A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits
Abstract
:1. Introduction
Contributions of the Paper
2. Literature Review
3. Materials and Methods
3.1. Dataset
3.2. Annotation
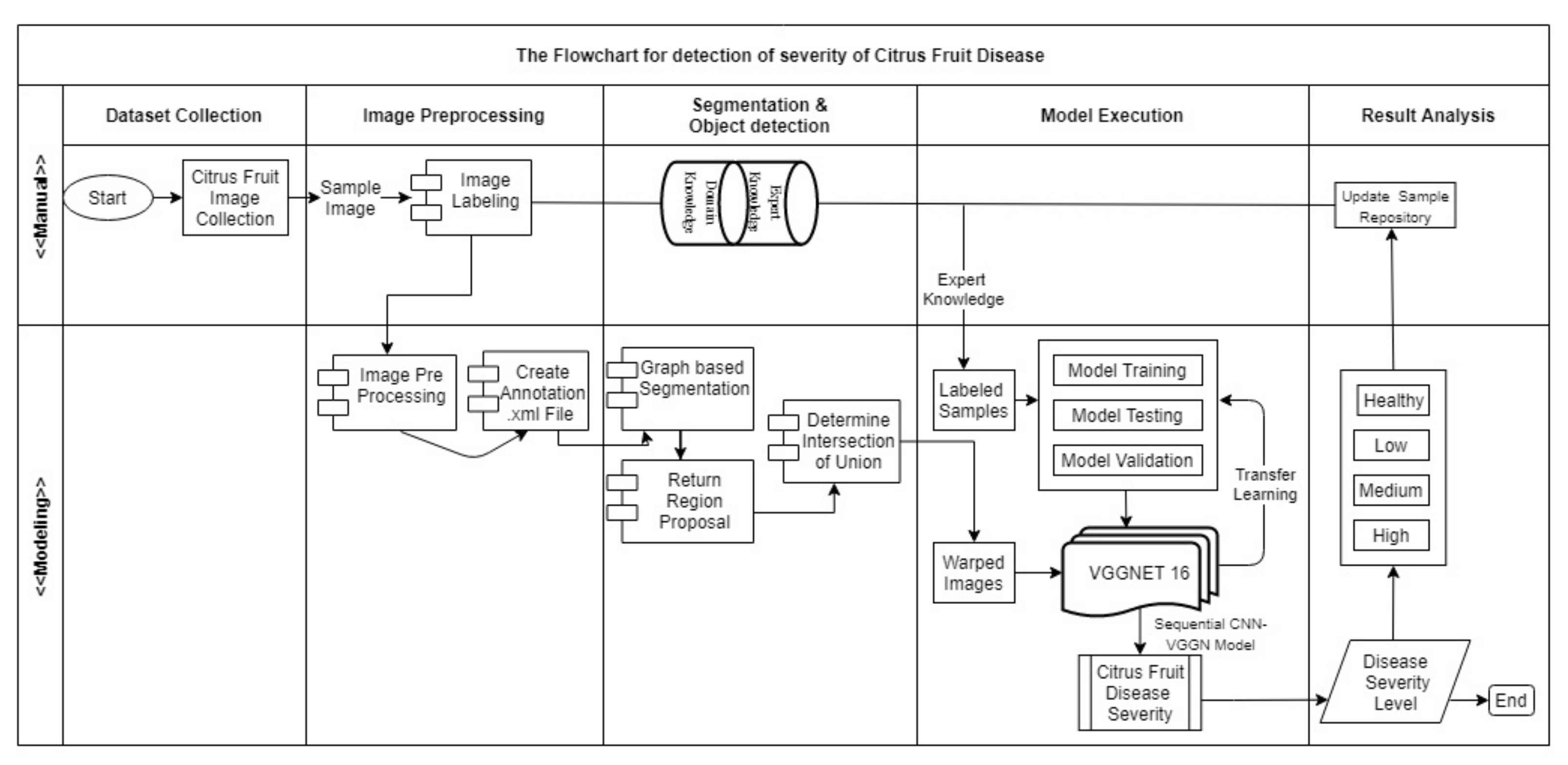
3.3. Proposed Algorithm for Detecting Severity Regions of the Citrus Diseases
- (1)
- Perform BoundingBox(Img) and annotate the image, i.e., Annotate(Img), where BoundingBox(Img) is used to create boundary coordinates on affected areas of the image and the Annotate(Img) function is used to create and extract the annotated image as an XML file for each image.
- (2)
- Create object for each category (i.e., healthy, low, medium and high).
- (3)
- Repeat step 5 for each object.
- (4)
- Repeat step 6 for each row of single object.
- (5)
- Extract Img_name and Img_url from object and perform preprocessing.
- (6)
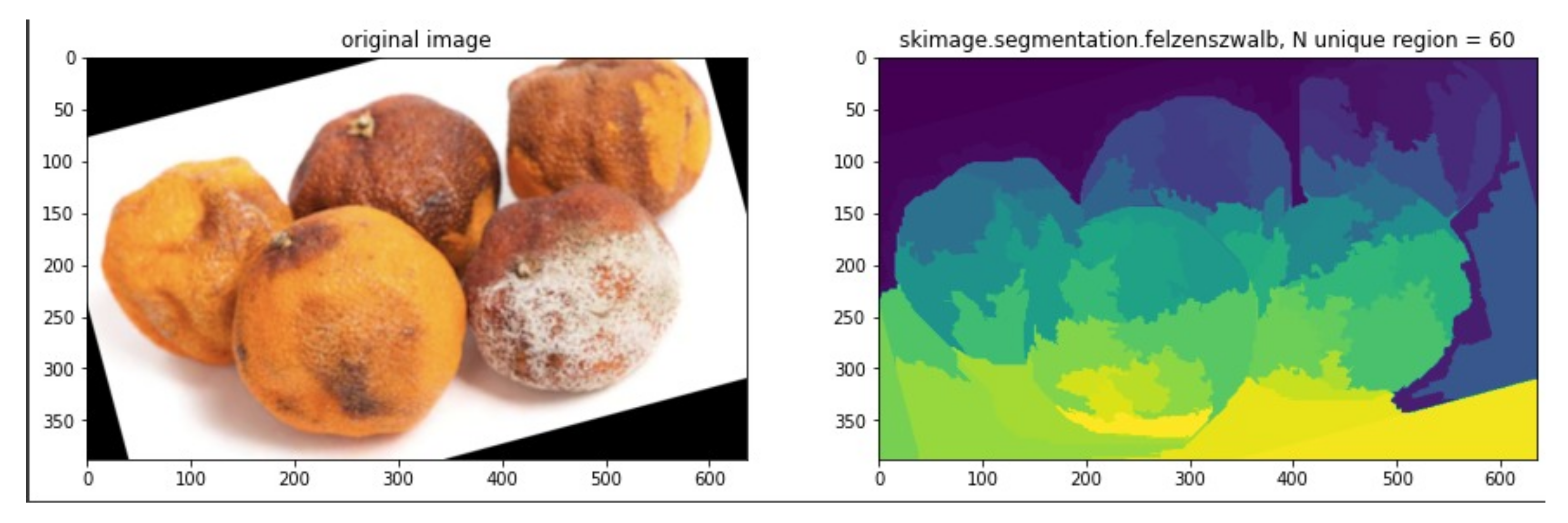
- Extract region using graph-based segmentation to determine the region proposal.
- (7)
- Repeat step 9-11 for each extracted segment region.
- (8)
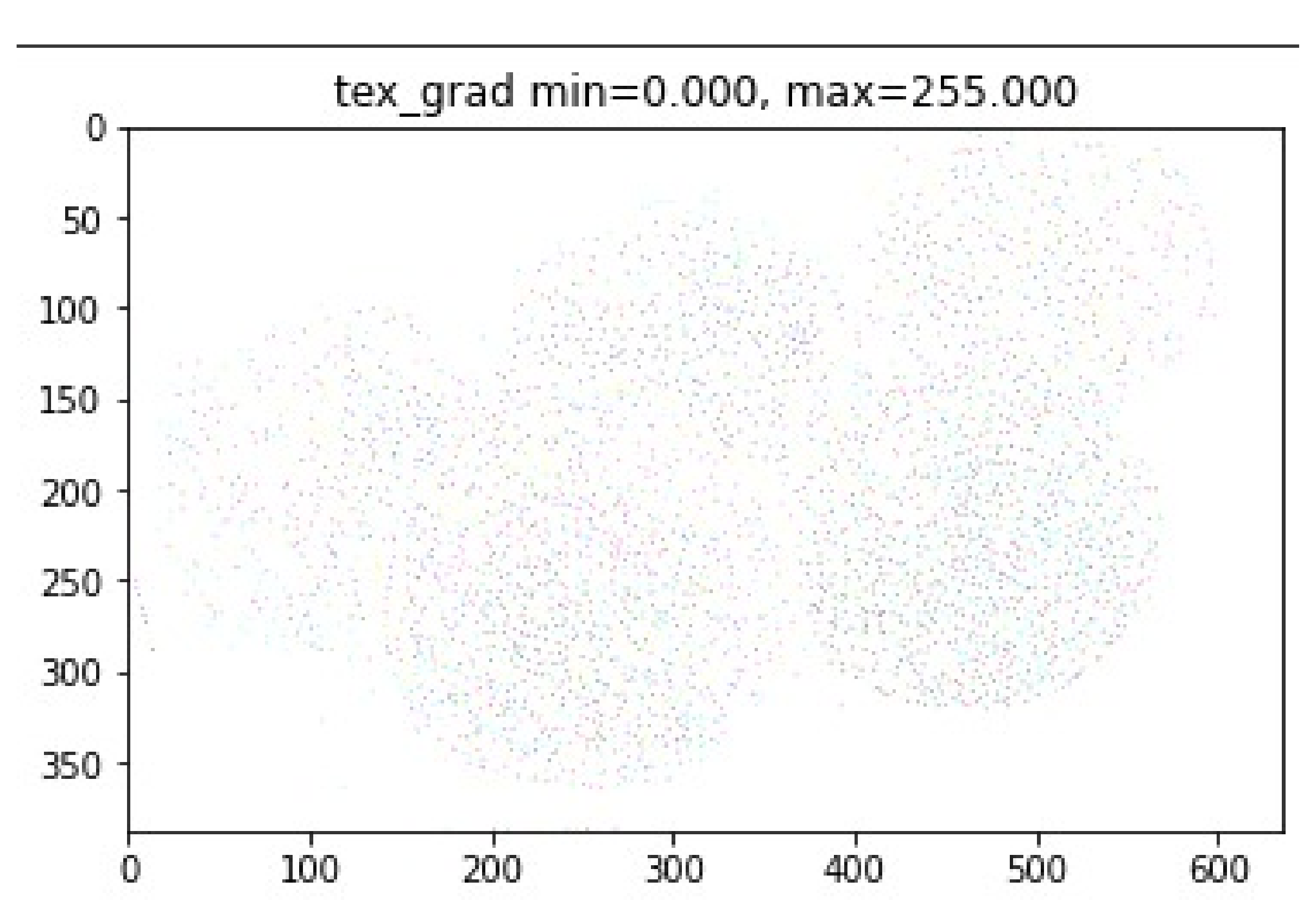
- Compute texture gradient of the image (using LBP).
- (9)
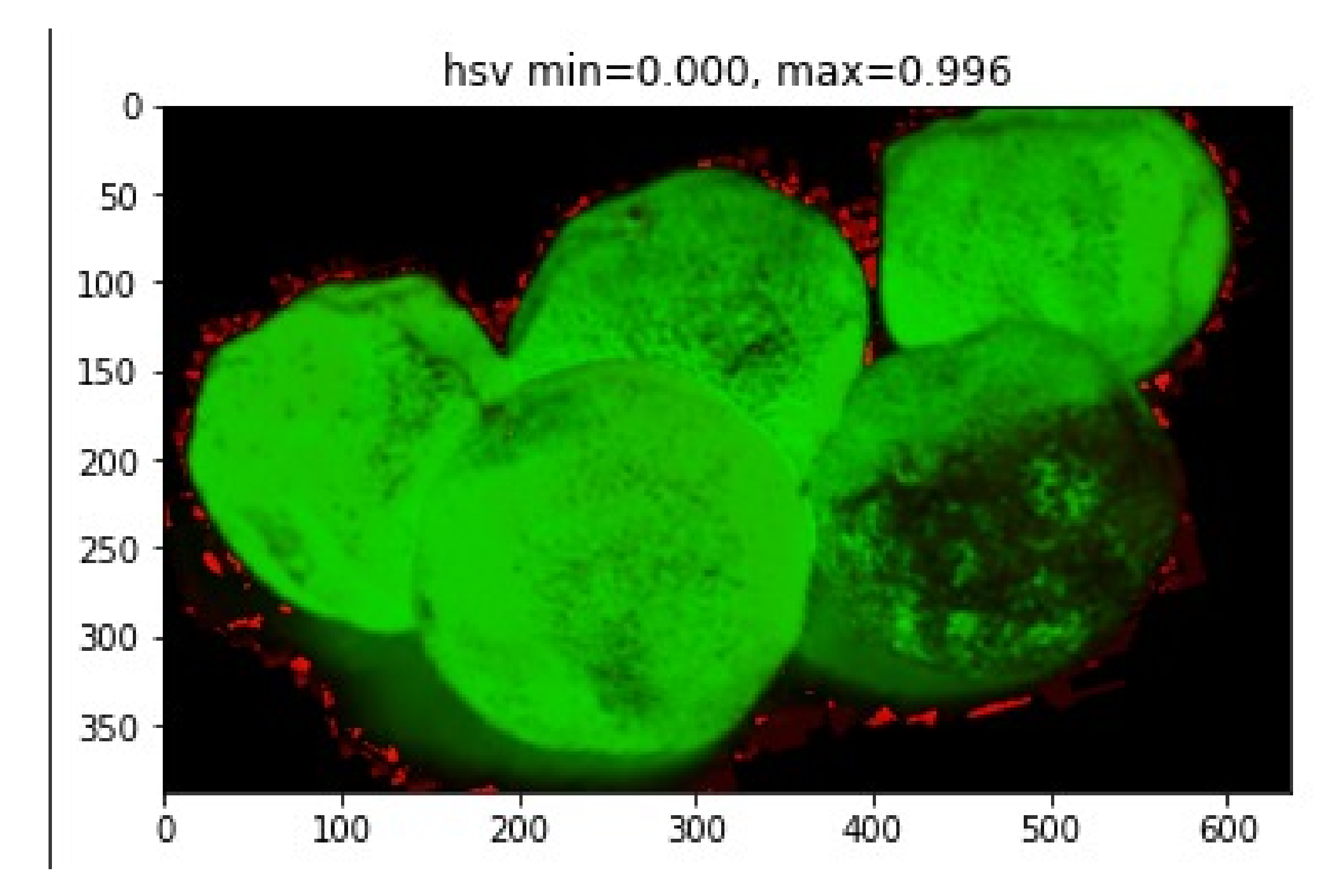
- Extract HSV for entire image using color histogram having COLOUR_CHANNELS (3)* bins with a total of 25 bins.
- (10)
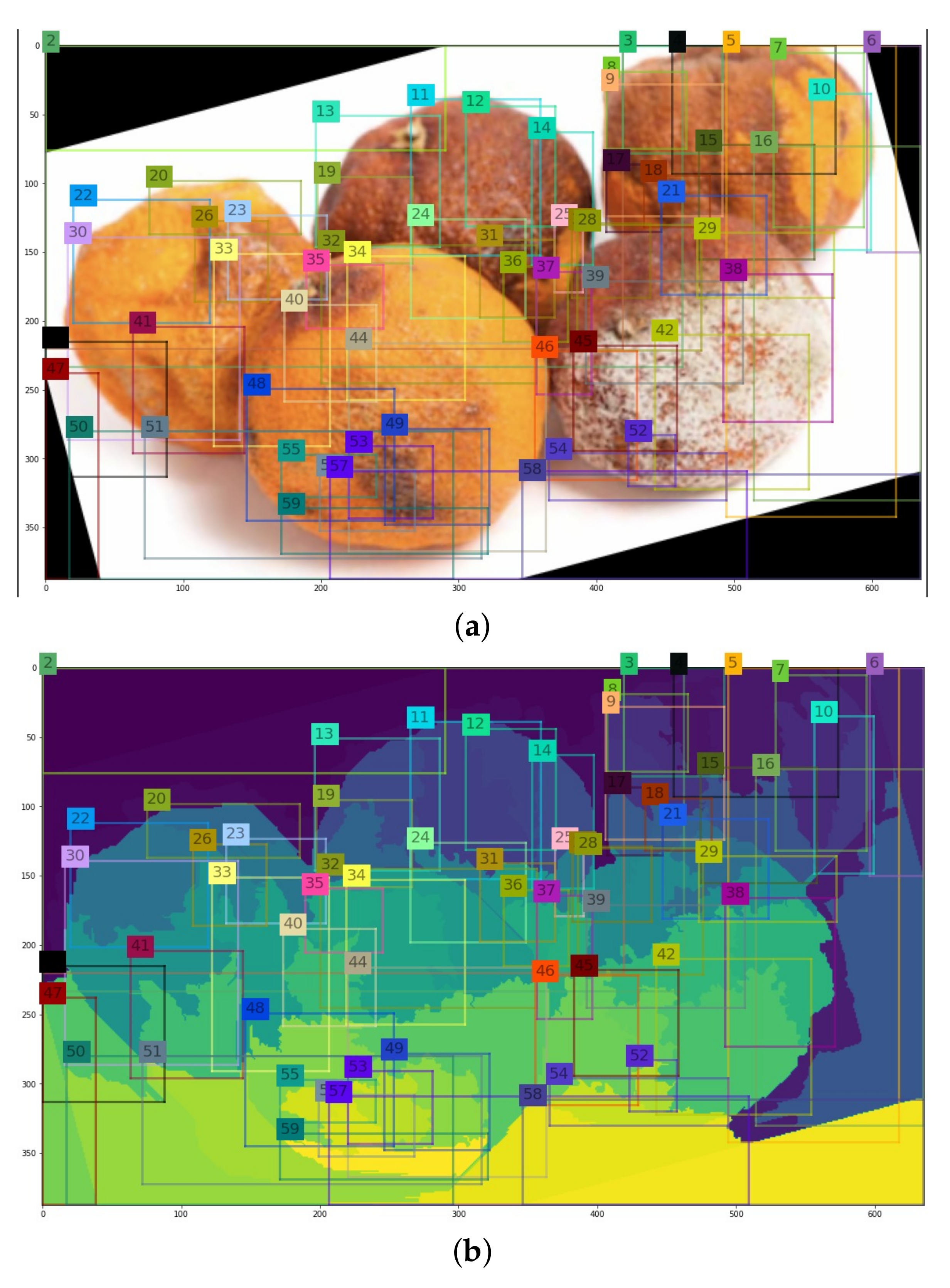
- Augment regions with histogram parameters and return region proposal.
- (11)
- Repeat step 13 and 14 for neighboring pair of regions .
- (12)
- (Compute similarity = colour similarity + texture similarity + + .
- (13)
- Merge regions, in order (, R).
- (14)
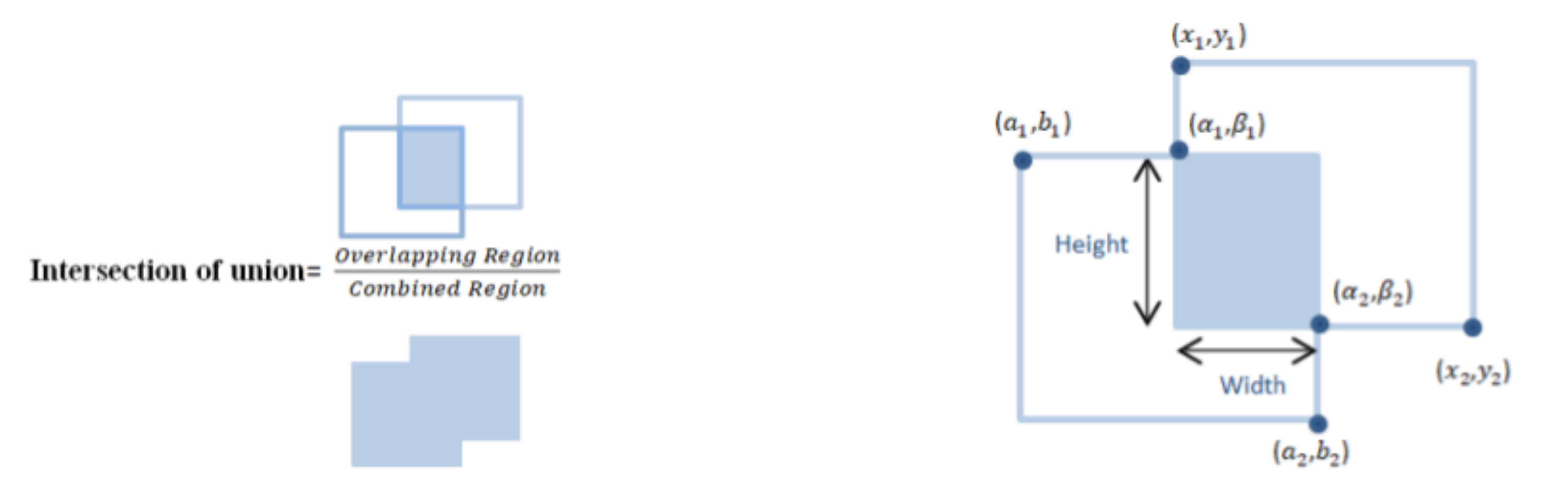
- Calculate IOU for regions.
3.4. Steps of Selective Search to Obtain the Region Proposal
- (1)
- Retrieve the pair of regions with the highest degree of similarity from the similarity dictionary.
- (2)
- Merge the region pairs and add them to the dictionary of regions.
- (3)
- Eliminate all pairs of regions from the similarity dictionary in which one of the regions is defined in step 1.
- (4)
- Determine the degree of similarity between the newly combined region and the regions and their intersecting regions (intersecting region is the region that is to be deleted).
3.5. Intersection of Union on Overlapped Region
3.6. Warp the Regions Proposed by the Selective Search
3.6.1. Feature Extraction
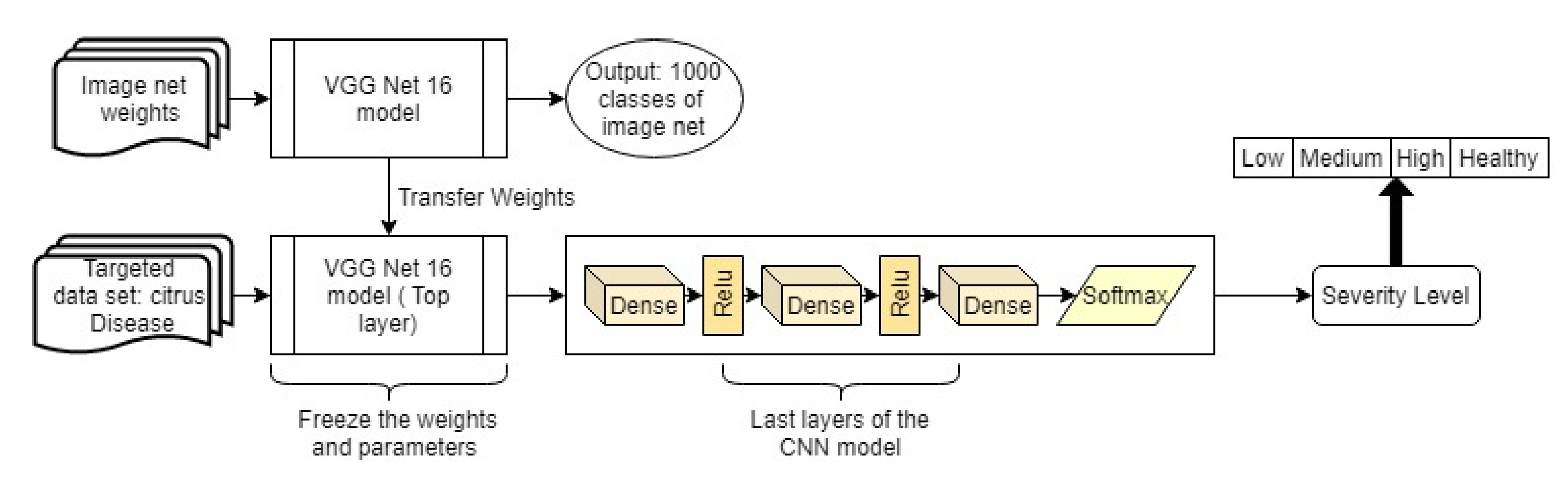
3.6.2. Transfer Learning
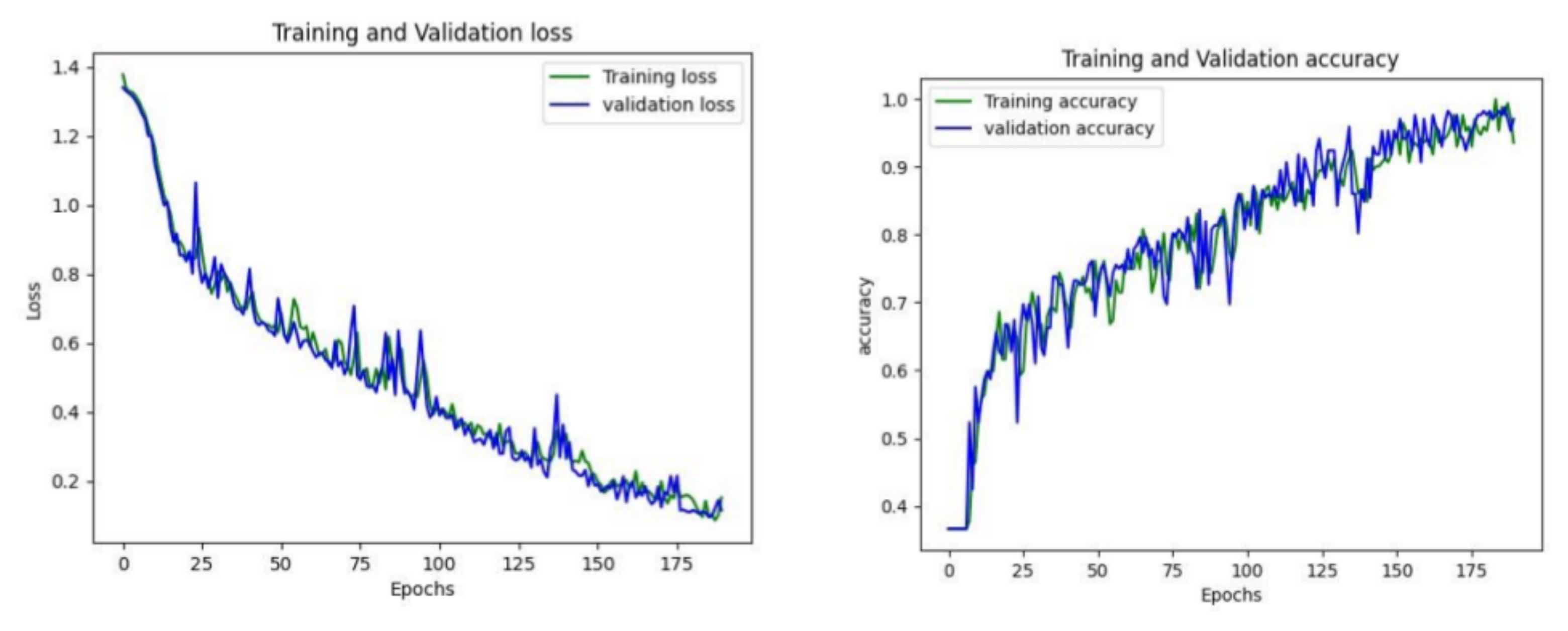
4. Result Analysis
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Food and Agriculture Organisation of the United States. Food and Agriculture Data. Available online: http://www.fao.org/faostat/en/#home (accessed on 15 July 2021).
- Cubero, S.; Lee, W.S.; Aleixos, N.; Albert, F.; Blasco, J. Automated Systems Based on Machine Vision for Inspecting Citrus Fruits from the Field to Postharvest-a Review. Food Bioprocess Technol. 2016, 9, 1623–1639. [Google Scholar] [CrossRef] [Green Version]
- Taylor, P. Critical Reviews in Plant Sciences Plant Disease Severity Estimated Visually, by Digital Photography and Image Analysis, and by Hyperspectral Imaging Plant Disease Severity Estimated Visually, by Digital Photography and Image Analysis, and by Hyperspe. CRC Crit. Rev. Plant Sci. 2010, 29, 37–41. [Google Scholar]
- Han, L.; Haleem, M.S.; Taylor, M. A Novel Computer Vision-based Approach to Automatic Detection and Severity Assessment of Crop Diseases. 2015. Available online: https://ieeexplore.ieee.org/document/7237209/ (accessed on 20 December 2021).
- Metin, M.; Adem, K. Automatic detection and classification of leaf spot disease in sugar beet using deep learning algorithms. Physica A 2019, 535, 122537. [Google Scholar]
- Lorente, D.; Escandell-Montero, P.; Cubero, S.; Gómez-Sanchis, J.; Blasco, J. Visible-NIR reflectance spectroscopy and manifold learning methods applied to the detection of fungal infections on citrus fruit. J. Food Eng. 2015, 163, 17–24. [Google Scholar] [CrossRef]
- Lan, Y. Comparison of machine learning methods for citrus greening detection on UAV multispectral images. Comput. Electron. Agric. 2020, 171, 105234. [Google Scholar] [CrossRef]
- Stegmayer, G.; Milone, D.H.; Garran, S.; Burdyn, L. Automatic recognition of quarantine citrus diseases. Expert Syst. Appl. 2013, 40, 3512–3517. [Google Scholar] [CrossRef] [Green Version]
- Jahanbakhshi, A.; Momeny, M.; Mahmoudi, M.; Zhang, Y.D. Classification of sour lemons based on apparent defects using stochastic pooling mechanism in deep convolutional neural networks. Sci. Hortic. 2020, 263, 109133. [Google Scholar] [CrossRef]
- Lopez, J.J.; Aguilera, E.; Cobos, M. Defect detection and classification in citrus using computer vision. In Proceedings of the International Conference on Neural Information Processing, Bangkok, Thailand, 1–5 December 2009; pp. 11–18. [Google Scholar]
- Lorente, D.; Aleixos, N.; Gómez-Sanchis, J.; Cubero, S.; Blasco, J. Selection of Optimal Wavelength Features for Decay Detection in Citrus Fruit Using the ROC Curve and Neural Networks. Food Bioprocess Technol. 2011, 6, 530–541. [Google Scholar] [CrossRef] [Green Version]
- Binary v1.8.1. Available online: https://github.com/tzutalin/labelImg/releases/tag/v1.8.1 (accessed on 20 December 2021).
- Behera, S.K.; Jena, L.; Rath, A.K.; Sethy, P.K. Disease Classification and Grading of Orange Using Machine Learning and Fuzzy Logic. In Proceedings of the IEEE International Conference on Communication and Signal Processing, Singapore, 28–30 September 2018; pp. 678–682. [Google Scholar] [CrossRef]
- Hafiz, T.R.; Basharat, A.S.; Ullah Lali, M.I.; Khan, A.; Sharif, M.; Chan Bukhari, S.A. A Citrus Fruits and Leaves Dataset for Detection and Classification of Citrus Diseases through Machine Learning. Available online: https://data.mendeley.com/datasets/3f83gxmv57/2 (accessed on 20 December 2021).
- Available online: https://www.kaggle.com/sriramr/fruits-fresh-and-rotten-for-classification (accessed on 20 December 2021).
- Yang, H.; Zhou, J.T.; Zhang, Y.; Gao, B.; Wu, J.; Cai, J. Exploit bounding box annotations for multi-label object recognition. In Proceedings of the IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Las Vegas, NV, USA, 27–30 June 2016; pp. 280–288. [Google Scholar]
- Garfinkel, S.L. Automating disk forensic processing with SleuthKit, XML and python. In Proceedings of the 2009 Fourth International IEEE Workshop on Systematic Approaches to Digital Forensic Engineering, Berkeley, CA, USA, 21 May 2009; pp. 73–84. [Google Scholar]
- Uijlings, A.W.; Sande, J.R.V.D.; Gevers, K.E.; Smeulders, T. Selective Search for Object Recognition. Int. J. Comput. Vis. 2012, 104, 154–171. [Google Scholar] [CrossRef] [Green Version]
- Ojala, T.; Pietikainen, M.; Maenpaa, T. Multiresolution Gray-Scale and Rotation Invariant Texture Classification with Local Binary Patterns. IEEE Trans. Pattern Anal. Mach. Intell. 2002, 24, 971–987. [Google Scholar] [CrossRef]
- Saravanan, G.; Yamuna, G.; Nandhini, S. Real time implementation of RGB to HSV/HSI/HSL and its reverse color space models. In Proceedings of the 2016 International Conference on Communication and Signal Processing (ICCSP), Melmaruvathur, India, 24 November 2016; pp. 462–466. [Google Scholar]
- Chen, J.W.; Lin, W.J.; Cheng, H.J.; Hung, C.L.; Lin, C.Y.; Chen, S.P. A smartphone-based application for scale pest detection using multiple-object detection methods. Electron 2021, 10, 1. [Google Scholar] [CrossRef]
- Jiang, B.; Luo, R.; Mao, J.; Xiao, T.; Jiang, Y. Acquisition of localization confidence for accurate object detection. In Proceedings of the European Conference on Computer Vision (ECCV), Munich, Germany, 8–14 September 2018; pp. 784–799. [Google Scholar]
- Girshick, R.; Donahue, J.; Darrell, T.; Malik, J.; Berkeley, U.C.; Malik, J. Rich feature hierarchies for accurate object detection and semantic segmentation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 23–28 June 2014; pp. 580–587. [Google Scholar]
- Wang, Q.; Qi, F.; Sun, M.; Qu, J.; Xue, J. Identification of Tomato Disease Types and Detection of Infected Areas Based on Deep Convolutional Neural Networks and Object Detection Techniques. Comput. Intell. Neurosci. 2019, 2019, 9142753. [Google Scholar] [CrossRef] [PubMed]
- Wimmer, G.; Vécsei, A.; Uhl, A. CNN transfer learning for the automated diagnosis of celiac disease. In Proceedings of the 2016 Sixth International Conference on Image Processing Theory, Tools and Applications (IPTA), Oulu, Finland, 12–15 December 2016. [Google Scholar] [CrossRef]
- Chen, J.; Chen, J.; Zhang, D.; Sun, Y.; Nanehkaran, Y.A. Using deep transfer learning for image-based plant disease identification. Comput. Electron. Agric 2020, 173, 105393. [Google Scholar] [CrossRef]
- Bock, S.; Weis, M. A Proof of Local Convergence for the Adam Optimizer. In Proceedings of the 2019 International Joint Conference on Neural Networks (IJCNN), Budapest, Hungary, 14–19 July 2019. [Google Scholar]

| Classes | Sample Count for Training | Sample Count for Testing |
|---|---|---|
| Healthy | 1173 | 293 |
| Low Severity | 737 | 184 |
| Middle Severity | 774 | 194 |
| High Severity | 625 | 156 |
| Layer | Layer Type | Kernel Size | Stride | Neuron Size | Maps | Param # |
|---|---|---|---|---|---|---|
| Block1_conv1 | Convolutional layer | 3 × 3 | 1 | 224 × 224 | 3 | 1792 |
| Block1_conv2 | Convolutional layer | 3 × 3 | 1 | 224 × 224 | 64 | 36,928 |
| Block1_pool | Pooling layer P1 | 2 × 2 | 2 | 112 × 112 | 64 | 0 |
| Block2_conv1 | Convolutional layer | 3 × 3 | 1 | 112 × 112 | 64 | 73,856 |
| Block2_conv2 | Convolutional layer C4 | 3 × 3 | 1 | 112 × 112 | 128 | 147,584 |
| Block2_pool | Pooling layer P2 | 2 × 2 | 2 | 56 × 56 | 128 | 0 |
| Block3_conv1 | Convolutional layer | 3 × 3 | 1 | 56 × 56 | 128 | 295,168 |
| Block3_conv2 | Convolutional layer | 3 × 3 | 1 | 56 × 56 | 256 | 590,080 |
| Block3_conv3 | Convolutional layer | 3 × 3 | 1 | 56 × 56 | 256 | 590,080 |
| Block3_pool | Pooling layer P3 | 2 × 2 | 2 | 28 × 28 | 256 | 0 |
| Block4_conv1 | Convolutional layer | 3 × 3 | 1 | 28 × 28 | 256 | 1,180,160 |
| Block4_conv2 | Convolutional layer | 3 × 3 | 1 | 28 × 28 | 512 | 23,598,038 |
| Block4_conv3 | Convolutional layer | 3 × 3 | 1 | 28 × 28 | 512 | 23,598,038 |
| Block4_pool | Pooling layer P4 | 2 × 2 | 2 | 14 × 14 | 512 | 0 |
| Block5_conv1 | Convolutional layer | 3 × 3 | 1 | 14 × 14 | 512 | 23,598,038 |
| Block5_conv2 | Convolutional layer | 3 × 3 | 1 | 14 × 14 | 512 | 23598038 |
| Block5_conv3 | Convolutional layer | 3 × 3 | 1 | 14 × 14 | 512 | 23,598,038 |
| Block5_pool | Pooling layer P5 | 2 × 2 | 2 | 7 × 7 | 512 | 0 |
| Flatten | Flatten | —– | —– | —— | 25,088 | 0 |
| Fc1 (Dense) | ——– | —– | —– | —— | 4096 | 102,764,544 |
| Dense (Dense) | Sequential CNN | —– | —– | —— | 32 | 131,104 |
| Dense_1 (Dense) | Sequential CNN | —— | —– | ——- | 32 | 1056 |
| Dense_2 (Dense) | Sequential CNN | —— | —– | —— | 4 | 132 |
| Output | Softmax | ——- | —— | Classifier | 4 | —- |
| Class | Healthy | Low | Medium | High |
|---|---|---|---|---|
| Healthy | 21 | 0 | 0 | 0 |
| Low | 0 | 25 | 0 | 1 |
| Medium | 3 | 0 | 25 | 0 |
| High | 1 | 0 | 0 | 24 |
| Class | Accuracy | Precision | Recall | F1 Score |
|---|---|---|---|---|
| Healthy | 96% | 100% | 84% | 91% |
| Low | 99% | 96% | 100% | 98% |
| Medium | 97% | 89% | 100% | 94% |
| High | 98% | 96% | 96% | 96% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dhiman, P.; Kukreja, V.; Manoharan, P.; Kaur, A.; Kamruzzaman, M.M.; Dhaou, I.B.; Iwendi, C. A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits. Electronics 2022, 11, 495. https://doi.org/10.3390/electronics11030495
Dhiman P, Kukreja V, Manoharan P, Kaur A, Kamruzzaman MM, Dhaou IB, Iwendi C. A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits. Electronics. 2022; 11(3):495. https://doi.org/10.3390/electronics11030495
Chicago/Turabian StyleDhiman, Poonam, Vinay Kukreja, Poongodi Manoharan, Amandeep Kaur, M. M. Kamruzzaman, Imed Ben Dhaou, and Celestine Iwendi. 2022. "A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits" Electronics 11, no. 3: 495. https://doi.org/10.3390/electronics11030495
APA StyleDhiman, P., Kukreja, V., Manoharan, P., Kaur, A., Kamruzzaman, M. M., Dhaou, I. B., & Iwendi, C. (2022). A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits. Electronics, 11(3), 495. https://doi.org/10.3390/electronics11030495










