J.; Nie, F. Spectral ensemble clustering from graph reconstruction with auto-weighted cluster. Pattern Recognit. Lett. 2025, 196, 243–249. [Google Scholar] [CrossRef]
- Zheng, X.; Lu, Y.; Wang, R.; Nie, F.; Li, X. Structured Graph-Based Ensemble Clustering. IEEE Trans. Knowl. Data Eng. 2025, 37, 3728–3738. [Google Scholar] [CrossRef]
- Zeng, L.; Yao, S.; Liu, X.; Xiao, L.; Qian, Y. A clustering ensemble algorithm for handling deep embeddings using cluster confidence. Comput. J. 2025, 68, 163–174. [Google Scholar] [CrossRef]
- Huang, D.; Chen, D.-H.; Chen, X.; Wang, C.-D.; Lai, J.-H. DeepCluE: Enhanced Deep Clustering via Multi-Layer Ensembles in Neural Networks. IEEE Trans. Emerg. Top. Comput. Intell. 2024, 8, 1582–1594. [Google Scholar] [CrossRef]
- Hao, Z.; Lu, Z.; Li, G.; Nie, F.; Wang, R.; Li, X. Ensemble Clustering with Attentional Representation. IEEE Trans. Knowl. Data Eng. 2023, 36, 581–593. [Google Scholar] [CrossRef]
- Shukla, P.K.; Veerasamy, B.D.; Alduaiji, N.; Addula, S.R.; Pandey, A.; Shukla, P.K. Fraudulent account detection in social media using hybrid deep transformer model and hyperparameter optimization. Sci. Rep. 2025, 15, 38447. [Google Scholar] [CrossRef]
- Miklautz, L.; Teuffenbach, M.; Weber, P.; Perjuci, R.; Durani, W.; Bohm, C.; Plant, C. Deep Clustering with Consensus Representations. In Proceedings of the 2022 IEEE International Conference on Data Mining (ICDM), Orlando, FL, USA, 28 November 2022–1 December 2022; pp. 1119–1124. [Google Scholar]
- Liang, C.; Dong, Z.; Yang, S.; Zhou, P. Jointly Learn the Base Clustering and Ensemble for Deep Image Clustering. In Proceedings of the 2024 IEEE International Conference on Multimedia and Expo, Niagara Falls, ON, Canada, 15–19 July 2024. [Google Scholar]
- Huang, D.; Lai, Q.; Wang, H.; Xu, Y.-K.; Guan, C.-B.; Wang, C.-D. REC-GCN: Robust ensemble clustering with graph convolutional networks. Pattern Recognit. 2026, 172, 112717. [Google Scholar] [CrossRef]
- Zhou, P.; Du, L.; Wang, H.; Shi, L.; Shen, Y.-D. Learning a Robust Consensus Matrix for Clustering Ensemble via Kullback-Leibler Divergence Minimization. In Proceedings of the Twenty-Fourth International Joint Conference on Artificial Intelligence, Buenos Aires, Argentina, 25–31 July 2015. [Google Scholar]
- Chen, M.; Lin, J.; Wang, C.; Xi, W.; Huang, D. On Regularizing Multiple Clusterings for Ensemble Clustering by Graph Tensor Learning. In Proceedings of the 31st ACM International Conference on Multimedia, Ottawa, ON, Canada, 29 October–3 November 2023. [Google Scholar]
- Zhou, P.; Du, L.; Liu, X.; Shen, Y.D.; Fan, M.; Li, X. Self-Paced Clustering Ensemble. IEEE Trans. Neural Netw. Learn. Syst. 2021, 32, 1497–1511. [Google Scholar] [CrossRef]
- Xu, J.; Li, T. Ensemble clustering with low-rank optimal Laplacian matrix learning. Appl. Soft Comput. 2024, 150, 111095. [Google Scholar] [CrossRef]
- Ren, Z.; Sun, Q.; Wei, D. Multiple Kernel Clustering with Kernel k-Means Coupled Graph Tensor Learning. In Proceedings of the Thirty-Fifth AAAI Conference on Artificial Intelligence, Online, 2–9 February 2021. [Google Scholar]
- Lu, C.; Feng, J.; Lin, Z.; Mei, T.; Yan, S. Subspace Clustering by Block Diagonal Representation. IEEE Trans. Pattern Anal. Mach. Intell. 2019, 41, 487–501. [Google Scholar] [CrossRef]
- Pan, E.; Kang, Z. High-order multi-view clustering for generic data. Inf. Fusion 2023, 100, 101947. [Google Scholar] [CrossRef]
- Wang, Z.; Li, L.; Ning, X.; Tan, W.; Liu, Y.; Song, H. Incomplete multi-view clustering via structure exploration and missing-view inference. Inf. Fusion 2024, 103, 102123. [Google Scholar] [CrossRef]
- Wang, Y.; Wu, N.-C.; Liu, Y.; Xiang, H. A generalized Nyström method with subspace iteration for low-rank approximations of large-scale nonsymmetric matrices. Appl. Math. Lett. 2025, 166, 109531. [Google Scholar] [CrossRef]
- Zhou, S.; Yang, M.; Wang, X.; Song, W. Anchor-based scalable multi-view subspace clustering. Inf. Sci. 2024, 666, 120374. [Google Scholar] [CrossRef]
- Zhou, J.; Zheng, H.; Pan, L. Ensemble clustering based on dense representation. Neurocomputing 2019, 357, 66–76. [Google Scholar] [CrossRef]
- Zhou, P.; Hu, B.; Yan, D.; Du, L. Clustering Ensemble via Diffusion on Adaptive Multiplex. IEEE Trans. Knowl. Data Eng. 2024, 36, 1463–1474. [Google Scholar] [CrossRef]
- Qin, Y.; Pu, N.; Sebe, N.; Feng, G. Latent Space Learning-Based Ensemble Clustering. IEEE Trans. Image Process. 2025, 34, 1259–1270. [Google Scholar] [CrossRef] [PubMed]
- Bian, Z.; Ishibuchi, H.; Wang, S. Joint Learning of Spectral Clustering Structure and Fuzzy Similarity Matrix of Data. IEEE Trans. Fuzzy Syst. 2019, 27, 31–44. [Google Scholar] [CrossRef]
- Bian, Z.; Qu, J.; Zhou, J.; Jiang, Z.; Wang, S. Weighted adaptively ensemble clustering method based on fuzzy Co-association matrix. Inf. Fusion 2024, 103, 102099. [Google Scholar] [CrossRef]
- Zhou, P.; Liu, X.; Du, L.; Li, X. Self-paced Adaptive Bipartite Graph Learning for Consensus Clustering. ACM Trans. Knowl. Discov. Data 2023, 17, 1–35. [Google Scholar] [CrossRef]
- Taheri, S.M.; Hesamian, G. A generalization of the Wilcoxon signed-rank test and its applications. Stat. Pap. 2012, 54, 457–470. [Google Scholar] [CrossRef]
- Bian, Z.; Yu, L.; Qu, J.; Deng, Z.; Wang, S. An ensemble clustering method via learning the CA matrix with fuzzy neighbors. Inf. Fusion 2025, 120, 103105. [Google Scholar] [CrossRef]






| Methods | WBC | WC | CSD | CD | BDS | OD |
|---|---|---|---|---|---|---|
| Bagherinia et al. [12] | √ | |||||
| Huang et al. [3] | √ | |||||
| Tao et al. [13] | Weak | Weak | ||||
| Zhou et al. [14] | √ | √ | √ | |||
| Jia et al. [7] | √ | Weak | ||||
| Zhou et al. [15] | √ | √ | ||||
| Gu et al. [16] | √ | √ | ||||
| Xu et al. [17] | √ | Weak | ||||
| Li et al. [10] | Weak | |||||
| Xu et al. [18] | √ | Weak | √ | |||
| Zheng et al. [20] | √ | √ | ||||
| Yang et al. [19] | √ | √ | ||||
| ECM-RCL | √ | √ | √ | √ | √ |
| Notations | Descriptions |
|---|---|
| The dataset. | |
| The CA matrix. | |
| -th element in | |
| The -th connective matrix. | |
| The -th error matrix. | |
| The identity matrix. | |
| The -th mapping matrix. | |
| The -th order graph structure. | |
| . | |
| The consensus matrix. | |
| -th element in . | |
| The -th base clustering result. | |
| The number of base clustering results. | |
| The maximum order of the multi-order graph structures. | |
| The number of samples. | |
| The number of ground-truth clusters. | |
| The number of clusters of the -th base clustering result. | |
| The number of features in . | |
| The -th sample. | |
| The Frobenius norm of . | |
| The trace of . |
| Datasets | Number of Samples | Number of Features | Number of Clusters |
|---|---|---|---|
| ALLAML | 72 | 7129 | 2 |
| CLL_SUB | 111 | 11,340 | 3 |
| GLIMA | 50 | 4434 | 4 |
| Heart | 270 | 13 | 2 |
| Iris | 150 | 4 | 3 |
| LM | 360 | 90 | 15 |
| Lymphoma | 96 | 4026 | 9 |
| MF | 2000 | 649 | 10 |
| Orlraws10p | 100 | 10,304 | 10 |
| USPS | 1854 | 256 | 10 |
| Vertebral | 310 | 6 | 3 |
| WarpPIE10p | 210 | 2420 | 10 |
| WDBC | 569 | 30 | 2 |
| Wine | 178 | 13 | 3 |
| Zoo | 101 | 16 | 7 |
| ISOLET | 7797 | 617 | 26 |
| LS | 6435 | 36 | 6 |
| ODR | 5620 | 64 | 10 |
| PD | 10,992 | 16 | 10 |
| Datasets | K-Means | LWEA | DREC | ECPCS-HC | EC-CMS | ECAR | CEAM | ECM-RCL |
|---|---|---|---|---|---|---|---|---|
| ALLAML | 0.6903 ± 0.0371 | 0.6833 ± 0.0667 | 0.6903 ± 0.0687 | 0.6569 ± 0.0365 | 0.6722 ± 0.0736 | 0.6806 ± 0.0000 | 0.5972 ± 0.0729 | 0.7153 ± 0.0150 |
| CLL_SUB | 0.5252 ± 0.0085 | 0.5279 ± 0.0047 | 0.5279 ± 0.0047 | 0.5279 ± 0.0047 | 0.5270 ± 0.0047 | 0.4775 ± 0.0000 | 0.4486 ± 0.0337 | 0.5459 ± 0.0047 |
| GLIOMA | 0.5600 ± 0.0794 | 0.5900 ± 0.0216 | 0.5860 ± 0.0232 | 0.5740 ± 0.0212 | 0.5900 ± 0.0368 | 0.3000 ± 0.0000 | 0.5680 ± 0.0473 | 0.6440 ± 0.0430 |
| Heart | 0.5904 ± 0.0019 | 0.5856 ± 0.0271 | 0.6037 ± 0.0287 | 0.5933 ± 0.0298 | 0.6100 ± 0.0243 | 0.6185 ± 0.0000 | 0.5837 ± 0.0379 | 0.6244 ± 0.0053 |
| Iris | 0.8560 ± 0.1041 | 0.8747 ± 0.0301 | 0.9020 ± 0.0336 | 0.8993 ± 0.0320 | 0.8987 ± 0.0320 | 0.6667 ± 0.0000 | 0.7300 ± 0.0668 | 0.9140 ± 0.0685 |
| LM | 0.4406 ± 0.0267 | 0.4503 ± 0.0153 | 0.4558 ± 0.0117 | 0.4292 ± 0.0184 | 0.4550 ± 0.0194 | 0.4611 ± 0.0000 | 0.4583 ± 0.0154 | 0.5200 ± 0.0140 |
| Lymphoma | 0.5427 ± 0.0546 | 0.5740 ± 0.0724 | 0.5646 ± 0.0576 | 0.6948 ± 0.0908 | 0.5198 ± 0.0700 | 0.5000 ± 0.0000 | 0.5677 ± 0.0256 | 0.6552 ± 0.0421 |
| MF | 0.5036 ± 0.0453 | 0.5493 ± 0.0298 | 0.5666 ± 0.0338 | 0.5447 ± 0.0359 | 0.6057 ± 0.0302 | 0.5200 ± 0.0000 | 0.5751 ± 0.0647 | 0.8004 ± 0.0096 |
| Orlraws10P | 0.6730 ± 0.0536 | 0.7960 ± 0.0481 | 0.7830 ± 0.0447 | 0.7220 ± 0.0377 | 0.7980 ± 0.0424 | 0.6500 ± 0.0000 | 0.8010 ± 0.0567 | 0.8470 ± 0.0408 |
| USPS | 0.6182 ± 0.0278 | 0.6836 ± 0.0254 | 0.6901 ± 0.0372 | 0.6859 ± 0.0560 | 0.7273 ± 0.0453 | 0.6208 ± 0.0000 | 0.6972 ± 0.0471 | 0.7573 ± 0.0000 |
| Vertebral | 0.5984 ± 0.0546 | 0.6177 ± 0.1135 | 0.5352 ± 0.0284 | 0.6110 ± 0.1337 | 0.5358 ± 0.0195 | 0.6516 ± 0.0000 | 0.5652 ± 0.0680 | 0.6703 ± 0.0866 |
| WarpPIE10p | 0.2605 ± 0.0182 | 0.2357 ± 0.0093 | 0.2552 ± 0.0202 | 0.2124 ± 0.0068 | 0.2386 ± 0.0182 | 0.1952 ± 0.0000 | 0.2676 ± 0.0214 | 0.4762 ± 0.0135 |
| WDBC | 0.8541 ± 0.0000 | 0.8035 ± 0.0876 | 0.8283 ± 0.0296 | 0.8738 ± 0.0411 | 0.8190 ± 0.0763 | 0.9244 ± 0.0000 | 0.8649 ± 0.0335 | 0.8903 ± 0.0191 |
| Wine | 0.6292 ± 0.0644 | 0.6713 ± 0.0651 | 0.6809 ± 0.0388 | 0.6494 ± 0.0746 | 0.6691 ± 0.0701 | 0.6404 ± 0.0000 | 0.6758 ± 0.0464 | 0.7247 ± 0.0000 |
| Zoo | 0.7347 ± 0.0800 | 0.7426 ± 0.0585 | 0.7455 ± 0.0469 | 0.7436 ± 0.0211 | 0.7188 ± 0.0725 | 0.7822 ± 0.0000 | 0.6267 ± 0.0841 | 0.7248 ± 0.0283 |
| ISOLET | 0.5255 ± 0.0284 | 0.5620 ± 0.0100 | 0.5624 ± 0.0163 | 0.5226 ± 0.0119 | 0.5508 ± 0.0015 | 0.5256 ± 0.0000 | 0.5319 ± 0.0107 | 0.5645 ± 0.0023 |
| LS | 0.6342 ± 0.0678 | 0.6276 ± 0.1915 | 0.6494 ± 0.0026 | 0.6709 ± 0.1789 | 0.6253 ± 0.0065 | 0.7052 ± 0.0000 | 0.5460 ± 0.1282 | 0.7239 ± 0.0036 |
| ODR | 0.7592 ± 0.0649 | 0.8391 ± 0.0316 | 0.9017 ± 0.0457 | 0.8669 ± 0.0025 | 0.9212 ± 0.0045 | 0.7194 ± 0.0000 | 0.8484 ± 0.0033 | 0.9790 ± 0.0000 |
| PD | 0.7039 ± 0.0494 | 0.7836 ± 0.0031 | 0.8088 ± 0.0519 | 0.7426 ± 0.0270 | 0.7832 ± 0.0060 | 0.6543 ± 0.0000 | 0.7212 ± 0.0384 | 0.8890 ± 0.0021 |
| Datasets | K-Means | LWEA | DREC | ECPCS-HC | EC-CMS | ECAR | CEAM | ECM-RCL |
|---|---|---|---|---|---|---|---|---|
| ALLAML | 0.0897 ± 0.0393 | 0.1119 ± 0.0333 | 0.1209 ± 0.0310 | 0.0417 ± 0.0452 | 0.0857 ± 0.0656 | 0.1003 ± 0.0000 | 0.0505 ± 0.0379 | 0.1448 ± 0.0173 |
| CLL_SUB | 0.1804 ± 0.0004 | 0.1805 ± 0.0003 | 0.1805 ± 0.0003 | 0.1805 ± 0.0003 | 0.1804 ± 0.0003 | 0.0967 ± 0.0000 | 0.0686 ± 0.0289 | 0.2627 ± 0.0005 |
| GLIOMA | 0.4257 ± 0.1202 | 0.4921 ± 0.0226 | 0.4943 ± 0.0267 | 0.4780 ± 0.0164 | 0.4938 ± 0.0293 | 0.0000 ± 0.0000 | 0.4044 ± 0.0449 | 0.5253 ± 0.0198 |
| Heart | 0.0187 ± 0.0007 | 0.0176 ± 0.0134 | 0.0280 ± 0.0136 | 0.0210 ± 0.0151 | 0.0295 ± 0.0133 | 0.0410 ± 0.0000 | 0.0214 ± 0.0156 | 0.0409 ± 0.0042 |
| Iris | 0.7204 ± 0.0669 | 0.7505 ± 0.0437 | 0.7915 ± 0.0528 | 0.7870 ± 0.0482 | 0.7862 ± 0.0483 | 0.7337 ± 0.0000 | 0.5398 ± 0.0878 | 0.8131 ± 0.0853 |
| LM | 0.5623 ± 0.0243 | 0.5906 ± 0.0129 | 0.5930 ± 0.0148 | 0.5612 ± 0.0176 | 0.5961 ± 0.0172 | 0.5801 ± 0.0000 | 0.5862 ± 0.0171 | 0.6460 ± 0.0082 |
| Lymphoma | 0.5607 ± 0.0424 | 0.6141 ± 0.0423 | 0.6161 ± 0.0483 | 0.6964 ± 0.0509 | 0.5851 ± 0.0358 | 0.5575 ± 0.0000 | 0.5926 ± 0.0174 | 0.6711 ± 0.0179 |
| MF | 0.5594 ± 0.0182 | 0.6045 ± 0.0252 | 0.6152 ± 0.0321 | 0.6151 ± 0.0245 | 0.6462 ± 0.0233 | 0.6184 ± 0.0000 | 0.5896 ± 0.0344 | 0.7644 ± 0.0083 |
| Orlraws10P | 0.7607 ± 0.0323 | 0.8484 ± 0.0337 | 0.8437 ± 0.0310 | 0.8112 ± 0.0304 | 0.8392 ± 0.0286 | 0.7846 ± 0.0000 | 0.8374 ± 0.0413 | 0.9207 ± 0.0158 |
| USPS | 0.6138 ± 0.0150 | 0.6757 ± 0.0210 | 0.6797 ± 0.0244 | 0.6655 ± 0.0180 | 0.6981 ± 0.0154 | 0.6405 ± 0.0000 | 0.6734 ± 0.0283 | 0.7998 ± 0.0000 |
| Vertebral | 0.3898 ± 0.0299 | 0.3658 ± 0.1729 | 0.4404 ± 0.1083 | 0.2954 ± 0.1938 | 0.2808 ± 0.1750 | 0.4209 ± 0.0000 | 0.3926 ± 0.0902 | 0.4970 ± 0.0576 |
| WarpPIE10p | 0.2448 ± 0.0322 | 0.2115 ± 0.0287 | 0.2412 ± 0.0291 | 0.1726 ± 0.0136 | 0.2210 ± 0.0321 | 0.1400 ± 0.0000 | 0.2676 ± 0.0242 | 0.5562 ± 0.0127 |
| WDBC | 0.4223 ± 0.0000 | 0.3262 ± 0.1703 | 0.3666 ± 0.0639 | 0.4696 ± 0.0911 | 0.3549 ± 0.1524 | 0.6064 ± 0.0000 | 0.4313 ± 0.0872 | 0.5045 ± 0.0441 |
| Wine | 0.4100 ± 0.0165 | 0.4118 ± 0.0264 | 0.4032 ± 0.0424 | 0.4083 ± 0.0317 | 0.4046 ± 0.0365 | 0.2973 ± 0.0000 | 0.3546 ± 0.0960 | 0.4225 ± 0.0116 |
| Zoo | 0.7179 ± 0.0521 | 0.7287 ± 0.0434 | 0.7254 ± 0.0348 | 0.7068 ± 0.0498 | 0.7166 ± 0.0497 | 0.7921 ± 0.0000 | 0.6485 ± 0.0578 | 0.7219 ± 0.0093 |
| ISOLET | 0.7114 ± 0.0123 | 0.7334 ± 0.0094 | 0.7468 ± 0.0139 | 0.7101 ± 0.0110 | 0.7362 ± 0.0067 | 0.6674 ± 0.0000 | 0.7232 ± 0.0002 | 0.7633 ± 0.0015 |
| LS | 0.5485 ± 0.0682 | 0.4937 ± 0.1476 | 0.6157 ± 0.0120 | 0.5290 ± 0.1467 | 0.5215 ± 0.0132 | 0.6366 ± 0.0000 | 0.4477 ± 0.1125 | 0.6732 ± 0.0004 |
| ODR | 0.7274 ± 0.0310 | 0.8206 ± 0.012 | 0.8565 ± 0.0298 | 0.8323 ± 0.0016 | 0.8628 ± 0.0048 | 0.7045 ± 0.0000 | 0.8154 ± 0.0052 | 0.9505 ± 0.0000 |
| PD | 0.6712 ± 0.0205 | 0.7581 ± 0.0443 | 0.7965 ± 0.0046 | 0.7327 ± 0.0137 | 0.7742 ± 0.0161 | 0.6945 ± 0.0000 | 0.7265 ± 0.0217 | 0.8522 ± 0.0051 |
| Datasets | K-Means | LWEA | DREC | ECPCS-HC | EC-CMS | ECAR | CEAM | ECM-RCL |
|---|---|---|---|---|---|---|---|---|
| ALLAML | 0.1333 ± 0.0596 | 0.1329 ± 0.0836 | 0.1451 ± 0.0868 | 0.0372 ± 0.0797 | 0.1044 ± 0.1106 | 0.1189 ± 0.0000 | 0.0413 ± 0.0650 | 0.1754 ± 0.0256 |
| CLL_SUB | 0.0933 ± 0.0209 | 0.0867 ± 0.0008 | 0.0867 ± 0.0008 | 0.0867 ± 0.0008 | 0.0866 ± 0.0008 | 0.0436 ± 0.0000 | 0.0270 ± 0.0227 | 0.1230 ± 0.0011 |
| GLIOMA | 0.3004 ± 0.1308 | 0.3588 ± 0.0348 | 0.3566 ± 0.0390 | 0.3876 ± 0.0382 | 0.3760 ± 0.0434 | 0.0000 ± 0.0000 | 0.2467 ± 0.0333 | 0.3854 ± 0.0311 |
| Heart | 0.0287 ± 0.0013 | 0.0266 ± 0.0207 | 0.0411 ± 0.0227 | 0.0318 ± 0.0245 | 0.0459 ± 0.0194 | 0.0527 ± 0.0000 | 0.0292 ± 0.0257 | 0.0584 ± 0.0052 |
| Iris | 0.6911 ± 0.0946 | 0.7016 ± 0.0517 | 0.7548 ± 0.0736 | 0.7489 ± 0.0688 | 0.7476 ± 0.0690 | 0.5681 ± 0.0000 | 0.4603 ± 0.1019 | 0.7975 ± 0.1174 |
| LM | 0.2930 ± 0.0273 | 0.3283 ± 0.0218 | 0.3262 ± 0.0183 | 0.3252 ± 0.0245 | 0.3432 ± 0.0245 | 0.3076 ± 0.0000 | 0.3123 ± 0.0223 | 0.3895 ± 0.0122 |
| Lymphoma | 0.3214 ± 0.0893 | 0.3547 ± 0.0733 | 0.3426 ± 0.0580 | 0.5594 ± 0.1150 | 0.3012 ± 0.0704 | 0.3270 ± 0.0000 | 0.3064 ± 0.0263 | 0.4119 ± 0.0433 |
| MF | 0.4242 ± 0.0232 | 0.4652 ± 0.0304 | 0.4762 ± 0.0409 | 0.4823 ± 0.0277 | 0.5128 ± 0.0307 | 0.4293 ± 0.0000 | 0.4436 ± 0.0361 | 0.6819 ± 0.0072 |
| Orlraws10P | 0.5703 ± 0.0541 | 0.7080 ± 0.0644 | 0.6986 ± 0.0607 | 0.6244 ± 0.0589 | 0.6959 ± 0.0419 | 0.4328 ± 0.0000 | 0.7094 ± 0.0732 | 0.8330 ± 0.0218 |
| USPS | 0.5182 ± 0.0228 | 0.5790 ± 0.0255 | 0.5760 ± 0.0411 | 0.5963 ± 0.0480 | 0.6354 ± 0.0488 | 0.5347 ± 0.0000 | 0.5790 ± 0.0451 | 0.7061 ± 0.0000 |
| Vertebral | 0.3145 ± 0.0089 | 0.3105 ± 0.2637 | 0.2952 ± 0.1225 | 0.2585 ± 0.3133 | 0.1374 ± 0.1809 | 0.3240 ± 0.0000 | 0.3198 ± 0.1127 | 0.4652 ± 0.1460 |
| WarpPIE10p | 0.0592 ± 0.0185 | 0.0381 ± 0.0161 | 0.0551 ± 0.0191 | 0.0208 ± 0.0067 | 0.0407 ± 0.0182 | 0.0133 ± 0.0000 | 0.0788 ± 0.0157 | 0.3181 ± 0.0129 |
| WDBC | 0.4914 ± 0.0000 | 0.3746 ± 0.2025 | 0.4199 ± 0.0812 | 0.5568 ± 0.1219 | 0.4100 ± 0.1872 | 0.7182 ± 0.0000 | 0.5294 ± 0.1024 | 0.6050 ± 0.0615 |
| Wine | 0.3575 ± 0.0126 | 0.3705 ± 0.0191 | 0.3569 ± 0.0297 | 0.3563 ± 0.0286 | 0.3608 ± 0.0414 | 0.2745 ± 0.0000 | 0.3320 ± 0.0871 | 0.4007 ± 0.0000 |
| Zoo | 0.6281 ± 0.1125 | 0.6390 ± 0.0789 | 0.6405 ± 0.0695 | 0.6226 ± 0.0501 | 0.6117 ± 0.0977 | 0.7178 ± 0.0000 | 0.5097 ± 0.1345 | 0.6282 ± 0.0154 |
| ISOLET | 0.4692 ± 0.0185 | 0.4921 ± 0.0268 | 0.5216 ± 0.0150 | 0.4827 ± 0.0404 | 0.5447 ± 0.0031 | 0.4270 ± 0.0000 | 0.4629 ± 0.0275 | 0.5320 ± 0.0022 |
| LS | 0.4572 ± 0.0856 | 0.4504 ± 0.2156 | 0.5274 ± 0.0073 | 0.5244 ± 0.1947 | 0.4632 ± 0.0152 | 0.5715 ± 0.0000 | 0.3090 ± 0.1029 | 0.6167 ± 0.0003 |
| ODR | 0.6410 ± 0.0608 | 0.7644 ± 0.0249 | 0.8262 ± 0.0511 | 0.7832 ± 0.0067 | 0.8404 ± 0.0070 | 0.6047 ± 0.0000 | 0.7560 ± 0.0062 | 0.9545 ± 0.0000 |
| PD | 0.5589 ± 0.0404 | 0.6492 ± 0.0399 | 0.6987 ± 0.0258 | 0.6105 ± 0.0247 | 0.6594 ± 0.0200 | 0.5298 ± 0.0000 | 0.6052 ± 0.0366 | 0.7971 ± 0.0040 |
| Methods | Rank | -Value | The Null Hypothesis |
|---|---|---|---|
| K-means | 6.0789 | 0 | rejected |
| LWEA | 4.6579 | ||
| DREC | 3.7105 | ||
| ECPCS-HC | 4.8421 | ||
| EC-CMS | 4.6053 | ||
| ECAR | 5.5263 | ||
| CEAM | 5.2105 | ||
| ECM-RCL | 1.3684 |
| Methods | -Value | The Null Hypothesis |
|---|---|---|
| ECM-RCL | 0.000143 | rejected |
| ECM-RCL | 0.000168 | rejected |
| ECM-RCL | 0.000271 | rejected |
| ECM-RCL | 0.00058 | rejected |
| ECM-RCL | 0.000121 | rejected |
| ECM-RCL | 0.000673 | rejected |
| ECM-RCL | 0.000121 | rejected |
| Methods | -Value | The Null Hypothesis | |||
|---|---|---|---|---|---|
| 7 | K-means ECM-RCL | 5.927282 | 0 | 0.007143 | rejected |
| 6 | ECAR ECM-RCL | 5.231903 | 0 | 0.008333 | rejected |
| 5 | CEAM ECM-RCL | 4.834543 | 0.000001 | 0.01 | rejected |
| 4 | ECPCS-HC ECM-RCL | 4.370957 | 0.000012 | 0.0125 | rejected |
| 3 | LWEA ECM-RCL | 4.138164 | 0.000035 | 0.0167 | rejected |
| 2 | EC-CMS ECM-RCL | 4.072937 | 0.000046 | 0.025 | rejected |
| 1 | DREC ECM-RCL | 2.947084 | 0.003208 | 0.05 | rejected |
| Methods | Indices | EECM-RCL | PECM-RCL | RECM-RCL | DECM-RCL | ECM-RCL | |
|---|---|---|---|---|---|---|---|
| Datasets | |||||||
| GLIMA | ACC | 0.6000 ± 0.0452 | 0.6080 ± 0.0551 | 0.6200 ± 0.0639 | 0.6320 ± 0.0567 | 0.6440 ± 0.0430 | |
| NMI | 0.5143 ± 0.0353 | 0.4986 ± 0.0380 | 0.5010 ± 0.0322 | 0.5171 ± 0.0183 | 0.5253 ± 0.0198 | ||
| ARI | 0.3763 ± 0.0563 | 0.3652 ± 0.0611 | 0.3601 ± 0.0366 | 0.3817 ± 0.0300 | 0.3854 ± 0.0311 | ||
| Heart | ACC | 0.6200 ± 0.0058 | 0.6189 ± 0.0064 | 0.6163 ± 0.0043 | 0.6222 ± 0.0000 | 0.6244 ± 0.0053 | |
| NMI | 0.0367 ± 0.0053 | 0.0375 ± 0.0033 | 0.0378 ± 0.0184 | 0.0408 ± 0.0000 | 0.0409 ± 0.0042 | ||
| ARI | 0.0539 ± 0.0059 | 0.0530 ± 0.0062 | 0.0505 ± 0.0041 | 0.0562 ± 0.0000 | 0.0584 ± 0.0052 | ||
| Iris | ACC | 0.9100 ± 0.0248 | 0.9080 ± 0.0208 | 0.9067 ± 0.0000 | 0.9067 ± 0.0000 | 0.9140 ± 0.0685 | |
| NMI | 0.8029 ± 0.0411 | 0.7985 ± 0.0305 | 0.7960 ± 0.0000 | 0.7960 ± 0.0000 | 0.8131 ± 0.0853 | ||
| ARI | 0.7787 ± 0.1101 | 0.7641 ± 0.0494 | 0.7592 ± 0.0000 | 0.7592 ± 0.0000 | 0.7975 ± 0.1174 | ||
| LM | ACC | 0.5122 ± 0.0096 | 0.5008 ± 0.0166 | 0.5169 ± 0.0070 | 0.5058 ± 0.0161 | 0.5200 ± 0.0140 | |
| NMI | 0.6412 ± 0.0057 | 0.6337 ± 0.0099 | 0.6403 ± 0.0055 | 0.6398 ± 0.0026 | 0.6460 ± 0.0082 | ||
| ARI | 0.3828 ± 0.0099 | 0.3705 ± 0.0162 | 0.3825 ± 0.0068 | 0.3697 ± 0.0180 | 0.3895 ± 0.0122 | ||
| Orlraws10p | ACC | 0.8470 ± 0.0432 | 0.8400 ± 0.0000 | 0.8430 ± 0.0067 | 0.8450 ± 0.0341 | 0.8470 ± 0.0408 | |
| NMI | 0.9113 ± 0.0310 | 0.9097 ± 0.0279 | 0.9172 ± 0.0159 | 0.9156 ± 0.0233 | 0.9207 ± 0.0158 | ||
| ARI | 0.8161 ± 0.0106 | 0.8148 ± 0.0502 | 0.8285 ± 0.0222 | 0.8208 ± 0.0452 | 0.8330 ± 0.0218 | ||
| WarpPIE10p | ACC | 0.4714 ± 0.0151 | 0.4671 ± 0.0163 | 0.4648 ± 0.0121 | 0.4648 ± 0.0181 | 0.4762 ± 0.0135 | |
| NMI | 0.5528 ± 0.0093 | 0.5512 ± 0.0141 | 0.5540 ± 0.0042 | 0.5512 ± 0.0083 | 0.5562 ± 0.0127 | ||
| ARI | 0.3132 ± 0.0159 | 0.3159 ± 0.0194 | 0.3121 ± 0.0112 | 0.3072 ± 0.0468 | 0.3181 ± 0.0129 | ||
| WDBC | ACC | 0.8896 ± 0.0190 | 0.8842 ± 0.0188 | 0.8989 ± 0.0124 | 0.8793 ± 0.0258 | 0.8903 ± 0.0191 | |
| NMI | 0.5026 ± 0.0420 | 0.4878 ± 0.0436 | 0.5218 ± 0.0481 | 0.4817 ± 0.0733 | 0.5045 ± 0.0441 | ||
| ARI | 0.6028 ± 0.0613 | 0.5862 ± 0.0597 | 0.6329 ± 0.0405 | 0.5706 ± 0.0825 | 0.6050 ± 0.0615 | ||
| Wine | ACC | 0.7247 ± 0.0000 | 0.7247 ± 0.0000 | 0.7253 ± 0.0018 | 0.7247 ± 0.0000 | 0.7247 ± 0.0000 | |
| NMI | 0.4155 ± 0.0146 | 0.4231 ± 0.0138 | 0.4210 ± 0.0171 | 0.4217 ± 0.0119 | 0.4225 ± 0.0116 | ||
| ARI | 0.4007 ± 0.0000 | 0.4007 ± 0.0000 | 0.4007 ± 0.0000 | 0.4007 ± 0.0000 | 0.4007 ± 0.0000 | ||
| Zoo | ACC | 0.7129 ± 0.0229 | 0.6960 ± 0.0959 | 0.7386 ± 0.0314 | 0.6366 ± 0.0247 | 0.7248 ± 0.0283 | |
| NMI | 0.7215 ± 0.0093 | 0.7015 ± 0.0201 | 0.7191 ± 0.0246 | 0.6217 ± 0.0137 | 0.7219 ± 0.0093 | ||
| ARI | 0.6214 ± 0.0732 | 0.6061 ± 0.0322 | 0.6220 ± 0.0410 | 0.4636 ± 0.0124 | 0.6282 ± 0.0154 | ||
| ALLAML | GLIOMA | Heart | Iris | Lymphoma | Orlraws10p | |
|---|---|---|---|---|---|---|
| 2nd | 0.7153 ± 0.0150 | 0.6440 ± 0.0430 | 0.6244 ± 0.0053 | 0.9140 ± 0.0685 | 0.6552 ± 0.0421 | 0.8470 ± 0.0408 |
| 3rd | 0.7139 ± 0.0199 | 0.6440 ± 0.0741 | 0.6148 ± 0.0000 | 0.9220 ± 0.0340 | 0.6406 ± 0.0403 | 0.8560 ± 0.0425 |
| 4th | 0.7139 ± 0.0360 | 0.6160 ± 0.0540 | 0.6111 ± 0.0000 | 0.9220 ± 0.0340 | 0.6365 ± 0.0356 | 0.8500 ± 0.0406 |
| 5th | 0.7153 ± 0.0410 | 0.6240 ± 0.0523 | 0.6078 ± 0.0012 | 0.9213 ± 0.0345 | 0.6542 ± 0.0343 | 0.8500 ± 0.0406 |
| Methods | LWEA | DREC | ECPCS-HC | EC-CMS | ECAR | CEAM | ECM-RCL | |
|---|---|---|---|---|---|---|---|---|
| Datasets | ||||||||
| ALLAML | 0.0278 | 1.1962 | 0.0076 | 0.0437 | 5.3731 | 0.2442 | 0.0124 | |
| CLL_SUB | 0.0023 | 0.0191 | 0.0019 | 0.0179 | 6.1449 | 0.2303 | 0.0317 | |
| GLIOMA | 0.0018 | 0.0248 | 0.0014 | 0.0014 | 4.7770 | 0.1304 | 0.0111 | |
| Heart | 0.0035 | 0.0264 | 0.0025 | 0.0381 | 4.6620 | 0.4803 | 0.1100 | |
| Iris | 0.0025 | 0.0201 | 0.0018 | 0.0183 | 4.3012 | 0.3162 | 0.1263 | |
| LM | 0.0055 | 0.0630 | 0.0052 | 0.0640 | 7.5051 | 0.7737 | 0.2029 | |
| Lymphoma | 0.0030 | 0.0544 | 0.0016 | 0.0080 | 5.9408 | 0.2820 | 0.1428 | |
| MF | 0.0293 | 0.1930 | 0.0942 | 6.1873 | 22.2389 | 9.7868 | 18.3646 | |
| Orlraws10P | 0.0028 | 0.0427 | 0.0023 | 0.0094 | 7.7261 | 0.2722 | 0.0563 | |
| USPS | 0.0278 | 0.3771 | 0.0770 | 4.9987 | 18.8030 | 9.6544 | 21.2204 | |
| Vertebral | 0.0045 | 0.0321 | 0.0042 | 0.0539 | 5.0715 | 0.5352 | 0.1206 | |
| WarpPIE10p | 0.0042 | 0.0534 | 0.0039 | 0.0247 | 7.0190 | 0.4849 | 0.0877 | |
| WDBC | 0.0057 | 0.0238 | 0.0111 | 0.4950 | 6.3375 | 1.8485 | 0.4830 | |
| Wine | 0.0024 | 0.0204 | 0.0019 | 0.0263 | 4.6208 | 0.3426 | 1.3352 | |
| Zoo | 0.0019 | 0.0368 | 0.0016 | 0.0128 | 4.7620 | 0.2212 | 0.0541 | |
| ISOLET | 0.2668 | 3.2181 | 2.0049 | 339.9318 | 316.5550 | 249.7655 | 1035.0218 | |
| LS | 0.2283 | 1.7801 | 1.1972 | 132.5164 | 100.9128 | 143.3467 | 872.7249 | |
| ODR | 0.1741 | 1.8478 | 0.8966 | 65.0205 | 95.9148 | 130.5054 | 448.3944 | |
| PD | 0.5057 | 2.2972 | 4.1880 | 732.8491 | 316.2745 | 442.8964 | 3118.2037 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Qu, J.; Dai, Q.; Bian, Z.; Zhou, J.; Jiang, Z. Ensemble Clustering Method via Robust Consensus Learning. Electronics 2025, 14, 4764. https://doi.org/10.3390/electronics14234764
Qu J, Dai Q, Bian Z, Zhou J, Jiang Z. Ensemble Clustering Method via Robust Consensus Learning. Electronics. 2025; 14(23):4764. https://doi.org/10.3390/electronics14234764
Chicago/Turabian StyleQu, Jia, Qidong Dai, Zekang Bian, Jie Zhou, and Zhibin Jiang. 2025. "Ensemble Clustering Method via Robust Consensus Learning" Electronics 14, no. 23: 4764. https://doi.org/10.3390/electronics14234764
APA StyleQu, J., Dai, Q., Bian, Z., Zhou, J., & Jiang, Z. (2025). Ensemble Clustering Method via Robust Consensus Learning. Electronics, 14(23), 4764. https://doi.org/10.3390/electronics14234764






