Highly Sensitive Detection and Differentiation of Endotoxins Derived from Bacterial Pathogens by Surface-Enhanced Raman Scattering
Abstract
1. Introduction
2. Materials and Methods
3. Results and Discussion
3.1. Meningococcal SERS Spectra
| Observed SERS Shift Δν (cm−1) | Vibrational Mode Assignment [22,27,28] |
|---|---|
| 550 | β(CH2) in ring |
| 680 | δ(C-O-C); fatty acid; |
| 850 | ν(C-O-C); saccharides (1,4 glycosidic link) |
| 988 | β(CH); carbohydrates |
| 1131 | ν(C-C); fatty acid |
| 1330 | δ(CH); phospholipid |
| 1450 | α(CH3/CH2), β(CH3/CH2) |
3.2. Enteric LPS SERS Spectra
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Appendix A
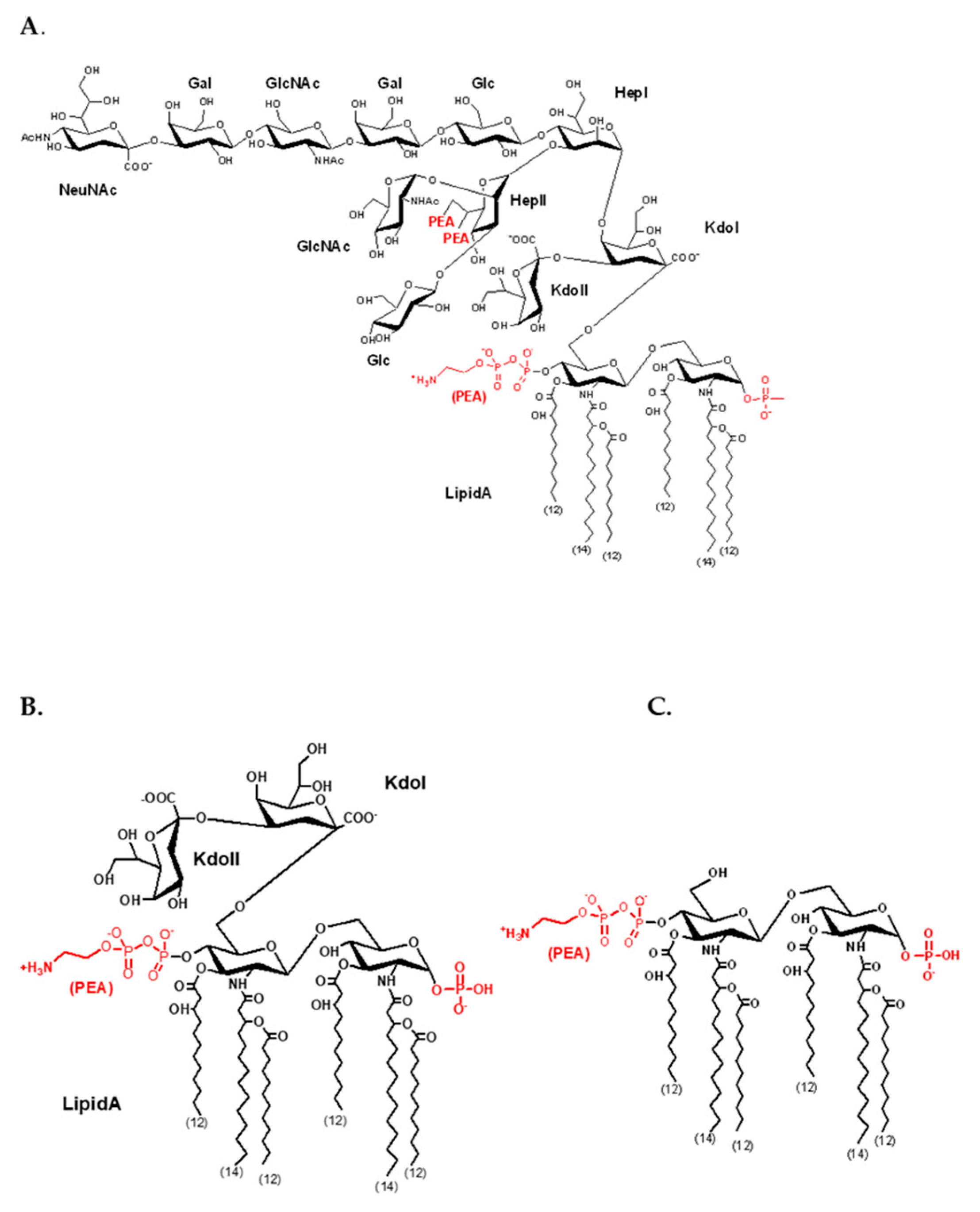
Appendix B
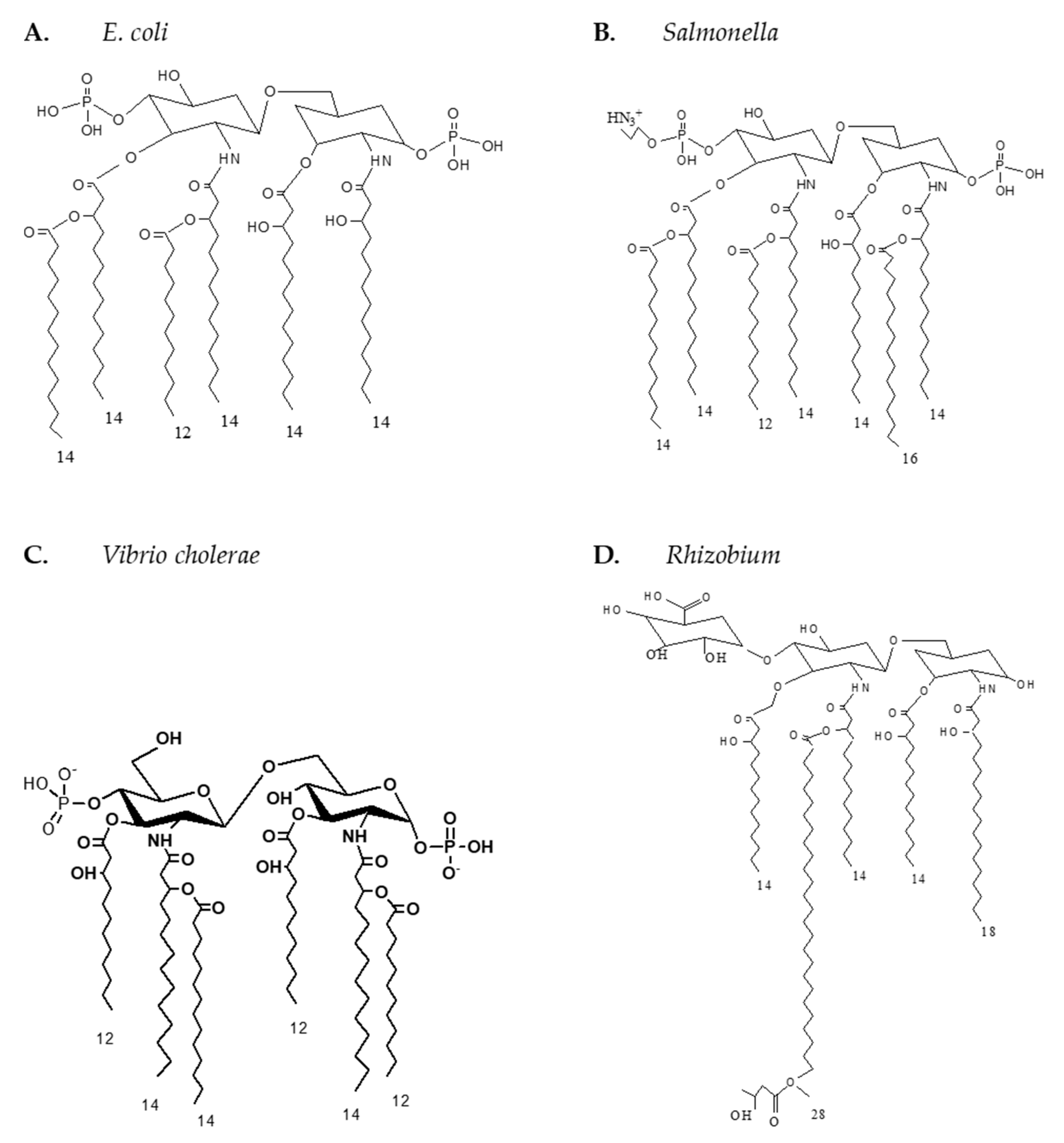
Appendix C
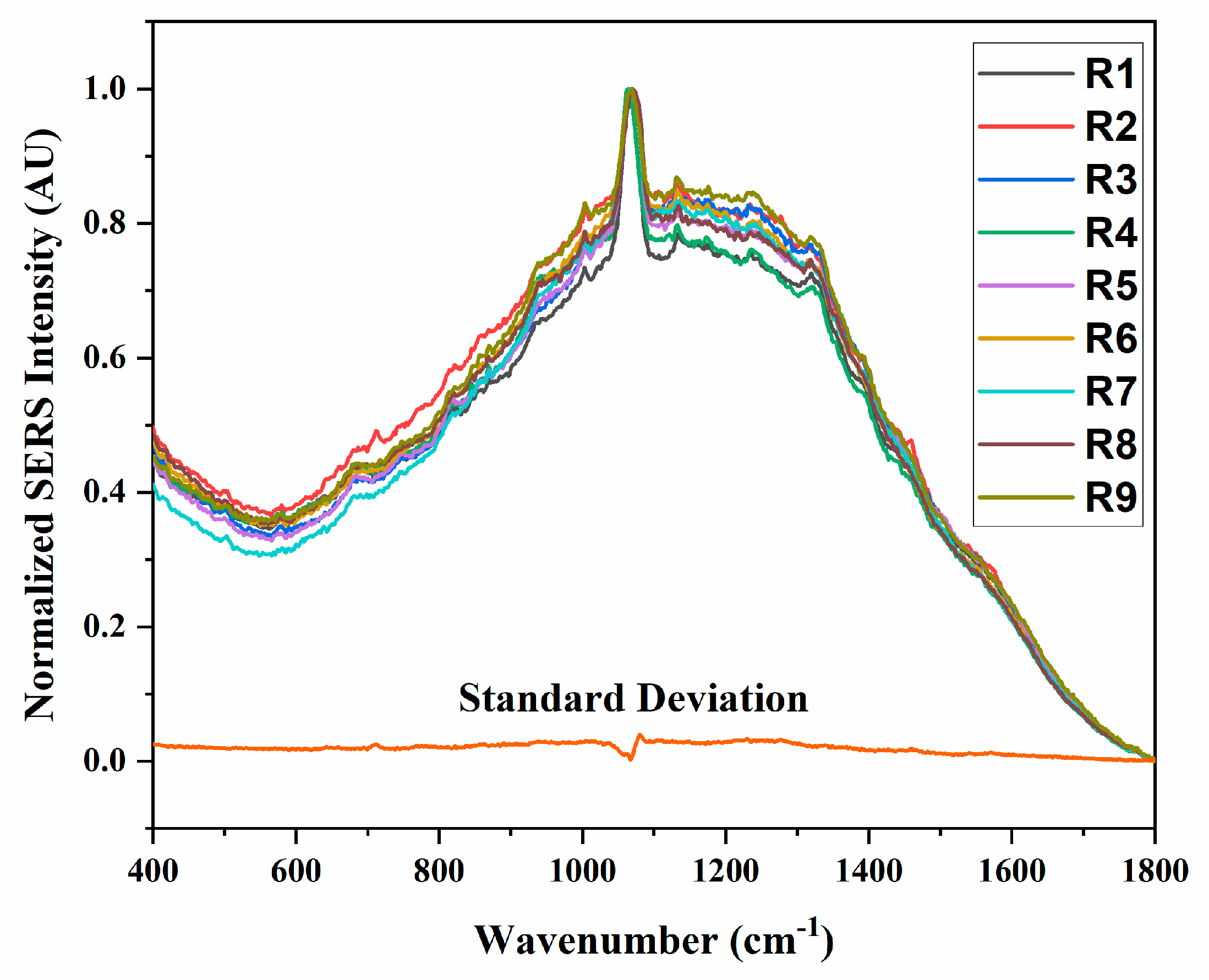
References
- Park, B.S.; Song, D.H.; Kim, H.M.; Choi, B.-S.; Lee, H.; Lee, J.-O. The structural basis of lipopolysaccharide recognition by the TLR4-MD-2 complex. Nature 2009, 458, 1191–1195. [Google Scholar] [CrossRef]
- Zughaier, S.M.; Tzeng, Y.-L.; Zimmer, S.M.; Datta, A.; Carlson, R.W.; Stephens, D.S. Neisseria meningitidis lipooligosaccharide structure-dependent activation of the macrophage CD14/Toll-like receptor 4 pathway. Infect. Immun. 2004, 72, 371–380. [Google Scholar] [CrossRef] [PubMed]
- Brandtzaeg, P.; Ovstebo, R.; Kierulf, P. Bacteremia and compartmentalization of LPS in meningococcal disease. Prog. Clin. Biol. Res. 1995, 392, 219–233. [Google Scholar] [PubMed]
- Zughaier, S.M.; Zimmer, S.M.; Datta, A.; Carlson, R.W.; Stephens, D.S. Differential induction of the toll-like receptor 4-MyD88-dependent and -independent signaling pathways by endotoxins. Infect. Immun. 2005, 73, 2940–2950. [Google Scholar] [CrossRef] [PubMed]
- Forsberg, L.S.; Carlson, R.W. The structures of the lipopolysaccharides from Rhizobium etli strains CE358 and CE359. The complete structure of the core region of R. etli lipopolysaccharides. J. Biol. Chem. 1998, 273, 2747–2757. [Google Scholar] [CrossRef]
- Iwanaga, S. Biochemical principle of Limulus test for detecting bacterial endotoxins. Proc. Jpn. Acad. Ser. B Phys. Biol. Sci. 2007, 83, 110–119. [Google Scholar] [CrossRef]
- Larrouy-Maumus, G.; Clements, A.; Filloux, A.; McCarthy, R.; Mostowy, S. Direct detection of lipid A on intact Gram-negative bacteria by MALDI-TOF mass spectrometry. J. Microbiol. Methods 2016, 120, 68–71. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Deng, P.; Tian, Y.; Feng, J.; Xiao, J.; Li, J.; Liu, J.; Li, G.; He, Q. Simultaneous and sensitive determination of ascorbic acid, dopamine and uric acid via an electrochemical sensor based on PVP-graphene composite. J. Nanobiotechnol. 2020, 18, 112. [Google Scholar] [CrossRef]
- Kim, S.-E.; Su, W.; Cho, M.; Lee, Y.; Choe, W.-S. Harnessing aptamers for electrochemical detection of endotoxin. Anal. Biochem. 2012, 424, 12–20. [Google Scholar] [CrossRef]
- Seth, R.; Ribeiro, M.; Romaschin, A.; Scott, J.A.; Manno, M.; Scott, J.A.; Liss, G.M.; Tarlo, S.M. Occupational endotoxin exposure and a novel luminol-enhanced chemiluminescence assay of nasal lavage neutrophil activation. J. Allergy Clin. Immunol. 2011, 127, 272–275. [Google Scholar] [CrossRef]
- Kneipp, J.; Kneipp, H.; Kneipp, K. SERS—A single-molecule and nanoscale tool for bioanalytics. Chem. Soc. Rev. 2008, 37, 1052–1060. [Google Scholar] [CrossRef]
- Choi, H.-K.; Lee, K.S.; Shin, H.-H.; Koo, J.-J.; Yeon, G.J.; Kim, Z.H. Single-Molecule Surface-Enhanced Raman Scattering as a Probe of Single-Molecule Surface Reactions: Promises and Current Challenges. Acc. Chem. Res. 2019, 52, 3008–3017. [Google Scholar] [CrossRef]
- Fang, W.; Jia, S.; Chao, J.; Wang, L.; Duan, X.; Liu, H.; Li, Q.; Zuo, X.; Wang, L.; Wang, L.; et al. Quantizing single-molecule surface-enhanced Raman scattering with DNA origami metamolecules. Sci. Adv. 2019, 5, eaau4506. [Google Scholar] [CrossRef]
- Xiang, S.; Ge, C.; Li, S.; Chen, L.; Wang, L.; Xu, Y. In Situ Detection of Endotoxin in Bacteriostatic Process by SERS Chip Integrated Array Microchambers within Bioscaffold Nanostructures and SERS Tags. ACS Appl. Mater. Interfaces 2020, 12, 28985–28992. [Google Scholar] [CrossRef]
- Driskell, J.D.; Shanmukh, S.; Liu, Y.; Chaney, S.; Hennigan, S.; Jones, L.; Krause, D.; Tripp, R.A.; Zhao, Y.-P.; Dluhy, R.A. Novel Nanoarray SERS Substrates Used for High Sensitivity Virus Biosensing and Classification. Nanosci. Nanotechnol. Chem. Biol. Def. 2009, 1016, 99–114. [Google Scholar]
- Wu, X.; Chen, J.; Park, B.; Huang, Y.W.; Zhao, Y. The Use of Silver Nanorod Array-Based Surface-Enhanced Raman Scattering Sensor for Food Safety Applications, in Advances in Applied Nanotechnology for Agriculture, in Advances in Applied Nanotechnology for Agriculture. Am. Chem. Soc. 2013, 1143, 85–108. [Google Scholar]
- Wu, X.; Chen, J.; Li, X.; Zhao, Y.; Zughaier, S.M. Culture-free diagnostics of Pseudomonas aeruginosa infection by silver nanorod array based SERS from clinical sputum samples. Nanomedicine 2014, 10, 1863–1870. [Google Scholar] [CrossRef]
- Liu, Y.J.; Chu, H.Y.; Zhao, Y.P. Silver Nanorod Array Substrates Fabricated by Oblique Angle Deposition: Morphological, Optical, and SERS Characterizations. J. Phys. Chem. C 2010, 114, 8176–8183. [Google Scholar] [CrossRef]
- Driskell, J.D.; Shanmukh, S.; Liu, Y.; Chaney, S.B.; Tang, X.J.; Zhao, Y.P.; Dluhy, A.R. The Use of Aligned Silver Nanorod Arrays Prepared by Oblique Angle Deposition as Surface Enhanced Raman Scattering Substrates. J. Phys. Chem. C 2008, 112, 895–901. [Google Scholar] [CrossRef]
- Zhao, Y.P.; Chaney, S.B.; Zhang, Z.Y. Absorbance spectra of aligned Ag nanorod arrays prepared by oblique angle deposition. J. Appl. Phys. 2006, 100, 063527. [Google Scholar] [CrossRef]
- Abell, J.; Driskell, J.; Dluhy, R.; Tripp, R.; Zhao, Y.-P. Fabrication and characterization of a multiwell array SERS chip with biological applications. Biosens. Bioelectron. 2009, 24, 3663–3670. [Google Scholar] [CrossRef] [PubMed]
- Czamara, K.; Majzner, K.; Pacia, M.Z.; Kochan, K.; Kaczor, A.; Baranska, M. Raman spectroscopy of lipids: A review. J. Raman Spectrosc. 2015, 46, 4–20. [Google Scholar] [CrossRef]
- Utzinger, U.; Heintzelman, D.L.; Mahadevan-Jansen, A.; Malpica, A.; Follen, M.; Richards-Kortum, R. Near-infrared Raman spectroscopy for in vivo detection of cervical precancers. Appl. Spectrosc. 2001, 55, 955–959. [Google Scholar] [CrossRef]
- Krafft, C.; Neudert, L.; Simat, T.; Salzer, R. Near infrared Raman spectra of human brain lipids. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2005, 61, 1529–1535. [Google Scholar] [CrossRef] [PubMed]
- Brandtzaeg, P.; Bryn, K.; Kierulf, P.; Ovstebø, R.; Namork, E.; Aase, B.; Jantzen, E. Meningococcal endotoxin in lethal septic shock plasma studied by gas chromatography, mass-spectrometry, ultracentrifugation, and electron microscopy. J. Clin. Investig. 1992, 89, 816–823. [Google Scholar] [CrossRef] [PubMed]
- Zughaier, S.M.; Lindner, B.; Howe, J.; Garidel, P.; Koch, M.H.; Brandenburg, K.; Stephens, D.S. Physicochemical characterization and biological activity of lipooligosaccharides and lipid A from Neisseria meningitidis. J. Endotoxin Res. 2007, 13, 343–357. [Google Scholar] [CrossRef]
- Wiercigroch, E.; Szafraniec, E.; Czamara, K.; Pacia, M.Z.; Majzner, K.; Kochan, K.; Kaczor, A.; Baranska, M.; Malek, K. Raman and infrared spectroscopy of carbohydrates: A review. Spectrochim. Acta Part A Mol. Biomol. Spectrosc. 2017, 185, 317–335. [Google Scholar] [CrossRef]
- Ruggiero, L.; Sodo, A.; Bruni, F.; Ricci, M.A. Hydration of monosaccharides studied by Raman scattering. J. Raman Spectrosc. 2018, 49, 1066–1075. [Google Scholar] [CrossRef]
- Notingher, I.; Green, C.; Dyer, C.; Perkins, E.; Hopkins, N.; Lindsay, C.; Hench, L.L. Discrimination between ricin and sulphur mustard toxicity in vitro using Raman spectroscopy. J. R. Soc. Interface 2004, 1, 79–90. [Google Scholar] [CrossRef]
- Malini, R.; Venkatakrishna, K.; Kurien, J.; Pai, K.M.; Rao, L.; Kartha, V.B.; Krishna, C.M. Discrimination of normal, inflammatory, premalignant, and malignant oral tissue: A Raman spectroscopy study. Biopolymers 2006, 81, 179–193. [Google Scholar] [CrossRef] [PubMed]
- Lu, X.; Al-Qadiri, H.M.; Lin, M.; Rasco, B.A. Application of Mid-infrared and Raman Spectroscopy to the Study of Bacteria. Food Bioprocess Technol. 2011, 4, 919–935. [Google Scholar] [CrossRef]
- Delgado-Coello, B.; Montalvan-Sorrosa, D.; Cruz-Rangel, A.; Sosa-Garrocho, M.; Hernández-Téllez, B.; Macías-Silva, M.; Castillo, R.; Mas-Oliva, J. Label-free surface-enhanced Raman spectroscopy of lipid-rafts from hepatocyte plasma membranes. J. Raman Spectrosc. 2017, 48, 659–667. [Google Scholar] [CrossRef]

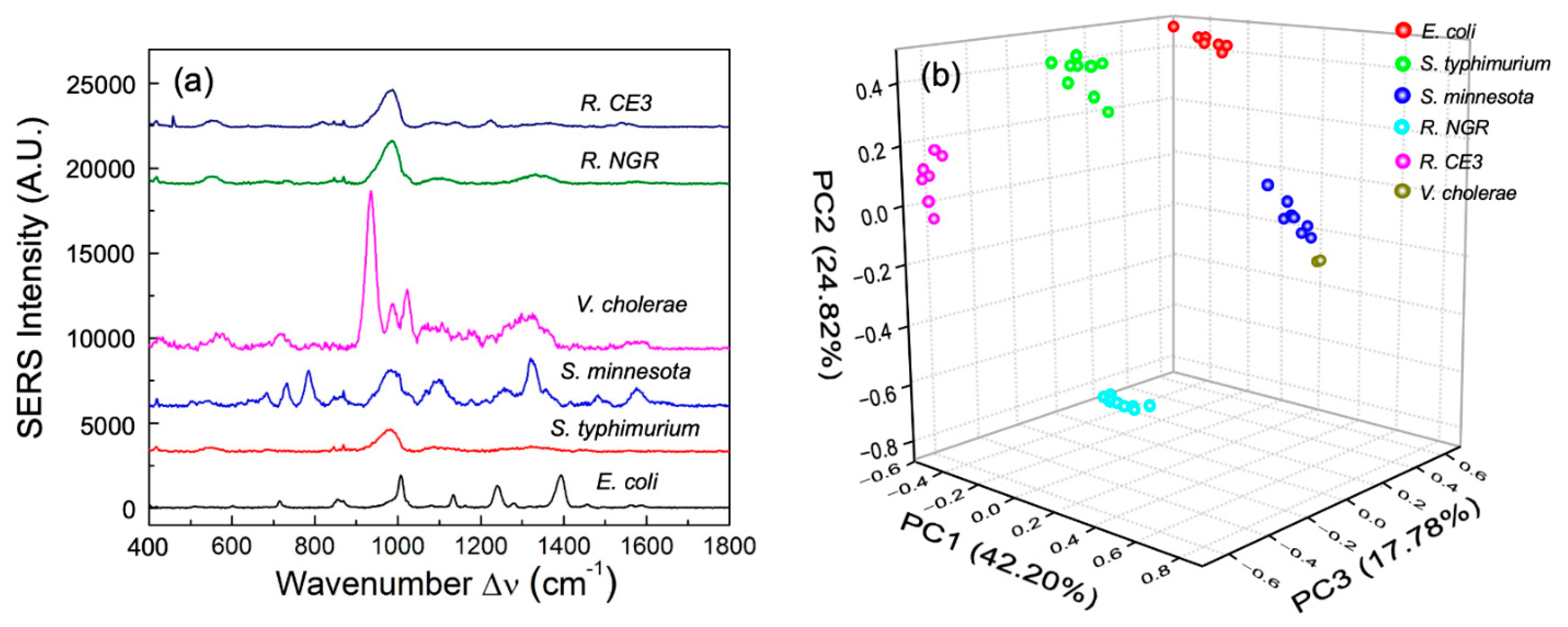
| Observed SERS Shift Δν (cm−1) | Vibrational Mode Assignment [22,27,30,31,32] |
|---|---|
| 543 | δ(C-O-C) in glycosidic ring |
| 552 | β(CH2) in ring |
| 715 | ν(C-N) |
| 733 | β(C-O-C) in carbohydrates |
| 787 | ν(C-O) in ring |
| 855 | δ(C-O-C) |
| 981 | β(CH) in lipid |
| 1025 | ν(CO) in carbohydrates |
| 1087 | ν(C-O) in lipid |
| 1131 | ν(C-C) in fatty acid |
| 1309 | τ(CH3/CH2) in lipid |
| 1330 | δ(CH) in phospholipid |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, X.; Zhao, Y.; Zughaier, S.M. Highly Sensitive Detection and Differentiation of Endotoxins Derived from Bacterial Pathogens by Surface-Enhanced Raman Scattering. Biosensors 2021, 11, 234. https://doi.org/10.3390/bios11070234
Wu X, Zhao Y, Zughaier SM. Highly Sensitive Detection and Differentiation of Endotoxins Derived from Bacterial Pathogens by Surface-Enhanced Raman Scattering. Biosensors. 2021; 11(7):234. https://doi.org/10.3390/bios11070234
Chicago/Turabian StyleWu, Xiaomeng, Yiping Zhao, and Susu M. Zughaier. 2021. "Highly Sensitive Detection and Differentiation of Endotoxins Derived from Bacterial Pathogens by Surface-Enhanced Raman Scattering" Biosensors 11, no. 7: 234. https://doi.org/10.3390/bios11070234
APA StyleWu, X., Zhao, Y., & Zughaier, S. M. (2021). Highly Sensitive Detection and Differentiation of Endotoxins Derived from Bacterial Pathogens by Surface-Enhanced Raman Scattering. Biosensors, 11(7), 234. https://doi.org/10.3390/bios11070234








