Breed-Specific Genetic Recombination Analysis in South African Bonsmara and Nguni Cattle Using Genomic Data
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and DNA Isolation
2.2. SNP Genotyping
2.3. Data Filtering and Quality Control
2.4. Haplotype Phasing
2.5. Detection of Recombination Crossover Events
2.6. Identification of Genes in Recombination Hotspot Regions
3. Results
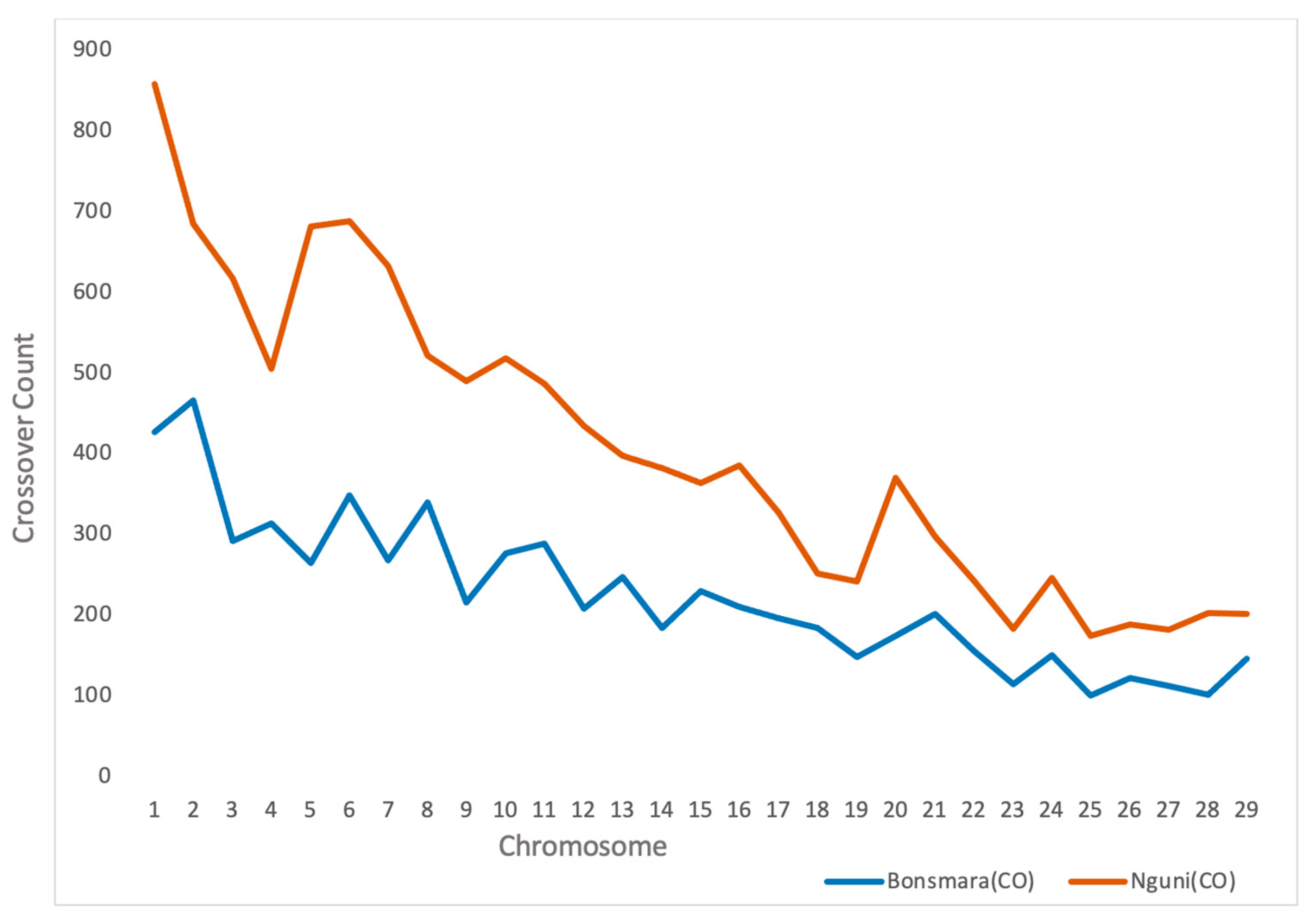
3.1. Crossover Events
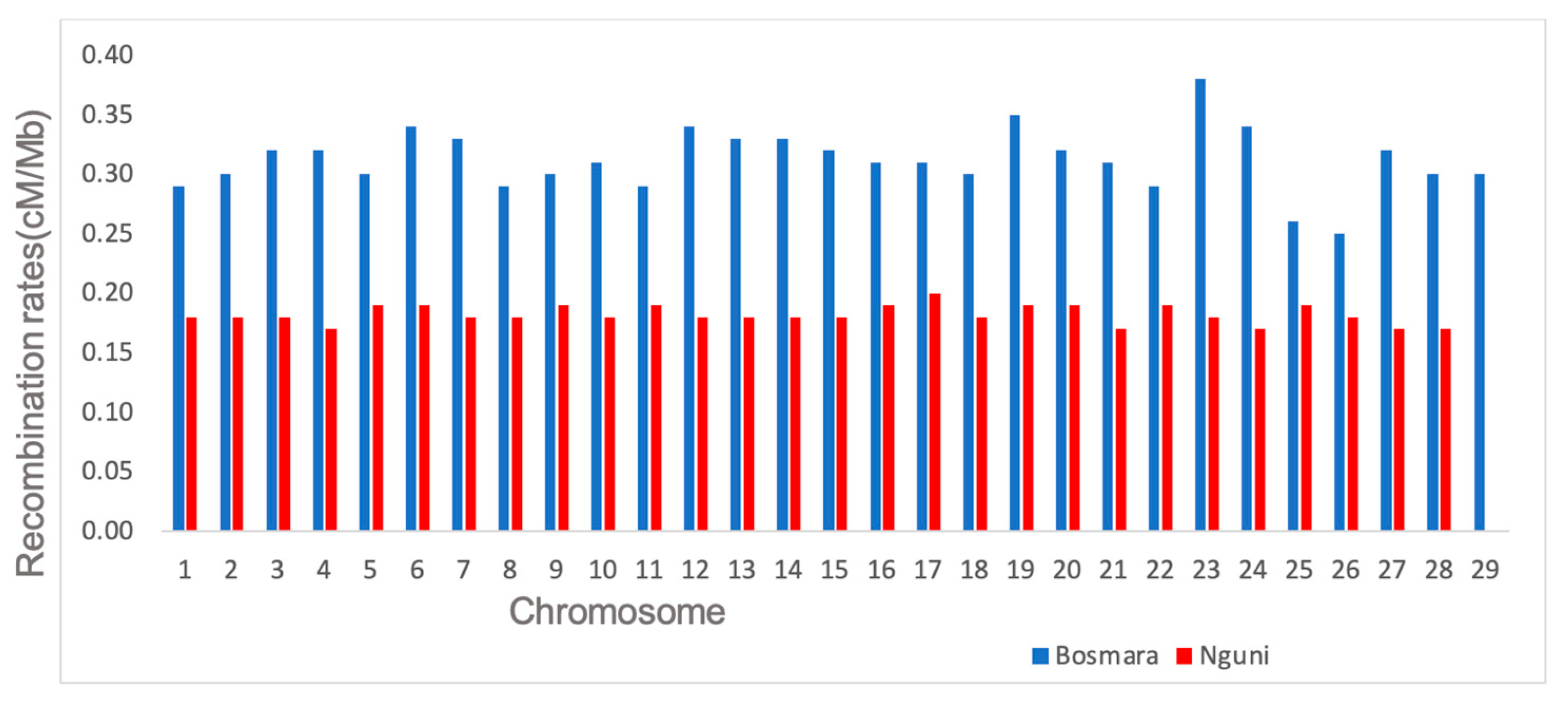
3.2. Determining Recombination Rates
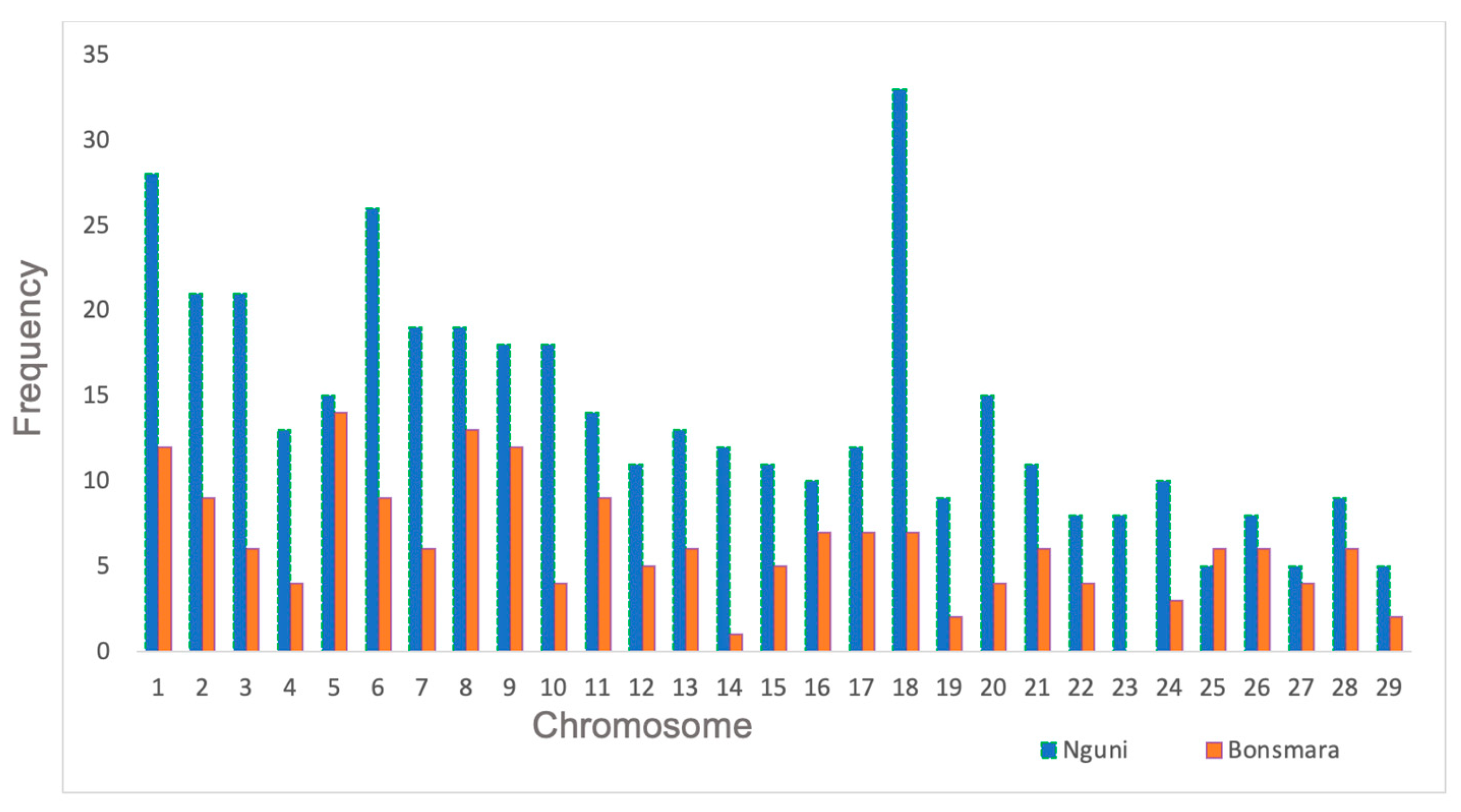
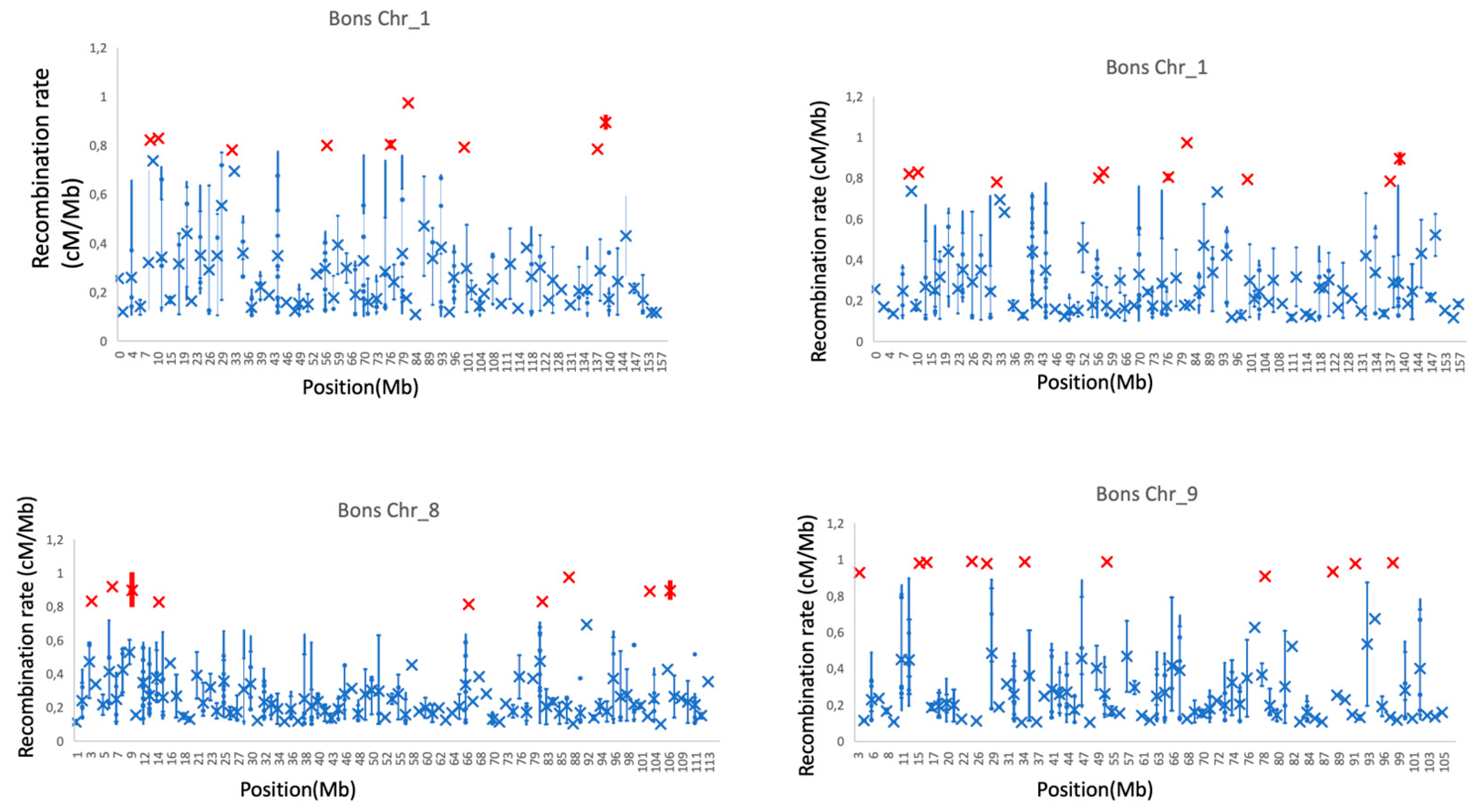
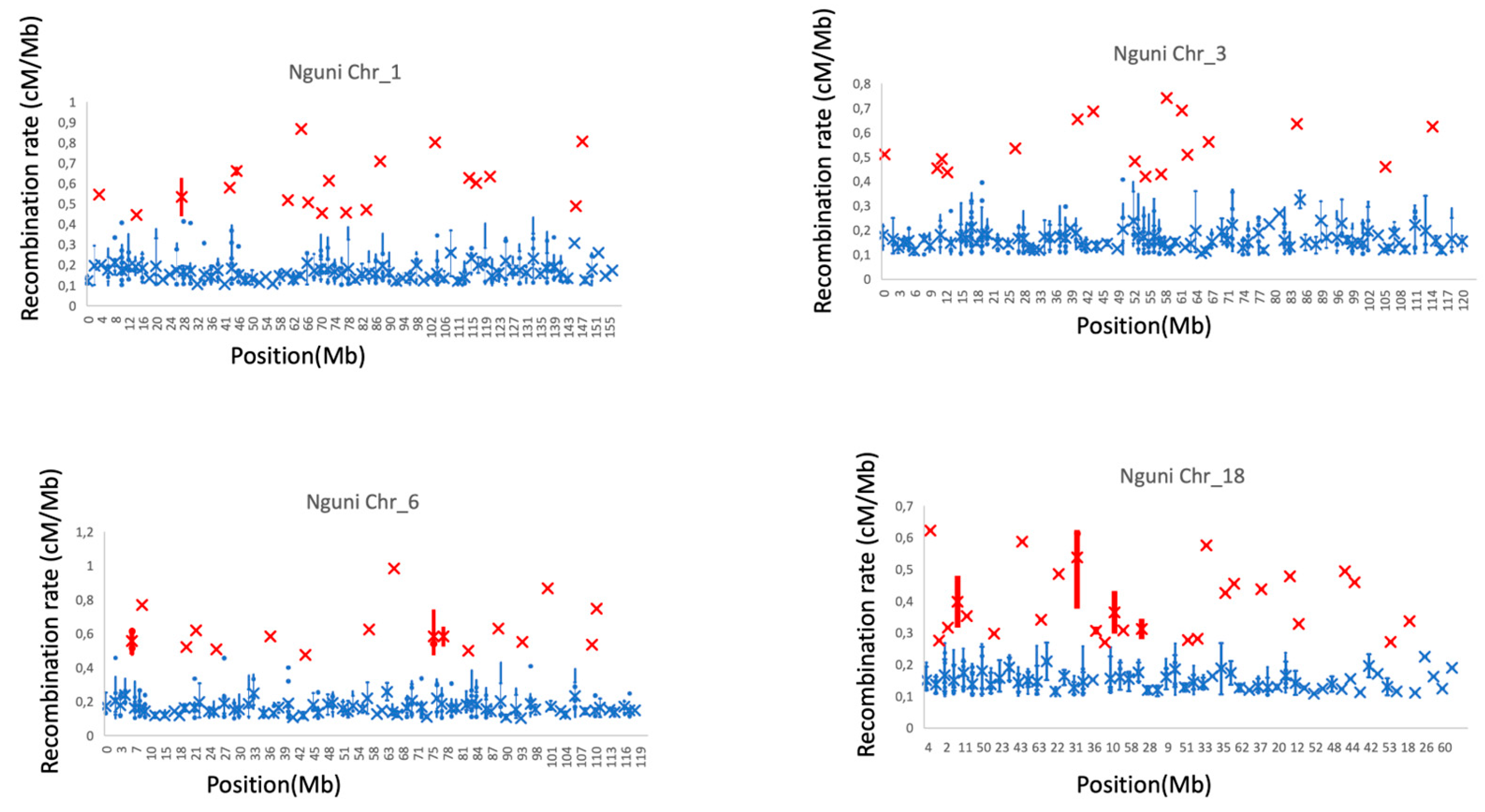
3.3. Recombination Hotspots Across Autosomes
3.4. Variation in Recombination Hotspots Along the Autosomes
3.5. Identification of Genes Associated with Recombination Rates and Hotspots
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Matjuda, L.E.; MacNeil, M.D.; Maiwashe, A.; Leesburg, V.R.; Malatje, M. Index-in-retrospect and breeding objectives characterizing genetic improvement programmes for South African Nguni cattle. S. Afr. J. Anim. Sci. 2014, 44, 161–172. [Google Scholar] [CrossRef]
- van Marle-Kӧster, E.; Pretorius, S.J.; Webb, E.C. Morphological and physiological characteristics of claw quality in South African Bonsmara cattle. S. Afr. J. Anim. Sci. 2019, 49, 966–976. [Google Scholar] [CrossRef]
- van Eeden, G.; Uren, C.; Möller, M.; Henn, B.M. Inferring recombination patterns in African populations. Hum. Mol. Genet. 2021, 30, R11–R16. [Google Scholar] [CrossRef] [PubMed]
- Capilla, L.; Garcia Caldés, M.; Ruiz-Herrera, A. Mammalian Meiotic Recombination: A Toolbox for Genome Evolution. Cytogenet. Genome Res. 2016, 150, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Baudat, F.; de Massy, B. Regulating double-stranded DNA break repair towards crossover or non-crossover during mammalian meiosis. Chromosome Res. 2007, 15, 565–577. [Google Scholar] [CrossRef]
- Myers, S.; Bottolo, L.; Freeman, C.; McVean, G.; Donnelly, P. A Fine-Scale Map of Recombination Rates and Hotspots Across the Human Genome. Science 2005, 310, 321–324. [Google Scholar] [CrossRef]
- Choi, K.; Henderson, I.R. Meiotic recombination hotspots—A comparative view. Plant J. 2015, 83, 52–61. [Google Scholar] [CrossRef]
- Singhal, S.; Leffler, E.M.; Sannareddy, K.; Turner, I.; Venn, O.; Hooper, D.M.; Strand, A.I.; Li, Q.; Raney, B.; Balakrishnan, C.N.; et al. Stable recombination hotspots in birds. Science 2015, 350, 928–932. [Google Scholar] [CrossRef]
- Baudat, F.; Imai, Y.; de Massy, B. Meiotic recombination in mammals: Localization and regulation. Nat. Rev. Genet. 2013, 14, 794–806. [Google Scholar] [CrossRef]
- Jensen-Seaman, M.I.; Furey, T.S.; Payseur, B.A.; Lu, Y.; Roskin, K.M.; Chen, C.-F.; Thomas, M.A.; Haussler, D.; Jacob, H.J. Comparative Recombination Rates in the Rat, Mouse, and Human Genomes. Genome Res. 2004, 14, 528–538. [Google Scholar] [CrossRef]
- Arnheim, N.; Calabrese, P.; Nordborg, M. Hot and Cold Spots of Recombination in the Human Genome: The Reason We Should Find Them and How This Can Be Achieved. Am. J. Hum. Genet. 2003, 73, 5–16. [Google Scholar] [CrossRef]
- Morris, G.P.; Ramu, P.; Deshpande, S.P.; Hash, C.T.; Shah, T.; Upadhyaya, H.D.; Riera-Lizarazu, O.; Brown, P.J.; Acharya, C.B.; Mitchell, S.E.; et al. Population genomic and genome-wide association studies of agroclimatic traits in sorghum. Proc. Natl. Acad. Sci. USA 2013, 110, 453–458. [Google Scholar] [CrossRef]
- Hohenlohe, P.A.; Funk, W.C.; Rajora, O.P. Population genomics for wildlife conservation and management. Mol. Ecol. 2021, 30, 62–82. [Google Scholar] [CrossRef]
- Kim, J.; Hanotte, O.; Mwai, O.A.; Dessie, T.; Bashir, S.; Diallo, B.; Agaba, M.; Kim, K.; Kwak, W.; Sung, S.; et al. The genome landscape of indigenous African cattle. Genome Biol. 2017, 18, 34. [Google Scholar] [CrossRef] [PubMed]
- Madilindi, M.A.; Banga, C.B.; Bhebhe, E.; Sanarana, Y.P.; Nxumalo, K.S.; Taela, M.G.; Magagula, B.S.; Mapholi, N.O. Genetic diversity and relationships among three Southern African Nguni cattle populations. Trop. Anim. Health Prod. 2020, 52, 753–762. [Google Scholar] [CrossRef] [PubMed]
- Hellsten, U.; Wright, K.M.; Jenkins, J.; Shu, S.; Yuan, Y.; Wessler, S.R.; Schmutz, J.; Willis, J.H.; Rokhsar, D.S. Fine-scale variation in meiotic recombination in Mimulus inferred from population shotgun sequencing. Proc. Natl. Acad. Sci. USA 2013, 110, 19478–19482. [Google Scholar] [CrossRef] [PubMed]
- Weng, Z.-Q.; Saatchi, M.; Schnabel, R.D.; Taylor, J.F.; Garrick, D.J. Recombination locations and rates in beef cattle assessed from parent-offspring pairs. Genet. Sel. Evol. 2014, 46, 34. [Google Scholar] [CrossRef]
- Ma, L.; O’Connell, J.R.; VanRaden, P.M.; Shen, B.; Padhi, A.; Sun, C.; Bickhart, D.M.; Cole, J.B.; Null, D.J.; Liu, G.E.; et al. Cattle Sex-Specific Recombination and Genetic Control from a Large Pedigree Analysis. PLoS Genet. 2015, 11, e1005387. [Google Scholar] [CrossRef]
- Auton, A.; McVean, G. Recombination rate estimation in the presence of hotspots. Genome Res. 2007, 17, 1219–1227. [Google Scholar] [CrossRef]
- Bhérer, C.; Campbell, C.L.; Auton, A. Refined genetic maps reveal sexual dimorphism in human meiotic recombination at multiple scales. Nat. Commun. 2017, 8, 14994. [Google Scholar] [CrossRef]
- Halldorsson, B.V.; Palsson, G.; Stefansson, O.A.; Jonsson, H.; Hardarson, M.T.; Eggertsson, H.P.; Gunnarsson, B.; Oddsson, A.; Halldorsson, G.H.; Zink, F.; et al. Characterizing mutagenic effects of recombination through a sequence-level genetic map. Science 2019, 363, eaau1043. [Google Scholar] [CrossRef]
- Kong, A.; Thorleifsson, G.; Gudbjartsson, D.F.; Masson, G.; Sigurdsson, A.; Jonasdottir, A.; Walters, G.B.; Jonasdottir, A.; Gylfason, A.; Kristinsson, K.T.; et al. Fine-scale recombination rate differences between sexes, populations and individuals. Nature 2010, 467, 1099–1103. [Google Scholar] [CrossRef]
- Peñalba, J.V.; Wolf, J.B.W. From molecules to populations: Appreciating and estimating recombination rate variation. Nat. Rev. Genet. 2020, 21, 476–492. [Google Scholar] [CrossRef]
- Fledel-Alon, A.; Leffler, E.M.; Guan, Y.; Stephens, M.; Coop, G.; Przeworski, M. Variation in Human Recombination Rates and Its Genetic Determinants. PLoS ONE 2011, 6, e20321. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A Tool Set for Whole-Genome Association and Population-Based Linkage Analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef] [PubMed]
- Delaneau, O.; Coulonges, C.; Zagury, J.F. Shape-IT: New rapid and accurate algorithm for haplotype inference. BMC Bioinform. 2008, 9, 540. [Google Scholar] [CrossRef] [PubMed]
- Yang, F.; Silber, S.; Leu, N.A.; Oates, R.D.; Marszalek, J.D.; Skaletsky, H.; Brown, L.G.; Rozen, S.; Page, D.C.; Wang, P.J. TEX11 is mutated in infertile men with azoospermia and regulates genome-wide recombination rates in mice. EMBO Mol. Med. 2015, 7, 1198–1210. [Google Scholar] [CrossRef]
| Chromosome | Bonsmara | Nguni | ||||
|---|---|---|---|---|---|---|
| #SNP | CO | RR (SD) | #SNP | CO | RR (SD) | |
| 1 | 2369 | 427 | 0.29 (0.20) | 2806 | 857 | 0.18 (0.10) |
| 2 | 2085 | 466 | 0.30 (0.20) | 2356 | 684 | 0.18 (0.08) |
| 3 | 1860 | 292 | 0.02 (0.24) | 2187 | 617 | 0.18 (0.009) |
| 4 | 1751 | 314 | 0.32 (0.26) | 2095 | 505 | 0.18 (0.10) |
| 5 | 1609 | 265 | 0.30 (0.23) | 1877 | 681 | 0.17 (0.08) |
| 6 | 2232 | 349 | 0.34 (0.22) | 2531 | 687 | 0.19 (0.11) |
| 7 | 1948 | 268 | 0.33 (0.23) | 2230 | 632 | 0.19 (0.10) |
| 8 | 1696 | 340 | 0.29 (0.19) | 1993 | 521 | 0.18 (0.10) |
| 9 | 1570 | 216 | 0.30 (0.24) | 1774 | 489 | 0.18 (0.10) |
| 10 | 1768 | 277 | 0.31 (0.22) | 2063 | 518 | 0.19 (0.11) |
| 11 | 1589 | 289 | 0.29 (0.19) | 1837 | 486 | 0.18 (0.09) |
| 12 | 1231 | 209 | 0.34 (0.25) | 1439 | 434 | 0.19 (0.09) |
| 13 | 1232 | 248 | 0.33 (0.25) | 1470 | 397 | 0.18 (0.10) |
| 14 | 1360 | 185 | 0.33 (0.24) | 1486 | 383 | 0.18 (0.09) |
| 15 | 1259 | 230 | 0.32 (0.22) | 1447 | 363 | 0.18 (0.11) |
| 16 | 1192 | 211 | 0.31 (0.22) | 1418 | 386 | 0.18 (0.11) |
| 17 | 1207 | 197 | 0.31 (0.22) | 1395 | 326 | 0.19 (0.12) |
| 18 | 962 | 185 | 0.30 (0.19) | 1116 | 252 | 0.18 (0.04) |
| 19 | 1052 | 149 | 0.35 (0.24) | 1171 | 242 | 0.20 (0.12) |
| 20 | 1199 | 175 | 0.32 (0.24) | 1394 | 371 | 0.19 (0.11) |
| 21 | 1073 | 202 | 0.31 (0.21) | 1234 | 298 | 0.17 (0.12) |
| 22 | 931 | 156 | 0.30 (0.20) | 1068 | 243 | 0.19 (0.19) |
| 23 | 840 | 115 | 0.38 (0.24) | 977 | 184 | 0.18 (0.20) |
| 24 | 898 | 151 | 0.34 (0.22) | 1064 | 247 | 0.17 (0.10) |
| 25 | 743 | 101 | 0.26 (0.19) | 787 | 175 | 0.19 (0.08) |
| 26 | 748 | 122 | 0.25 (0.20) | 889 | 189 | 0.18 (0.13) |
| 27 | 723 | 113 | 0.32 (0.23) | 807 | 182 | 0.18 (0.11) |
| 28 | 674 | 102 | 0.30 (0.25) | 749 | 203 | 0.17 (0.08) |
| 29 | 761 | 147 | 0.30 (0.20) | 865 | 202 | 0.17 (0.10) |
| Chr | Position | Gene | Gene ID | Function |
|---|---|---|---|---|
| 1 | 8.35 | AAP | ENSSBTG00000001534 | Regulates synopsis formation. Receptor-binding activity. Growth releasing hormone. |
| 1 | 30.74 | ROBO1 | ENSBTAG00000009851 | Enables protein binding. Enables axon guidance receptor. |
| 1 | 80.0 | BRWD1 | ENSBTAG00000025910 | Enables molecular function. Regulates transcription by RNA polymerase II. Enables gametogenesis and oocyte embryo transition (Mice). Epigenomic mediator of normal neurodevelopment (mice). |
| 1 | 75.69 | ZBTB38 | ENSBTAG00000040061 | Regulates cytokine production. Regulates immune system process. Enables protein binding. Negative regulation of transcription by RNA polymerase II. Target gene for body measurement trait. |
| 5 | 28.76 | PDE1B | ENSBTAG00000004337 | Target gene for carcass traits. |
| 5 | 4.60 | LOC617565 LINE-1 | Determinant for DNA target structure. | |
| 8 | 65.57 | Prnp | ENSBTAG00000048903 | Regulation of DNA-binding transcription factor. Regulation of long-term synaptic potentiation. ATP independent protein, amyloid and copper binding. |
| 8 | 103.28 | AMBP | ENSBTAG00000015676 | Supports ovulation. Repairs micro molecules. |
| 9 | 91.09 | CNKSR3 | ENSBTAG00000010581 | Regulates sodium ion transport. Maintenance of transepithelial sodium transport in the kidneys in humans. |
| 9 | 87.84 | SCAF8 | ENSBTAT00000045257 | Enables mRNA processing and RNA transcription. Enables domain specific binding activity. |
| 9 | 87.84 | TIAM2 | ENSBTAG00000049782 | Encodes Guanine nucleotide exchange factor. Regulates lymphoma. Promotes lung and pancreatic tumour progression in humans. |
| Chr | Position (Mb) | Gene | Gene ID | Function |
|---|---|---|---|---|
| 1 | 6.35 | PAK2 | ENSBTAG00000038996 | Chromosomal alignment in metaphase I. |
| 1 | 25.33 | SLC51A | ENSBTAG00000022314 | Enables transporter activity of bile acid and bile salt. |
| 1 | 117.62 | BRWD1 | ENSBTAG00000023175 | Enables molecular function. Regulation of transcription by RNA polymerase II. Enables gametogenesis and oocyte embryo transition in mice. Epigenomic mediator of normal neurodevelopment in mice. |
| 1 | 62.28 | LLP | ENSBTAG00000046064 | Responsible for cell adhesion and cell motility. |
| 3 | 62 | FP | ENSBTAG00000018542 | Milk fat percentage. |
| 3 | 38.84 | COL241A1 | ENSBTAG00000024802 | Enables extracellular structure constituent. |
| 3 | 12.42 | CLCA2 | ENSBTAG00000035503 | Enables intracellular calcium activated chloride channel activity. |
| 6 | 42.78 | NCAPG | ENSBTAG00000025573 | Mitotic chromosome condensation. |
| 6 | 77.34 | ADGRL3 | ENSBTAG00000032411 | Enables G-protein coupled receptor activity. |
| 18 | 51.42 | Prnp | ENSBTAG00000048903 | Regulation of DNA-binding transcription factor. Regulation of long-term synaptic potentiation. ATP independent protein, amyloid and copper binding. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Magagula, N.A.; Mtileni, B.; Ncube, K.T.; Khanyile, K.S.; Zwane, A.A. Breed-Specific Genetic Recombination Analysis in South African Bonsmara and Nguni Cattle Using Genomic Data. Agriculture 2025, 15, 1846. https://doi.org/10.3390/agriculture15171846
Magagula NA, Mtileni B, Ncube KT, Khanyile KS, Zwane AA. Breed-Specific Genetic Recombination Analysis in South African Bonsmara and Nguni Cattle Using Genomic Data. Agriculture. 2025; 15(17):1846. https://doi.org/10.3390/agriculture15171846
Chicago/Turabian StyleMagagula, Nozipho A., Bohani Mtileni, Keabetswe T. Ncube, Khulekani S. Khanyile, and Avhashoni A. Zwane. 2025. "Breed-Specific Genetic Recombination Analysis in South African Bonsmara and Nguni Cattle Using Genomic Data" Agriculture 15, no. 17: 1846. https://doi.org/10.3390/agriculture15171846
APA StyleMagagula, N. A., Mtileni, B., Ncube, K. T., Khanyile, K. S., & Zwane, A. A. (2025). Breed-Specific Genetic Recombination Analysis in South African Bonsmara and Nguni Cattle Using Genomic Data. Agriculture, 15(17), 1846. https://doi.org/10.3390/agriculture15171846






