Nanoparticles Surface Chemistry Influence on Protein Corona Composition and Inflammatory Responses
Abstract
1. Introduction
2. Materials and Methods
2.1. Nanoparticle Design and Characterization
2.1.1. Functionalization of Silica Nanoparticles
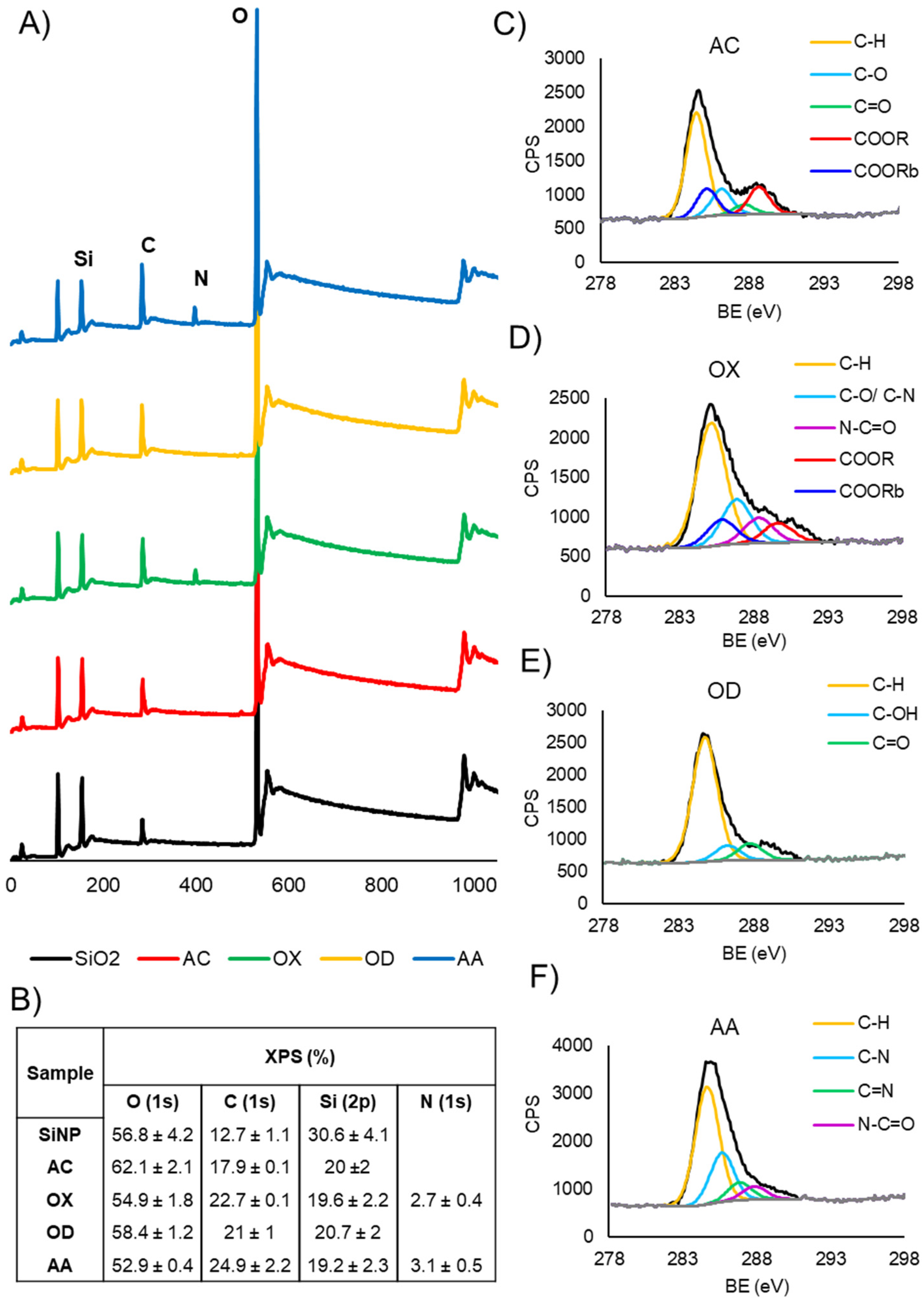
2.1.2. X-ray Photoelectron Spectroscopy (XPS)
2.1.3. Time of Flight Secondary Ion Mass Spectrometry (ToF-SIMS)
2.1.4. Zeta-Potential
2.1.5. Fluorescence Imaging
2.1.6. Transmission Electron Microscopy
2.2. Protein Corona Study
2.2.1. Protein Corona Preparation
2.2.2. SDS Polyacrylamide Gel Electrophoresis (SDS-PAGE)
2.2.3. Digestion of the Protein Corona for Mass Spectrometry Analysis
2.2.4. Liquid Chromatography Mass Spectrometry (LC-MS) Analysis
2.3. Cell Viability
2.4. Immune Response Studies
3. Results and Discussion
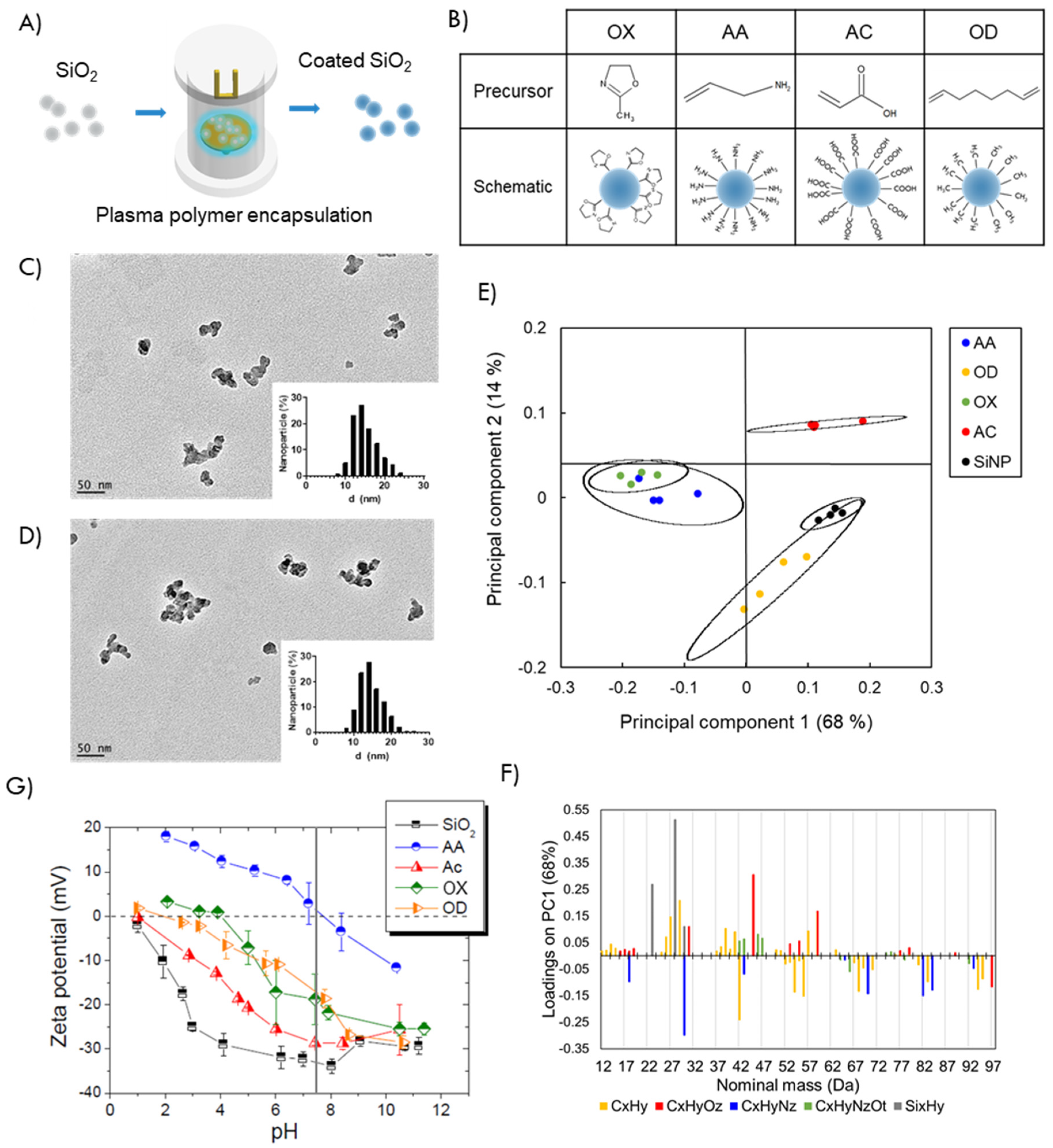
3.1. Tuning Surface Chemistry of Silica Nanoparticles
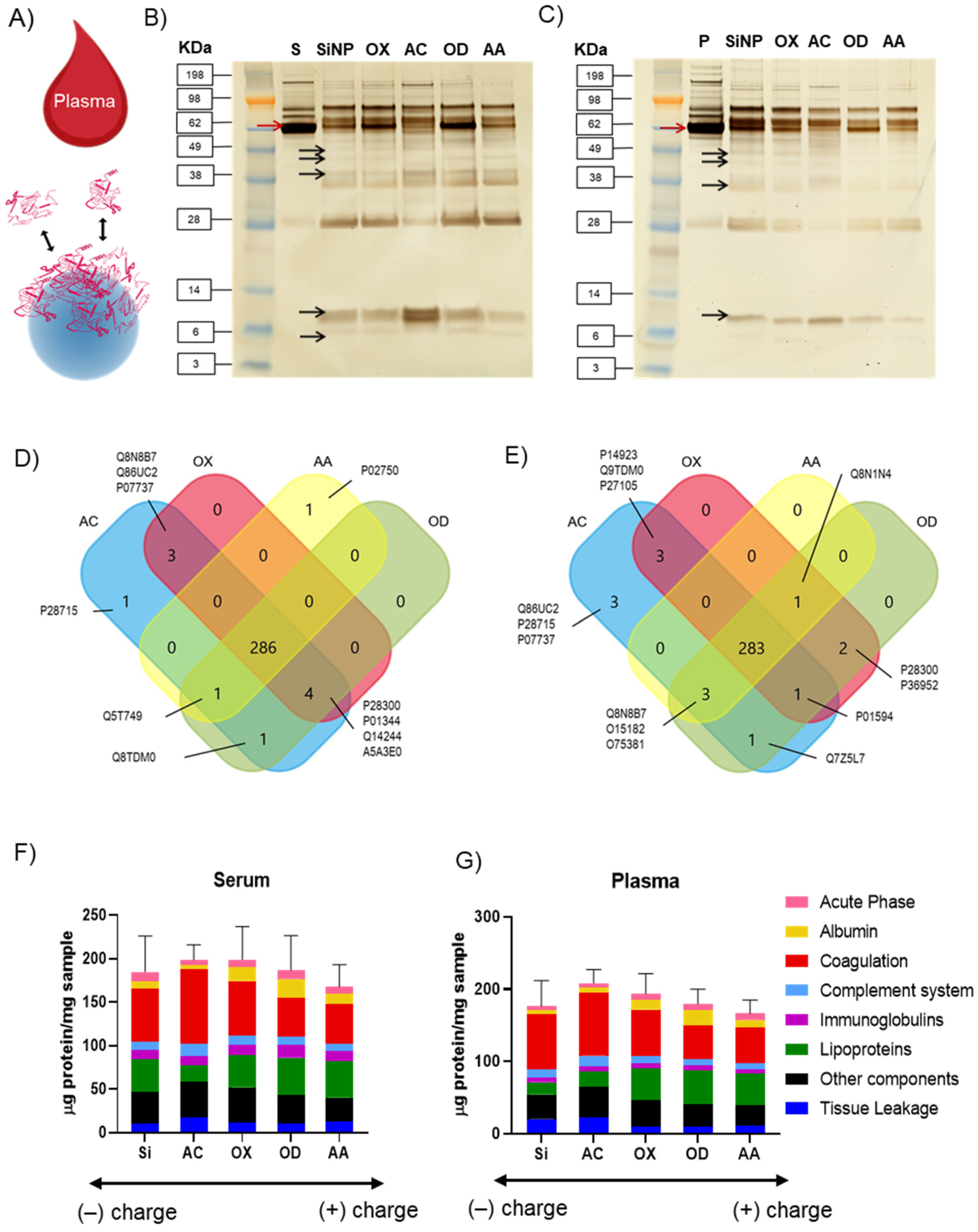
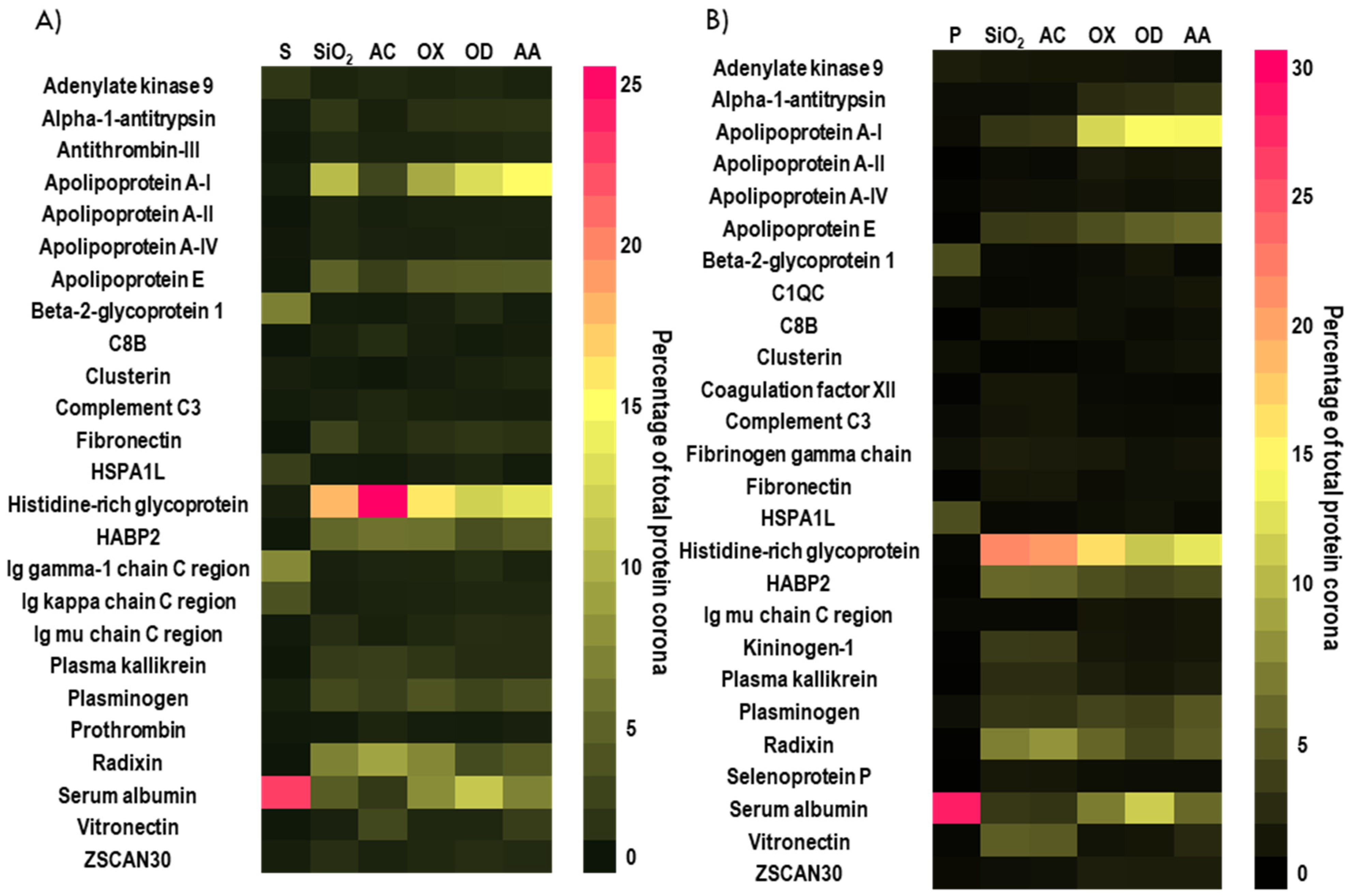
3.2. Influence of Different Plasma Polymer Chemistries on Protein Corona Formation
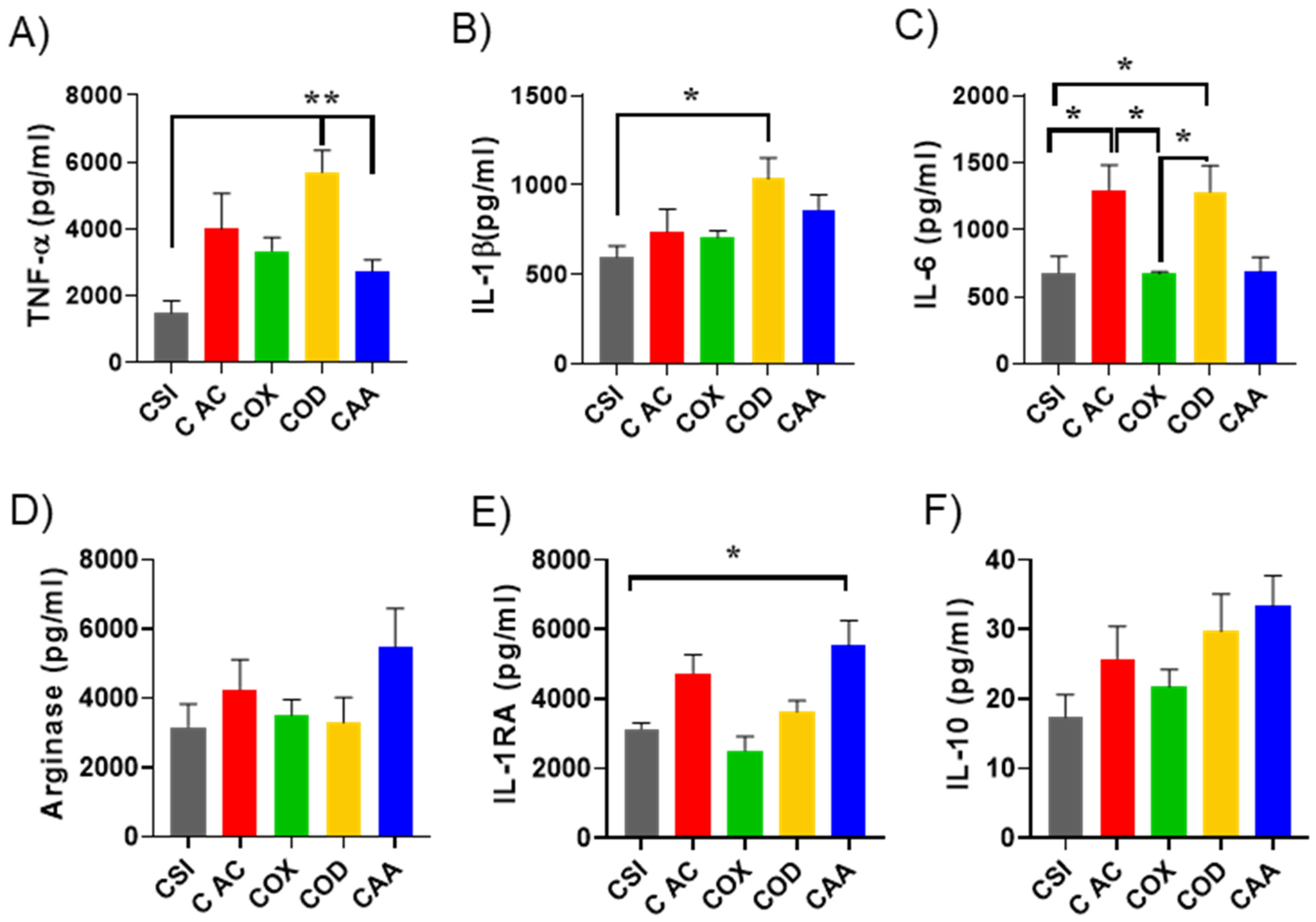
3.3. Effect of Plasma Polymer Surface Modification on Immunomodulatory Properties of Silica Nanoparticles
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Aftab, S.; Shah, A.; Nadhman, A.; Kurbanoglu, S.; Ozkan, S.A.; Dionysiou, D.D.; Shukla, S.S.; Aminabhavi, T.M. Nanomedicine: An effective tool in cancer therapy. Int. J. Pharm. 2018, 540, 132–149. [Google Scholar] [CrossRef]
- Fenton, O.S.; Olafson, K.N.; Pillai, P.S.; Mitchell, M.J.; Langer, R. Advances in Biomaterials for Drug Delivery. Adv. Mater. 2018, 30, e1705328. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, M.F.; Jennings, G.T. Vaccine delivery: A matter of size, geometry, kinetics and molecular patterns. Nat. Rev. Immunol. 2010, 10, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Apostolopoulos, V.; Thalhammer, T.; Tzakos, A.G.; Stojanovska, L. Targeting antigens to dendritic cell receptors for vaccine development. J. Drug Deliv. 2013, 2013, 869718. [Google Scholar] [CrossRef]
- Gheibi Hayat, S.M.; Darroudi, M. Nanovaccine: A novel approach in immunization. J. Cell. Physiol. 2019, 234, 12530–12536. [Google Scholar] [CrossRef] [PubMed]
- Sahdev, P.; Ochyl, L.J.; Moon, J.J. Biomaterials for Nanoparticle Vaccine Delivery Systems. Pharm. Res. 2014, 31, 2563–2582. [Google Scholar] [CrossRef]
- Carnovale, C.; Bryant, G.; Shukla, R.; Bansal, V. Identifying Trends in Gold Nanoparticle Toxicity and Uptake: Size, Shape, Capping Ligand, and Biological Corona. ACS Omega 2019, 4, 242–256. [Google Scholar] [CrossRef]
- Kopp, M.; Kollenda, S.; Epple, M. Nanoparticle-Protein Interactions: Therapeutic Approaches and Supramolecular Chemistry. Acc. Chem. Res. 2017, 50, 1383–1390. [Google Scholar] [CrossRef]
- Tenzer, S.; Docter, D.; Kuharev, J.; Musyanovych, A.; Fetz, V.; Hecht, R.; Schlenk, F.; Fischer, D.; Kiouptsi, K.; Reinhardt, C.; et al. Rapid formation of plasma protein corona critically affects nanoparticle pathophysiology. Nat. Nanotechnol. 2013, 8, 772–781. [Google Scholar] [CrossRef]
- Tonigold, M.; Simon, J.; Estupiñán, D.; Kokkinopoulou, M.; Reinholz, J.; Kintzel, U.; Kaltbeitzel, A.; Renz, P.; Domogalla, M.P.; Steinbrink, K.; et al. Pre-adsorption of antibodies enables targeting of nanocarriers despite a biomolecular corona. Nat. Nanotechnol. 2018, 13, 862–869. [Google Scholar] [CrossRef]
- Schottler, S.; Becker, G.; Winzen, S.; Steinbach, T.; Mohr, K.; Landfester, K.; Mailander, V.; Wurm, F.R. Protein adsorption is required for stealth effect of poly(ethylene glycol)- and poly(phosphoester)-coated nanocarriers. Nat. Nanotechnol. 2016, 11, 372–377. [Google Scholar] [CrossRef]
- García-Álvarez, R.; Hadjidemetriou, M.; Sánchez-Iglesias, A.; Liz-Marzán, L.M.; Kostarelos, K. In vivo formation of protein corona on gold nanoparticles. The effect of their size and shape. Nanoscale 2018, 10, 1256–1264. [Google Scholar] [CrossRef] [PubMed]
- McNamara, L.E.; McMurray, R.J.; Biggs, M.J.P.; Kantawong, F.; Oreffo, R.O.C.; Dalby, M.J. Nanotopographical control of stem cell differentiation. J. Tissue Eng. 2010, 1, 120623. [Google Scholar] [CrossRef]
- Lundqvist, M.; Augustsson, C.; Lilja, M.; Lundkvist, K.; Dahlback, B.; Linse, S.; Cedervall, T. The nanoparticle protein corona formed in human blood or human blood fractions. PLoS ONE 2017, 12, e0175871. [Google Scholar] [CrossRef]
- Mirshafiee, V.; Kim, R.; Mahmoudi, M.; Kraft, M.L. The importance of selecting a proper biological milieu for protein corona analysis in vitro: Human plasma versus human serum. Int. J. Biochem. Cell Biol. 2016, 75, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Brun, E.; Sicard-Roselli, C. Could nanoparticle corona characterization help for biological consequence prediction? Cancer Nanotechnol. 2014, 5, 7. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Hu, Y.; Yu, C.; Boerhan, R.; Jiang, G. Charged surface groups of nanoparticles and the adsorbed proteins codetermine the fate of nanoparticles upon interacting with cells. RSC Adv. 2016, 6, 58315–58324. [Google Scholar] [CrossRef]
- Yue, Z.-G.; Wei, W.; Lv, P.-P.; Yue, H.; Wang, L.-Y.; Su, Z.-G.; Ma, G.-H. Surface Charge Affects Cellular Uptake and Intracellular Trafficking of Chitosan-Based Nanoparticles. Biomacromolecules 2011, 12, 2440–2446. [Google Scholar] [CrossRef]
- Kakizawa, Y.; Lee, J.S.; Bell, B.; Fahmy, T.M. Precise manipulation of biophysical particle parameters enables control of proinflammatory cytokine production in presence of TLR 3 and 4 ligands. Acta Biomater. 2017, 57, 136–145. [Google Scholar] [CrossRef]
- Hwang, T.L.; Aljuffali, I.A.; Lin, C.F.; Chang, Y.T.; Fang, J.Y. Cationic additives in nanosystems activate cytotoxicity and inflammatory response of human neutrophils: Lipid nanoparticles versus polymeric nanoparticles. Int. J. Nanomed. 2015, 10, 371–385. [Google Scholar] [CrossRef]
- Madathiparambil Visalakshan, R.; González García, L.E.; Benzigar, M.R.; Ghazaryan, A.; Simon, J.; Mierczynska-Vasilev, A.; Michl, T.D.; Vinu, A.; Mailänder, V.; Morsbach, S.; et al. The Influence of Nanoparticle Shape on Protein Corona Formation. Small (Weinh. Der Bergstr. Ger.) 2020, 16, 2000285. [Google Scholar] [CrossRef] [PubMed]
- Yan, Y.; Gause, K.T.; Kamphuis, M.M.J.; Ang, C.S.; O’Brien-Simpson, N.M.; Lenzo, J.C.; Reynolds, E.C.; Nice, E.C.; Caruso, F. Differential Roles of the Protein Corona in the Cellular Uptake of Nanoporous Polymer Particles by Monocyte and Macrophage Cell Lines. Acs Nano 2013, 7, 10960–10970. [Google Scholar] [CrossRef] [PubMed]
- Saptarshi, S.R.; Duschl, A.; Lopata, A.L. Interaction of nanoparticles with proteins: Relation to bio-reactivity of the nanoparticle. J. Nanobiotechnol. 2013, 11, 26. [Google Scholar] [CrossRef] [PubMed]
- Anselmo, A.C.; Mitragotri, S. Nanoparticles in the clinic: An update post COVID-19 vaccines. Bioeng. Transl. Med. 2021, 6, e10246. [Google Scholar] [CrossRef]
- Kharlamov, A.N.; Feinstein, J.A.; Cramer, J.A.; Boothroyd, J.A.; Shishkina, E.V.; Shur, V. Plasmonic photothermal therapy of atherosclerosis with nanoparticles: Long-term outcomes and safety in NANOM-FIM trial. Future Cardiol. 2017, 13, 345–363. [Google Scholar] [CrossRef] [PubMed]
- Niu, Y.; Popat, A.; Yu, M.; Karmakar, S.; Gu, W.; Yu, C. Recent advances in the rational design of silica-based nanoparticles for gene therapy. Ther. Deliv. 2012, 3, 1217–1237. [Google Scholar] [CrossRef]
- Tang, L.; Cheng, J. Nonporous silica nanoparticles for nanomedicine application. Nano Today 2013, 8, 290–312. [Google Scholar] [CrossRef]
- Minko, S. Grafting on Solid Surfaces: “Grafting to” and “Grafting from” Methods. In Polymer Surfaces and Interfaces: Characterization, Modification and Applications; Stamm, M., Ed.; Springer: Berlin, Heidelberg, 2008; pp. 215–234. [Google Scholar]
- Mierczynska-Vasilev, A.; Boyer, P.; Vasilev, K.; Smith, P.A. A novel technology for the rapid, selective, magnetic removal of pathogenesis-related proteins from wines. Food Chem. 2017, 232, 508–514. [Google Scholar] [CrossRef]
- Cavallaro, A.; Vasilev, K. Controlled and sustained release of pharmaceuticals via single step solvent-free encapsulation. Chem. Commun. 2015, 51, 1838–1841. [Google Scholar] [CrossRef][Green Version]
- Bachhuka, A.; Christo, S.N.; Cavallaro, A.; Diener, K.R.; Mierczynska, A.; Smith, L.E.; Marian, R.; Manavis, J.; Hayball, J.D.; Vasilev, K. Hybrid core/shell microparticles and their use for understanding biological processes. J. Colloid Interface Sci. 2015, 457, 9–17. [Google Scholar] [CrossRef]
- Goreham, R.V.; Mierczynska, A.; Pierce, M.; Short, R.D.; Taheri, S.; Bachhuka, A.; Cavallaro, A.; Smith, L.E.; Vasilev, K. A substrate independent approach for generation of surface gradients. Thin Solid Film. 2013, 528, 106–110. [Google Scholar] [CrossRef]
- Christo, S.N.; Bachhuka, A.; Diener, K.R.; Mierczynska, A.; Hayball, J.D.; Vasilev, K. The Role of Surface Nanotopography and Chemistry on Primary Neutrophil and Macrophage Cellular Responses. Adv. Healthc. Mater. 2016, 5, 956–965. [Google Scholar] [CrossRef] [PubMed]
- Macgregor-Ramiasa, M.N.; Cavallaro, A.A.; Vasilev, K. Properties and reactivity of polyoxazoline plasma polymer films. J. Mat. Chem. B 2015, 3, 6327–6337. [Google Scholar] [CrossRef] [PubMed]
- Cavallaro, A.; Mierczynska, A.; Barton, M.; Majewski, P.; Vasilev, K. Influence of immobilized quaternary ammonium group surface density on antimicrobial efficacy and cytotoxicity. Biofouling 2016, 32, 13–24. [Google Scholar] [CrossRef]
- Michl, T.D.; Coad, B.R.; Hüsler, A.; Vasilev, K.; Griesser, H.J. Laboratory Scale Systems for the Plasma Treatment and Coating of Particles. Plasma Process. Polym. 2015, 12, 305–313. [Google Scholar] [CrossRef]
- Vasilev, K. Nanoengineered Plasma Polymer Films for Biomaterial Applications. Plasma Chem. Plasma Process. 2014, 34, 545–558. [Google Scholar] [CrossRef]
- Hernandez-Lopez, J.L.; Bauer, R.E.; Chang, W.S.; Glasser, G.; Grebel-Koehler, D.; Klapper, M.; Kreiter, M.; Leclaire, J.; Majoral, J.P.; Mittler, S.; et al. Functional polymers as nanoscopic building blocks. Mater. Sci. Eng. C 2003, 23, 267–274. [Google Scholar] [CrossRef]
- Vasilev, K.; Michelmore, A.; Griesser, H.J.; Short, R.D. Substrate influence on the initial growth phase of plasma-deposited polymer films. Chem. Commun. 2009, 3600–3602. [Google Scholar] [CrossRef]
- Mierczynska-Vasilev, A.; Mierczynski, P.; Maniukiewicz, W.; Visalakshan, R.M.; Vasilev, K.; Smith, P.A. Magnetic separation technology: Functional group efficiency in the removal of haze-forming proteins from wines. Food Chem. 2019, 275, 154–160. [Google Scholar] [CrossRef]
- Sze, A.; Erickson, D.; Ren, L.; Li, D. Zeta-potential measurement using the Smoluchowski equation and the slope of the current–time relationship in electroosmotic flow. J. Colloid Interface Sci. 2003, 261, 402–410. [Google Scholar] [CrossRef]
- Mei, K.C.; Ghazaryan, A.; Teoh, E.Z.; Summers, H.D.; Li, Y.; Ballesteros, B.; Piasecka, J.; Walters, A.; Hider, R.C.; Mailänder, V. Protein-Corona-by-Design in 2D: A Reliable Platform to Decode Bio–Nano Interactions for the Next-Generation Quality-by-Design Nanomedicines. Adv. Mater. 2018, 30, 1802732. [Google Scholar] [CrossRef] [PubMed]
- Tenzer, S.; Docter, D.; Rosfa, S.; Wlodarski, A.; Kuharev, J.; Rekik, A.; Knauer, S.K.; Bantz, C.; Nawroth, T.; Bier, C.; et al. Nanoparticle size is a critical physicochemical determinant of the human blood plasma corona: A comprehensive quantitative proteomic analysis. ACS Nano 2011, 5, 7155–7167. [Google Scholar] [CrossRef]
- Chanput, W.; Mes, J.J.; Savelkoul, H.F.J.; Wichers, H.J. Characterization of polarized THP-1 macrophages and polarizing ability of LPS and food compounds. Food Funct. 2013, 4, 266–276. [Google Scholar] [CrossRef] [PubMed]
- Hopp, I.; MacGregor, M.N.; Doherty, K.; Visalakshan, R.M.; Vasilev, K.; Williams, R.L.; Murray, P. Plasma Polymer Coatings To Direct the Differentiation of Mouse Kidney-Derived Stem Cells into Podocyte and Proximal Tubule-like Cells. ACS Biomater. Sci. Eng. 2019, 5, 2834–2845. [Google Scholar] [CrossRef] [PubMed]
- Ramiasa, M.N.; Cavallaro, A.A.; Mierczynska, A.; Christo, S.N.; Gleadle, J.M.; Hayball, J.D.; Vasilev, K. Plasma polymerised polyoxazoline thin films for biomedical applications. Chem. Commun. 2015, 51, 4279–4282. [Google Scholar] [CrossRef] [PubMed]
- Kessel, A.; Ben-Tal, N. Introduction to Proteins: Structure, Function, and Motion, SECOND EDITION (Chapman & Hall/CRC Mathematical and Computational Biology); CRC Press: Boca Raton, FL, USA, 2018. [Google Scholar]
- Avadiar, L.; Leong, Y.-K.J.C.; Science, P. Interactions of PEI (polyethylenimine)–silica particles with citric acid in dispersions. Colloid Polym. Sci. 2011, 289, 237–245. [Google Scholar] [CrossRef]
- Liu, L.; Guo, Z.; Huang, Z.; Zhuang, J.; Yang, W. Size-selective separation of DNA fragments by using lysine-functionalized silica particles. Sci. Rep. 2016, 6, 22029. [Google Scholar] [CrossRef]
- Mierczynska, A.; Michelmore, A.; Tripathi, A.; Goreham, R.V.; Sedev, R.; Vasilev, K. pH-tunable gradients of wettability and surface potential. Soft Matter 2012, 8, 8399–8404. [Google Scholar] [CrossRef]
- Macgregor, M.N.; Michelmore, A.; Safizadeh Shirazi, H.; Whittle, J.; Vasilev, K. Secrets of Plasma-Deposited Polyoxazoline Functionality Lie in the Plasma Phase. Chem. Mater. 2017, 29, 8047–8051. [Google Scholar] [CrossRef]
- Visalakshan, R.M.; MacGregor, M.N.; Sasidharan, S.; Ghazaryan, A.; Mierczynska-Vasilev, A.M.; Morsbach, S.; Mailander, V.; Landfester, K.; Hayball, J.D.; Vasilev, K. Biomaterial Surface Hydrophobicity-Mediated Serum Protein Adsorption and Immune Responses. ACS Appl. Mater. Interfaces 2019, 11, 27615–27623. [Google Scholar] [CrossRef]
- Hamerli, P.; Weigel, T.; Groth, T.; Paul, D. Surface properties of and cell adhesion onto allylamine-plasma-coated polyethylenterephtalat membranes. Biomaterials 2003, 24, 3989–3999. [Google Scholar] [CrossRef]
- Bullett, N.A.; Whittle, J.D.; Short, R.D.; Douglas, C.W.I. Adsorption of immunoglobulin G to plasma-co-polymer surfaces of acrylic acid and 1,7-octadiene. J. Mater. Chem. 2003, 13, 1546–1553. [Google Scholar] [CrossRef]
- Parry, K.L.; Shard, A.G.; Short, R.D.; White, R.G.; Whittle, J.D.; Wright, A. ARXPS characterisation of plasma polymerised surface chemical gradients. Surf. Interface Anal. 2006, 38, 1497–1504. [Google Scholar] [CrossRef]
- Nguyen, V.H.; Lee, B.J. Protein corona: A new approach for nanomedicine design. Int. J. Nanomed. 2017, 12, 3137–3151. [Google Scholar] [CrossRef] [PubMed]
- Lundqvist, M.; Stigler, J.; Elia, G.; Lynch, I.; Cedervall, T.; Dawson, K.A. Nanoparticle size and surface properties determine the protein corona with possible implications for biological impacts. Proc. Natl. Acad. Sci. USA 2008, 105, 14265–14270. [Google Scholar] [CrossRef] [PubMed]
- Kurtz-Chalot, A.; Villiers, C.; Pourchez, J.; Boudard, D.; Martini, M.; Marche, P.N.; Cottier, M.; Forest, V. Impact of silica nanoparticle surface chemistry on protein corona formation and consequential interactions with biological cells. Mater. Sci. Eng. C 2017, 75, 16–24. [Google Scholar] [CrossRef]
- Maiorano, G.; Sabella, S.; Sorce, B.; Brunetti, V.; Malvindi, M.A.; Cingolani, R.; Pompa, P.P. Effects of Cell Culture Media on the Dynamic Formation of Protein-Nanoparticle Complexes and Influence on the Cellular Response. Acs Nano 2010, 4, 7481–7491. [Google Scholar] [CrossRef]
- Clemments, A.M.; Botella, P.; Landry, C.C. Protein Adsorption From Biofluids on Silica Nanoparticles: Corona Analysis as a Function of Particle Diameter and Porosity. ACS Appl. Mater. Interfaces 2015, 7, 21682–21689. [Google Scholar] [CrossRef]
- Simon, J.; Christmann, S.; Mailander, V.; Wurm, F.R.; Landfester, K. Protein Corona Mediated Stealth Properties of Biocompatible Carbohydrate-based Nanocarriers. Isr. J. Chem. 2018, 58, 1363–1372. [Google Scholar] [CrossRef]
- Gessner, A.; Lieske, A.; Paulke, B.R.; Müller, R.H. Influence of surface charge density on protein adsorption on polymeric nanoparticles: Analysis by two-dimensional electrophoresis. Eur. J. Pharm. Biopharm. 2002, 54, 165–170. [Google Scholar] [CrossRef]
- Visalakshan, R.M.; Cavallaro, A.A.; MacGregor, M.N.; Lawrence, E.P.; Koynov, K.; Hayball, J.D.; Vasilev, K. Nanotopography-Induced Unfolding of Fibrinogen Modulates Leukocyte Binding and Activation. Adv. Funct. Mater. 2019, 29, 1807453. [Google Scholar] [CrossRef]
- Mahmoudi, M.; Lynch, I.; Ejtehadi, M.R.; Monopoli, M.P.; Bombelli, F.B.; Laurent, S. Protein−Nanoparticle Interactions: Opportunities and Challenges. Chem. Rev. 2011, 111, 5610–5637. [Google Scholar] [CrossRef] [PubMed]
- Gessner, A.; Lieske, A.; Paulke, B.R.; Muller, R.H. Functional groups on polystyrene model nanoparticles: Influence on protein adsorption. J. Biomed. Mater. Res. Part A 2003, 65A, 319–326. [Google Scholar] [CrossRef] [PubMed]
- Kozlowski, L.P. Isoelectric point for Swissprot proteins Dec 2015. RepOD 2016. [Google Scholar] [CrossRef]
- Luo, Z.; Hu, Y.; Xin, R.L.; Zhang, B.L.; Li, J.H.; Ding, X.W.; Hou, Y.H.; Yang, L.; Cai, K.Y. Surface functionalized mesoporous silica nanoparticles with natural proteins for reduced immunotoxicity. J. Biomed. Mater. Res. Part A 2014, 102, 3781–3794. [Google Scholar] [CrossRef] [PubMed]
- Digiacomo, L.; Pozzi, D.; Palchetti, S.; Zingoni, A.; Caracciolo, G. Impact of the protein corona on nanomaterial immune response and targeting ability. Wiley Interdiscip. Rev. Nanomed. Nanobiotechnol. 2020, 12, e1615. [Google Scholar] [CrossRef]
- Bachhuka, A.; Hayball, J.; Smith, L.E.; Vasilev, K. Effect of Surface Chemical Functionalities on Collagen Deposition by Primary Human Dermal Fibroblasts. ACS Appl. Mater. Interfaces 2015, 7, 23767–23775. [Google Scholar] [CrossRef]
- Fedeli, C.; Segat, D.; Tavano, R.; Bubacco, L.; De Franceschi, G.; de Laureto, P.P.; Lubian, E.; Selvestrel, F.; Mancin, F.; Papini, E. The functional dissection of the plasma corona of SiO2-NPs spots histidine rich glycoprotein as a major player able to hamper nanoparticle capture by macrophages. Nanoscale 2015, 7, 17710–17728. [Google Scholar] [CrossRef]
- Jansen, R.W.; Molema, G.; Harms, G.; Kruijt, J.K.; van Berkel, T.J.C.; Hardonk, M.J.; Meijer, D.K.F. Formaldehyde treated albumin contains monomeric and polymeric forms that are differently cleared by endothelial and kupffer cells of the liver: Evidence for scavenger receptor heterogeneity. Biochem. Biophys. Res. Commun. 1991, 180, 23–32. [Google Scholar] [CrossRef]
- Mortimer, G.M.; Butcher, N.J.; Musumeci, A.W.; Deng, Z.J.; Martin, D.J.; Minchin, R.F. Cryptic Epitopes of Albumin Determine Mononuclear Phagocyte System Clearance of Nanomaterials. ACS Nano 2014, 8, 3357–3366. [Google Scholar] [CrossRef]
- Zhang, X.H.; Kimura, Y.; Fang, C.Y.; Zhou, L.; Sfyroera, G.; Lambris, J.D.; Wetsel, R.A.; Miwa, T.; Song, W.C. Regulation of Toll-like receptor-mediated inflammatory response by complement in vivo. Blood 2007, 110, 228–236. [Google Scholar] [CrossRef]
- Vuilleumier, N.; Dayer, J.M.; von Eckardstein, A.; Roux-Lombard, P. Pro- or anti-inflammatory role of apolipoprotein A-1 in high-density lipoproteins? Swiss Med. Wkly. 2013, 143, w13781. [Google Scholar] [CrossRef] [PubMed]
- Umemoto, T.; Han, C.Y.; Mitra, P.; Averill, M.M.; Tang, C.; Goodspeed, L.; Omer, M.; Subramanian, S.; Wang, S.; Den Hartigh, L.J.; et al. Apolipoprotein AI and high-density lipoprotein have anti-inflammatory effects on adipocytes via cholesterol transporters: ATP-binding cassette A-1, ATP-binding cassette G-1, and scavenger receptor B-1. Circ. Res. 2013, 112, 1345–1354. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez Garcia, L.E.; MacGregor-Ramiasa, M.; Visalakshan, R.M.; Vasilev, K. Protein Interactions with Nanoengineered Polyoxazoline Surfaces Generated via Plasma Deposition. Langmuir 2017, 33, 7322–7331. [Google Scholar] [CrossRef] [PubMed]
- Saboohi, S.; Griesser, H.J.; Coad, B.R.; Short, R.D.; Michelmore, A. Promiscuous hydrogen in polymerising plasmas. Phys. Chem. Chem. Phys. 2018, 20, 7033–7042. [Google Scholar] [CrossRef]
- Bachhuka, A.; Heng, S.; Vasilev, K.; Kostechi, R.; Abell, A.; Ebendorff-Heidepriem, H. Surface functionalization of exposed core glass optical fiber for metal ion sensing. Sensors 2019, 19, 1829. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
González-García, L.E.; MacGregor, M.N.; Visalakshan, R.M.; Lazarian, A.; Cavallaro, A.A.; Morsbach, S.; Mierczynska-Vasilev, A.; Mailänder, V.; Landfester, K.; Vasilev, K. Nanoparticles Surface Chemistry Influence on Protein Corona Composition and Inflammatory Responses. Nanomaterials 2022, 12, 682. https://doi.org/10.3390/nano12040682
González-García LE, MacGregor MN, Visalakshan RM, Lazarian A, Cavallaro AA, Morsbach S, Mierczynska-Vasilev A, Mailänder V, Landfester K, Vasilev K. Nanoparticles Surface Chemistry Influence on Protein Corona Composition and Inflammatory Responses. Nanomaterials. 2022; 12(4):682. https://doi.org/10.3390/nano12040682
Chicago/Turabian StyleGonzález-García, Laura E., Melanie N. MacGregor, Rahul M. Visalakshan, Artur Lazarian, Alex A. Cavallaro, Svenja Morsbach, Agnieszka Mierczynska-Vasilev, Volker Mailänder, Katharina Landfester, and Krasimir Vasilev. 2022. "Nanoparticles Surface Chemistry Influence on Protein Corona Composition and Inflammatory Responses" Nanomaterials 12, no. 4: 682. https://doi.org/10.3390/nano12040682
APA StyleGonzález-García, L. E., MacGregor, M. N., Visalakshan, R. M., Lazarian, A., Cavallaro, A. A., Morsbach, S., Mierczynska-Vasilev, A., Mailänder, V., Landfester, K., & Vasilev, K. (2022). Nanoparticles Surface Chemistry Influence on Protein Corona Composition and Inflammatory Responses. Nanomaterials, 12(4), 682. https://doi.org/10.3390/nano12040682






