Wild Animals Are Reservoirs and Sentinels of Staphylococcus aureus and MRSA Clones: A Problem with “One Health” Concern
Abstract
:1. Introduction
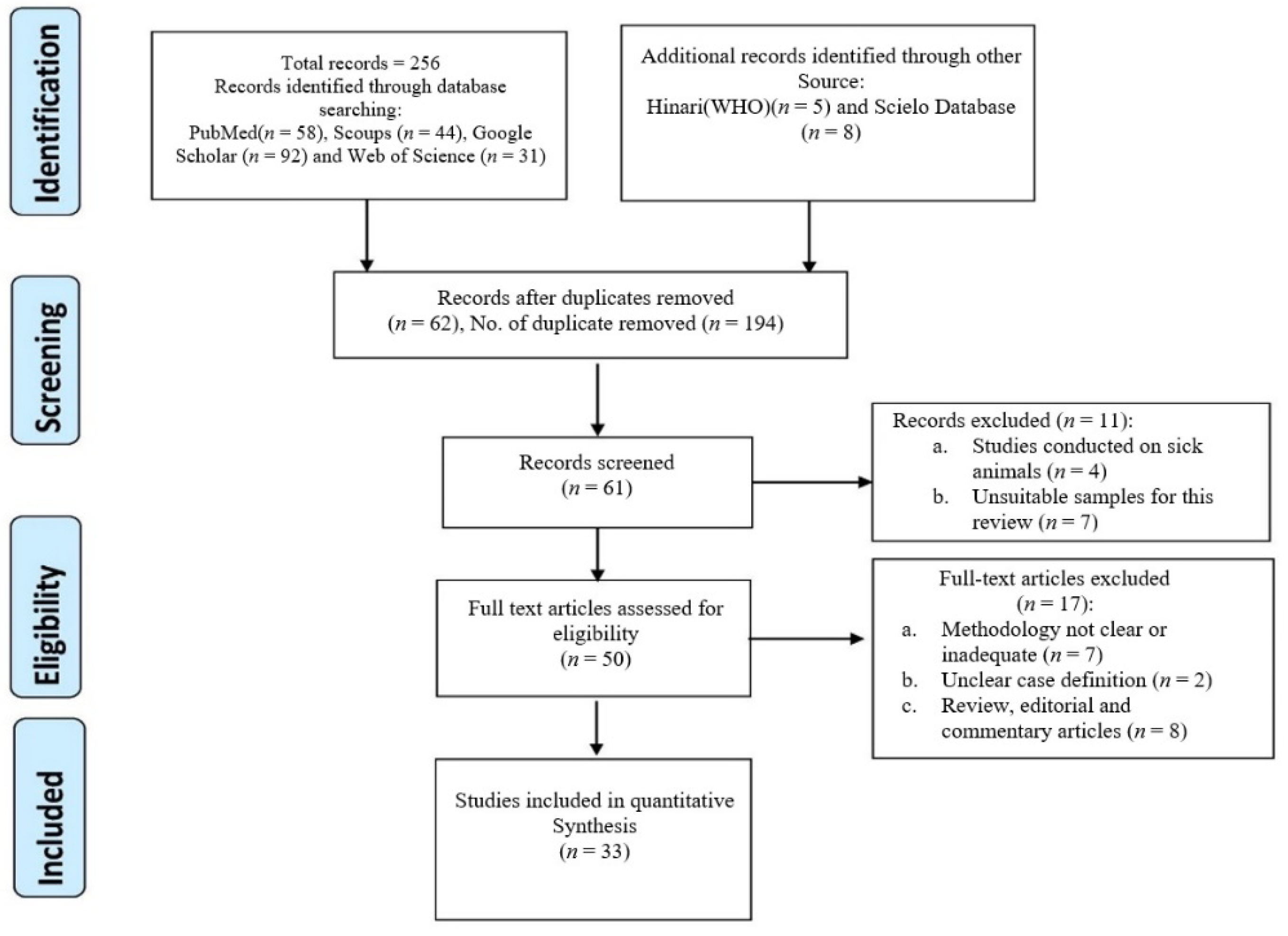
2. Methodology
2.1. Study Design
2.2. Articles Search Strategy
2.3. Inclusion Criteria
- Wild mammals (WM) are comprised of wild boars, red deer, Iberian ibex, deer, lynx, wild rabbits, hedgehogs, European mouflons, red foxes, common genets, bats, shrews and mustelids (otters, European badgers, beech martens, American minks and least weasels), among others. Rodents and primates were excluded from this group. This category of wild mammals has almost absolute confinement to the wildlife. However, it is hypothesised that these animals could contract MRSA from the predation of infected rodents and, in turn, spread them to humans who hunt wild animals. This is particularly possible in geographical locations with abundant forests and poor or no wildlife anti-poaching laws.
- Wild birds (WB) are comprised of storks, vultures and other birds that are naturally found in the wild.
- Non-human primates (NHP) are comprised of chimpanzees, monkeys, gorillas, lemurs and apes. These mammals have significant physiological and microbiota similarities to those found in humans.
- Wild rodents (WR) are comprised of mice and rats, both those confined to forests and those with proximity to human settlements and agricultural farms. These animals are also mammals (small) and originate from bushes or the wild. Importantly, it is expected that they could frequently relocate and transverse into human settlements, households, farms and vice versa. Hence, they are separated from other mammals.
2.4. Exclusion Criteria
2.5. Data Extraction
2.6. Statistical Analysis
3. Main Findings
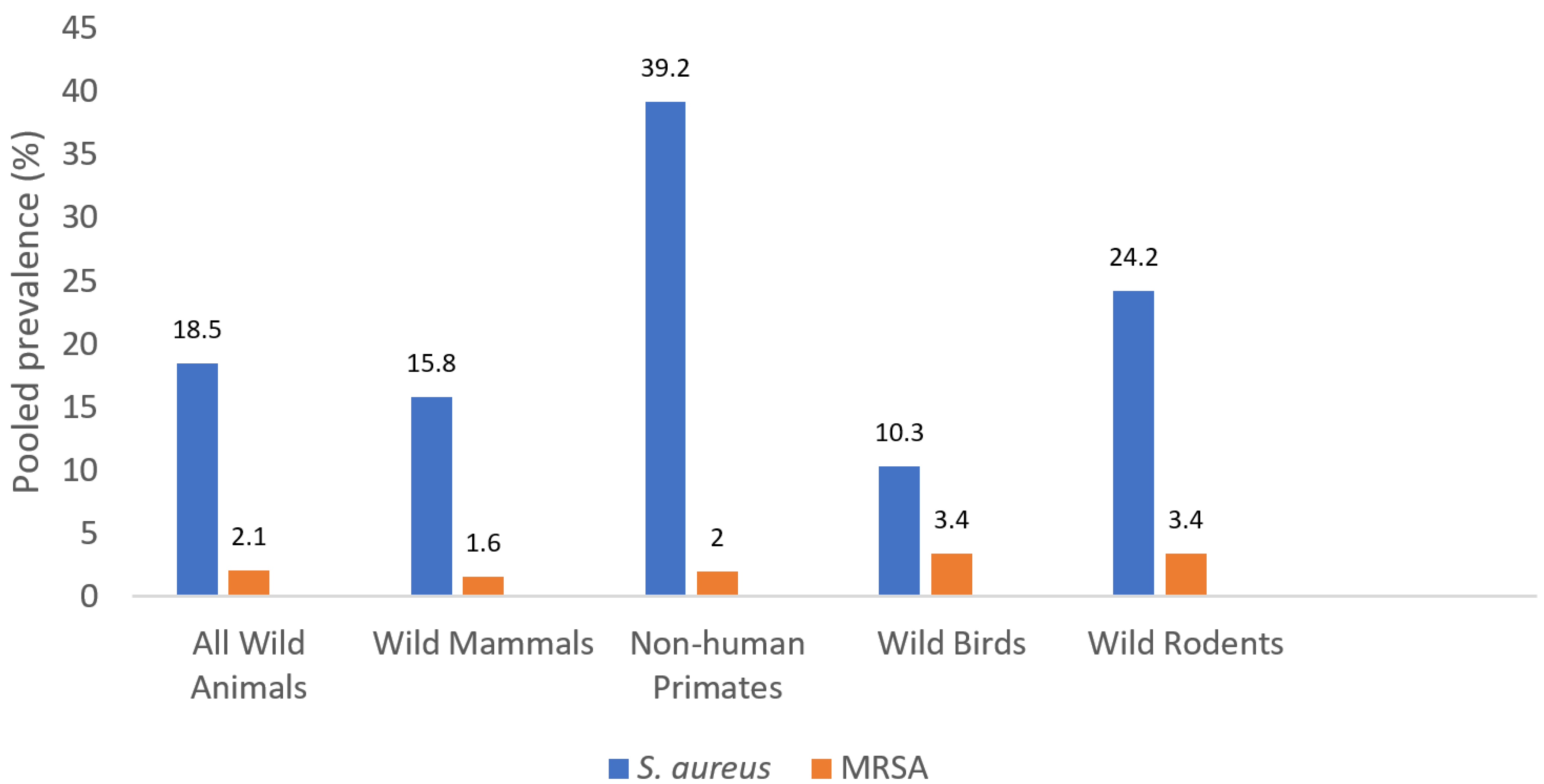
3.1. The Pooled Prevalence of S. aureus and MRSA Isolates
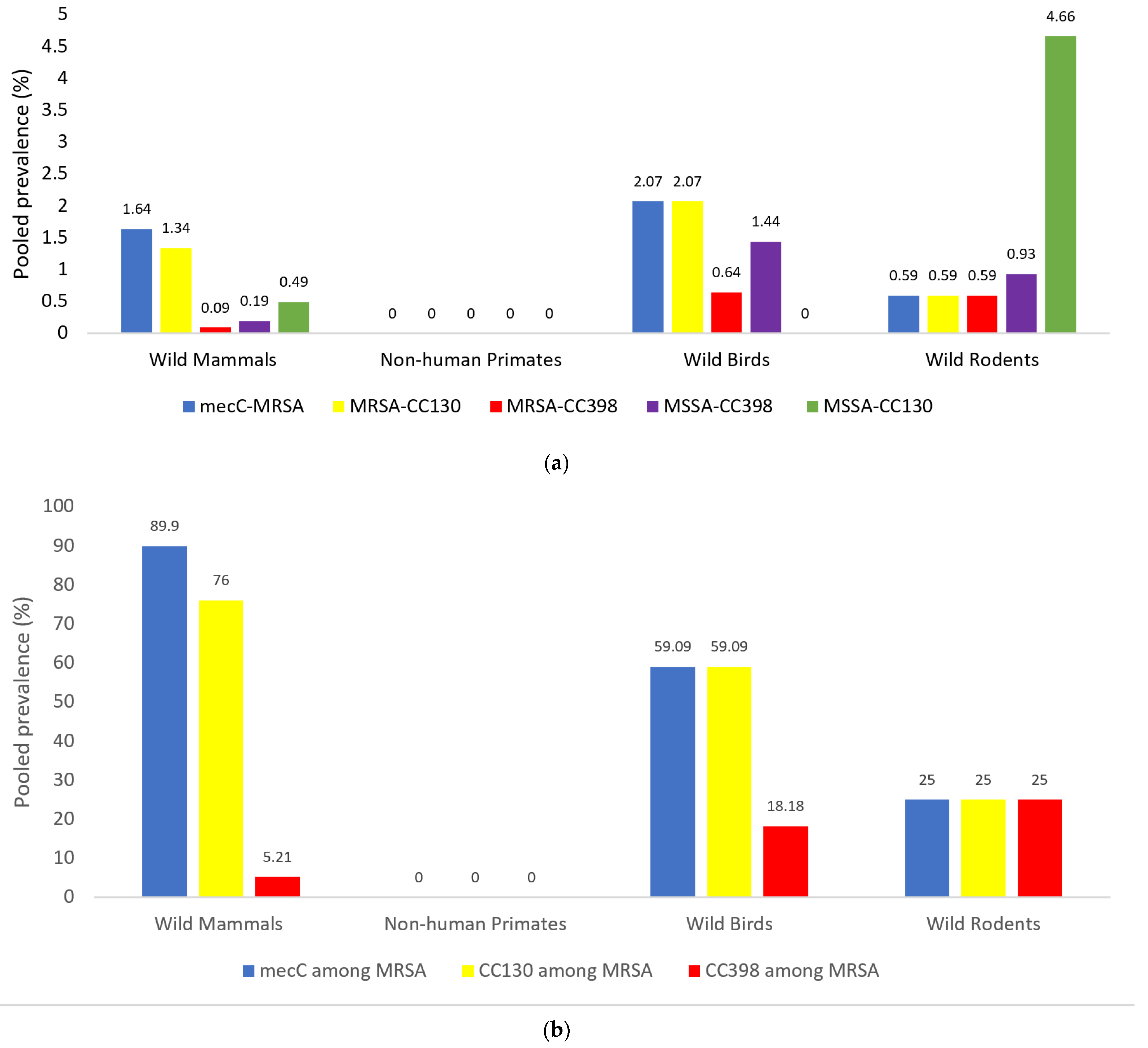
3.2. Prevalence of mecC-MRSA Isolates and Specific Genetic Lineages (CC398, CC130) in the Four Groups of Wild Animals
3.3. Characteristics of mecC MRSA Isolates from NTO Cavities of Wild Animals
3.4. Characteristics of S. aureus-CC398 Isolates Detected from NTO Cavities of Wild Animals
3.5. Characteristics of Other S. aureus Lineages Detected from NTO Cavities of Wild Animals
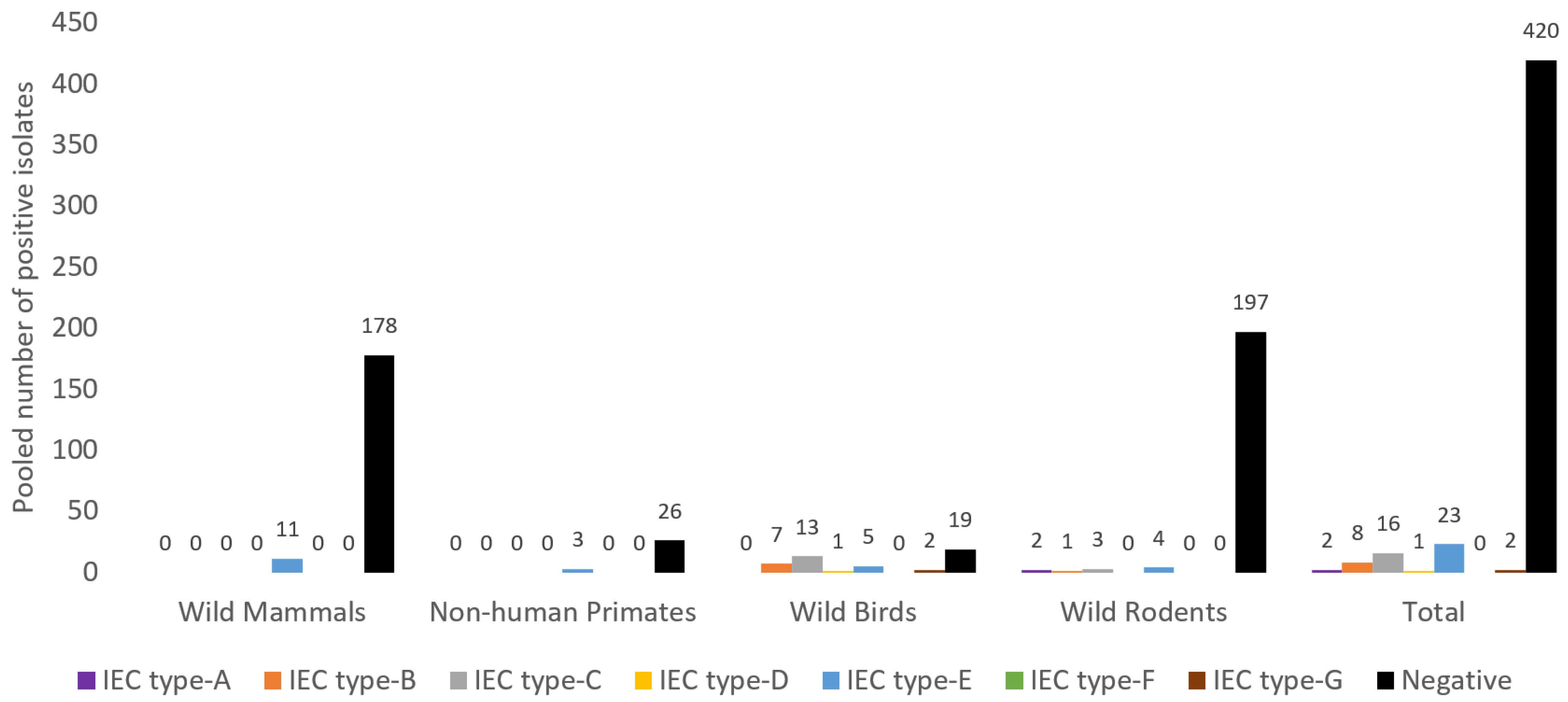
3.6. Antibiotic Resistance and Virulence Genes Detected from S. aureus of NTO Cavities of Wild Animals
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rossi, C.C.; Pereira, M.F.; Giambiagi-Demarval, M. Underrated Staphylococcus species and their role in antimicrobial resistance spreading. Genet. Mol. Biol. 2020, 43, e20190065. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Silveira, D.R.; De Moraes, T.P.; Kaefer, K.; Bach, L.G.; Barbosa, A.D.O.; Moretti, V.D.; De Menezes, P.Q.; De Medeiros, U.S.; Da Silva, T.T.; Bandarra, P.M.; et al. MRSA and enterobacteria of One Health concern in wild animals undergoing rehabilitation. Res. Soc. Dev. 2021, 10, e34810111809. [Google Scholar] [CrossRef]
- Swift, B.M.; Bennett, M.; Waller, K.; Dodd, C.; Murray, A.; Gomes, R.L.; Humphreys, B.; Hobman, J.L.; Jones, M.A.; Whitlock, S.E.; et al. Anthropogenic environmental drivers of antimicrobial resistance in wildlife. Sci. Total Environ. 2018, 649, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Penna, B.; Silva, M.B.; Soares, A.E.R.; Vasconcelos, A.T.R.; Ramundo, M.S.; Ferreira, F.A.; Silva-Carvalho, M.C.; de Sousa, V.S.; Rabello, R.F.; Bandeira, P.T.; et al. Comparative genomics of MRSA strains from human and canine origins reveals similar virulence gene repertoire. Sci. Rep. 2021, 11, 4724. [Google Scholar] [CrossRef]
- Porrero, M.C.; Mentaberre, G.; Sánchez, S.; Fernández-Llario, P.; Casas-Díaz, E.; Mateos, A.; Vidal, M.D.; Lavín, S.; Fernandez-Garayzabal, J.F.; Domínguez, L. Carriage of Staphylococcus aureus by free-living wild animals in Spain. Appl. Environ. Microbiol. 2014, 80, 4865–4870. [Google Scholar] [CrossRef] [Green Version]
- Rousham, E.K.; Unicomb, L.; Islam, M.A. Human, animal and environmental contributors to antibiotic resistance in low-resource settings: Integrating behavioural, epidemiological and One Health approaches. Proc. R. Soc. B Boil. Sci. 2018, 285, 20180332. [Google Scholar] [CrossRef]
- Becker, K. Methicillin-Resistant Staphylococci and Macrococci at the interface of human and animal health. Toxins 2021, 13, 61. [Google Scholar] [CrossRef]
- Dube, F.; Söderlund, R.; Salomonsson, M.L.; Troell, K.; Börjesson, S. Benzylpenicillin-producing Trichophyton erinacei and methicillin resistant Staphylococcus aureus carrying the mecC gene on European hedgehogs—A pilot-study. BMC Microbiol. 2021, 21, 212. [Google Scholar] [CrossRef]
- Zarazaga, M.; Gómez, P.; Ceballos, S.; Torres, C. Molecular epidemiology of Staphylococcus aureus lineages in the animal-human interface. In Staphylococcus aureus; Fetsch, A., Ed.; Academic Press: Cambridge, MA, USA, 2018; pp. 189–214. [Google Scholar]
- Gómez-Sanz, E.; Benito, D.; Lozano, C.; Gómez, P.; Ceballos, S.; López-Vazquez, M.; Zarazaga, M.; Torres, C. Factores de virulencia en Staphylococcus aureus. Tipos y caracterización genotípica. In Producción y Calidad de la Leche; Velázquez Ordóñez, V., Ed.; 2015; pp. 275–296. [Google Scholar]
- Gómez, P.; Ruiz-Ripa, L.; Fernández-Fernández, R.; Gharsa, H.; Ben, S.K.; Höfle, U.; Zarazaga, M.; Holmes, M.A.; Torres, C. Genomic analysis of Staphylococcus aureus of the lineage CC130, including mecC-carrying MRSA and MSSA isolate recovered of animal, human, and environmental origins. Front. Microbiol. 2021, 12, e655994. [Google Scholar] [CrossRef]
- Raafat, D.; Mrochen, D.M.; Al’Sholui, F.; Heuser, E.; Ryll, R.; Pritchett-Corning, K.R.; Jacob, J.; Walther, B.; Matuschka, F.-R.; Richter, D.; et al. Molecular Epidemiology of Methicillin-Susceptible and Methicillin-Resistant Staphylococcus aureus in Wild, Captive and Laboratory Rats: Effect of Habitat on the Nasal S. aureus Population. Toxins 2020, 12, 80. [Google Scholar] [CrossRef] [Green Version]
- García-Álvarez, L.; Holden, M.; Lindsay, H.; Webb, C.; Brown, D.F.; Curran, M.D.; Walpole, E.; Brooks, K.; Pickard, D.J.; Teale, C.; et al. Meticillin-resistant Staphylococcus aureus with a novel mecA homologue in human and bovine populations in the UK and Denmark: A descriptive study. Lancet Infect. Dis. 2011, 11, 595–603. [Google Scholar] [CrossRef] [Green Version]
- World Health Organization. Regional Office for South-East Asia. A Brief Guide to Emerging Infectious Diseases and Zoonoses. Available online: https://apps.who.int/iris/rest/bitstreams/909329/retrieve (accessed on 10 September 2021).
- Rahman, T.; Sobur, A.; Islam, S.; Ievy, S.; Hossain, J.; El Zowalaty, M.E.; Rahman, A.T.; Ashour, H.M. Zoonotic Diseases: Etiology, Impact, and Control. Microorganisms 2020, 8, 1405. [Google Scholar] [CrossRef]
- Morand, S.; McIntyre, K.M.; Baylis, M. Domesticated animals and human infectious diseases of zoonotic origins: Domestication time matters. Infect. Genet. Evol. 2014, 24, 76–81. [Google Scholar] [CrossRef] [Green Version]
- Silva, V.; Gabriel, S.; Borrego, S.; Tejedor-Junco, M.; Manageiro, V.; Ferreira, E.; Reis, L.; Caniça, M.; Capelo, J.; Igrejas, G.; et al. Antimicrobial Resistance and Genetic Lineages of Staphylococcus aureus from Wild Rodents: First Report of mecC-Positive Methicillin-Resistant S. aureus (MRSA) in Portugal. Animals 2021, 11, 1537. [Google Scholar] [CrossRef]
- Sousa, M.; Silva, N.; Manageiro, V.; Ramos, S.; Coelho, A.; Gonçalves, D.; Caniça, M.; Torres, C.; Igrejas, G.; Poeta, P. First report on MRSA CC398 recovered from wild boars in the north of Portugal. Are we facing a problem? Sci. Total Environ. 2017, 596–597, 26–31. [Google Scholar] [CrossRef]
- Gómez, P.; Lozano, C.; González-Barrio, D.; Zarazaga, M.; Ruiz-Fons, F.; Torres, C. High prevalence of methicillin-resistant Staphylococcus aureus (MRSA) carrying the mecC gene in a semi-extensive red deer (Cervus elaphus hispanicus) farm in Southern Spain. Vet. Microbiol. 2015, 177, 326–331. [Google Scholar] [CrossRef]
- Meemken, D.; Blaha, T.; Hotzel, H.; Strommenger, B.; Klein, G.; Ehricht, R.; Monecke, S.; Kehrenberg, C. Genotypic and Phenotypic Characterization of Staphylococcus aureus Isolates from Wild Boars. Appl. Environ. Microbiol. 2012, 79, 1739–1742. [Google Scholar] [CrossRef] [Green Version]
- Porrero, M.C.; Mentaberre, G.; Sánchez, S.; Fernández-Llario, P.; Gomez-Barrero, S.; Navarro-Gonzalez, N.; Serrano, E.; Casas-Díaz, E.; Marco, I.; Fernandez-Garayzabal, J.F.; et al. Methicillin resistant Staphylococcus aureus (MRSA) carriage in different free-living wild animal species in Spain. Vet. J. 2013, 198, 127–130. [Google Scholar] [CrossRef]
- Mrochen, D.M.; Schulz, D.; Fischer, S.; Jeske, K.; El Gohary, H.; Reil, D.; Imholt, C.; Trübe, P.; Suchomel, J.; Tricaud, E.; et al. Wild rodents and shrews are natural hosts of Staphylococcus aureus. Int. J. Med. Microbiol. 2018, 308, 590–597. [Google Scholar] [CrossRef]
- Ruiz-Ripa, L.; Alcalá, L.; Simón, C.; Gómez, P.; Mama, O.M.; Rezusta, A.; Zarazaga, M.; Torres, C. Diversity of Staphylococcus aureus clones in wild mammals in Aragon, Spain, with detection of MRSA ST130-mecC in wild rabbits. J. Appl. Microbiol. 2019, 127, 284–291. [Google Scholar] [CrossRef]
- Wardyn, S.E.; Kauffman, L.K.; Smith, T.C. Methicillin-resistant Staphylococcus aureus in Central Iowa Wildlife. J. Wildl. Dis. 2012, 48, 1069–1073. [Google Scholar] [CrossRef] [Green Version]
- Mama, O.M.; Ruiz-Ripa, L.; Fernández-Fernández, R.; González-Barrio, D.; Ruiz-Fons, F.; Torres, C. High frequency of coagulase-positive staphylococci carriage in healthy wild boar with detection of MRSA of lineage ST398-t011. FEMS Microbiol. Lett. 2019, 366, fny292. [Google Scholar] [CrossRef]
- Martínez, L.S.; Viana, D.; Arenas, J.M.C. Staphylococcus aureus nasal carriage could be a risk for development of clinical infections in rabbits. World Rabbit Sci. 2015, 23, 181. [Google Scholar] [CrossRef] [Green Version]
- García, L.A.; Torres, C.; López, A.R.; Rodríguez, C.O.; Espinosa, J.O.; Valencia, C.S. Staphylococcus spp. from wild mammals in Aragón (Spain): Antibiotic resistance status. J. Vet. Res. 2020, 64, 373–379. [Google Scholar] [CrossRef] [PubMed]
- Plaza-Rodríguez, C.; Alt, K.; Grobbel, M.; Hammerl, J.A.; Irrgang, A.; Szabo, I.; Stingl, K.; Schuh, E.; Wiehle, L.; Pfefferkorn, B.; et al. Wildlife as Sentinels of Antimicrobial Resistance in Germany? Front. Vet. Sci. 2021, 7, 1251. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Grúa, E.; Pérez-Fuentes, S.; Viana, D.; Cardells, J.; Lizana, V.; Aguiló, J.; Selva, L.; Corpa, J.M. Marked Presence of Methicillin-Resistant Staphylococcus aureus in Wild Lagomorphs in Valencia, Spain. Animals 2020, 10, 1109. [Google Scholar] [CrossRef] [PubMed]
- Pérez, J.R.; Rosa, L.Z.; Sánchez, A.G.; Salcedo, J.H.D.M.; Rodríguez, J.A.; Horrillo, R.C.; Zurita, S.; Gil Molino, M. Multiple Antimicrobial Resistance in Methicillin-Resistant Staphylococcus sciuri Group Isolates from Wild Ungulates in Spain. Antibiotics 2021, 10, 920. [Google Scholar] [CrossRef] [PubMed]
- Loncaric, I.; Kübber-Heiss, A.; Posautz, A.; Stalder, G.L.; Hoffmann, D.; Rosengarten, R.; Walzer, C. Characterization of methicillin-resistant Staphylococcus spp. carrying the mecC gene, isolated from wildlife. J. Antimicrob. Chemother. 2013, 68, 2222–2225. [Google Scholar] [CrossRef] [Green Version]
- Sousa, M.; Silva, V.; Silva, A.; Silva, N.; Ribeiro, J.; Tejedor-Junco, M.T.; Capita, R.; Chenouf, N.S.; Alonso-Calleja, C.; Rodrigues, T.M.; et al. Staphylococci among Wild European Rabbits from the Azores: A Potential Zoonotic Issue? J. Food Prot. 2020, 83, 1110–1114. [Google Scholar] [CrossRef]
- Seinige, D.; Von Altrock, A.; Kehrenberg, C. Genetic diversity and antibiotic susceptibility of Staphylococcus aureus isolates from wild boars. Comp. Immunol. Microbiol. Infect. Dis. 2017, 54, 7–12. [Google Scholar] [CrossRef]
- Bengtsson, B.; Persson, L.; Ekström, K.; Unnerstad, H.E.; Uhlhorn, H.; Börjesson, S. High occurrence of mecC -MRSA in wild hedgehogs (Erinaceus europaeus) in Sweden. Vet. Microbiol. 2017, 207, 103–107. [Google Scholar] [CrossRef]
- Held, J.; Gmeiner, M.; Mordmüller, B.; Matsiégui, P.-B.; Schaer, J.; Eckerle, I.; Weber, N.; Matuschewski, K.; Bletz, S.; Schaumburg, F. Bats are rare reservoirs of Staphylococcus aureus complex in Gabon. Infect. Genet. Evol. 2017, 47, 118–120. [Google Scholar] [CrossRef]
- Sousa, M.; Silva, N.; Igrejas, G.; Silva, F.; Sargo, R.; Alegria, N.; Benito, D.; Gómez, P.; Lozano, C.; Gómez-Sanz, E.; et al. Antimicrobial resistance determinants in Staphylococcus spp. recovered from birds of prey in Portugal. Vet. Microbiol. 2014, 171, 436–440. [Google Scholar] [CrossRef]
- Gómez, P.; Lozano, C.; Camacho, M.C.; Lima-Barbero, J.-F.; Hernández, J.-M.; Zarazaga, M.; Höfle, U.; Torres, C. Detection of MRSA ST3061-t843-mecC and ST398-t011-mecA in white stork nestlings exposed to human residues: Table 1. J. Antimicrob. Chemother. 2015, 71, 53–57. [Google Scholar] [CrossRef] [Green Version]
- Ruiz-Ripa, L.; Gómez, P.; Alonso, C.A.; Camacho, M.C.; De La Puente, J.; Fernández-Fernández, R.; Ramiro, Y.; Quevedo, M.A.; Blanco, J.M.; Zarazaga, M.; et al. Detection of MRSA of Lineages CC130-mecC and CC398-mecA and Staphylococcus delphini-lnu(A) in Magpies and Cinereous Vultures in Spain. Microb. Ecol. 2019, 78, 409–415. [Google Scholar] [CrossRef]
- Ge, J.; Zhong, X.-S.; Xiong, Y.-Q.; Qiu, M.; Huo, S.-T.; Chen, X.-J.; Mo, Y.; Cheng, M.-J.; Chen, Q. Methicillin-resistant Staphylococcus aureus among urban rodents, house shrews, and patients in Guangzhou, Southern China. BMC Vet. Res. 2019, 15, 260. [Google Scholar] [CrossRef]
- Lee, M.J.; Byers, K.; Donovan, C.M.; Zabek, E.; Stephen, C.; Patrick, D.M.; Himsworth, C.G. Methicillin-resistant Staphylococcus aureus in urban Norway rat (Rattus norvegicus) populations: Epidemiology and the impacts of kill-trapping. Zoonoses Public Health 2018, 66, 343–348. [Google Scholar] [CrossRef]
- John-Bosco, K.; Valeria, N.; Peter, S. Nasal Carriage of Methicillin-Resistant Staphylococcus aureus among Sympatric Free Ranging Domestic Pigs and Wild Chlorocebus pygerythrus in a Rural African Setting. Available online: https://www.researchsquare.com/article/rs-524223/v1 (accessed on 10 September 2021).
- Schaumburg, F.; Pauly, M.; Anoh, E.; Mossoun, A.; Wiersma, L.; Schubert, G.; Flammen, A.; Alabi, A.S.; Muyembe-Tamfum, J.-J.; Grobusch, M.P.; et al. Staphylococcus aureus complex from animals and humans in three remote African regions. Clin. Microbiol. Infect. 2015, 21, 345.e1–345.e8. [Google Scholar] [CrossRef] [Green Version]
- Schaumburg, F.; Mugisha, L.; Peck, B.; Becker, K.; Gillespie, T.R.; Peters, G.; Leendertz, F.H. Drug-Resistant Human Staphylococcus aureus in Sanctuary Apes Pose a Threat to Endangered Wild Ape Populations. Am. J. Primatol. 2012, 74, 1071–1075. [Google Scholar] [CrossRef]
- Hoefer, A.; Boyen, F.; Beierschmitt, A.; Moodley, A.; Roberts, M.; Butaye, P. Methicillin-Resistant and Methicillin-Susceptible Staphylococcus from Vervet Monkeys (Chlorocebus sabaeus) in Saint Kitts. Antibiotics 2021, 10, 290. [Google Scholar] [CrossRef]
- Schaumburg, F.; Mugisha, L.; Kappeller, P.; Fichtel, C.; Köck, R.; Köndgen, S.; Becker, K.; Boesch, C.; Peters, G.; Leendertz, F. Evaluation of Non-Invasive Biological Samples to Monitor Staphylococcus aureus Colonization in Great Apes and Lemurs. PLoS ONE 2013, 8, e78046. [Google Scholar] [CrossRef]
- Schaumburg, F.; Alabi, A.S.; Köck, R.; Mellmann, A.; Kremsner, P.G.; Boesch, C.; Becker, K.; Leendertz, F.H.; Peters, G. Highly divergent Staphylococcus aureus isolates from African non-human primates. Environ. Microbiol. Rep. 2011, 4, 141–146. [Google Scholar] [CrossRef]
- Roberts, M.C.; Joshi, P.R.; Greninger, A.L.; Melendez, D.; Paudel, S.; Acharya, M.; Bimali, N.K.; Koju, N.P.; No, D.; Chalise, M.; et al. The human clone ST22 SCCmec IV methicillin-resistant Staphylococcus aureus isolated from swine herds and wild primates in Nepal: Is man the common source? FEMS Microbiol. Ecol. 2018, 94, fiy052. [Google Scholar] [CrossRef] [Green Version]
- Gongal, G. One Health Approach in the South East Asia Region: Opportunities and challenges. In One Health: The Human-Animal-Environment Interfaces in Emerging Infectious Diseases; Mackenzie, J.S., Jeggo, M., Daszak, P., Richt, J.A., Eds.; Springer: Berlin/Heidelberg, Germany, 2013; pp. 113–122. [Google Scholar] [CrossRef]
- Silva, V.; Capelo, J.L.; Igrejas, G.; Poeta, P. Molecular Epidemiology of Staphylococcus aureus Lineages in Wild Animals in Europe: A Review. Antibiotics 2020, 9, 122. [Google Scholar] [CrossRef] [Green Version]
- Abdullahi, I.; Lozano, C.; Ruiz-Ripa, L.; Fernández-Fernández, R.; Zarazaga, M.; Torres, C. Ecology and Genetic Lineages of Nasal Staphylococcus aureus and MRSA Carriage in Healthy Persons with or without Animal-Related Occupational Risks of Colonization: A Review of Global Reports. Pathogens 2021, 10, 1000. [Google Scholar] [CrossRef]
- Bierowiec, K.; Płoneczka-Janeczko, K.; Rypuła, K. BioMed Res. Int. 2016, 2016, 3070524. [CrossRef] [Green Version]
- Ruiz-Ripa, L.; Gómez, P.; Alonso, C.A.; Camacho, M.C.; Ramiro, Y.; De La Puente, J.; Fernández-Fernández, R.; Quevedo, M.; Blanco, J.M.; Báguena, G.; et al. Frequency and Characterization of Antimicrobial Resistance and Virulence Genes of Coagulase-Negative Staphylococci from Wild Birds in Spain. Detection of tst-Carrying S. sciuri Isolates. Microorganisms 2020, 8, 1317. [Google Scholar] [CrossRef]
- Jourdain, E.; Gauthier-Clerc, M.; Bicout, D.J.; Sabatier, P. Bird Migration Routes and Risk for Pathogen Dispersion into Western Mediterranean Wetlands. Emerg. Infect. Dis. 2007, 13, 365–372. [Google Scholar] [CrossRef]
- Gómez-Sanz, E.; Torres, C.; Lozano, C.; Fernández-Pérez, R.; Aspiroz, C.; Larrea, F.R.; Zarazaga, M. Detection, Molecular Characterization, and Clonal Diversity of Methicillin-Resistant Staphylococcus aureus CC398 and CC97 in Spanish Slaughter Pigs of Different Age Groups. Foodborne Pathog. Dis. 2010, 7, 1269–1277. [Google Scholar] [CrossRef]
- Lozano, C.; Fernández-Fernández, R.; Ruiz-Ripa, L.; Gómez, P.; Zarazaga, M.; Torres, C. Human mecC-Carrying MRSA: Clinical Implications and Risk Factors. Microorganisms 2020, 8, 1615. [Google Scholar] [CrossRef]
- Loncaric, I.; Kübber-Heiss, A.; Posautz, A.; Stalder, G.L.; Hoffmann, D.; Rosengarten, R.; Walzer, C. mecC- and mecA-positive meticillin-resistant Staphylococcus aureus (MRSA) isolated from livestock sharing habitat with wildlife previously tested positive for mecC-positive MRSA. Vet. Dermatol. 2014, 25, 147–148. [Google Scholar] [CrossRef] [PubMed]
- Monecke, S.; Jatzwauk, L.; Müller, E.; Nitschke, H.; Pfohl, K.; Slickers, P.; Reissig, A.; Ruppelt-Lorz, A.; Ehricht, R. Diversity of SCCmec Elements in Staphylococcus aureus as Observed in South-Eastern Germany. PLoS ONE 2016, 11, e0162654. [Google Scholar] [CrossRef] [PubMed]
- Aklilu, E.; Chia, H.Y. First mecC and mecA Positive Livestock-Associated Methicillin Resistant Staphylococcus aureus (mecC MRSA/LA-MRSA) from Dairy Cattle in Malaysia. Microorganisms 2020, 8, 147. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Porrero, M.C.; Harrison, E.; Fernández-Garayzábal, J.F.; Paterson, G.K.; Díez-Guerrier, A.; Holmes, M.A.; Domínguez, L. Detection of mecC-Methicillin-resistant Staphylococcus aureus isolates in river water: A potential role for water in the environmental dissemination. Environ. Microbiol. Rep. 2014, 6, 705–708. [Google Scholar] [CrossRef]
- Heaton, C.J.; Gerbig, G.R.; Sensius, L.D.; Patel, V.; Smith, T.C. Staphylococcus aureus Epidemiology in Wildlife: A Systematic Review. Antibiotics 2020, 9, 89. [Google Scholar] [CrossRef] [Green Version]
- Harrison, E.M.; Coll, F.; Toleman, M.S.; Blane, B.; Brown, N.M.; Torok, E.; Parkhill, J.; Peacock, S.J. Genomic surveillance reveals low prevalence of livestock-associated methicillin-resistant Staphylococcus aureus in the East of England. Sci. Rep. 2017, 7, 7406. [Google Scholar] [CrossRef] [Green Version]
- Gómez, P.; González-Barrio, D.; Benito, D.; García, J.T.; Viñuela, J.; Zarazaga, M.; Ruiz-Fons, F.; Torres, C. Detection of methicillin-resistant Staphylococcus aureus (MRSA) carrying the mecC gene in wild small mammals in Spain. J. Antimicrob. Chemother. 2014, 69, 2061–2064. [Google Scholar] [CrossRef] [Green Version]
- Ceballos, S.; Aspiroz, C.; Ruiz-Ripa, L.; Reynaga, E.; Azcona-Gutiérrez, J.M.; Rezusta, A.; Seral, C.; Antoñanzas, F.; Torres, L.; López, A.R.; et al. Epidemiology of MRSA CC398 in hospitals located in Spanish regions with different pig-farming densities: A multicentre study. J. Antimicrob. Chemother. 2019, 74, 2157–2161. [Google Scholar] [CrossRef]
- Price, L.B.; Stegger, M.; Hasman, H.; Aziz, M.; Larsen, J.; Andersen, P.S.; Pearson, T.; Waters, A.E.; Foster, J.; Schupp, J.; et al. Staphylococcus aureus CC398: Host Adaptation and Emergence of Methicillin Resistance in Livestock. mBio 2012, 3, e00305-11. [Google Scholar] [CrossRef] [Green Version]
- Mama, O.M.; Aspiroz, C.; Ruiz-Ripa, L.; Ceballos, S.; Iñiguez-Barrio, M.; Cercenado, E.; Azcona, J.M.; López-Cerero, L.; Seral, C.; López-Calleja, A.I.; et al. Prevalence and Genetic Characteristics of Staphylococcus aureus CC398 Isolates From Invasive Infections in Spanish Hospitals, Focusing on the Livestock-Independent CC398-MSSA Clade. Front. Microbiol. 2021, 12, 623108. [Google Scholar] [CrossRef]
- Salgueiro, V.; Manageiro, V.; Bandarra, N.M.; Ferreira, E.; Clemente, L.; Caniça, M. Genetic Relatedness and Diversity of Staphylococcus aureus from Different Reservoirs: Humans and Animals of Livestock, Poultry, Zoo, and Aquaculture. Microorganisms 2020, 8, 1345. [Google Scholar] [CrossRef]
- Monecke, S.; Gavier-Widén, D.; Hotzel, H.; Peters, M.; Guenther, S.; Lazaris, A.; Loncaric, I.; Müller, E.; Reissig, A.; Ruppelt-Lorz, A.; et al. Diversity of Staphylococcus aureus Isolates in European Wildlife. PLoS ONE 2016, 11, e0168433. [Google Scholar] [CrossRef] [Green Version]
- Michalik, M.; Kosecka-Strojek, M.; Wolska, M.; Samet, A.; Podbielska-Kubera, A.; Międzobrodzki, J. First Case of Staphylococci Carrying Linezolid Resistance Genes from Laryngological Infections in Poland. Pathogens 2021, 10, 335. [Google Scholar] [CrossRef]
- Zheng, Y.; Qin, C.; Zhang, X.; Zhu, Y.; Li, A.; Wang, M.; Tang, Y.; Kreiswirth, B.N.; Chen, L.; Zhang, H.; et al. The tst gene associated Staphylococcus aureus pathogenicity island facilitates its pathogenesis by promoting the secretion of inflammatory cytokines and inducing immune suppression. Microb. Pathog. 2019, 138, 103797. [Google Scholar] [CrossRef]
- Novick, R.P.; Ram, G. Staphylococcal pathogenicity islands—Movers and shakers in the genomic firmament. Curr. Opin. Microbiol. 2017, 38, 197–204. [Google Scholar] [CrossRef]

| (a) | |||||||||||||
| Study Groups | Number of S. aureus Studies Included | Total Number | Pooled S. aureus Carriage Rate (%) (Range) | OR (95% CI) | p Value | Number of MRSA Studies Included | Total Number | Pooled MRSA Carriage Rate (%) (Range) | OR (95% CI) | p Value | Total Number of Studies Included a | ||
| Animals | S. aureus | Animals | MRSA | ||||||||||
| Wild Mammals (excluding rodents and NHP) | 13 | 3031 | 479 | 15.8 (0.0–36.9) | Referent | Referent | 17 | 6110 | 99 | 1.6 (0.0–63.6) | Referent | Referent | 18 |
| Wild Rodents | 4 | 856 | 207 | 24.2 (15.3–41.0) | 1.69 (1.41–2.04) | <0.0001 | 5 | 1452 | 49 | 3.4 (0.3–4.7) | 2.12 (1.49–3.00) | <0.0001 | 5 |
| Non-human Primates | 7 | 403 | 158 | 39.2 (0.0–100.0) | 3.44 (2.78–4.29) | <0.0001 | 7 | 403 | 8 | 2.0 (0.0–26.7) | 1.23 (0.59–2.55) | 0.578 | 7 |
| Wild Birds | 5 | 586 | 60 | 10.3 (5.0–34.8) | 0.61 (0.46–0.81) | 0.0006 | 6 | 626 | 21 | 3.4 (0.0–4.0) | 2.11 (1.31–3.40) | 0.002 | 6 |
| Total Wild Animals | 29 | 4876 | 905 | 18.5 (0.0–100) | NA | NA | 35 a | 8601 | 177 | 2.1 (0.0–63.9) | NA | NA | 36 a |
| (b) | |||||||||||||
| Study Groups | Number of S. aureus Studies Included | Total Number | Pooled S. aureus Carriage Rate (%) (Range) | OR (95% CI) | p Value | Number of MRSA Studies Included | Total Number | Pooled MRSA Carriage Rate (%) (Range) | OR (95% CI) | p Value | Total Number of Studies Included a | ||
| Animals | S. aureus | Animals | MRSA | ||||||||||
| Wild Animals (excluding wild birds) | 24 | 4290 | 844 | 19.7 (0.0–100.0) | Referent | Referent | 29 | 7965 | 156 | 1.9 (0.0–63.6) | Referent | Referent | 30 |
| Wild Birds | 5 | 586 | 60 | 10.3 (5.0–34.8) | 0.46 (0.35–0.61) | <0.0001 | 6 | 626 | 21 | 3.4 (0.0–4.0) | 1.74 (1.09–2.76) | 0.019 | 6 |
| Reference | Animal Species (Location) | No. of Animals Tested | No. of S. aureus | No. of mecC-MRSA (% Colonised Animals) | spa-type/ST/CC (Number of Isolates) | AMR Phenotype for Non-Beta-Lactams of mecC-MRSA | IEC-type in mecC-MRSA (Number Isolates, spa) | Other Virulence Genes in mecC-MRSA Isolates (Number Strains) |
|---|---|---|---|---|---|---|---|---|
| [12] | Wild free-living rodents (Germany) | 145 | 37 | 1 (0.7) | t843 (1)/CC130 (1) | Susceptible (all) | IEC-negative (1) | NT |
| [17] | Wild rodents (Portugal) | 204 | 38 | 3 (1.5) | t1525 (3)/ST1945 (3)/ CC130 (3) | Susceptible (all) | IEC-E (3, t1535) | Negative (for lukS/F-PV, hla, hlb, eta, etb, tst) (3) |
| [19] | Red deer (Spain) | 65 | 16 | 11 (16.9) | t843 (4), t1535 (7)/CC130 (11) | Susceptible (all) | IEC-E (11, t843, t1535) | etd2 (11) |
| [22] | Wild rodents and shrews (Germany, Czech and France) | 295 | 45 | 1 (0.3) | t843 (1)/CC130 (1) | Susceptible (all) | IEC-negative (1) | Negative (for lukS/F-PV, sea-seu, tst, eta, etd) (1) |
| [23] | European hedgehog, European rabbit, red deer, wild boar, European mouflon (Spain) | 103 | 23 | 3 (2.9) | t843 (3)/ST130 (3)/CC130 (3) | Susceptible (all) | IEC-negative (3) | seg (1), seh (1). Negative for lukS/F-PV, tst, eta, etb, and 18 enterotoxin genes |
| [29] | Rabbit and hare (Spain) | 363 | 70 | 34 (9.3) | ST1945 (33), ST5823 (1)/CC130 (34) | Susceptible (all) | IEC-negative (34) | NT |
| [31] | European brown hare, European otter, European hedgehog, Eurasian lynx (Germany) | 40 | 5 | 5 (12.5) | t843 (2), t10513 (1), t3256 (1), t4335 (1)/ST2620 (1). ST130 (4)/CC130 (5) | NT | NT | NT |
| [34] | Wild hedgehog (Sweden) | 55 | 35 | 35 (63.6) | t843 (17), t10751, t978 (3), t9111 (3), t15312 (4), t3391 (5), t10893 (1), t11015 (1)/CC130 (20), CC2361 (15) | CIP (5), CLI (6), ERY (5), GEN (7), KAN (5), TET (2) | NT | NT |
| [37] | Stork (Spain) | 92 | 32 | 1 (1.1) | t843 (1)/ST3061 (1)/CC130 (1) | Susceptible (all) | IEC-negative (1) | etd2 (1) |
| [38] | Cinereous vulture and magpie (Spain) | 324 | 15 | 12 (3.7) | t843 (11), t1535 (1)/CC130 (12) | Susceptible (all) | IEC-E (4, t843) IEC-negative (8, t843 and t1535) | Negative (for lukS/F-PV, tst, eta, etb, and etd) (12) |
| Reference | Animal Species (Location) | No. of MRSA-CC398 | spa/ST of MRSA-CC398 (Number of Strains) | IEC-type (Number of Strains) in MRSA-CC398 | AMR Phenotypes/Genes (Number Strains) of MRSA-CC398 | Other Virulence Genes (Number Strains) of MRSA-CC398 | No. of MSSA-CC398 (%) | spa/ST of MSSA-CC398 (Number of Strains) | IEC-type (Number of Strains) in MSSA-CC398 | AMR Phenotypes/Genes (Number Strains) in MSSA-CC398 | Other Virulence Genes (Number Strains) in MSSA-CC398 |
|---|---|---|---|---|---|---|---|---|---|---|---|
| [5] | Iberian ibex, red deer and wild boars (Spain) | 0 | NA | NA | NA | NA | 3 | t034 (2), t571 (1)/ST398/CC398 | NT | TET (3) | NT |
| [17] | Wild rodents (Portugal) | 0 | NA | NA | NA | NA | 6 | t1451 (5), t571 (1)/ST398 (4), ST5926 (2) | IEC-C (2), IEC-negative (4) | Susceptible (all) | hld (all) |
| [18] | Wild boars (Portugal) | 1 | t899/ST398 | IEC-B (1) | TET, PEN FOX, OXA, CIP/mecA | NT | 0 | NA | NA | NA | NA |
| [21] a | Wild mammals (Spain) | 3 | t011 (2), t1451 (1)/ST398 (3) | NT | TET (3), CIP (2), ERY (1), CLI (1) | NT | 0 | NA | NA | NA | NA |
| [21] a | Eurasian griffon vulture (Spain) | 2 | t011 (2)/ST398 (2) | NT | TET (2), CIP (1), ERY (1), CLI (1) | NT | 0 | NA | NA | NA | NA |
| [25] | Wild boar (Spain) | 1 | t011/ST398 (1) | IEC-negative (1) | PEN- FOX- TET/blaZ, mecA, tet(M), tet(K) | Negative (for lukS/F-PV, tst, eta, and etb) (1) | 0 | NA | NA | NA | NA |
| [33] | Wild boar (Germany) | 0 | NA | NA | NA | NA | 1 | t571 (1)/ST804 (1) | NT | AMP (1), ERY (1)/blaZ | Negative (for sea, seb, sec, sed, see, she, eta, etb, tst, luk-S/F-PV) (1) |
| [37] | Stork (Spain) | 1 | t011 (1)/ST398 (1) | IEC-negative (1) | PEN, OXA, FOX, TET/mecA, tetK, tetM | cna (1) | 9 | t571 (5), t6606 (3), t3625 (1)/ / ST398 (8), ST2377 (1) | IEC-C (9) | PEN (all), ERY (all), CLI (all)/ blaZ, erm(T) | cna (all) |
| [38] | Cinereous vulture (Spain) | 1 | t011 (1)/ST398 (1) | IEC-negative (1) | PEN, FOX, ERI, CLI, TET/mecA, blaZ, erm(C), vga(A), tetK, tetM | Negative (for lukS/F-PV, tst, eta, etb, and etd) (1) | 0 | NA | NA | NA | NA |
| [39] | Rodents (China) | 5 | t034 (1), t011 (1), t4552 (1), t2582 (2)/ST398 (4), ST1232 (1) | IEC-E (1), IEC-A (1) IEC-negative (3) | TET (2), AZM (1), CLI (1) | lukS/F-PV (1, spa t034) | 0 | NA | NA | NA | NA |
| (a) | |||||||||
| Reference | Animal Species | No. of Animals Tested/S. aureus/MRSA | No. of tst (%) in MRSA | No. of tst (%) in MSSA | No. of lukS/F-PV (%) in MRSA | No. of lukS/F-PV (%) in MSSA | No. Strains with IEC (%) in MSSA | No. Strains with IEC (%) in MRSA | No. of S. aureus IEC-Negative (%) |
| [12] | Wild free-living rodents | 145/37/2 | 1 (50.0) | 0 | 0 | 0 | 0 | IEC-E, 1 (50.0) | 36 (97.3) |
| [17] | Wild rodents | 208/38/6 | 0 | 0 | 0 | 0 | IEC-E, 1 (3.1) | IEC-E, 3 (50.0) | 31 (81.6) |
| IEC-C, 2 (6.2) | IEC-A, 1 (16.7) | ||||||||
| [18] | Wild boars | 45/15/1 | NT | NT | NT | NT | NT | IEC-B, 1 (100.0) | 14 (93.3) |
| [19] | Red deer | 65/16/11 | 0 | NT | 0 | NT | NT | IEC-E, 11 (100.0) | 5 (31.3) |
| [23] | Wild mammals | 103/23/4 | 0 | 1 (5.3) | 0 | 0 | 0 | 0 | 23 (100.0) |
| [37] | Storks | 92/32/3 | 0 (0.0) | 2 (6.9) | 0 | 0 | IEC-B, 6 (20.7) | 0 | 5 (15.6) |
| IEC-C, 11 (37.9) | |||||||||
| IEC-D, 1 (3.4) | |||||||||
| IEC-G, 9 (31.0) | |||||||||
| [38] | Wild birds | 324/15/13 | 0 | 0 | 0 | 0 | IEC-E, 1 (50.0) | IEC-E, 4 (30.8) | 10 (66.7) |
| [39] | Urban rodents | 212/87/11 | NT | NT | 2 (18.2%) | NT | NT | IEC-E, 1 (9.1), IEC-IEC-G, 1 (9.1) | NT |
| IEC-B, 1 (9.1) | |||||||||
| IEC-A, 2 (18.2) | |||||||||
| [42] | NHP | 132/15/0 | ND | Present but number not specified | Present but number not specified | 0 | 0 | NT | NT |
| [43] | NHP | 62/36/0 | NT | 0 | 0 | 10 (27.8) | NT | NT | NT |
| [46] | NHP | 95/58/0 | NT | 0 | 0 | 2 (3.4) | NT | NT | NT |
| [47] | NHP | 59/29/4 | 3 (75.0) | NT | 3 (75.0) | 0 | 0 | IEC-E, 3 (75.0) | 26 (89.7) |
| (b) | |||||||||
| Reference | Origin of Isolates | Spa-type/ST/CC of Positive Isolates (No. of Isolates) | Virulence Gene | Methicillin Resistance Phenotype | IEC-type (Number of Strains) | ||||
| [12] | Germany/Rodent/Nasal | t684/CC30 (1) | tst | MRSA | E (1) | ||||
| [23] | Spain/Wild boar/Nasal | t1534/CC522 (1) | tst | MSSA | IEC-negative | ||||
| [37] | Spain/Storks/Trachea | t012/CC30 (2) | tst | MSSA | D (1) | ||||
| [47] | Nepal/NHP/Oral | ST22 (3) | tst | MRSA | E (3) | ||||
| [37] | Spain/Stork/Trachea | t209/CC5 (1) | eta | MSSA | B (1) | ||||
| [39] | China/Wild Rodents/Nasal | t034/ST1232/CC398 (1) | luk-SF-PV | MRSA | G (1) | ||||
| t127/ST1/CC5 (1) | |||||||||
| [43] | Zambia and Uganda/NHP/Nasal | ST80 (9) | luk-SF-PV | MSSA | NT | ||||
| ST2178 (1) | |||||||||
| [46] | Gabon and Cote d’ Ivoire)/ NHP/Nasal | ST1855 (1) | luk-SF-PV | MSSA | NT | ||||
| ST 1928 (1) | |||||||||
| [47] | Nepal/NHP/Oral | ST22 (3) | luk-SF-PV | MRSA | E (3) | ||||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abdullahi, I.N.; Fernández-Fernández, R.; Juárez-Fernández, G.; Martínez-Álvarez, S.; Eguizábal, P.; Zarazaga, M.; Lozano, C.; Torres, C. Wild Animals Are Reservoirs and Sentinels of Staphylococcus aureus and MRSA Clones: A Problem with “One Health” Concern. Antibiotics 2021, 10, 1556. https://doi.org/10.3390/antibiotics10121556
Abdullahi IN, Fernández-Fernández R, Juárez-Fernández G, Martínez-Álvarez S, Eguizábal P, Zarazaga M, Lozano C, Torres C. Wild Animals Are Reservoirs and Sentinels of Staphylococcus aureus and MRSA Clones: A Problem with “One Health” Concern. Antibiotics. 2021; 10(12):1556. https://doi.org/10.3390/antibiotics10121556
Chicago/Turabian StyleAbdullahi, Idris Nasir, Rosa Fernández-Fernández, Guillermo Juárez-Fernández, Sandra Martínez-Álvarez, Paula Eguizábal, Myriam Zarazaga, Carmen Lozano, and Carmen Torres. 2021. "Wild Animals Are Reservoirs and Sentinels of Staphylococcus aureus and MRSA Clones: A Problem with “One Health” Concern" Antibiotics 10, no. 12: 1556. https://doi.org/10.3390/antibiotics10121556
APA StyleAbdullahi, I. N., Fernández-Fernández, R., Juárez-Fernández, G., Martínez-Álvarez, S., Eguizábal, P., Zarazaga, M., Lozano, C., & Torres, C. (2021). Wild Animals Are Reservoirs and Sentinels of Staphylococcus aureus and MRSA Clones: A Problem with “One Health” Concern. Antibiotics, 10(12), 1556. https://doi.org/10.3390/antibiotics10121556







