Elucidating the Rhizosphere Associated Bacteria for Environmental Sustainability
Abstract
1. Introduction
2. The Rhizosphere Soil as a Treasure Trove for Bacterial Community Concentration
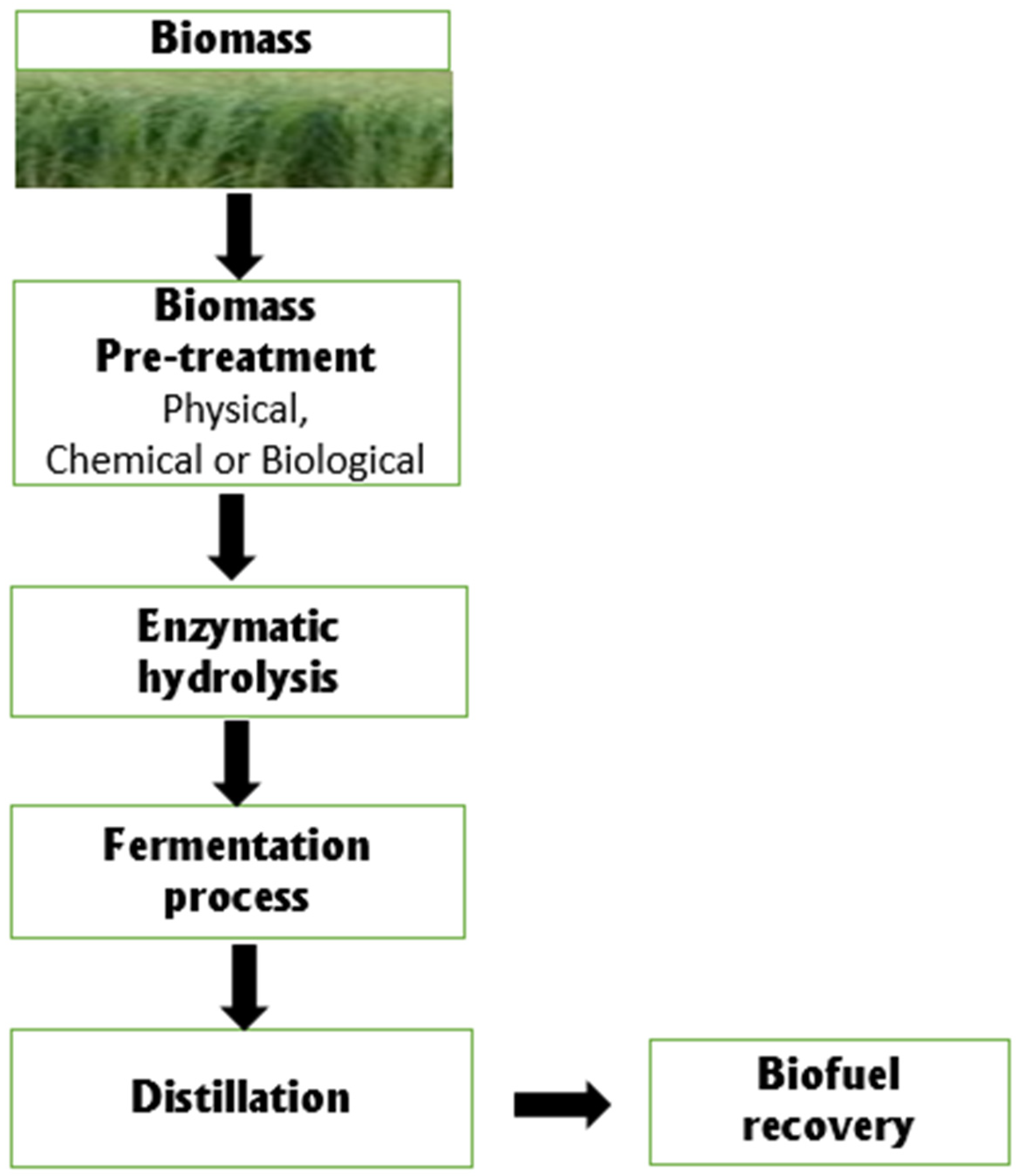
3. Biodegradation of Lignocellulose for Biofuel Production by Rhizospheric Bacteria
4. Biofertilization: The Use of Rhizosphere Bacteria as a Soil Amendment for Plant Growth Promotion
5. The Role of Rhizosphere Soil and Its Bacteria for Bioremediation and Biofiltration
6. Concluding Remarks and Future Perspective
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Ethical Approval
References
- Backer, R.; Rokem, J.S.; Ilangumaran, G.; Lamont, J.; Praslickova, D.; Ricci, E.; Subramanian, S.; Smith, D.L. Plant growth-promoting rhizobacteria: Context, mechanisms of action, and roadmap to commercialization of biostimulants for sustainable agriculture. Front. Plant Sci. 2018, 9, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, A.A.; Häggblom, M.M.; Babalola, O.O. Sustainable agriculture in Africa: Plant growth-promoting rhizobacteria (PGPR) to the rescue. Sci. Afr. 2020, 9, 1–14. [Google Scholar] [CrossRef]
- Zhu, Y. International trade and food security: Conceptual discussion, WTO and the case of China. China Agric. Eco. Rev. 2016, 8, 399–411. [Google Scholar] [CrossRef]
- Igiehon, N.O.; Babalola, O.O. Rhizosphere microbiome modulators: Contributions of nitrogen fixing bacteria towards sustainable agriculture. Int. J. Environ. Res. Public Health 2018, 15, 1–25. [Google Scholar]
- Enebe, M.C.; Babalola, O.O. The influence of plant growth-promoting rhizobacteria in plant tolerance to abiotic stress: A survival strategy. Appl. Microbiol. Biotechnol. 2018, 102, 7821–7835. [Google Scholar] [CrossRef]
- Baghaie, A.H.; Aghilizefreei, A. Neighbor presence of plant growth-promoting rhizobacteria (PGPR) and arbuscular mycorrhizal fungi (AMF) can increase sorghum phytoremediation efficiency in a soil treated with Pb polluted cow manure. J. Hum. Environ. Health Promot. 2019, 5, 153–159. [Google Scholar] [CrossRef]
- Babalola, O.O.; Fadiji, A.E.; Enagbonma, B.J.; Alori, E.T.; Ayilara, M.S.; Ayangbenro, A.S. The Nexus Between Plant and Plant Microbiome: Revelation of the Networking Strategies. Front. Microbiol. 2020, 11, 1–16. [Google Scholar] [CrossRef]
- Yan, Y.; Kuramae, E.E.; de Hollander, M.; Klinkhamer, P.G.; van Veen, J.A. Functional traits dominate the diversity-related selection of bacterial communities in the rhizosphere. Int. Soc. Microb. Ecol. J. 2017, 11, 56–66. [Google Scholar] [CrossRef]
- Keswani, C.; Dilnashin, H.; Birla, H.; Singh, S.P. Regulatory barriers to agricultural research commercialization: A case study of biopesticides in India. Rhizosphere 2019, 11, 1–4. [Google Scholar] [CrossRef]
- Ju, W.; Jin, X.; Liu, L.; Shen, G.; Zhao, W.; Duan, C.; Fang, L. Rhizobacteria inoculation benefits nutrient availability for phytostabilization in copper contaminated soil: Drivers from bacterial community structures in rhizosphere. Appl. Soil Ecol. 2020, 150, 1–13. [Google Scholar] [CrossRef]
- Nuccio, E.E.; Starr, E.; Karaoz, U.; Brodie, E.L.; Zhou, J.; Tringe, S.G.; Malmstrom, R.R.; Woyke, T.; Banfield, J.F.; Firestone, M.K. Niche differentiation is spatially and temporally regulated in the rhizosphere. ISME J. 2020, 14, 999–1014. [Google Scholar] [CrossRef] [PubMed]
- Igiehon, N.O.; Babalola, O.O.; Aremu, B.R. Genomic insights into plant growth promoting rhizobia capable of enhancing soybean germination under drought stress. BMC Microbiol. 2019, 19, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Garrido-Oter, R.; Nakano, R.T.; Dombrowski, N.; Ma, K.-W.; Team, T.A.; McHardy, A.C.; Schulze-Lefert, P. Modular traits of the rhizobiales root microbiota and their evolutionary relationship with symbiotic rhizobia. Cell. Host Microbe 2018, 24, 155–167. [Google Scholar] [CrossRef] [PubMed]
- Ambrosini, A.; Sant’Anna, F.H.; Heinzmann, J.; de Carvalho Fernandes, G.; Bach, E.; Passaglia, L.M.P. Paenibacillus helianthi sp. nov., a nitrogen-fixing species isolated from the rhizosphere of Helianthus annuus L. Antoni. Van Leeuwen. 2018, 111, 2463–2471. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.-F.; Chaparro, J.M.; Reardon, K.F.; Zhang, R.; Shen, Q.; Vivanco, J.M. Rhizosphere interactions: Root exudates, microbes, and microbial communities. Botany 2014, 92, 267–275. [Google Scholar] [CrossRef]
- Yurgel, S.N.; Nearing, J.T.; Douglas, G.M.; Langille, M.G.I. Metagenomic functional shifts to plant induced environmental changes. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Bukhat, S.; Imran, A.; Javaid, S.; Shahid, M.; Majeed, A.; Naqqash, T. Communication of plants with microbial world: Exploring the regulatory networks for PGPR mediated defense signaling. Microbiol. Res. 2020, 238, 126486. [Google Scholar] [CrossRef]
- Odelade, K.A.; Babalola, O.O. Bacteria, fungi and archaea domains in rhizospheric soil and their effects in enhancing agricultural productivity. Int. J. Environ. Res. Public Health 2019, 16, 3873. [Google Scholar] [CrossRef]
- De Luna, L.Z.; Stubbs, T.L.; Kennedy, A.C.; Kremer, R.J. Deleterious bacteria in the rhizosphere. In Roots and Soil Management: Interactions between Roots and the Soil; American Society of Agronomy, Crop Science Society of America, Soil Science Society of America: Madison, WI, USA, 2005; Volume 48, pp. 233–261. [Google Scholar]
- Long, X.E.; Yao, H.; Huang, Y.; Wei, W.; Zhu, Y.G. Phosphate levels influence the utilisation of rice rhizodeposition carbon and the phosphate-solubilising microbial community in a paddy soil. Soil Biol. Biochem. 2018, 118, 103–114. [Google Scholar] [CrossRef]
- Tian, T.; Reverdy, A.; She, Q.; Sun, B.; Chai, Y. The role of rhizodeposits in shaping rhizomicrobiome. Environ. Microbiol. Rep. 2020, 12, 160–172. [Google Scholar] [CrossRef]
- Bakker, M.G.; Manter, D.K.; Sheflin, A.M.; Weir, T.L.; Vivanco, J.M. Harnessing the rhizosphere microbiome through plant breeding and agricultural management. Plant Soil 2012, 360, 1–13. [Google Scholar] [CrossRef]
- Dennis, P.G.; Miller, A.J.; Hirsch, P.R. Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbiol. Ecol. 2010, 72, 313–327. [Google Scholar] [CrossRef] [PubMed]
- Sasse, J.; Martinoia, E.; Northen, T. Feed your friends: Do plant exudates shape the root microbiome? Trends Plant Sci. 2018, 23, 25–41. [Google Scholar] [CrossRef] [PubMed]
- Lettice, E.P. The rhizosphere: Measuring the zone of interaction. Ann. Plant Rev. Online 2018, 2, 219–236. [Google Scholar]
- Lu, G.-H.; Hua, X.-M.; Liang, L.; Wen, Z.-L.; Du, M.-H.; Meng, F.-F.; Pang, Y.-J.; Qi, J.-L.; Tang, C.-Y.; Yang, Y.-H. Identification of major rhizobacterial taxa affected by a glyphosate-tolerant soybean line via shotgun metagenomic approach. Genes 2018, 9, 214. [Google Scholar] [CrossRef]
- Kari, A.; Nagymáté, Z.; Romsics, C.; Vajna, B.; Kutasi, J.; Puspán, I.; Kárpáti, É.; Kovács, R.; Márialigeti, K. Monitoring of soil microbial inoculants and their impact on maize (Zea mays L.) rhizosphere using T-RFLP molecular fingerprint method. Appl. Soil Ecol. 2019, 138, 233–244. [Google Scholar] [CrossRef]
- Bais, H.P.; Weir, T.L.; Perry, L.G.; Gilroy, S.; Vivanco, J.M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 2006, 57, 233–266. [Google Scholar] [CrossRef]
- Phour, M.; Sehrawat, A.; Sindhu, S.S.; Glick, B.R. Interkingdom signaling in plant-rhizomicrobiome interactions for sustainable agriculture. Microbiol. Res. 2020, 241, 1–19. [Google Scholar] [CrossRef]
- Naylor, D.; Coleman-Derr, D. Drought stress and root-associated bacterial communities. Front. Plant Sci. 2018, 8, 1–16. [Google Scholar] [CrossRef]
- Sarhan, M.S.; Patz, S.; Hamza, M.A.; Youssef, H.H.; Mourad, E.F.; Fayez, M.; Murphy, B.; Ruppel, S.; Hegazi, N.A. G3 PhyloChip analysis confirms the promise of plant-based culture media for unlocking the composition and diversity of the maize root microbiome and for recovering unculturable candidate divisions/phyla. Microbes Environ. 2018, 33, 1–9. [Google Scholar] [CrossRef]
- Palansooriya, K.N.; Shaheen, S.M.; Chen, S.S.; Tsang, D.C.; Hashimoto, Y.; Hou, D.; Bolan, N.S.; Rinklebe, J.; Ok, Y.S. Soil amendments for immobilization of potentially toxic elements in contaminated soils: A critical review. Environ. Int. 2020, 134, 1–29. [Google Scholar] [CrossRef] [PubMed]
- dos Santos, J.J.; Maranho, L.T. Rhizospheric microorganisms as a solution for the recovery of soils contaminated by petroleum: A review. J. Environ. Manag. 2018, 210, 104–113. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Ge, T.; Liu, S.; Hu, Y.; Ye, R.; Xiao, M.; Tong, C.; Kuzyakov, Y.; Wu, J. Rice rhizodeposits affect organic matter priming in paddy soil: The role of N fertilization and plant growth for enzyme activities, CO2 and CH4 emissions. Soil Biol. Biochem. 2018, 116, 369–377. [Google Scholar] [CrossRef]
- Gómez Expósito, R.; de Bruijn, I.; Postma, J.; Raaijmakers, J.M. Current insights into the role of rhizosphere bacteria in disease suppressive soils. Front. Microbiol. 2017, 8, 1–12. [Google Scholar] [CrossRef]
- Chaudhary, T.; Shukla, P. Bioinoculants for bioremediation applications and disease resistance: Innovative perspectives. Indian J. Microbiol. 2019, 59, 129–136. [Google Scholar] [CrossRef]
- Zhang, Y.; Gao, X.; Wang, S.; Zhu, C.; Li, R.; Shen, Q. Application of Bacillus velezensis NJAU-Z9 enhanced plant growth associated with efficient rhizospheric colonization monitored by qPCR with primers designed from the whole genome sequence. Curr. Microbiol. 2018, 75, 1574–1583. [Google Scholar] [CrossRef]
- Wang, X.; Bei, Q.; Yang, W.; Zhang, H.; Hao, J.; Qian, L.; Feng, Y.; Xie, Z. Unveiling of active diazotrophs in a flooded rice soil by combination of NanoSIMS and 15 N 2-DNA-stable isotope probing. Biol. Fert. Soils 2020, 56, 1189–1199. [Google Scholar] [CrossRef]
- Matthews, A.; Pierce, S.; Hipperson, H.; Raymond, B. Rhizobacterial community assembly patterns vary between crop species. Front. Microbiol. 2019, 10, 581. [Google Scholar] [CrossRef]
- Zhong, Y.; Yang, Y.; Liu, P.; Xu, R.; Rensing, C.; Fu, X.; Liao, H. Genotype and rhizobium inoculation modulate the assembly of soybean rhizobacterial communities. Plant Cell Environ. 2019, 42, 2028–2044. [Google Scholar] [CrossRef]
- Kumar, P.; Thakur, S.; Dhingra, G.; Singh, A.; Pal, M.K.; Harshvardhan, K.; Dubey, R.; Maheshwari, D. Inoculation of siderophore producing rhizobacteria and their consortium for growth enhancement of wheat plant. Biocatal. Agric. Biotechnol. 2018, 15, 264–269. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, W.; Zhou, D.; Jing, T.; Li, K.; Zhao, Y.; Tang, W.; Qi, D.; Zhang, M.; Zang, X.; et al. Biodegradation of lignocellulosic agricultural residues by a newly isolated Fictibacillus sp. YS-26 improving carbon metabolic properties and functional diversity of the rhizosphere microbial community. Bioresour. Technol. 2020, 310, 123381. [Google Scholar] [CrossRef] [PubMed]
- Sharma, H.K.; Xu, C.; Qin, W. Biological pretreatment of lignocellulosic biomass for biofuels and bioproducts: An overview. Waste Biomass Valoriz. 2019, 10, 235–251. [Google Scholar] [CrossRef]
- Enagbonma, B.J.; Babalola, O.O. Potentials of termite mound soil bacteria in ecosystem engineering for sustainable agriculture. Ann. Microbiol. 2019, 69, 211–219. [Google Scholar] [CrossRef]
- Mathews, S.L.; Pawlak, J.; Grunden, A.M. Bacterial biodegradation and bioconversion of industrial lignocellulosic streams. Appl. Microbiol. Biotechnol. 2015, 99, 2939–2954. [Google Scholar] [CrossRef]
- Zwetsloot, M.J.; Ucros, J.M.; Wickings, K.; Wilhelm, R.C.; Sparks, J.; Buckley, D.H.; Bauerle, T.L. Prevalent root-derived phenolics drive shifts in microbial community composition and prime decomposition in forest soil. Soil. Biol. Biochem. 2020, 145, 1–13. [Google Scholar] [CrossRef]
- Harun, R.; Danquah, M.K.; Forde, G.M. Microalgal biomass as a fermentation feedstock for bioethanol production. J. Chem. Technol. Biotechnol. 2010, 85, 199–203. [Google Scholar] [CrossRef]
- Morgan Jr, H.M.; Bu, Q.; Liang, J.; Liu, Y.; Mao, H.; Shi, A.; Lei, H.; Ruan, R. A review of catalytic microwave pyrolysis of lignocellulosic biomass for value-added fuel and chemicals. Biores. Technol. 2017, 230, 112–121. [Google Scholar] [CrossRef]
- Velvizhi, G.; Balakumar, K.; Dharanidharan, S. Sequestering of CO2 to value-added products through various biological processes. In Next Generation Biomanufacturing Technologies; American Chemical Society Publications: Washington, DC, USA, 2019; pp. 261–284. [Google Scholar]
- Dou, T.-Y.; Liu, W.-J.; Chen, J. Isolation, substrate specificity, and subunit characterization of the Xylanosomes Produced by Oerskovia turbata JCM 3160. Curr. Microbiol. 2020, 77, 924–930. [Google Scholar] [CrossRef]
- Liu, H.; Sun, J.; Chang, J.S.; Shukla, P. Engineering microbes for direct fermentation of cellulose to bioethanol. Crit. Rev. Biotechnol. 2018, 38, 1089–1105. [Google Scholar] [CrossRef]
- Umadevi, P.; Anandaraj, M.; Srivastav, V.; Benjamin, S. Trichoderma harzianum MTCC 5179 impacts the population and functional dynamics of microbial community in the rhizosphere of black pepper (Piper nigrum L.). Braz. J. Microbiol. 2018, 49, 463–470. [Google Scholar] [CrossRef]
- Mood, S.H.; Golfeshan, A.H.; Tabatabaei, M.; Jouzani, G.S.; Najafi, G.H.; Gholami, M.; Ardjmand, M. Lignocellulosic biomass to bioethanol, a comprehensive review with a focus on pretreatment. Renew. Sustain. Energy Rev. 2013, 27, 77–93. [Google Scholar] [CrossRef]
- Alvira, P.; Tomás-Pejó, E.; Ballesteros, M.; Negro, M. Pretreatment technologies for an efficient bioethanol production process based on enzymatic hydrolysis: A review. Biores. Technol. 2010, 101, 4851–4861. [Google Scholar] [CrossRef] [PubMed]
- Xin, F.; Chen, T.; Jiang, Y.; Dong, W.; Zhang, W.; Zhang, M.; Wu, H.; Ma, J.; Jiang, M. Strategies for improved isopropanol–butanol production by a Clostridium strain from glucose and hemicellulose through consolidated bioprocessing. Biotechnol. Biofuels 2017, 10, 118. [Google Scholar] [CrossRef] [PubMed]
- Wierzbicka-Woś, A.; Henneberger, R.; Batista-García, R.A.; Martínez-Ávila, L.; Jackson, S.A.; Kennedy, J.; Dobson, A.D.W. Biochemical characterization of a novel monospecific endo-β-1,4-glucanase belonging to GH family 5 from a rhizosphere metagenomic library. Front. Microbiol. 2019, 10, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Rai, M.; Ingle, A.P.; Pandit, R.; Paralikar, P.; Biswas, J.K.; da Silva, S.S. Emerging role of nanobiocatalysts in hydrolysis of lignocellulosic biomass leading to sustainable bioethanol production. Catal. Rev. 2019, 61, 1–26. [Google Scholar] [CrossRef]
- Valliammai, M.G.; Gopal, N.O.; Anandham, R. Elucidation of microbial diversity and lignocellulolytic enzymes for the degradation of lignocellulosic biomass in the forest soils of Eastern and Western Ghats of Tamil Nadu, India. Biofuels Bioprod. Bioref. 2021, 15, 47–60. [Google Scholar] [CrossRef]
- Artzi, L.; Bayer, E.A.; Moraïs, S. Cellulosomes: Bacterial nanomachines for dismantling plant polysaccharides. Nat. Rev. Microbiol. 2017, 15, 83. [Google Scholar] [CrossRef]
- Hyeon, J.E.; You, S.K.; Kang, D.H.; Ryu, S.-H.; Kim, M.; Lee, S.-S.; Han, S.O. Enzymatic degradation of lignocellulosic biomass by continuous process using laccase and cellulases with the aid of scaffoldin for ethanol production. Process Biochem. 2014, 49, 1266–1273. [Google Scholar] [CrossRef]
- Prasad, R.K.; Chatterjee, S.; Mazumder, P.B.; Gupta, S.K.; Sharma, S.; Vairale, M.G.; Datta, S.; Dwivedi, S.K.; Gupta, D.K. Bioethanol production from waste lignocelluloses: A review on microbial degradation potential. Chemosphere 2019, 231, 588–606. [Google Scholar] [CrossRef]
- Silva, C.O.; Vaz, R.P.; Filho, E.X. Bringing plant cell wall-degrading enzymes into the lignocellulosic biorefinery concept. Biofuels Bioprod. Bioref. 2018, 12, 277–289. [Google Scholar] [CrossRef]
- Behera, B.; Sethi, B.; Mishra, R.; Dutta, S.; Thatoi, H. Microbial cellulases–Diversity and biotechnology with reference to mangrove environment: A review. J. Genet. Eng. Biotechnol. 2017, 15, 197–210. [Google Scholar] [CrossRef] [PubMed]
- Dou, T.-Y.; Chen, J.; Liu, W.-J.; Wang, L. Isolation and subunit structure of the xylanosome complex produced by Actinotalea fermentans JCM9966. Biotechnol. Lett. 2020, 42, 143–149. [Google Scholar] [CrossRef] [PubMed]
- Dobrovolskaya, T.; Golovchenko, A.; Lysak, L.; Yurchenko, E. Taxonomic structure of bacterial communities of rhizospheric soil under bogs’ plants. Mosc. Univ. Soil Sci. Bull. 2020, 75, 93–100. [Google Scholar] [CrossRef]
- Bao, Y.; Dolfing, J.; Wang, B.; Chen, R.; Huang, M.; Li, Z.; Lin, X.; Feng, Y. Bacterial communities involved directly or indirectly in the anaerobic degradation of cellulose. Biol. Fert. Soils. 2019, 55, 201–211. [Google Scholar] [CrossRef]
- Taubert, M.; Stähly, J.; Kolb, S.; Küsel, K. Divergent microbial communities in groundwater and overlying soils exhibit functional redundancy for plant-polysaccharide degradation. PLoS ONE 2019, 14, e0212937. [Google Scholar] [CrossRef] [PubMed]
- Costa, O.Y.; Pijl, A.; Kuramae, E.E. Dynamics of active potential bacterial and fungal interactions in the assimilation of acidobacterial EPS in soil. Soil Biol. Biochem. 2020, 148, 107916. [Google Scholar] [CrossRef]
- Saleh, D.; Jarry, J.; Rani, M.; Aliferis, K.; Seguin, P.; Jabaji, S.H. Diversity, distribution and multi-functional attributes of bacterial communities associated with the rhizosphere and endosphere of timothy (Phleum pratense L.). J. Appl. Microbiol. 2019, 127, 794–811. [Google Scholar] [CrossRef]
- Mukhtar, S.; Mehnaz, S.; Mirza, M.S.; Malik, K.A. Isolation and characterization of bacteria associated with the rhizosphere of halophytes (Salsola stocksii and Atriplex amnicola) for production of hydrolytic enzymes. Braz. J. Microbiol. 2019, 50, 85–97. [Google Scholar] [CrossRef]
- Adegboye, M.F.; Lobb, B.; Babalola, O.O.; Doxey, A.C.; Ma, K. Draft genome sequences of two novel cellulolytic Streptomyces strains isolated from South African rhizosphere soil. Genome Announc. 2018, 6, e00632–e1618. [Google Scholar] [CrossRef]
- Lasa, A.V.; Mašínová, T.; Baldrian, P.; Fernández-López, M. Bacteria from the endosphere and rhizosphere of Quercus spp. use mainly cell wall-associated enzymes to decompose organic matter. PLoS ONE 2019, 14, e0214422. [Google Scholar] [CrossRef]
- Yadav, A.N.; Verma, P.; Kumar, S.; Kumar, V.; Kumar, M.; Sugitha, T.C.K.; Singh, B.P.; Saxena, A.K.; Dhaliwal, H.S. Actinobacteria from rhizosphere: Molecular diversity, distributions, and potential biotechnological applications. In New and Future Developments in Microbial Biotechnology and Bioengineering; Elsevier: Amsterdam, The Netherlands, 2018; pp. 13–41. [Google Scholar]
- Patel, V.; Sharma, A.; Lal, R.; Al-Dhabi, N.A.; Madamwar, D. Response and resilience of soil microbial communities inhabiting in edible oil stress/contamination from industrial estates. BMC Microbiol. 2016, 16, 50. [Google Scholar] [CrossRef] [PubMed]
- Kour, D.; Rana, K.L.; Yadav, N.; Yadav, A.N.; Kumar, A.; Meena, V.S.; Singh, B.; Chauhan, V.S.; Dhaliwal, H.S.; Saxena, A.K. Rhizospheric microbiomes: Biodiversity, mechanisms of plant growth promotion, and biotechnological applications for sustainable agriculture. In Plant Growth Promoting Rhizobacteria for Agricultural Sustainability; Springer: Singapore, 2019; pp. 19–65. [Google Scholar]
- Song, C.W.; Park, J.M.; Chung, S.C.; Lee, S.Y.; Song, H. Microbial production of 2,3-butanediol for industrial applications. J. Indust. Microbiol. Biotechnol. 2019, 46, 1583–1601. [Google Scholar] [CrossRef] [PubMed]
- Wieczorek, A.S.; Schmidt, O.; Chatzinotas, A.; von Bergen, M.; Gorissen, A.; Kolb, S. Ecological functions of agricultural soil bacteria and microeukaryotes in chitin degradation: A case study. Front. Microbiol. 2019, 10. [Google Scholar] [CrossRef]
- Gkarmiri, K.; Mahmood, S.; Ekblad, A.; Alström, S.; Högberg, N.; Finlay, R. Identifying the active microbiome associated with roots and rhizosphere soil of oilseed rape. Appl. Environ. Microbiol. 2017, 83, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Kielak, A.M.; Castellane, T.C.; Campanharo, J.C.; Colnago, L.A.; Costa, O.Y.; Da Silva, M.L.C.; Van Veen, J.A.; Lemos, E.G.; Kuramae, E.E. Characterization of novel Acidobacteria exopolysaccharides with potential industrial and ecological applications. Sci. Rep. 2017, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; Wang, X.; Cao, P.; Gao, Z.; Chen, A.J.; Han, J. Microbial community changes in the rhizosphere soil of healthy and rusty Panax ginseng and discovery of pivotal fungal genera associated with rusty roots. BioMed. Res. Int. 2020, 2020, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Antar, M.; Gopal, P.; Msimbira, L.A.; Naamala, J.; Nazari, M.; Overbeek, W.; Backer, R.; Smith, D.L. Inter-organismal signaling in the rhizosphere. In Rhizosphere Biology: Interactions between Microbes and Plants; Springer: Singapore, 2020; pp. 255–293. [Google Scholar]
- Itelima, J.U.; Bang, W.J.; Onyimba, I.A.; Oj, E. A review: Biofertilizer; a key player in enhancing soil fertility and crop productivity. J. Microbiol. Biotechnol. Rep. 2018, 2, 22–28. [Google Scholar]
- Hamid, Y.; Tang, L.; Sohail, M.I.; Cao, X.; Hussain, B.; Aziz, M.Z.; Usman, M.; He, Z.-l.; Yang, X. An explanation of soil amendments to reduce cadmium phytoavailability and transfer to food chain. Sci. Total Environ. 2019, 660, 80–96. [Google Scholar] [CrossRef]
- Ghori, Z.; Iftikhar, H.; Bhatti, M.F.; Sharma, I.; Kazi, A.G.; Ahmad, P. Phytoextraction: The use of plants to remove heavy metals from soil. In Plant Metal Interaction; Elsevier: Amsterdam, The Netherlands, 2016; pp. 385–409. [Google Scholar]
- Cui, Y.; Bing, H.; Fang, L.; Wu, Y.; Yu, J.; Shen, G.; Jiang, M.; Wang, X.; Zhang, X. Diversity patterns of the rhizosphere and bulk soil microbial communities along an altitudinal gradient in an alpine ecosystem of the eastern Tibetan Plateau. Geoderma 2019, 338, 118–127. [Google Scholar] [CrossRef]
- Fan, K.; Weisenhorn, P.; Gilbert, J.A.; Shi, Y.; Bai, Y.; Chu, H. Soil pH correlates with the co-occurrence and assemblage process of diazotrophic communities in rhizosphere and bulk soils of wheat fields. Soil Biol.Biochem. 2018, 121, 185–192. [Google Scholar] [CrossRef]
- Angst, G.; Messinger, J.; Greiner, M.; Häusler, W.; Hertel, D.; Kirfel, K.; Kögel-Knabner, I.; Leuschner, C.; Rethemeyer, J.; Mueller, C.W. Soil organic carbon stocks in topsoil and subsoil controlled by parent material, carbon input in the rhizosphere, and microbial-derived compounds. Soil Biol. Biochem. 2018, 122, 19–30. [Google Scholar] [CrossRef]
- Maseko, I.; Beletse, Y.G.; Nogemane, N.; du Plooy, C.P.; Musimwa, T.R.; Mabhaudhi, T. Productivity of non-heading Chinese cabbage (Brassica rapa subsp. chinensis) under different agronomic management factors. S. Afr. J. Plant Soil 2017, 34, 275–282. [Google Scholar] [CrossRef]
- Deng, S.; Wipf, H.M.-L.; Pierroz, G.; Raab, T.K.; Khanna, R.; Coleman-Derr, D. A plant growth-promoting microbial soil amendment dynamically alters the strawberry root bacterial microbiome. Sci. Rep. 2019, 9, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Calvo, P.; Zebelo, S.; McNear, D.; Kloepper, J.; Fadamiro, H. Plant growth-promoting rhizobacteria induce changes in Arabidopsis thaliana gene expression of nitrate and ammonium uptake genes. J. Plant Interact. 2019, 14, 224–231. [Google Scholar] [CrossRef]
- Saleem, M.; Asghar, H.N.; Zahir, Z.A.; Shahid, M. Impact of lead tolerant plant growth promoting rhizobacteria on growth, physiology, antioxidant activities, yield and lead content in sunflower in lead contaminated soil. Chemosphere 2018, 195, 606–614. [Google Scholar] [CrossRef] [PubMed]
- Grobelak, A.; Kokot, P.; Hutchison, D.; Grosser, A.; Kacprzak, M. Plant growth-promoting rhizobacteria as an alternative to mineral fertilizers in assisted bioremediation-sustainable land and waste management. J. Environ. Manag. 2018, 227, 1–9. [Google Scholar] [CrossRef]
- Ren, H.; Huang, B.; Fernández-García, V.; Miesel, J.; Yan, L.; Lv, C. Biochar and rhizobacteria amendments improve several soil properties and bacterial diversity. Microorganisms 2020, 8, 502. [Google Scholar] [CrossRef]
- Chenniappan, C.; Narayanasamy, M.; Daniel, G.; Ramaraj, G.; Ponnusamy, P.; Sekar, J.; Ramalingam, P.V. Biocontrol efficiency of native plant growth promoting rhizobacteria against rhizome rot disease of turmeric. Biol. Cont. 2019, 129, 55–64. [Google Scholar] [CrossRef]
- Timmusk, S.; Behers, L.; Muthoni, J.; Muraya, A.; Aronsson, A.C. Perspectives and challenges of microbial application for crop improvement. Front. Plant Sci. 2017, 8, 49. [Google Scholar] [CrossRef]
- Singh, S.K.; Singh, P.P.; Gupta, A.; Singh, A.K.; Keshri, J. Tolerance of heavy metal toxicity using PGPR strains of Pseudomonas species. In PGPR Amelioration in Sustainable Agriculture; Woodhead publishing, Elsevier: Duxford, UK, 2019; Volume 140, pp. 239–252. [Google Scholar]
- Correa-García, S.; Rheault, K.; Tremblay, J.; Séguin, A.; Yergeau, E. Soil characteristics constrain the response of microbial communities and associated hydrocarbon degradation genes during phytoremediation. Appl. Environ. Microbiol. 2020. [Google Scholar] [CrossRef]
- Kalaimurugan, D.; Balamuralikrishnan, B.; Durairaj, K.; Vasudhevan, P.; Shivakumar, M.; Kaul, T.; Chang, S.; Ravindran, B.; Venkatesan, S. Isolation and characterization of heavy-metal-resistant bacteria and their applications in environmental bioremediation. Int. J. Environ. Sci. Technol. 2020, 17, 1455–1462. [Google Scholar] [CrossRef]
- Asilian, E.; Ghasemi-Fasaei, R.; Ronaghi, A.; Sepehri, M.; Niazi, A. Chemical-and microbial-enhanced phytoremediation of cadmium-contaminated calcareous soil by maize. Toxicol. Indust. Health 2019, 35, 378–386. [Google Scholar] [CrossRef] [PubMed]
- Mourato, M.P.; Moreira, I.N.; Leitão, I.; Pinto, F.R.; Sales, J.R.; Martins, L.L. Effect of heavy metals in plants of the genus Brassica. Int. J. Mol. Sci. 2015, 16, 17975–17998. [Google Scholar] [CrossRef] [PubMed]
- Tangahu, B.V.; Sheikh Abdullah, S.R.; Basri, H.; Idris, M.; Anuar, N.; Mukhlisin, M. A review on heavy metals (As, Pb, and Hg) uptake by plants through phytoremediation. Int. J.Chem. Eng. 2011, 939161. [Google Scholar] [CrossRef]
- Kumar, P.; Fulekar, M.H. Research article rhizosphere bioremediation of heavy metals (copper and lead) by Cenchrus ciliaris. J. Environ. Sci. 2018, 12, 166–176. [Google Scholar]
- Xu, S.; Xing, Y.; Liu, S.; Hao, X.; Chen, W.; Huang, Q. Characterization of Cd2+ biosorption by Pseudomonas sp. strain 375, a novel biosorbent isolated from soil polluted with heavy metals in Southern China. Chemosphere 2020, 240, 124893. [Google Scholar] [CrossRef]
- Canizo, B.V.; Agostini, E.; Oller, A.L.W.; Dotto, G.L.; Vega, I.A.; Escudero, L.B. Removal of crystal violet from natural water and effluents through biosorption on bacterial biomass isolated from rhizospheric soil. Water Air Soil Poll. 2019, 230, 210. [Google Scholar] [CrossRef]
- Yuan, J.; Yuan, Y.; Zhu, Y.; Cao, L. Effects of different fertilizers on methane emissions and methanogenic community structures in paddy rhizosphere soil. Sci. Total Environ. 2018, 627, 770–781. [Google Scholar] [CrossRef]
- Aimen, H.; Khan, A.S.; Kanwal, N. Methanotrophs: The natural way to tackle greenhouse effect. J. Biorem. Biodeg. 2018, 9, 432. [Google Scholar] [CrossRef]
- Davamani, V.; Parameswari, E.; Arulmani, S. Mitigation of methane gas emissions in flooded paddy soil through the utilization of methanotrophs. Sci. Total Environ. 2020, 726, 138570. [Google Scholar] [CrossRef]
- Viesser, J.A.; Sugai-Guerios, M.H.; Malucelli, L.C.; Pincerati, M.R.; Karp, S.G.; Maranho, L.T. Petroleum-tolerant rhizospheric bacteria: Isolation, characterization and bioremediation potential. Sci. Rep. 2020, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Chang, Y.C.; Reddy, M.V.; Umemoto, H.; Kondo, S.; Choi, D. Biodegradation of alkylphenols by rhizosphere microorganisms isolated from the roots of Hosta undulata. J. Environ. Chem. Eng. 2020, 8, 103771. [Google Scholar] [CrossRef]
- Jiao, S.; Li, Q.; Zai, X.; Gao, X.; Wei, G.; Chen, W. Complexity of bacterial communities within the rhizospheres of legumes drives phenanthrene degradation. Geoderma 2019, 353, 1–10. [Google Scholar] [CrossRef]
- Toyama, T.; Furukawa, T.; Maeda, N.; Inoue, D.; Sei, K.; Mori, K.; Kikuchi, S.; Ike, M. Accelerated biodegradation of pyrene and benzo [a] pyrene in the Phragmites australis rhizosphere by bacteria–root exudate interactions. Water Res. 2011, 45, 1629–1638. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Dong, G.; Liu, X.; Zhang, S.; Li, C.; Lu, X.; Xia, T. Poly-γ-glutamic acid-producing bacteria reduced Cd uptake and effected the rhizosphere microbial communities of lettuce. J. Hazard. Mater. 2020, 398, 123146. [Google Scholar] [CrossRef]
- Chaturvedi, S.; Chandra, R.; Rai, V. Isolation and characterization of Phragmites australis (L.) rhizosphere bacteria from contaminated site for bioremediation of colored distillery effluent. Ecol. Eng. 2006, 27, 202–207. [Google Scholar] [CrossRef]
- Roman-Ponce, B.; Reza-Vázquez, D.M.; Gutierrez-Paredes, S.; María de Jesús, D.E.; Maldonado-Hernandez, J.; Bahena-Osorio, Y.; Estrada-De los Santos, P.; Wang, E.T.; Vásquez-Murrieta, M.S. Plant growth-promoting traits in rhizobacteria of heavy metal-resistant plants and their effects on Brassica nigra seed germination. Pedosphere 2017, 27, 511–526. [Google Scholar] [CrossRef]
- Ahmad, I.; Akhtar, M.J.; Mehmood, S.; Akhter, K.; Tahir, M.; Saeed, M.F.; Hussain, M.B.; Hussain, S. Combined application of compost and Bacillus sp. CIK-512 ameliorated the lead toxicity in radish by regulating the homeostasis of antioxidants and lead. Ecotoxicol. Environ. Saf. 2018, 148, 805–812. [Google Scholar] [CrossRef]
| Technique Used | Bacteria Reported | Plant | Reference |
|---|---|---|---|
| Denaturing Gradient Gel Electrophoresis (DGGE) | Sphingobacteriales, Flavobacteriaceae, Xanthomonadaceae, Cyanobacteria | Lettuce, soybean, potato, maize | [35,36] |
| Quantitative PCR (qPCR) analysis | Bacillus velezensis NJAU-Z9 | Pepper | [37] |
| G3 PhyloChip microarray analyses | Atribacteria, Dependentiae, TM6, Latescibacteria WS3 Marinimicrobia, SAR406; Omnitrophica, OP3; BRC1. Acidobacteria, Gemmatimonadetes, and Tenericutes | Wheat, barley | [31] |
| Restriction Fragment Length Polymorphism (RFLP) | Azospirillum, Pseudomonas chlororaphis, P. frederiksbergensis, Bacillus aryabhattai, and Paenibacillus peoriae | Maize | [27] |
| DNA-Stable Isotope Probing (DNA-SIP) | Nostocales, Stigonematales, Streptomyces Bacillus, Alicyclobacillus, Clostridium. Rhizobiales, Rhodospirillale, Myxococcales, and Actinomycetales | Rice | [20,38] |
| 16S amplicon sequencing | Proteobacteria, Actinobacteria, Bacteroidetes, Acidobacteria, Firmicutes, Verrucomicrobia, Planctomycetes, Actinobacteria, Cyanobacteria, and Gemmatimonadetes | Wheat, maize, potato, soybean | [39,40] |
| Shotgun sequencing | Stenotrophomonas, Rahnella, Sphingomonas, Janthinobacterium Luteibacter, Arthrobacter, Streptomyces, Bradyrhizobium, Methylobacterium, Ramlibacter, Nitrospira, Nocardioides, Geodermatophilus, and Burkholderia | Soybean, sunflower, sugar beet | [26] |
| Culture-based | Bacillus, Pseudomonas, Ochrobactrium, Providencia, Achromobacter, Burkholderia, and Enterobacter | Wheat | [41] |
| Rhizobacteria | Plant | Country | Reference |
|---|---|---|---|
| Arthrobacter, Brevibacterium, Bacillus, Chryseobacterium, Stenotrophomonas, Streptomyces, Pseudomonas, Xanthomonas, Paenibacillus | Phleum pretense L. | Canada | [69] |
| Bacillus, Pseudomonas, Kocuria | Salsola stocksii and Atriplex amnicola | Pakistan | [70] |
| Streptomyces | Zea mays | South Africa | [71] |
| Arthrobacter, Pseudomonas | Quercus sp. | Spain | [72] |
| Bacterial Phylum | Plant Rhizosphere | Biotechnological Product and Application | Industrial Application | Reference |
|---|---|---|---|---|
| Actinobacteria | Helianthus annuus, Zea mays, Triticum aestivum, Glycine max | Kanamycin enhanced shoot growth. | Actinobacteria secrete cellulases suitable for cellulosic biofuel production. | [73,74] |
| Proteobacteria | Zea mays, Oryza sativa, Saccharum officinarum, and Glycine max | Bioinoculants—significantly increased crop yield, biomass dry weight, nodulation, phosphorus, and nitrogen uptake. Bioprotectants—protect plant from phytopathogens. | Bioremediation strategies for the degradation of oil spill contamination (edible such as fats and lipids; and crude oil) as well as carbamate and organophosphate insecticides. Zymomonas—produces an abundance of alcohol for industrial use. Acetic acid bacteria can be employed for the production of acetic acid, vinegar, ascorbic (vitamin c), glucoronic, galactonic, arabonic acids, and sorbose. | [75] |
| Firmicutes | Triticum aestivum and Vitis vinifera | Nitrogen-fixing ability, enhance soil porosity and produce compound similar in activity to indole-3-acetic acid with the capacity to stimulate plant growth. Biocontrol activity effective against a wide range of phytopathogens, such as Fusaricidin identified as a potential antifungal agent, has been identified from P. polymyxa E68. In addition, control Fusarium oxysporum. Flocculants production. | Paenibacillus polymyxa produces 2, 3-butanediol (BDL) forms methyl ethyl ketone used as a liquid fuel additive by dehydration. Produce cell wall degrading enzymes (proteases, β-1,3-glucanases, xylanase, chitinases, and cellulases) available in detergent formulations, leather processing, food industry (starter culture for yogurt production, additives, and beer production), waste management and chemical synthesis. Lactobacillus pentosus has been applied in sulfite waste liquor fermentation. Flocculating or flocking agents used for water treatment. | [76] |
| Bacteroidetes | Brassica napus | Use alternative enzymatic mechanisms to solubilize biopolymers apart from glycosidic hydrolases, the so-called “polysaccharide-utilizer”. Degrade complex polysaccharides in soils and contributes to synergistic breakdown of solubilized chitin oligosaccharides. | Produce enzymes exhibiting activities such as degrading cellulose, lignin or chitin. In addition, various lipids, polysaccharides, or proteins used in industries such as leather processing, detergent, paper, and shoe production Used for biofuel production. Used in phytoextraction of heavy metals from polluted soil. | [77,78] |
| Acidobacteria | Castanea crenata, Saccharum officinarum, Vigna mungo and Solanum lycopersicum L. | Produce exopolysaccharide (EPS), which provide protection against environmental stress and enable bacterial survival under unfavorable soil conditions. Form soil matrix, serve to sequester water and nutrition, and are involved in bacterial cell-surface adherence and soil aggregate formation. In addition, they produce plant growth-promoting traits and phytohormones. Produce biofilms, which enhance rhizobacterial root colonization by holding moisture and protect plant roots from phytopathogens. | EPS possess physical and chemical properties such as thickening, gelling, stabilizing, suspending, emulsifying, texture-enhancing, and coagulating. Although some of these bacterial products (e.g., gellan gum, dextran, alginate, and xanthan) have been commercialized successfully in the food and fodder production industries, EPSs are used as gelling, thickening, and suspending agents. For instance, xanthan (from Xanthomonas campestris) is used as a food additive. EPSs are bioemulsifiers that are used in the cosmetic and chemical (e.g., pesticide) industries. Use in environmental technologies, such as phytoremediation and bioremediation in soil and water by enhancing oil and heavy metal recovery. In addition, use in human health and chemical industries. | [79] |
| Nitrospirae | Panax ginseng Meyer | Plant growth promoter: Nitrite-oxidizing bacteria (NOB) are involved in nitrification, including the oxidation processes of ammonia and nitrite. In addition, they are known to convert nitrite to nitrate, improve shoot/root biomass, improve nutrient uptake, alleviate cold stress in plants, and serve as a biocontrol agent. | Potential in the petroleum industry for the exploration of petroleum, clean-up of oil spills both in situ and ex situ conditions and enhance microbial oil recovery. Bioconversion of food waste and activated sewage sludge into useful products. Biohydrometallurgy (microbial recovery of minerals from ores), used for fuel production and for clean-up of oil spills, and deterioration of petroleum products. | [80] |
| Plant | Impact of Rhizobacteria on the Plant | Reference |
|---|---|---|
| Capsicum annuum L. | The soil amended with Bacillus velezensis improves seedling height, stem diameter, and yields compared to those pepper plants grown on un-amended soil | [37] |
| Arabidopsis thaliana | Combined mixture of rhizosphere soil or soil-like substrates and Bacillus mixtures resulted in a significant increase in plant root fresh weight, shoot fresh weight, nutrient uptake, chlorophyll content, and plant diameter. In addition, the transcript levels of ammonium and nitrate uptake genes in the plant were increased | [90] |
| Helianthus annuus | Pseudomonas fluorescens A506, P. gessardii strain BLP141, and P. fluorescens strain LMG 2189 improved plant growth, yield, physiology, proline, antioxidant activities, and reduced the malondialdehyde content in inoculated soil | [91] |
| Ocimum basilicum L. | Rhizobacteria consortium (Bacillus lentus, Pseudomonas sp. and Azospirillum brasilens) had positive effects on the antioxidant activity and chlorophyll pigment content under water-induced and salinity stress | [5] |
| Festuca rubra | Bacterial consortium immobilized in a mixture of perlite and sawdust (ratio 1:1:1 v/v) led to a substantial improvement of plant roots, stem length, and stem biomass, as well as influencing the elongation of the plants in all soil treated. Soil additives (phosphate fertilizer and sewage sludge) and an immobilized consortium of microorganism had a positive effect on plant growth (longer root, stem length, and stem biomass) compared to the control | [92] |
| Eucalyptus globulus | Co-application of biochar (20 t hm−2) and PGPB (5 × 1010 CFU mL−1) amendments significantly decreased the concentrations of soil total P and NH4+-N, whereas they advanced total K, NO3-N, and soil water content, and hence maintained soil sustainability in eucalyptus plantation | [93] |
| Curcuma longa | The Curcuma longa soil amended with B. subtilis MML2490 and P. aeruginosa MML2424 enhanced plant growth promotion and management of turmeric rhizome rot disease, and thus appeared promising for commercialization | [94] |
| Rhizobacteria | Isolation Source | Pollutant | Reference |
|---|---|---|---|
| Bacillus thurigiensis, B. pumilus and Rhodococcus hoagii | Panicum aquaticum | Petroluem | [108] |
| Lysinbacilus fusiformis L8, Bacillus weihenstephanensis UT11, Paenibacillus sp. M10-6, | Hosta undulata | Alkylphenol | [109] |
| Ensifer, Novosphingobium, Norcardioides, Streptomyces, Rhizobium | Coronilla varia, Vigna unguiculata | Phenantrene | [110] |
| Mycobacterium gilvum | Phragmites australis | Pyrene, benzo[a]pyrene | [111] |
| Bacillus subtilis, Bacillus amyloliquefaciens | Lactuca sativa L. | Cadmium | [112] |
| Microbacterium hydrocarbonoxydans, Achromobacter xylosoxidans, Bacillus subtili, B. megaterium, Alcaligens faecalis, Pseudomonas migulae | Phragmites australis | Colored distillery effluent | [113] |
| Alcaligenes, Bacillus, Curtobacterium, Microbacterium | Prosopis laevigata, Spharealce aangustifolia | As(V), Pb(II), Cu(II), Zn(II) | [114] |
| Bacillussp. CIK-512 | Zea mays | Pb | [115] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nwachukwu, B.C.; Ayangbenro, A.S.; Babalola, O.O. Elucidating the Rhizosphere Associated Bacteria for Environmental Sustainability. Agriculture 2021, 11, 75. https://doi.org/10.3390/agriculture11010075
Nwachukwu BC, Ayangbenro AS, Babalola OO. Elucidating the Rhizosphere Associated Bacteria for Environmental Sustainability. Agriculture. 2021; 11(1):75. https://doi.org/10.3390/agriculture11010075
Chicago/Turabian StyleNwachukwu, Blessing Chidinma, Ayansina Segun Ayangbenro, and Olubukola Oluranti Babalola. 2021. "Elucidating the Rhizosphere Associated Bacteria for Environmental Sustainability" Agriculture 11, no. 1: 75. https://doi.org/10.3390/agriculture11010075
APA StyleNwachukwu, B. C., Ayangbenro, A. S., & Babalola, O. O. (2021). Elucidating the Rhizosphere Associated Bacteria for Environmental Sustainability. Agriculture, 11(1), 75. https://doi.org/10.3390/agriculture11010075







