1. Introduction
Within the machine learning field, the term “wide data” [
1] refers to datasets containing a much greater number of features than instances. This type of data is common in bioinformatics [
2,
3], and usually presents two main problems that affect the performance of learning algorithms: the curse of dimensionality and data imbalance.
The curse of dimensionality [
4] refers to the difficulty of accurately generalizing problems with high-dimensional datasets when using machine learning algorithms. This increases both the processing time and required space, as well as the risk of overfitting, as it makes it difficult to distinguish meaningful patterns from noise.
Considering the low number of instances, wide data are prone to imbalance caused by the large difference in the number of instances per class [
5]. The algorithms trained with this data may be biased towards the majority class, making it difficult to accurately classify data belonging to the minority classes.
One of the solutions that may mitigate these problems is the use of preprocessing techniques. In particular, the curse of dimensionality can be addressed with feature selection (FS) [
6] and feature reduction (FR) [
7] methods. FS methods identify and select the most informative and relevant features from a given dataset, discarding the noisy or redundant data. In contrast, FR methods transform the original feature space into a lower-dimensional one using the information present in the original features.
Resampling methods [
8] solve the imbalanced data problem through removing instances from the majority class or creating new ones for the minority class.
There are many application examples of these methods in different areas. For example, in medicine, FR improves the accuracy of epilepsy diagnosis through analyzing electroencephalography signals, avoiding invasive techniques [
9]. FS has been also used for breast cancer detection [
10] analyzing microarray data. Regarding engineering, the authors of [
11] used Principal Component Analysis (PCA) in several predictors to remove noise from building energy consumption datasets. In another example, in fault diagnosis, FS was applied to select the best features extracted from magnet DC motors [
12] or rotating machinery [
13]. Furthermore, FR techniques are valuable in text mining tasks, such as document classification, e.g., in [
14] PCA and Latent Semantic Indexing (LSI) were used to extract useful features for an SVM classifier.
This study aims to find the best strategies to process wide datasets, composed of combinations of FS or FR, resampling, and classifiers, through evaluating their performance and computation time. In the literature, studies comparing dimensionality reduction techniques with resampling methods have been limited to the use of FS [
15]. In this case, new FR experiments are compared to the best results from [
16], which extensively compared various FS, resampling methods, and classifiers on a wide dataset.
As mentioned in
Section 2, the use of FR techniques with wide datasets requires thorough research to identify compatible approaches. For example, applying nonlinear transformations requires estimation to handle out-of-sample instances.
The scope of this study excludes the use of wrapper FS methods. As detailed in
Section 3, their high computational cost is a crucial factor when processing wide data, as it presents a large number of features. Both the FS and FR algorithms used require the dimensionality to be set, which simplifies the comparison and visibility of the results, as the dimensionality can be set to be the same. The obtained results are analyzed to address the following objectives of this study:
- 1.
To find an FR method that is compatible with wide data and provides a means to perform nonlinear transformations over out-of-data instances.
- 2.
To compare the two previously mentioned types of preprocessing techniques (FR and FS) and determine which is more suitable to use on wide datasets.
- 3.
To determine whether balancing is important while using FR methods and, if so, whether it is more convenient to use it before or after the FR step.
- 4.
To determine the best FR method for each classifier.
While previous studies have included some comparisons of FS and FR algorithms over the same datasets [
17], these evaluations are not particularly exhaustive. They predominantly focus on basic and widely used FR techniques, avoiding the use of nonlinear approaches. Furthermore, the lack of use of wide datasets makes it impossible to discern which techniques are optimal in such cases, as not all algorithms are compatible with such data.
Performing a large number of experiments to compare FS and FR approaches for wide datasets is one of the main novelties of this study. Due to the high presence of the imbalance problem in wide data, this study also focuses on resampling techniques in combination with FR and FS preprocessing methods. The code for all of the algorithms, allowing for their standardized use, can be found on GitHub (
https://github.com/Ismael-rp/feature_reduction_feature_selection_wide_data_comparison, accessed on 15 May 2024).
The remainder of this paper is organized as follows. First, in
Section 2,
Section 3 and
Section 4 the background for all the preprocessing methods used (i.e., FR, FS, and resampling) is provided. In
Section 5, the experimental setup is detailed, while the results are shown in
Section 6. Finally, the conclusions, limitations, and future work are presented in
Section 7,
Section 8 and
Section 9, respectively. Finally, the conclusions and future work are presented in
Section 7.
2. Feature Reduction
Feature reduction (FR) or manifold learning [
7] methods are preprocessing techniques used to reduce the number of dimensions of high-dimensional datasets. This is useful for improving the performance of learning algorithms, enhancing data visualization, and facilitating feature extraction from images. The present study focuses on FR for wide data classification.
As previously explained, high dimensionality poses a significant challenge to classification algorithms, as it complicates the distinction between useful and noisy features. FR methods attempt to solve this problem through creating a new dataset with the desired dimensionality, combining all the original features. A good FR method should be able to determine the structure of the original dataset (manifold) and preserve it in a lower dimensional representation. This structure is divided into local and global structures. Preserving the local structure refers to preserving the distance of all individual points to their nearest neighbors, whereas the global structure refers to the rest of the further points. Preserving both structures simultaneously is difficult [
18], and in FR methods, usually, only one is well retained.
There are some taxonomies that can be used to discriminate FR algorithms according to their behavior, and some of them can be very extensive, such as the one presented in the study [
7]. The present manuscript divides FR methods according to two properties: supervised/unsupervised and linear/nonlinear. Unsupervised methods ignore the data labels when creating the new dataset, which makes them useful for clustering problems. On the other hand, supervised methods utilize data labels, allowing classes to be separated more effectively and, therefore, making these methods more suitable for classification problems.
Linear FR methods transform the data using a linear transformation which minimizes or maximizes some criteria and, at the same time, reduces the dimensionality as desired. As shown in Equation (
1), matrix
A with dimensions of
is reduced to a
B matrix with
k dimensions using a kernel or linear transformation
K.
Non-linear transformations are required to uncover the hidden manifold in nonlinear data. As most of these algorithms are unsupervised, all of the methods used in this study are unsupervised. Unlike their linear alternatives, due to their intrinsic behavior, nonlinear methods do not provide a way to reproduce the transformation on out-of-sample data; however, the authors of [
19] presented a generalized and accurate approximation to solve this issue.
Based on the fact that every point in the space is linearly relocated to a new position in the lower-dimensional space under a nonlinear transformation, this linear transformation can be approximated through the following three steps. (1) Retrieve the K out-of-sample nearest neighbor instances from the training dataset. As the authors recommend, the K value was set to 5. (2) Reduce this neighbor sub-dataset to the desired dimensionality using Principal Component Analysis (PCA). (3) Using linear regression, obtain the linear projection to transpose the neighbor sub-dataset into the final positions obtained with the FR method.
The PCA and linear regression models are applied to the out-of-sample instance to be transformed. This process is repeated for each out-of-sample instance.
Unlike traditional datasets, wide data has a much greater number of columns than rows (), preventing some of the most popular linear and nonlinear FR algorithms from being able to calculate the projection.
The FR methods used in this study are listed below, following the taxonomy mentioned above:
Linear
- -
Unsupervised
- ∗
Principal Component Analysis (PCA) [
20] is the most popular FR method, which reduces the feature dimensionality while maintaining the maximum data variance.
- ∗
Locality Pursuit Embedding (LPE) [
21] respects the local structure through maximizing the variance of each local patch according to Euclidean distances (unlike PCA, which preserves the global structure).
- ∗
Parameter-Free Locality Preserving Projection (PFLPP) [
22] is a parameter-free version of the Locality Preserving Projection (LPP) algorithm [
23], which is a linear version of the nonlinear graph-based Laplacian Eigenmaps method [
24].
- ∗
Random Projection (RNDPROJ) [
25] projects the data into a new random spherical hyperplane that is randomly selected using the origin. It is not a trivial computation problem.
- -
Supervised
- ∗
Fisher Score (FSCORE) [
26] finds the projection that maximizes the ratio between each feature mean and the standard deviation of each class.
- ∗
Locality Sensitive Laplacian Score (LSLS) [
27] is based on the Laplacian score FS method [
28]. It adjusts the Laplacian graph using the class label to simultaneously minimize the local within-class information and maximize the local between-class information.
- ∗
Local Fisher Discriminant Analysis (LFDA) [
29] is an improved version of the FDA-supervised FR method, which is suitable for reducing datasets in which individual classes are separated into several clusters.
- ∗
Maximum Margin Criterion (MMC) [
30] projects the data while maximizing the average margin between classes.
- ∗
Sliced Average Variance Estimation (SAVE) [
31] calculates the projection matrix by averaging the covariance of the data of each slice in which the whole dataset has been divided.
- ∗
Supervised Locality Pursuit Embedding (SLPE) [
32] is a supervised version of the LPE algorithm, which enhances the model using label data.
Non-linear
- -
Classical Multidimensional Scaling (MDS) [
33] computes the dissimilarities between pairs of objects (assuming Euclidean distance). This matrix serves as the input for the algorithm that outputs a coordinate that minimizes a loss function called
strain.
- -
Metric Multidimensional Scaling (MMDS) [
34] is a superset of the previous method. It iteratively updates the weights given by the MDS using the SMACOF algorithm, in order to minimize a stress function such as the residual sum of squares.
- -
Locally Linear Embedding (LLE) [
35] bases its performance on producing low-dimensional vectors that best reconstruct the original objects through computing the
kNN and using this information to weight them.
- -
Neighborhood Preserving Embedding (NPE) [
36] first identifies the structure of the data neighborhood in the original space, then determines a linear subspace minimizing the reconstruction error of the local neighborhood structure [
37].
- -
Locally Embedded Analysis (LEA) [
34] aims to preserve the local structure of the original data in the computed embedding space.
- -
Stochastic Neighbor Embedding (SNE) [
38] is a probabilistic approach that places the data in a low-dimensional space that optimally preserves the neighborhood of the original space.
- -
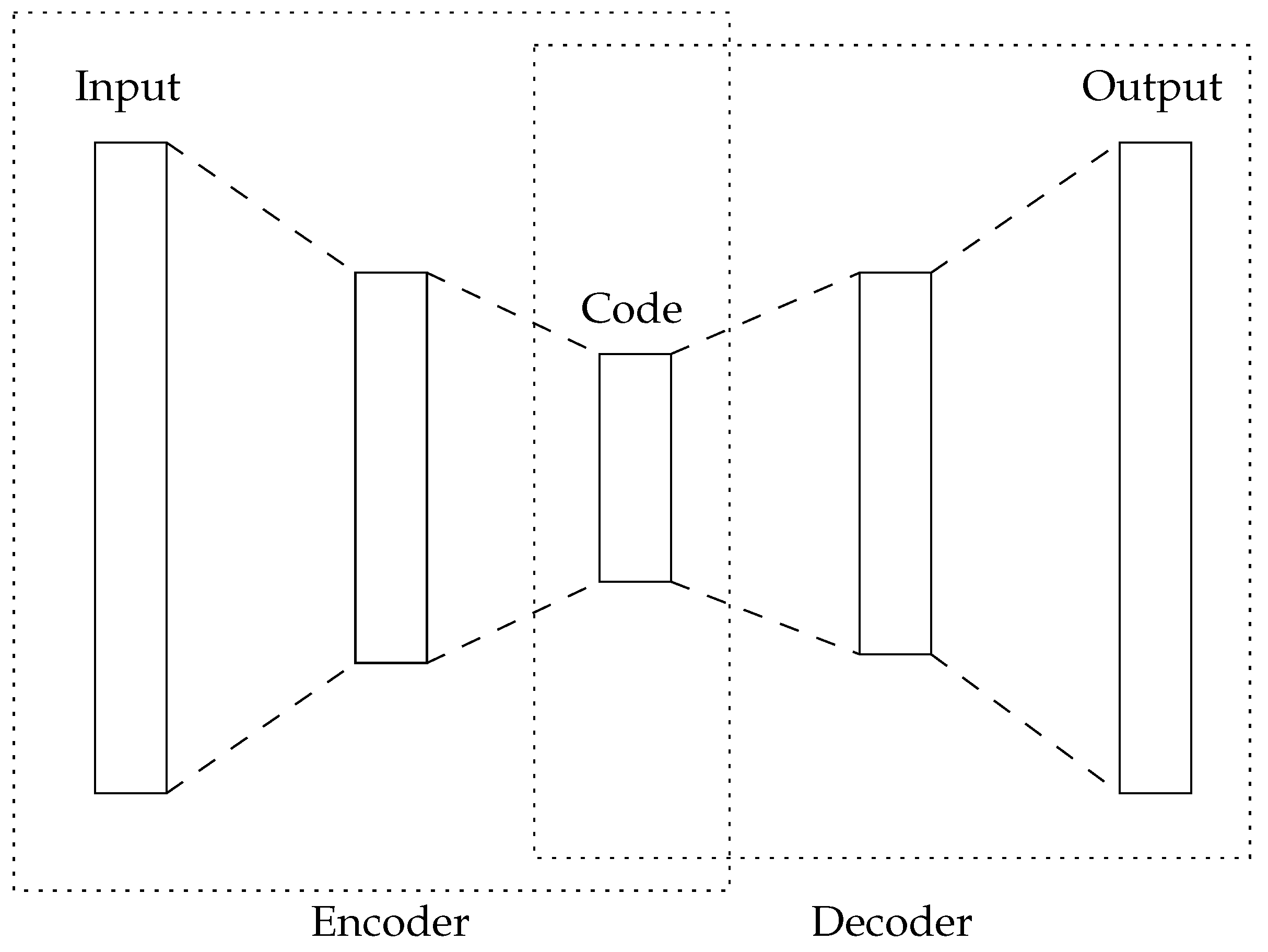
An Autoencoder [
39,
40] is a kind of artificial neural network that is trained in an unsupervised manner. The aim of the autoencoder is to capture the hidden information in the high-dimensional input space of the dataset. Autoencoders have the same number of artificial neurons in their first (input) and last (output) layers, while having less in their center layers (see
Figure 1). During training, Autoencoders attempt to generate the same information in the output layer that is presented in the input layer. Therefore, the center layer aims to capture the intrinsic information of the dataset and, thus, can be used for feature reduction.
Author Contributions
Conceptualization, I.R.-P.; software, I.R.-P.; writing—original draft preparation, I.R.-P., J.M.-R. and Á.A.-G.; writing—review and editing, I.R.-P., J.A.B.-A., A.C.-O., J.M.-R. and Á.A.-G.; supervision, J.M.-R. and Á.A.-G. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the Junta de Castilla y León under project BU055P20 (JCyL/FEDER, UE) and by the Ministry of Science and Innovation under project PID2020-119894GB-I00, co-financed through European Union FEDER funds. Ismael Ramos-Pérez is funded through a pre-doctoral grant by the Universidad de Burgos.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
All datasets used are referred to in the paper.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Lai, K.; Twine, N.; O’brien, A.; Guo, Y.; Bauer, D. Artificial intelligence and machine learning in bioinformatics. Encycl. Bioinform. Comput. Biol. ABC Bioinform. 2018, 1, 272–286. [Google Scholar]
- Hao, Z.; Lv, D.; Ge, Y.; Shi, J.; Weijers, D.; Yu, G.; Chen, J. RIdeogram: Drawing SVG graphics to visualize and map genome-wide data on the idiograms. PeerJ Comput. Sci. 2020, 6, e251. [Google Scholar] [CrossRef] [PubMed]
- Salesi, S.; Cosma, G.; Mavrovouniotis, M. TAGA: Tabu Asexual Genetic Algorithm embedded in a filter/filter feature selection approach for high-dimensional data. Inf. Sci. 2021, 565, 105–127. [Google Scholar] [CrossRef]
- Keogh, E.J.; Mueen, A. Curse of dimensionality. Encycl. Mach. Learn. Data Min. 2017, 2017, 314–315. [Google Scholar]
- He, H.; Garcia, E.A. Learning from imbalanced data. IEEE Trans. Knowl. Data Eng. 2009, 21, 1263–1284. [Google Scholar]
- Saeys, Y.; Inza, I.; Larrañaga, P. A review of feature selection techniques in bioinformatics. Bioinformatics 2007, 23, 2507–2517. [Google Scholar] [CrossRef] [PubMed]
- Ayesha, S.; Hanif, M.K.; Talib, R. Overview and comparative study of dimensionality reduction techniques for high dimensional data. Inf. Fusion 2020, 59, 44–58. [Google Scholar] [CrossRef]
- Mohammed, R.; Rawashdeh, J.; Abdullah, M. Machine learning with oversampling and undersampling techniques: Overview study and experimental results. In Proceedings of the 2020 11th International Conference on Information and Communication Systems (ICICS), Irbid, Jordan, 7–9 April 2020; IEEE: Piscataway, NJ, USA, 2020; pp. 243–248. [Google Scholar]
- Wijayanto, I.; Humairani, A.; Hadiyoso, S.; Rizal, A.; Prasanna, D.L.; Tripathi, S.L. Epileptic seizure detection on a compressed EEG signal using energy measurement. Biomed. Signal Process. Control 2023, 85, 104872. [Google Scholar] [CrossRef]
- Sachdeva, R.K.; Bathla, P.; Rani, P.; Kukreja, V.; Ahuja, R. A Systematic Method for Breast Cancer Classification using RFE Feature Selection. In Proceedings of the 2022 2nd International Conference on Advance Computing and Innovative Technologies in Engineering, ICACITE 2022, Greater Noida, India, 28–29 April 2022; pp. 1673–1676. [Google Scholar] [CrossRef]
- Parhizkar, T.; Rafieipour, E.; Parhizkar, A. Evaluation and improvement of energy consumption prediction models using principal component analysis based feature reduction. J. Clean. Prod. 2021, 279, 123866. [Google Scholar] [CrossRef]
- Wang, W.; Lu, L.; Wei, W. A Novel Supervised Filter Feature Selection Method Based on Gaussian Probability Density for Fault Diagnosis of Permanent Magnet DC Motors. Sensors 2022, 22, 7121. [Google Scholar] [CrossRef] [PubMed]
- Zhao, X.; Jia, M. Fault diagnosis of rolling bearing based on feature reduction with global-local margin Fisher analysis. Neurocomputing 2018, 315, 447–464. [Google Scholar] [CrossRef]
- Ayadi, R.; Maraoui, M.; Zrigui, M. LDA and LSI as a dimensionality reduction method in arabic document classification. Commun. Comput. Inf. Sci. 2015, 538, 491–502. [Google Scholar] [CrossRef]
- Pes, B. Learning from High-Dimensional and Class-Imbalanced Datasets Using Random Forests. Information 2021, 12, 286. [Google Scholar] [CrossRef]
- Ramos-Pérez, I.; Arnaiz-González, A.; Rodríguez, J.J.; García-Osorio, C. When is resampling beneficial for feature selection with imbalanced wide data? Expert Syst. Appl. 2022, 188, 116015. [Google Scholar] [CrossRef]
- Mendes Junior, J.J.A.; Freitas, M.L.; Siqueira, H.V.; Lazzaretti, A.E.; Pichorim, S.F.; Stevan, S.L. Feature selection and dimensionality reduction: An extensive comparison in hand gesture classification by sEMG in eight channels armband approach. Biomed. Signal Process. Control 2020, 59, 101920. [Google Scholar] [CrossRef]
- Muntasa, A.; Sirajudin, I.A.; Purnomo, M.H. Appearance global and local structure fusion for face image recognition. TELKOMNIKA (Telecommun. Comput. Electron. Control) 2011, 9, 125–132. [Google Scholar] [CrossRef][Green Version]
- Yang, Y.; Nie, F.; Xiang, S.; Zhuang, Y.; Wang, W. Local and global regressive mapping for manifold learning with out-of-sample extrapolation. In Proceedings of the AAAI Conference on Artificial Intelligence, Atlanta, GA, USA, 11–13 October 2010; pp. 649–654. [Google Scholar]
- Pearson, K. LIII. On lines and planes of closest fit to systems of points in space. Lond. Edinb. Dublin Philos. Mag. J. Sci. 1901, 2, 559–572. [Google Scholar] [CrossRef]
- Min, W.; Lu, K.; He, X. Locality pursuit embedding. Pattern Recognit. 2004, 37, 781–788. [Google Scholar] [CrossRef]
- Dornaika, F.; Assoum, A. Enhanced and parameterless Locality Preserving Projections for face recognition. Neurocomputing 2013, 99, 448–457. [Google Scholar] [CrossRef]
- He, X.; Niyogi, P. Locality Preserving Projections. Adv. Neural Inf. Process. Syst. 2003, 16. [Google Scholar]
- Belkin, M.; Niyogi, P. Laplacian eigenmaps for dimensionality reduction and data representation. Neural Comput. 2003, 15, 1373–1396. [Google Scholar] [CrossRef]
- Achlioptas, D. Database-friendly random projections: Johnson-Lindenstrauss with binary coins. J. Comput. Syst. Sci. 2003, 66, 671–687. [Google Scholar] [CrossRef]
- Fisher, R.A. The use of multiple measurements in taxonomic problems. Ann. Eugen. 1936, 7, 179–188. [Google Scholar] [CrossRef]
- Liao, B.; Jiang, Y.; Liang, W.; Zhu, W.; Cai, L.; Cao, Z. Gene selection using locality sensitive Laplacian score. IEEE/ACM Trans. Comput. Biol. Bioinform. 2014, 11, 1146–1156. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Cai, D.; Niyogi, P. Laplacian score for feature selection. Adv. Neural Inf. Process. Syst. 2005, 18. [Google Scholar]
- Sugiyama, M. Local fisher discriminant analysis for supervised dimensionality reduction. ACM Int. Conf. Proceeding Ser. 2006, 148, 905–912. [Google Scholar] [CrossRef]
- Li, H.; Jiang, T.; Zhang, K. Efficient and robust feature extraction by maximum margin criterion. IEEE Trans. Neural Netw. 2006, 17, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Dennis Cook, R. SAVE: A method for dimension reduction and graphics in regression. Commun.-Stat.-Theory Methods 2000, 29, 2109–2121. [Google Scholar] [CrossRef]
- Zheng, Z.; Yang, F.; Tan, W.; Jia, J.; Yang, J. Gabor feature-based face recognition using supervised locality preserving projection. Signal Process. 2007, 87, 2473–2483. [Google Scholar] [CrossRef]
- Kruskal, J.B. Multidimensional scaling by optimizing goodness of fit to a nonmetric hypothesis. Psychometrika 1964, 29, 1–27. [Google Scholar] [CrossRef]
- Borg, I.; Groenen, P.J. Modern Multidimensional Scaling: Theory and Applications; Springer: New York, NY, USA, 2005. [Google Scholar]
- Roweis, S.T.; Saul, L.K. Nonlinear dimensionality reduction by locally linear embedding. Science 2000, 290, 2323–2326. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Cai, D.; Yan, S.; Zhang, H.J. Neighborhood preserving embedding. In Proceedings of the Tenth IEEE International Conference on Computer Vision (ICCV’05) Volume 1, Beijing, China, 17–21 October 2005; pp. 1208–1213. [Google Scholar] [CrossRef]
- Yao, C.; Guo, Z. Revisit Neighborhood Preserving Embedding: A New Criterion for Measuring the Manifold Similarity in Dimension Reduction. Available online: https://ssrn.com/abstract=4349051 (accessed on 7 April 2024).
- Hinton, G.E.; Roweis, S. Stochastic neighbor embedding. Adv. Neural Inf. Process. Syst. 2002, 15. [Google Scholar]
- Rumelhart, D.E.; Hinton, G.E.; Williams, R.J. Learning Internal Representations by Error Propagation; Technical Report; California Univ San Diego La Jolla Inst for Cognitive Science: La Jolla, CA, USA, 1985. [Google Scholar]
- Hinton, G.E.; Salakhutdinov, R.R. Reducing the dimensionality of data with neural networks. Science 2006, 313, 504–507. [Google Scholar] [CrossRef]
- Bommert, A.; Sun, X.; Bischl, B.; Rahnenführer, J.; Lang, M. Benchmark for filter methods for feature selection in high-dimensional classification data. Comput. Stat. Data Anal. 2020, 143, 106839. [Google Scholar] [CrossRef]
- Kohavi, R.; John, G.H. Wrappers for feature subset selection. Artif. Intell. 1997, 97, 273–324. [Google Scholar] [CrossRef]
- Lal, T.N.; Chapelle, O.; Weston, J.; Elisseeff, A. Embedded methods. In Feature Extraction: Foundations and Applications; Springer: Berlin/Heidelberg, Germany, 2006; pp. 137–165. [Google Scholar]
- Guyon, I.; Weston, J.; Barnhill, S.; Vapnik, V. Gene selection for cancer classification using support vector machines. Mach. Learn. 2002, 46, 389–422. [Google Scholar] [CrossRef]
- Luque, A.; Carrasco, A.; Martín, A.; de las Heras, A. The impact of class imbalance in classification performance metrics based on the binary confusion matrix. Pattern Recognit. 2019, 91, 216–231. [Google Scholar] [CrossRef]
- Japkowicz, N. The Class Imbalance Problem: Significance and Strategies. In Proceedings of the 2000 International Conference on Artificial Intelligence (ICAI), Vancouver, BC, Canada, 13–15 November 2000; pp. 111–117. [Google Scholar]
- Chawla, N.V.; Bowyer, K.W.; Hall, L.O.; Kegelmeyer, W.P. SMOTE: Synthetic minority over-sampling technique. J. Artif. Intell. Res. 2002, 16, 321–357. [Google Scholar] [CrossRef]
- Dietterich, T.G. Approximate Statistical Tests for Comparing Supervised Classification Learning Algorithms. Neural Comput. 1998, 10, 1895–1923. [Google Scholar] [CrossRef] [PubMed]
- García, V.; Sánchez, J.S.; Mollineda, R.A. On the effectiveness of preprocessing methods when dealing with different levels of class imbalance. Knowl.-Based Syst. 2012, 25, 13–21. [Google Scholar] [CrossRef]
- Orriols-Puig, A.; Bernadó-Mansilla, E. Evolutionary rule-based systems for imbalanced datasets. Soft Comput. 2009, 13, 213–225. [Google Scholar] [CrossRef]
- Zhu, Z.; Ong, Y.S.; Dash, M. Markov blanket-embedded genetic algorithm for gene selection. Pattern Recognit. 2007, 40, 3236–3248. [Google Scholar] [CrossRef]
- Li, J.; Cheng, K.; Wang, S.; Morstatter, F.; Trevino, R.P.; Tang, J.; Liu, H. Feature selection: A data perspective. ACM Comput. Surv. (CSUR) 2018, 50, 1–45. [Google Scholar] [CrossRef]
- Bolón-Canedo, V.; Alonso-Betanzos, A. Recent Advances in Ensembles for Feature Selection; Springer: Berlin/Heidelberg, Germany, 2018; Volume 147. [Google Scholar] [CrossRef]
- Galar, M.; Fernandez, A.; Barrenechea, E.; Bustince, H.; Herrera, F. A review on ensembles for the class imbalance problem: Bagging-, boosting-, and hybrid-based approaches. IEEE Trans. Syst. Man Cybern. Part C (Appl. Rev.) 2011, 42, 463–484. [Google Scholar] [CrossRef]
- Matthews, B. Comparison of the predicted and observed secondary structure of T4 phage lysozyme. Biochim. Biophys. Acta (BBA)-Protein Struct. 1975, 405, 442–451. [Google Scholar] [CrossRef]
- Baldi, P.; Brunak, S.; Chauvin, Y.; Andersen, C.A.F.; Nielsen, H. Assessing the accuracy of prediction algorithms for classification: An overview. Bioinformatics 2000, 16, 412–424. [Google Scholar] [CrossRef] [PubMed]
- Chicco, D.; Jurman, G. The advantages of the Matthews correlation coefficient (MCC) over F1 score and accuracy in binary classification evaluation. BMC Genom. 2020, 21, 6. [Google Scholar] [CrossRef]
- Chicco, D.; Jurman, G. The Matthews correlation coefficient (MCC) should replace the ROC AUC as the standard metric for assessing binary classification. Biodata Min. 2023, 16, 4. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Demšar, J. Statistical comparisons of classifiers over multiple datasets. J. Mach. Learn. Res. 2006, 7, 1–30. [Google Scholar]
- Garcia, S.; Herrera, F. An Extension on “Statistical Comparisons of Classifiers over Multiple Data Sets” for all Pairwise Comparisons. J. Mach. Learn. Res. 2008, 9, 2677–2694. [Google Scholar]
- Benavoli, A.; Corani, G.; Mangili, F.; Zaffalon, M.; Ruggeri, F. A Bayesian Wilcoxon signed-rank test based on the Dirichlet process. In Proceedings of the International Conference on Machine Learning, Beijing, China, 22–24 June 2014; PMLR; pp. 1026–1034. [Google Scholar]
- Kuncheva, L.I.; Matthews, C.E.; Arnaiz-González, A.; Rodríguez, J.J. Feature selection from high-dimensional data with very low sample size: A cautionary tale. arXiv 2020, arXiv:2008.12025. [Google Scholar]
- Van Engelen, J.E.; Hoos, H.H. A survey on semi-supervised learning. Mach. Learn. 2020, 109, 373–440. [Google Scholar] [CrossRef]
| Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).