4.1. Standard Model: ACE Simplex
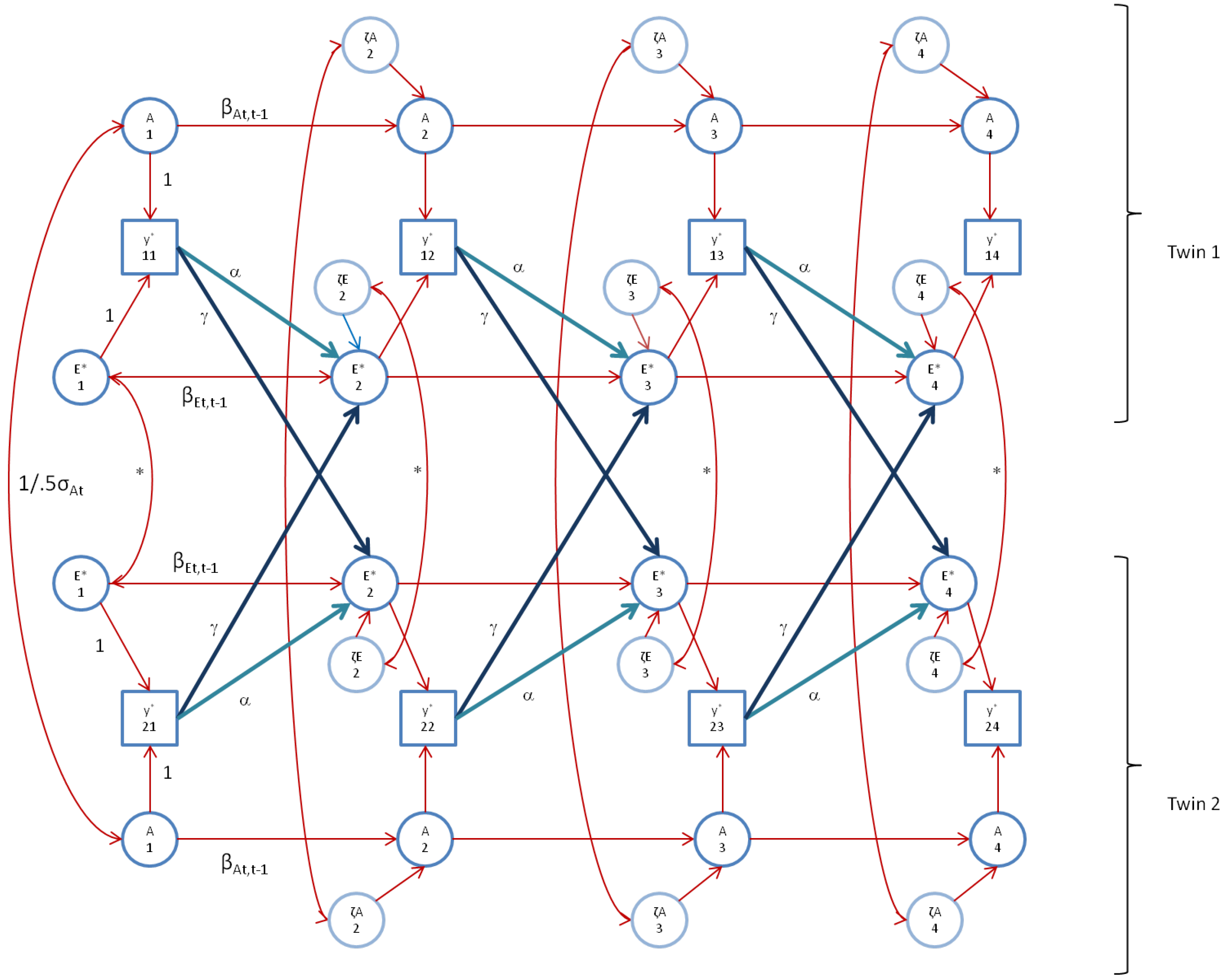
We first fitted the standard genetic simplex model, as depicted in
Figure 1 (note that, to avoid clutter,
Figure 1 does not include occasion specific effects). As mentioned above, we constrained the occasion specific genetic and environmental variances to be equal over time (var(a
t) = var(a), var(c
t) = var(c), var(e
t) = var(e), t=1,...,4). For FSIQ, the standard simplex model fitted the data well (χ
2(60) = 76.8; TLI = 0.99; adjBIC = 21,470; BIC = 21,559; RMSEA = 0.032 (CI95: 0.0–0.051)). The time-specific genetic variance (denoted var(a) above) (1 parameter) and the variances var(ζ
A3) and var(ζ
A4) (two parameters) were estimated at zero. They were therefore fixed to zero, and the model refitted: χ
2(63) = 76.8; TLI = 0.99; adjBIC = 21,460; BIC = 21,540; RMSEA = 0.028 (CI95: 0.0–0.048). The fact that the χ
2 is unchanged is consistent with the fact that the fixed variances were estimated as zero. Although some parameters were not significant (at α = 0.01, say) in this model, we made no effort to further prune the model. In the analysis of perfIQ, we obtained similar results. The standard simplex model fitted the data well (χ
2(60) = 56.9; TLI > 0.99; adjBIC = 18,496; BIC = 18,585; RMSEA = 0.0 (CI95: 0.0–0.035)). The time-specific genetic variance was zero (one parameter), as were var(ζ
A2), var(ζ
A3) (two parameters), and var(ζ
C2), var(ζ
C3), var(ζ
C4), and var(ζ
E3) (four parameters). We refitted the model with these parameters fixed to zero, and obtained χ
2(67) = 56.9, TLI>0.99, adjBIC = 18,475, BIC = 18,541, and RMSEA = 0.0 (CI95: 0.0–0.025).
Based on the parameter estimates, we calculated genetic and environmental covariance matrices, and the standardized variance components (
Table 1). These results are quite consistent with the literature [
32,
33,
34,
35,
36]. For instance, the relative contribution of additive genetic influences to the phenotypic (co-) variance increases (FSIQ: 0.27–0.54; perfIQ: 0.29–0.53), and that of shared environmental influences decreases (FSIQ: 0.50–0.24 perfIQ: 0.36–0.18). Genetic influences are highly stable (genetic correlations 0.801–1) in contrast to unshared environmental influences (correlations 0.002–0.205). We note that the results obtained with FSIQ and perfIQ are quite similar.
Table 1.
Standard ACE simplex model results of full scale IQ (FISQ) and performance IQ (perfIQ): Correlations among A, E, and C, standard deviations, variances and standardized variance. Note in this model A, C, and E are uncorrelated.
Table 1.
Standard ACE simplex model results of full scale IQ (FISQ) and performance IQ (perfIQ): Correlations among A, E, and C, standard deviations, variances and standardized variance. Note in this model A, C, and E are uncorrelated.
| | FSIQ | perfIQ |
|---|
| Age in years: | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 |
|---|
| A | A |
| Correlations | 1.000 | | | | 1.000 | | | |
| 0.801 | 1.000 | | | 1.000 | 1.000 | | |
| 0.801 | 1.000 | 1.000 | | 1.000 | 1.000 | 1.000 | |
| 0.801 | 1.000 | 1.000 | 1.000 | 0.959 | 0.959 | 0.959 | 1.000 |
| Standard deviation | 7.69 | 10.21 | 11.84 | 10.52 | 7.58 | 11.67 | 12.38 | 12.37 |
| Variance | 59.1 | 104.3 | 140.1 | 110.8 | 57.4 | 136.2 | 153.3 | 152.9 |
| Standardized variance | 0.27 | 0.48 | 0.60 | 0.54 | 0.29 | 0.50 | 0.60 | 0.53 |
| E | E |
| Correlations | 1.000 | | | | 1.000 | | | |
| 0.112 | 1.000 | | | 0.077 | 1.000 | | |
| 0.056 | 0.212 | 1.000 | | 0.048 | 0.205 | 1.000 | |
| 0.030 | 0.115 | 0.119 | 1.000 | 0.002 | 0.008 | 0.005 | 1.000 |
| Standard deviation | 7.02 | 7.93 | 6.80 | 6.78 | 8.35 | 9.41 | 8.28 | 9.18 |
| Variance | 49.3 | 62.9 | 46.2 | 46.0 | 69.6 | 88.5 | 68.5 | 84.2 |
| Standardized variance | 0.23 | 0.29 | 0.20 | 0.22 | 0.35 | 0.32 | 0.27 | 0.29 |
| C | C |
| Correlations | 1.000 | | | | 1.000 | | | |
| 0.756 | 1.000 | | | 0.760 | 1.000 | | |
| 0.457 | 0.478 | 1.000 | | 0.697 | 0.658 | 1.000 | |
| 0.400 | 0.418 | 0.675 | 1.000 | 0.768 | 0.724 | 0.665 | 1.000 |
| Standard deviation | 10.39 | 7.08 | 6.78 | 7.05 | 8.39 | 6.96 | 5.88 | 7.16 |
| Variance | 107.8 | 50.1 | 46.0 | 49.8 | 70.5 | 48.5 | 34.6 | 51.3 |
| Standardized variance | 0.50 | 0.23 | 0.20 | 0.24 | 0.36 | 0.18 | 0.14 | 0.18 |
4.2. Alternative Model: AE* Simplex with Ph->E Transmission
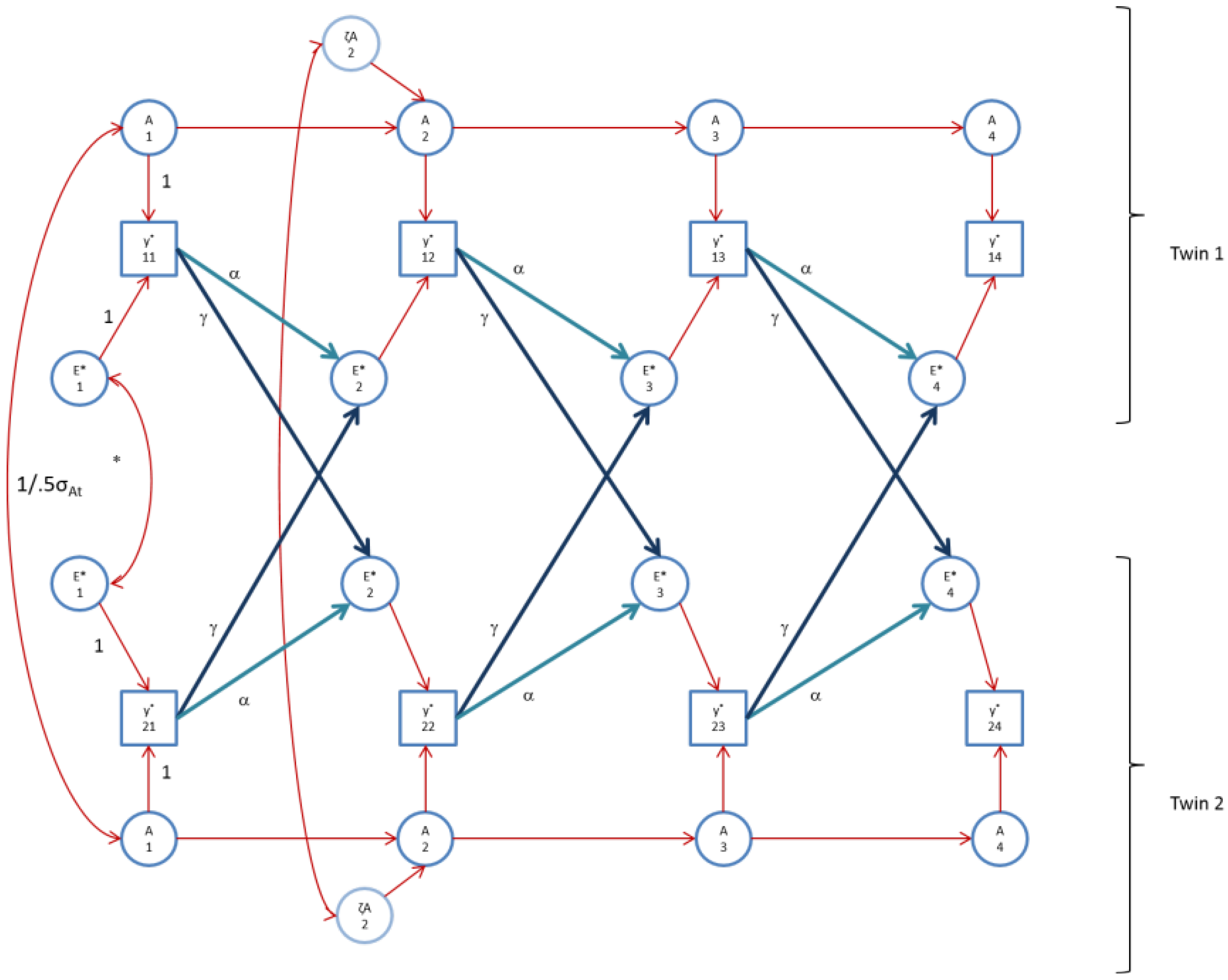
We fitted the alternative model, the AE* simplex model with Ph->E transmission, as depicted in
Figure 2 (
Figure 2 does not display the occasion specific effects). As above, we constrained the occasion specific genetic and environmental variances to be equal over time. The alternative simplex model fitted the FSIQ data well (χ
2(61) = 73.3; TLI>0.99; adjBIC = 21,463; BIC = 21,549; RMSEA = 0.027 (CI95: 0.0–0.047)). However, in this model, some variances were estimated at zero. We reduced the model in the following way. First, as in the standard model, the variance var(a), and the variances var(ζ
A3) and var(ζ
A4) were estimated at zero, so we fixed these to zero, (three parameters). In addition, the unshared environmental auto-regression coefficients (β
Et,t-1) were near (one time) zero or (two times) negative (
i.e., hard to interpret), and none were significant, so we fixed these to zero (three parameters). Finally, the correlations between ζ
E*1t and ζ
E*2t at t = 2, 3, 4 were not significant, so we fixed these to zero (three parameters). The goodness of fit indices of the resulting model are χ
2(70) = 74.9, LI>0.99, adjBIC = 21,463, BIC = 21,494, and RMSEA = 0.016 (CI95: 0.0–0.039). The small increase in χ
2(9) =1.6) underlines the non-significance of the dropped parameters. The genetic autoregressive coefficients equaled 0.761 (s.e. 0.276), 1.062 (s.e. 0.133), and 0.772 (s.e. 0.124). We constrained these to be equal merely to facilitate subsequent calculations (see below). This resulted in χ
2(72) = 77.379, TLI>0.99, adjBIC = 21,432, BIC = 21,483, and RMSEA = 0.016 (CI95: 0–0.039), and a test for the equality constraints of χ
2(2) = 2.48 (
p = 0.29), which suggests that the equality constraint is tenable.
Table 2 contains the parameter estimates of this model,
Table 3 contains the standardized variance components, and
Table 4 contains the covariance matrices of A and E*, based on the estimates in
Table 2.
Figure 3 shows the model as fitted. This final model gives rise to an AE* correlation equaling 0 (t = 1), 0.32 (t = 2), 0.56 (t = 3) and 0.66 (t = 4). As formulated, at time point 1, we assume GE-cov = 0. However, within the final model, the within twin member correlation between A
i1 and E*
i1 is identified. To evaluate whether our initial assumption is correct, we refitted the model to estimate cor(A
1,E*
1). Based on the Wald test, this parameter showed not to be significant (cor(A
1,E*
1) = −0.018, s.e. 0.032).
Table 2.
FIML parameters and standard errors (s.e.) of the AE* model with Ph->E transmission as fitted to FSIQ and perfIQ. 0* indicates that the parameter value was fixed to zero.
Table 2.
FIML parameters and standard errors (s.e.) of the AE* model with Ph->E transmission as fitted to FSIQ and perfIQ. 0* indicates that the parameter value was fixed to zero.
| Parameter | FSIQ ML Estimate (s.e.) | perfIQ ML Estimate (s.e.) |
|---|
| α1 = α2 = α3 | 0.385 (0.13) | 0.482 (0.147) |
| γ1 = γ2 = γ3 | 0.070 (0.03) | 0.037 (0.028) |
| var(ζE*1)0.5 = var(E*1)0.5 | 10.18 (0.74) | 5.68 (1.301) |
| var(ζE*2)0.5 | 5.644 (1.39) | 5.46 (1.010) |
| var(ζE*3)0.5 | 4.197 (2.00) | 0* |
| var(ζE*4)0.5 | 3.857 (2.00) | 4.819 (0.974) |
| cor(E*11 E*21) | 0.804 (0.12) | 0.765 (0.249) |
| var(ζA1)0.5 = var(A1)0.5 | 7.964 (0.80) | 9.041 (0.813) |
| var(ζA2)0.5 | 4.185 (1.10) | 0* |
| var(ζA3)0.5 | 0* | 0* |
| var(ζA4)0.5 | 0* | 4.482 (1.471) |
| βAt,t-1 (t = 2,3,4) | 0.895 (0.06) | 0.877 (0.095) |
| var(e)0.5 | 5.350 (1.32) | 7.763 (0.340) |
| var(c)0.5 | 4.277 (0.35) | 3.620 (0.620) |
Table 3.
Decomposition of phenotypic variance into A, E*, AE*-cov and C components. The standardized components are given in parentheses. Note that here var(E*) includes both var(ζE) and var(e) and C is limited to time specific effects (var(c)). Additive genetic time specific effects are absent.
Table 3.
Decomposition of phenotypic variance into A, E*, AE*-cov and C components. The standardized components are given in parentheses. Note that here var(E*) includes both var(ζE) and var(e) and C is limited to time specific effects (var(c)). Additive genetic time specific effects are absent.
| Age | var(FSIQ) | var(A) | var(E*) | var(C) (Time Specific) | 2cov(AE*) | 2cov(AE*) + var(A) |
|---|
| 5.5 years | 214.1 | 63.43 (0.30) | 132.40 (0.62) | 18.29 (0.09) | 0 (0) | 63.43 (0.30) |
| 6.8 years | 226.5 | 68.3 (0.30) | 92.29 (0.41) | 18.29 (0.08) | 47.6 (0.21) | 115.9 (0.51) |
| 7.9 years | 221.3 | 54.7 (0.25) | 78.63 (0.36) | 18.29 (0.08) | 69.6 (0.31) | 124.3 (0.56) |
| 12.2 years | 205.5 | 43.8 (0.21) | 75.39 (0.37) | 18.29 (0.09) | 67.8 (0.33) | 111.6 (0.54) |
| Age | var(perfIQ) | var(A) | var(E*) | var(C) (Time Specific) | 2cov(AE*) | 2cov(AE*) + var(A) |
| 5.5 years | 187.4 | 81.7 (0.44) | 92.6 (0.49) | 13.1 (0.07) | 0 (0) | 81.7 (0.44) |
| 6.8 years | 266.8 | 62.8 (0.24) | 119.1 (0.45) | 13.1 (0.05) | 71.7 (0.27) | 134.5 (0.51) |
| 7.9 years | 256.9 | 48.3 (0.19) | 108.7 (0.42) | 13.1 (0.05) | 86.8 (0.34) | 135.1 (0.53) |
| 12.2 years | 281.1 | 57.3 (0.20) | 130.0 (0.46) | 13.1 (0.05) | 80.7 (0.29) | 138.0 (0.49) |
In the analysis of perfIQ, we fitted the model with the time-specific genetic variances (one parameter), and var(ζ
A2), var(ζ
A3) (two parameters), and the correlations between ζ
E*1t and ζ
E*2t at t = 2,3,4 fixed to zero. The model fitted well (χ
2(67) = 59.7; TLI > 0.99; adjBIC = 18,477; BIC = 18,544;RMSEA = 0.0 (CI95: 0.0–0.029)). However, as in FSIQ, the unshared environmental auto-regression coefficients (β
Et,t-1) were not significant, and var(ζ
E3) was zero. We dropped these parameters to arrive at χ
2(71) = 60.43, TLI > 0.99, adjBIC = 18,466, BIC = 18,520, and RMSEA = 0.0 (CI95: 0.0–0.024). The genetic autoregressive coefficients equaled 0.791 (s.e. 0.189), 0.787 (s.e. 0.194), and 0.977 (s.e. 0.178). As above, we constrained these to be equal to facilitate subsequent calculations (see below). This resulted in χ
2(73) = 61.1, TLI>0.99, adjBIC = 18,460, BIC = 18,508, and RMSEA = 0.0 (CI95: 0–0.022), and a test for the equality constraints of χ
2(2) = 0.75 (
p = 0.69). The results are given in
Table 2,
Table 3, and
Table 5.
Figure 4 depicts the model as fitted. This final model gives rise to an AE* correlation equaling 0 (t = 1), 0.41 (t = 2), 0.60 (t = 3) and 0.47 (t = 4). We estimated cor(A
1, E*
1), and found this correlation to be zero (cor(A
1, E*
1) = −0.002, s.e. 0.078).
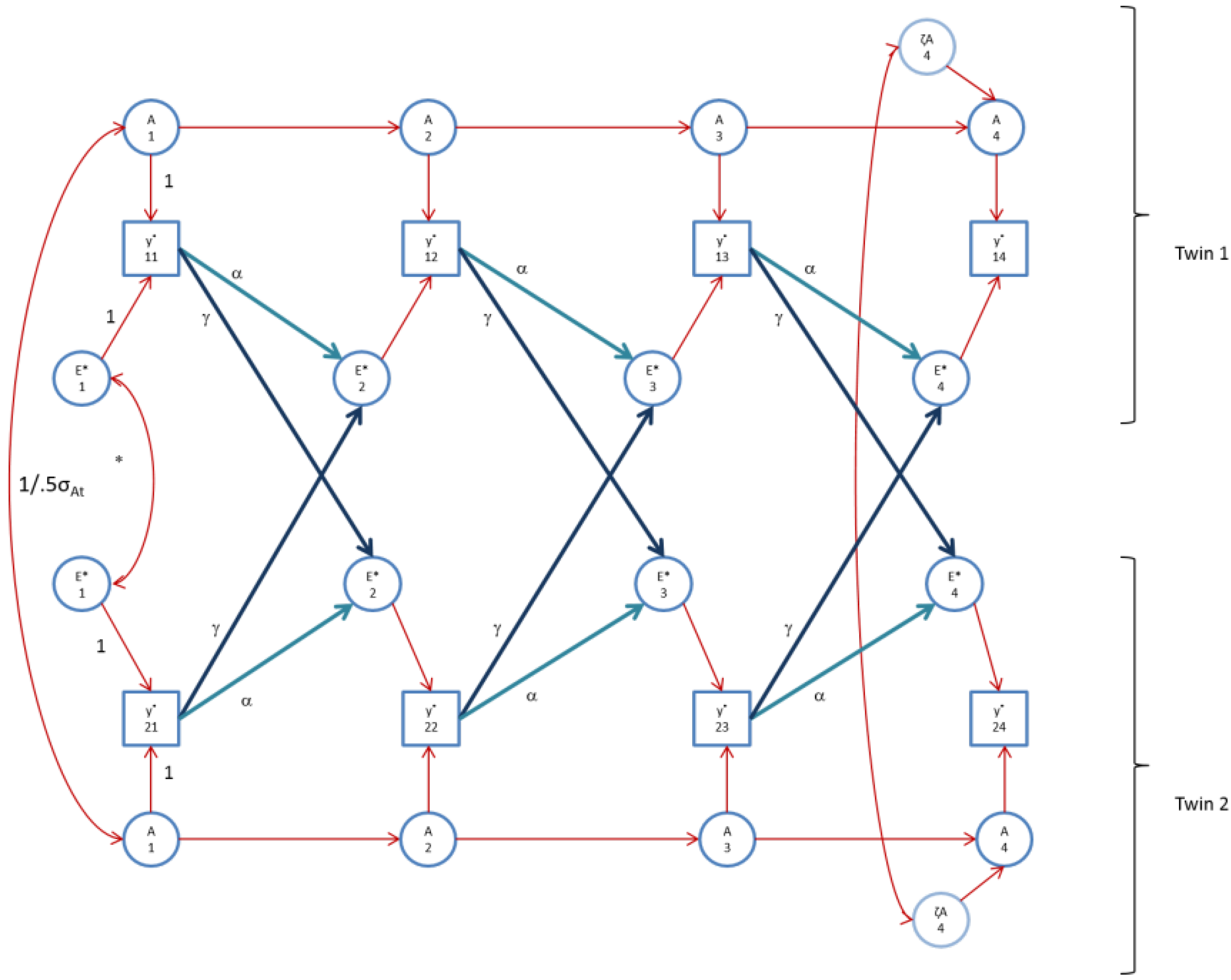
Figure 3.
The AE* simplex model with Ph->E transmission as fitted to FSIQ. Time specific effects are not shown. Note that in FSIQ, time specific effects were found to be environmental (time specific genetic variance was zero).
Figure 3.
The AE* simplex model with Ph->E transmission as fitted to FSIQ. Time specific effects are not shown. Note that in FSIQ, time specific effects were found to be environmental (time specific genetic variance was zero).
Figure 4.
The AE* simplex model with Ph->E transmission as fitted to perfIQ. Time specific effects are not shown. Note that in perfIQ, time specific effects were found to be environmental (time specific genetic variance was zero).
Figure 4.
The AE* simplex model with Ph->E transmission as fitted to perfIQ. Time specific effects are not shown. Note that in perfIQ, time specific effects were found to be environmental (time specific genetic variance was zero).
The standard ACE simplex model and the AE* simplex with Ph->E transmission both fitted well judging by the fit indices. The information criteria BIC and adjBIC favored the alternative model, although the differences are not large, and the models are not nested (FSIQ: adjBIC: 21,460 vs. 21,432 and 21,540 vs. 21,483; perfIQ: 18,475 vs. 18,460 and 18,541 vs. 18,508). Based on the similarity between the fit indices, we conclude that the extended AE* model fits the data adequately relative to the standard ACE model, and therefore is compatible with the standard ACE model.
Comparison of the standardized variance components as given in
Table 1 and
Table 3 shows that the two models also decompose the phenotypic variance in a similar fashion. Consider the decomposition of FSIQ at age 12.2 years. In the standard model, the standardized components are 0.54 (A), 0.22 (E), and 0.24 (C). In the alternative model, the standard components are 0.21 (A), 0.37 (E*), 0.09 (C; occasion specific), and 0.33 (2∙cov(AE*)). While the models differ in the details of the decomposition, they are identical in terms of the total environmental variance and total genetic variance. The total environmental variance in the standard model equals 0.22 (E) + 0.24 (C) = 0.46 whereas in the alternative model the total environmental variance equals 0.37 (E*) + 0.09 (C occasion specific) = 0.46. Similarly, the total genetic variance within standard model equals 0.54 (A), whereas in the alternative model this variance equals 0.21 (A) + 0.33 (2∙cov(AE*)) = 0.54, if we treat 2∙cov(AE*) as genetic variance [
37].
Additionally, we see that the correlation between the environments of the twins shows a similar pattern as the standardized variance component C. For example, consider the correlations in E* for FSIQ which are 0.63 (5.5 years), 0.29 (6.8 years), 0.30 (9.7 years) and 0.32 (12.2 years), whereas the standardized C components in the standard model 0.50 (5.5 years), 0.23 (6.8 years), 0.20 (9.7 years) and 0.24 (12.2 years). It should be noted that in the alternative model, after 5.5 years the correlation in E* is influenced by the initial correlation in environment at time point 1 plus the influence of Ph->E transmission over time. The influence this transmission (i.e., the AE correlation in FSIQ) becomes more prominent over time: 0 (5.5 years), 0.30 (6.8 years), 0.53 (9.7 years), and 0.57 (12.2 years). These observations show that traditional heritability estimates can be decomposed in direct and indirect genetic effects. As such, it raises the question of which decomposition (the standard ACE or the AE* with Ph->E transmission) leads to greater insight in the developmental process. Here, we provisionally accept the alternative model, as we can utilize the results to answer the hypothetical question: What increases in phenotypic mean at age 12.2 years can we expect from changes in environmental means in combination with changes in Ph->E transmission?
Table 4.
MZ covariance matrix of A and E* in the AE* model of FSIQ with Ph->E transmission. These include the AE* covariance and AE* correlation within twins (bold) and between twins (bold italics) and the covariance and correlations among the twin environments (italics). Age is given in years.
Table 4.
MZ covariance matrix of A and E* in the AE* model of FSIQ with Ph->E transmission. These include the AE* covariance and AE* correlation within twins (bold) and between twins (bold italics) and the covariance and correlations among the twin environments (italics). Age is given in years.
| | Correlation | Covariance |
|---|
| | A Twin 1 | E* Twin 1 | A Twin 2 | E* Twin 2 |
|---|
| Age in years | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 |
|---|
| A twin 1 | 1 | | | | | | | | 31.71 | | | | | | | |
| 0.86 | 1 | | | | | | | 28.38 | 34.16 | | | | | | |
| 0.86 | 1.00 | 1 | | | | | | 25.40 | 30.57 | 27.36 | | | | | |
| 0.86 | 1.00 | 1.00 | 1 | | | | | 22.74 | 27.36 | 24.49 | 21.92 | | | | |
| E twin 1 | 0.00 | 0.00 | 0.00 | 0.00 | 1 | | | | 0.00 | 0.00 | 0.00 | 0.00 | 83.44 | | | |
| 0.35 | 0.30 | 0.30 | 0.30 | 0.41 | 1 | | | 16.65 | 14.90 | 13.34 | 11.94 | 39.39 | 26.64 | | |
| 0.50 | 0.53 | 0.53 | 0.53 | 0.20 | 0.43 | 1 | | 23.18 | 25.34 | 22.68 | 20.30 | 18.37 | 22.12 | 23.56 | |
| 0.53 | 0.59 | 0.59 | 0.59 | 0.09 | 0.30 | 0.47 | 1 | 24.73 | 28.53 | 25.53 | 22.85 | 8.50 | 17.71 | 23.74 | 24.14 |
| A twin 2 | 0.50 | | | | | | | | 63.43 | | | | | | | |
| 0.43 | 0.50 | | | | | | | 56.77 | 68.32 | | | | | | |
| 0.43 | 0.50 | 0.50 | | | | | | 50.81 | 61.15 | 54.73 | | | | | |
| 0.43 | 0.50 | 0.50 | 0.50 | | | | | 45.47 | 54.73 | 48.98 | 43.84 | | | | |
| E twin 2 | 0.00 | 0.00 | 0.00 | 0.00 | 0.63 | | | | 0.00 | 0.00 | 0.00 | 0.00 | 132.40 | | | |
| 0.22 | 0.19 | 0.19 | 0.19 | 0.36 | 0.29 | | | 26.64 | 23.84 | 21.34 | 19.10 | 45.79 | 92.29 | | |
| 0.33 | 0.35 | 0.35 | 0.35 | 0.18 | 0.26 | 0.30 | | 35.26 | 38.92 | 34.83 | 31.17 | 20.39 | 36.60 | 78.63 | |
| 0.36 | 0.40 | 0.40 | 0.40 | 0.09 | 0.21 | 0.31 | 0.32 | 36.54 | 42.44 | 37.98 | 33.99 | 9.14 | 24.79 | 35.90 | 75.39 |
Table 5.
DZ covariance matrix of A and E* in the AE* model of perfIQ with Ph->E transmission. These include the AE* covariance and AE* correlation within twins (bold italics) and between twins (bold italics) and the covariance and correlations among the twin environments (italics), which mimics the presence of shared variance. Age is given in years.
Table 5.
DZ covariance matrix of A and E* in the AE* model of perfIQ with Ph->E transmission. These include the AE* covariance and AE* correlation within twins (bold italics) and between twins (bold italics) and the covariance and correlations among the twin environments (italics), which mimics the presence of shared variance. Age is given in years.
| | Correlation | Covariance |
|---|
| | A Twin 1 | E* Twin 1 | A Twin 2 | E* Twin 2 |
|---|
| Age in years | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 | 5.5 | 6.8 | 9.7 | 12.2 |
|---|
| A twin 1 | 1.00 | | | | | | | | 40.87 | | | | | | | |
| 1.00 | 1.00 | | | | | | | 35.84 | 31.43 | | | | | | |
| 1.00 | 1.00 | 1.00 | | | | | | 31.43 | 27.57 | 24.18 | | | | | |
| 0.81 | 0.81 | 0.81 | 1.00 | | | | | 27.57 | 24.18 | 21.20 | 28.64 | | | | |
| E twin 1 | 0.00 | 0.00 | 0.00 | 0.00 | 1.00 | | | | 0.00 | 0.00 | 0.00 | 0.00 | 24.74 | | | |
| 0.41 | 0.41 | 0.41 | 0.33 | 0.16 | 1.00 | | | 22.72 | 19.93 | 17.48 | 15.33 | 13.12 | 19.40 | | |
| 0.60 | 0.60 | 0.60 | 0.48 | 0.08 | 0.41 | 1.00 | | 32.40 | 28.41 | 24.92 | 21.85 | 6.94 | 22.46 | 28.10 | |
| 0.58 | 0.58 | 0.58 | 0.47 | 0.04 | 0.32 | 0.39 | 1.00 | 35.18 | 30.85 | 27.06 | 23.73 | 3.66 | 22.16 | 28.95 | 30.41 |
| A twin 2 | 0.50 | | | | | | | | 81.74 | | | | | | | |
| 0.50 | 0.50 | | | | | | | 71.69 | 62.87 | | | | | | |
| 0.50 | 0.50 | 0.50 | | | | | | 62.87 | 55.14 | 48.35 | | | | | |
| 0.40 | 0.40 | 0.40 | 0.50 | | | | | 55.14 | 48.35 | 42.41 | 57.28 | | | | |
| E twin 2 | 0.00 | 0.00 | 0.00 | 0.00 | 0.27 | | | | 0.00 | 0.00 | 0.00 | 0.00 | 92.61 | | | |
| 0.23 | 0.23 | 0.23 | 0.19 | 0.12 | 0.16 | | | 40.91 | 35.88 | 31.47 | 27.60 | 16.50 | 119.1 | | |
| 0.34 | 0.34 | 0.34 | 0.28 | 0.07 | 0.20 | 0.26 | | 56.44 | 49.50 | 43.41 | 38.07 | 8.44 | 47.12 | 108.7 | |
| 0.34 | 0.34 | 0.34 | 0.27 | 0.03 | 0.18 | 0.24 | 0.23 | 59.87 | 52.50 | 46.05 | 40.38 | 4.32 | 39.36 | 46.24 | 130.0 |