Recombinase Polymerase Amplification Combined with Lateral Flow Strip for Rapid Detection of OXA-48-like Carbapenemase Genes in Enterobacterales
Abstract
1. Introduction
2. Results
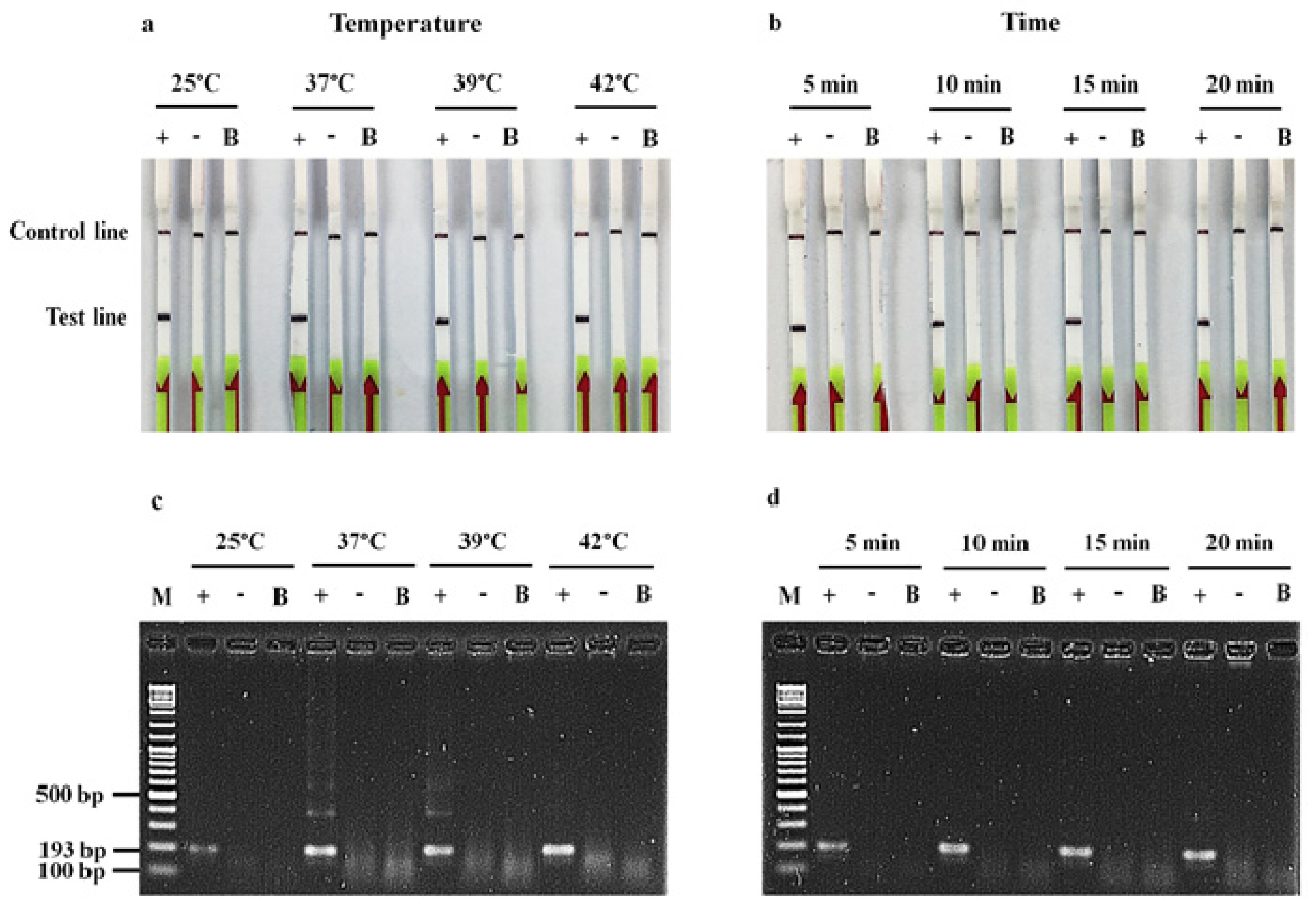
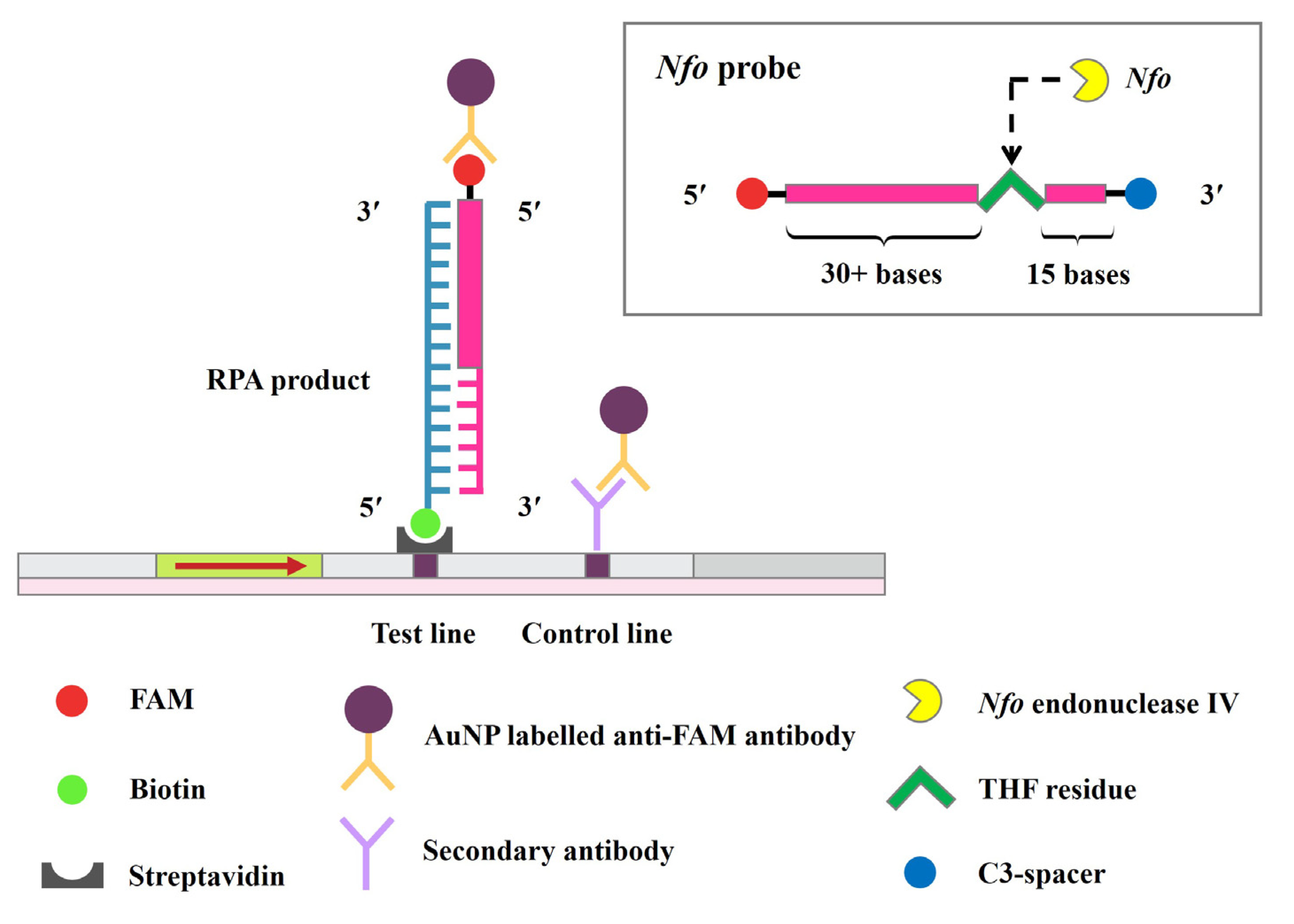
2.1. Detection of blaOXA-48-like by RPA-LF Assay
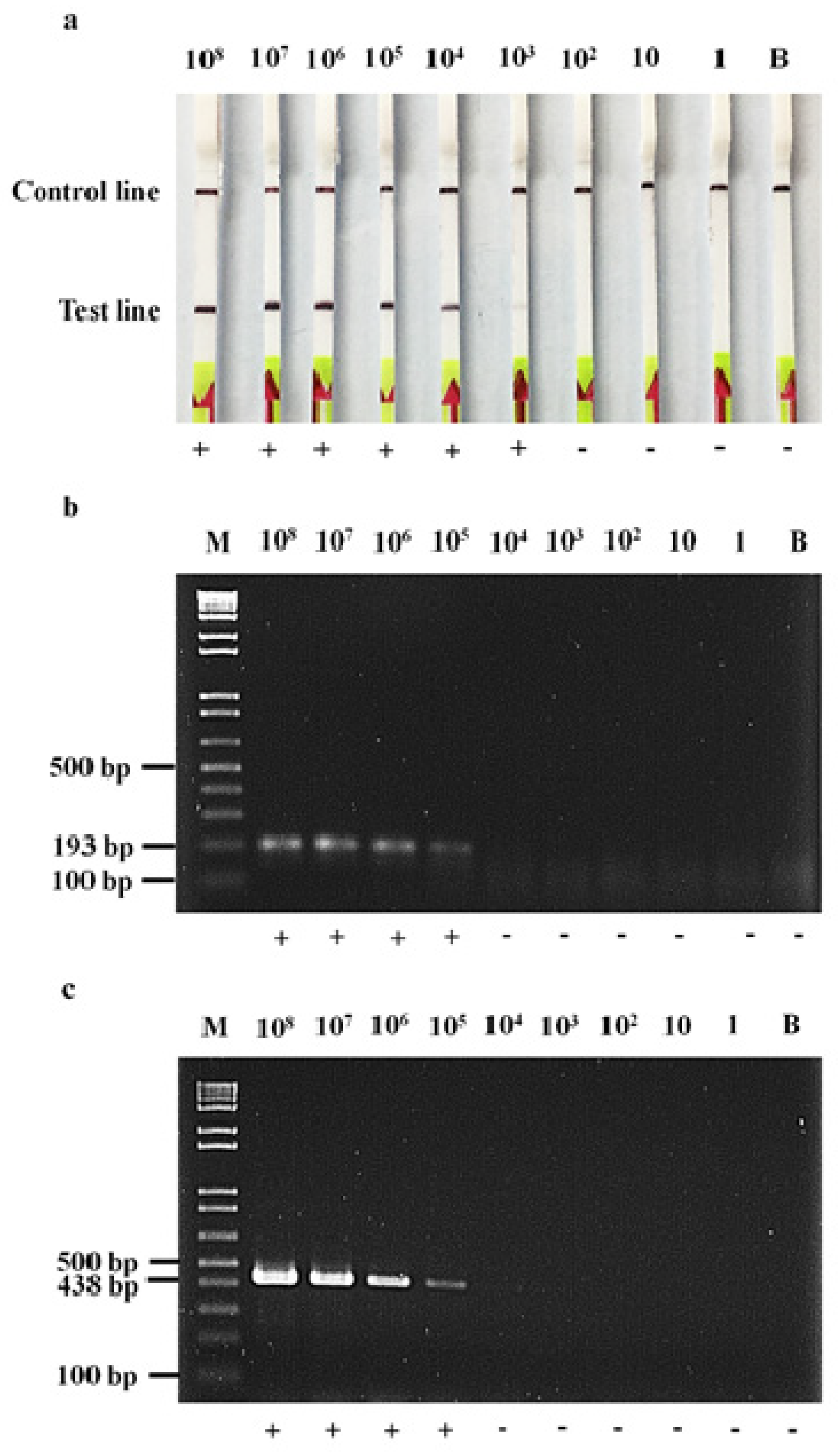
2.2. Detection Limit of RPA-LF, RPA-AGE and PCR-AGE Assays
2.3. Assessment of RPA-LF Assay
3. Discussion
4. Materials and Methods
4.1. Bacterial Isolates
4.2. Genomic DNA Extraction
4.3. Conventional PCR Method
4.4. RPA Primer and Probe Design
4.5. RPA-LF and RPA-AGE Assays
4.6. Detection Limit of RPA-LF, RPA-AGE and PCR-AGE Assays
4.7. Assessment of RPA-LF Assay
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- CDC. Diseases and organisms in healthcare settings. Available online: https://www.cdc.gov/hai/organisms/organisms.html (accessed on 25 March 2021).
- Codjoe, F.S.; Donkor, E.S. Carbapenem resistance: A review. Med. Sci. 2018, 6, 1. [Google Scholar] [CrossRef]
- Doi, Y. Treatment options for carbapenem-resistant Gram-negative bacterial infections. Clin. Infect. Dis. 2019, 69, 565–575. [Google Scholar] [CrossRef]
- Nordmann, P.; Poirel, L. Epidemiology and diagnostics of carbapenem resistance in Gram-negative bacteria. Clin. Infect. Dis. 2019, 69, S521–S528. [Google Scholar] [CrossRef]
- Jean, S.-S.; Harnod, D.; Hsueh, P.-R. Global threat of carbapenem-resistant Gram-negative bacteria. Front. Cell Infect. Microbiol. 2022, 12, 823684. [Google Scholar] [CrossRef]
- Andrade, F.F.; Silva, D.; Rodrigues, A.; Pina-Vaz, C. Colistin update on its mechanism of action and resistance, present and future challenges. Microorganisms 2020, 8, 1716. [Google Scholar] [CrossRef]
- Pogue, J.M.; Bonomo, R.A.; Kaye, K.S. Ceftazidime/avibactam, meropenem/vaborbactam, or both? Clinical and formulary considerations. Clin. Infect. Dis. 2019, 68, 519–524. [Google Scholar] [CrossRef]
- Tamma, P.D.; Aitken, S.L.; Bonomo, R.A.; Mathers, A.J.; van Duin, D.; Clancy, C.J. Infectious Diseases Society of America Guidance on the treatment of extended-spectrum β-lactamase producing Enterobacterales (ESBL-E), carbapenem-resistant Enterobacterales (CRE), and Pseudomonas aeruginosa with difficult-to-treat resistance (DTR-P. aeruginosa). Clin. Infect. Dis. 2021, 72, e169. [Google Scholar]
- Pitout, J.D.D.; Peirano, G.; Kock, M.M.; Strydom, K.A.; Matsumura, Y. The global ascendency of OXA-48-type carbapenemases. Clin. Microbiol. Rev. 2020, 33, e00102-19. [Google Scholar] [CrossRef]
- Dabos, L.; Oueslati, S.; Bernabeu, S.; Bonnin, R.A.; Dortet, L.; Naas, T. To be or not to be an OXA-48 carbapenemase. Microorganisms. 2022, 10, 258. [Google Scholar] [CrossRef]
- Srisrattakarn, A.; Lulitanond, A.; Charoensri, N.; Wonglakorn, L.; Kenprom, S.; Sukkasem, C.; Kuwatjanakul, W.; Piyapatthanakul, S.; Luanphairin, O.; Phukaw, W.; et al. Multi evaluation of modified GoldNano Carb test for carbapenemase detection in clinical isolates of Gram-negative bacilli. Antibiotics. 2022, 11, 684. [Google Scholar] [CrossRef]
- Clinical and Laboratory Standards Institute (CLSI). Performance Standards for Antimicrobial Susceptibility Testing, 31st ed.; CLSI Guideline M100; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2021. [Google Scholar]
- Gallagher, L.C.; Roundtree, S.S.; Lancaster, D.P.; Rudin, S.D.; Bard, J.D.; Roberts, A.L.; Marshall, S.H.; Bonomo, R.A.; Sullivan, K.V. Performance of the CLSI Carba NP and the Rosco Carb Screen assays using North American carbapenemase-producing Enterobacteriaceae and Pseudomonas aeruginosa isolates. J. Clin. Microbiol. 2015, 53, 3370–3373. [Google Scholar] [CrossRef] [PubMed]
- Pierce, V.M.; Simner, P.J.; Lonsway, D.R.; Roe-Carpenter, D.E.; Johnson, J.K.; Brasso, W.B.; Bobenchik, A.M.; Lockett, Z.C.; Charnot-Katsikas, A.; Ferraro, M.J.; et al. Modified carbapenem inactivation method for phenotypic detection of carbapenemase production among Enterobacteriaceae. J. Clin. Microbiol. 2017, 55, 2321–2333. [Google Scholar] [CrossRef]
- Gelmez, G.A.; Can, B.; Hasdemir, U.; Soyletir, G. Evaluation of phenotypic tests for detection of carbapenemases: New modifications with new interpretation. J. Infect. Chemother. 2021, 27, 226–231. [Google Scholar] [CrossRef]
- Dortet, L.; Jousset, A.; Sainte-Rose, V.; Cuzon, G.; Naas, T. Prospective evaluation of the OXA-48 K-SeT assay, an immunochromatographic test for the rapid detection of OXA-48-type carbapenemases. J. Antimicrob. Chemother. 2016, 71, 1834–1840. [Google Scholar] [CrossRef] [PubMed]
- Fauconnier, C.; Dodemont, M.; Depouhon, A.; Anantharajah, A.; Verroken, A.; Rodriguez-Villalobos, H. Lateral flow immunochromatographic assay for rapid screening of faecal carriage of carbapenemase-producing Enterobacteriaceae. J. Antimicrob. Chemother. 2019, 1, 357–359. [Google Scholar] [CrossRef] [PubMed]
- Wareham, D.W.; Shah, R.; Betts, J.W.; Phee, L.M.; Momin, M.H.F.A. Evaluation of an immunochromatographic lateral flow assay (OXA-48 K-SeT) for rapid detection of OXA-48-like carbapenemases in Enterobacteriaceae. J. Clin. Microbiol. 2016, 54, 471–473. [Google Scholar] [CrossRef] [PubMed]
- Poirel, L.; Walsh, T.R.; Cuvillier, V.; Nordmann, P. Multiplex PCR for detection of acquired carbapenemase genes. Diagn. Microbiol. Infect. Dis. 2011, 70, 119–123. [Google Scholar] [CrossRef]
- Notomi, T.; Okayama, H.; Masubuchi, H.; Yonekawa, T.; Watanabe, K.; Amino, N.; Hase, T. Loop-mediated isothermal amplification of DNA. Nucleic Acids Res. 2000, 28, e63. [Google Scholar] [CrossRef]
- Vincent, M.; Xu, Y.; Kong, H. Helicase-dependent isothermal DNA amplification. EMBO Rep. 2004, 5, 795–800. [Google Scholar] [CrossRef]
- Piepenburg, O.; Williams, C.H.; Stemple, D.L.; Armes, N.A. DNA detection using recombination proteins. PLoS Biol. 2006, 4, e204. [Google Scholar] [CrossRef]
- Wharam, S.D.; Hall, M.J.; Wilson, W.H. Detection of virus mRNA within infected host cells using an isothermal nucleic acid amplification assay: Marine cyanophage gene expression within Synechococcus sp. Virol. J. 2007, 4, 52. [Google Scholar] [CrossRef]
- Daher, R.K.; Stewart, G.; Boissinot, M.; Bergeron, M.G. Recombinase polymerase amplification for diagnostic applications. Clin. Chem. 2016, 62, 947–958. [Google Scholar] [CrossRef]
- Tröger, V.; Niemann, K.; Gärtig, C.; Kuhlmeier, D. Isothermal amplification and quantification of nucleic acids and its use in microsystems. J. Nanomed. Nanotechnol. 2015, 6, 1000282. [Google Scholar]
- Xu, J.; Wang, X.; Yang, L.; Kan, B.; Lu, X. Rapid detection of mcr-1 by recombinase polymerase amplification. J. Med. Microbiol. 2018, 67, 1682–1688. [Google Scholar] [CrossRef]
- Nelson, M.M.; Waldron, C.L.; Bracht, J.R. Rapid molecular detection of macrolide resistance. BMC Infect. Dis. 2019, 19, 144. [Google Scholar] [CrossRef]
- Srisrattakarn, A.; Tippayawat, P.; Chanawong, A.; Tavichakorntrakool, R.; Daduang, J.; Wonglakorn, L.; Sooksongsoontorn, P.; Lulitanond, A. Direct detection of methicillin-resistant in Staphylococcus spp. in positive blood culture by isothermal recombinase polymerase amplification combined with lateral flow dipstick assay. World J. Microbiol. Biotechnol. 2020, 36, 162. [Google Scholar] [CrossRef]
- Kanokudom, S.; Assawakongkarat, T.; Akeda, Y.; Ratthawongjirakul, P.; Chuanchuen, R.; Chaichanawongsaroj, N. Rapid detection of extended spectrum β-lactamase producing Escherichia coli isolated from fresh pork meat and pig cecum samples using multiplex recombinase polymerase amplification and lateral flow strip analysis. PLoS ONE 2021, 16, e0248536. [Google Scholar] [CrossRef]
- Panpru, P.; Srisrattakarn, A.; Panthasri, N.; Tippayawat, P.; Chanawong, A.; Tavichakorntrakool, R.; Daduang, J.; Wonglakorn, L.; Lulitanond, A. Rapid detection of Enterococcus and vancomycin resistance using recombinase polymerase amplification. PeerJ 2021, 9, e12561. [Google Scholar] [CrossRef]
- Srisrattakarn, A.; Panpru, P.; Tippayawat, P.; Chanawong, A.; Tavichakorntrakool, R.; Daduang, J.; Wonglakorn, L.; Lulitanond, A. Rapid detection of methicillin-resistant Staphylococcus aureus in positive blood-cultures by recombinase polymerase amplification combined with lateral flow strip. PLoS ONE 2022, 17, e0270686. [Google Scholar] [CrossRef]
- Xu, G.; Hu, L.; Zhong, H.; Wang, H.; Yusa, S.I.; Weiss, T.C.; Romaniuk, P.J.; Pickerill, S.; You, Q. Cross priming amplification: Mechanism and optimization for isothermal DNA amplification. Sci. Rep. 2012, 2, 246. [Google Scholar] [CrossRef]
- Wang, Y.; Wang, Y.; Ma, A.J.; Li, D.X.; Luo, L.J.; Liu, D.X.; Jin, D.; Liu, K.; Ye, C.Y. Rapid and sensitive isothermal detection of nucleic-acid sequence by multiple cross displacement amplification. Sci. Rep. 2015, 5, 11902. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, R.; Srivastava, N.; Kumar, S.; Saritha, R.K.; Sharma, S.K.; Jain, R.K.; Baranwal, V.K. Development of a recombinase polymerase amplification assay for the diagnosis of banana bunchy top virus in different banana cultivars. Arch. Virol. 2017, 162, 2791–2796. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Macdonald, J.; von Stetten, F. Review: A comprehensive summary of a decade development of the recombinase polymerase amplification. Analyst 2019, 144, 31–67. [Google Scholar] [CrossRef] [PubMed]
- Sutton, M.; Turner, C.; Allen, C. Method for detecting carbapenemase-producing bacteria. WO2017/187178A1, 2 November 2017. Available online: https://patents.google.com/patent/WO2017187178A1/en (accessed on 29 September 2022).
- Kuşkucu, M.A.; Aygün, G.; Karakullukçu, A.; İmamova, N.; Küçükbasmacı, Ö.; Midilli, K. Development of a rapid molecular test format based on isothermal recombinase polymerase amplification technique and lateral flow method for detection of blaOXA-48. Klimik Derg. 2018, 31, 185–189, (English abstract). [Google Scholar] [CrossRef]
- Wang, F.; Wang, L.; Chen, H.; Li, N.; Wang, Y.; Li, Y.; Liang, W. Rapid detection of blaKPC, blaNDM, blaOXA-48-like and blaIMP carbapenemases in Enterobacterales using recombinase polymerase amplification combined with lateral flow strip. Front. Cell. Infect. Microbiol. 2021, 11, 772966. [Google Scholar] [CrossRef]
- Srisrattakarn, A.; Lulitanond, A.; Wilailuckana, C.; Charoensri, N.; Wonglakorn, L.; Saenjamla, P.; Chaimanee, P.; Daduang, J.; Chanawong, A. Rapid and simple identification of carbapenemase genes, blaNDM, blaOXA-48, blaVIM and blaIMP-14 groups, in Gram-negative bacilli by in-house loop-mediated isothermal amplification with hydroxynaphthol blue dye. World J. Microbiol. Biotechnol. 2017, 33, 130. [Google Scholar] [CrossRef]
- Peirano, G.; Matsumura, Y.; Nobrega, D.; Pitout, J.D.D. A cost-effective method for identifying Enterobacterales with OXA-181. J. Clin. Microbiol. 2019, 57, e01281-19. [Google Scholar] [CrossRef]
- Hu, J.; Huang, R.; Sun, Y.; Wei, X.; Wang, Y.; Jiang, C.; Geng, Y.; Sun, X.; Jing, J.; Gao, H.; et al. Sensitive and rapid visual detection of Salmonella Typhimurium in milk based on recombinase polymerase amplification with lateral flow dipsticks. J. Microbiol. Methods. 2019, 158, 25–32. [Google Scholar] [CrossRef]
- Ngudsuntia, A.; Lunha, K.; Lulitanond, A.; Tippayawat, P.; Sukkasem, C.; Charoensri, N.; Chanawong, A. Colistin susceptibility testing by rapid colistin disk elution test among Enterobacteriaceae in low-resource setting. Microb. Drug Resist. 2021, 12, 1685–1691. [Google Scholar] [CrossRef]
- Sanders, E.R. Aseptic laboratory techniques: Plating methods. J. Vis. Exp. 2012, 63, e3064. [Google Scholar] [CrossRef]
- Dortet, L.; Tandé, D.; de Briel, D.; Bernabeu, S.; Lasserre, C.; Gregorowicz, G.; Jousset, A.B.; Naas, T. MALDI-TOF for the rapid detection of carbapenemase-producing Enterobacteriaceae: Comparison of the commercialized MBT STAR®-Carba IVD Kit with two in-house MALDI-TOF techniques and the RAPIDEC® CARBA NP. J. Antimicrob. Chemother. 2018, 73, 2352–2359. [Google Scholar] [CrossRef]

| β-Lactamase Genes (n) | Species (n) | No. Isolates Tested by RPA-LF or PCR-AGE | |
|---|---|---|---|
| Positive | Negative | ||
| Carbapenemase genes (114) | |||
| Ambler class A (4) | |||
| blaKPC-2 (1) | K. pneumoniae ATCC BAA-1705 (1) | - | 1 |
| blaIMI-1 (2) blaIMI (1) | E. cloacae (2) E. cloacae (1) | - - | 2 1 |
| Ambler class B (32) | |||
| blaNDM (23) | E. coli (11) | - | 11 |
| K. pneumoniae (8) | - | 8 | |
| K. oxytoca (1) | - | 1 | |
| E. cloacae (1) | - | 1 | |
| C. freundii (2) | - | 2 | |
| blaIMP-14a (3) | P. aeruginosa (2) | - | 2 |
| K. pneumoniae (1) | - | 1 | |
| blaIMP (2) | P. aeruginosa (2) | - | 2 |
| blaVIM-2 (3) | P. aeruginosa (3) | - | 3 |
| blaVIM (1) | P. aeruginosa (1) | - | 1 |
| Ambler class D (76) | |||
| blaOXA-48-like (51) | K. pneumoniae (45) | 45 | - |
| E. coli (5) | 5 | - | |
| E. cloacae (1) | 1 | - | |
| blaOXA-48 (4) | K. pneumoniae (3) | 3 | - |
| E. coli (1) | 1 | - | |
| blaOXA-181 (5) | E. coli (3) | 3 | - |
| K. pneumoniae (1) | 1 | - | |
| E. cloacae (1) | 1 | - | |
| blaOXA-23-like (6) | A. baumannii (6) | - | 6 |
| blaOXA-23 (2) | A. baumannii (2) | - | 2 |
| blaOXA-72 (4) | A. baumannii (3) | - | 3 |
| A. pittii (1) | - | 1 | |
| blaOXA-58 (3) | A. baumannii (3) | - | 3 |
| blaOXA-23-like, blaOXA-58 (1) | A. baumannii (1) | - | 1 |
| Ambler class B and D (2) | |||
| blaNDM, blaOXA-48 (1) | K. pneumoniae (1) | 1 | - |
| blaNDM, blaOXA-23-like, blaOXA-58 (1) | A. baumannii (1) | - | 1 |
| Non-carbapenemase genes (17) | |||
| ESBL (9) blaCTX-M-15, blaTEM | E. coli (1) | - | 1 |
| blaCTX-M-1-like | K. pneumoniae (1) | - | 1 |
| blaCTX-M-1-like | E. cloacae complex (1) | - | 1 |
| blaVEB-like | E. coli (1) | - | 1 |
| blaSHV | E. cloacae complex (1) | - | 1 |
| blaCTX-M-9, blaSHV-12, blaVEB-like | K. pneumoniae (1) | - | 1 |
| blaCTX-M-1-like, blaSHV | K. pneumoniae (1) | - | 1 |
| blaCTX-M-1-like, blaVEB | E. coli (1) | - | 1 |
| blaVEB, blaSHV | K. pneumoniae (1) | - | 1 |
| pAmpC (2) | |||
| blaCMY-2 | E. coli (1) | - | 1 |
| blaDHA-1 | E. coli (1) | - | 1 |
| ESBL and pAmpC (4) | |||
| blaCTX-M-1-like, blaCMY-2 | E. coli (1) | - | 1 |
| blaCTX-M-15, blaCMY-2 | E. coli (1) | - | 1 |
| blaVEB-like, blaCMY-8b | E. coli (1) | - | 1 |
| blaVEB-like, blaMOX-2-like | E. coli (1) | - | 1 |
| Non-ESBL and non-pAmpC (2) | E. coli ATCC 25922 (1) | - | 1 |
| K. pneumoniae ATCC BAA-1706 (1) | - | 1 | |
| Total (131) | 61 | 70 | |
| Assays | Primers | Sequences (5′→3′) | Product Sizes (bp) | Reference |
|---|---|---|---|---|
| PCR-AGE | OXA-48-F | GCGTGGTTAAGGATGAACAC | 438 | [19] |
| OXA-48-R | CATCAAGTTCAACCCAACCG | |||
| RPA-AGE | OXA-48-F_RPA | GATTATCGGAATGCCWGCGGTAGCAAAGGAATG | 193 | This study |
| OXA-48-R_RPA | CAAGCTATTGGGAATTTTAAAGGTAGATGCGGG | This study | ||
| RPA-LF | OXA-48-R_RPA-LF | Biotin-CAAGCTATTGGGAATTTTAAAGGTAG ATGCGGG | This study | |
| OXA-48-probe | FAM-GTTGGAATGCTCACTTTACTGAACATA AAT-[THF]-ACAGGGCGTAGTTGT-C3-Spacer | This study |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hemwaranon, P.; Srisrattakarn, A.; Lulitanond, A.; Tippayawat, P.; Tavichakorntrakool, R.; Wonglakorn, L.; Daduang, J.; Chanawong, A. Recombinase Polymerase Amplification Combined with Lateral Flow Strip for Rapid Detection of OXA-48-like Carbapenemase Genes in Enterobacterales. Antibiotics 2022, 11, 1499. https://doi.org/10.3390/antibiotics11111499
Hemwaranon P, Srisrattakarn A, Lulitanond A, Tippayawat P, Tavichakorntrakool R, Wonglakorn L, Daduang J, Chanawong A. Recombinase Polymerase Amplification Combined with Lateral Flow Strip for Rapid Detection of OXA-48-like Carbapenemase Genes in Enterobacterales. Antibiotics. 2022; 11(11):1499. https://doi.org/10.3390/antibiotics11111499
Chicago/Turabian StyleHemwaranon, Phatsarawadee, Arpasiri Srisrattakarn, Aroonlug Lulitanond, Patcharaporn Tippayawat, Ratree Tavichakorntrakool, Lumyai Wonglakorn, Jureerut Daduang, and Aroonwadee Chanawong. 2022. "Recombinase Polymerase Amplification Combined with Lateral Flow Strip for Rapid Detection of OXA-48-like Carbapenemase Genes in Enterobacterales" Antibiotics 11, no. 11: 1499. https://doi.org/10.3390/antibiotics11111499
APA StyleHemwaranon, P., Srisrattakarn, A., Lulitanond, A., Tippayawat, P., Tavichakorntrakool, R., Wonglakorn, L., Daduang, J., & Chanawong, A. (2022). Recombinase Polymerase Amplification Combined with Lateral Flow Strip for Rapid Detection of OXA-48-like Carbapenemase Genes in Enterobacterales. Antibiotics, 11(11), 1499. https://doi.org/10.3390/antibiotics11111499






