Is Hyperuricemia, an Early-Onset Metabolic Disorder, Causally Associated with Cardiovascular Disease Events in Han Chinese?
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Onset Sequence Study
2.3. UA Mendelian Randomization Study
2.3.1. Han-Chinese SUA-SNP Selection with a Two-Stage GWAS
2.3.2. SUA-SNPs from Literature Review
2.3.3. Calculating the WGRS
2.3.4. Association between SUA and CVD Events
2.4. UAMR Study: Sensitivity Analyses
3. Results
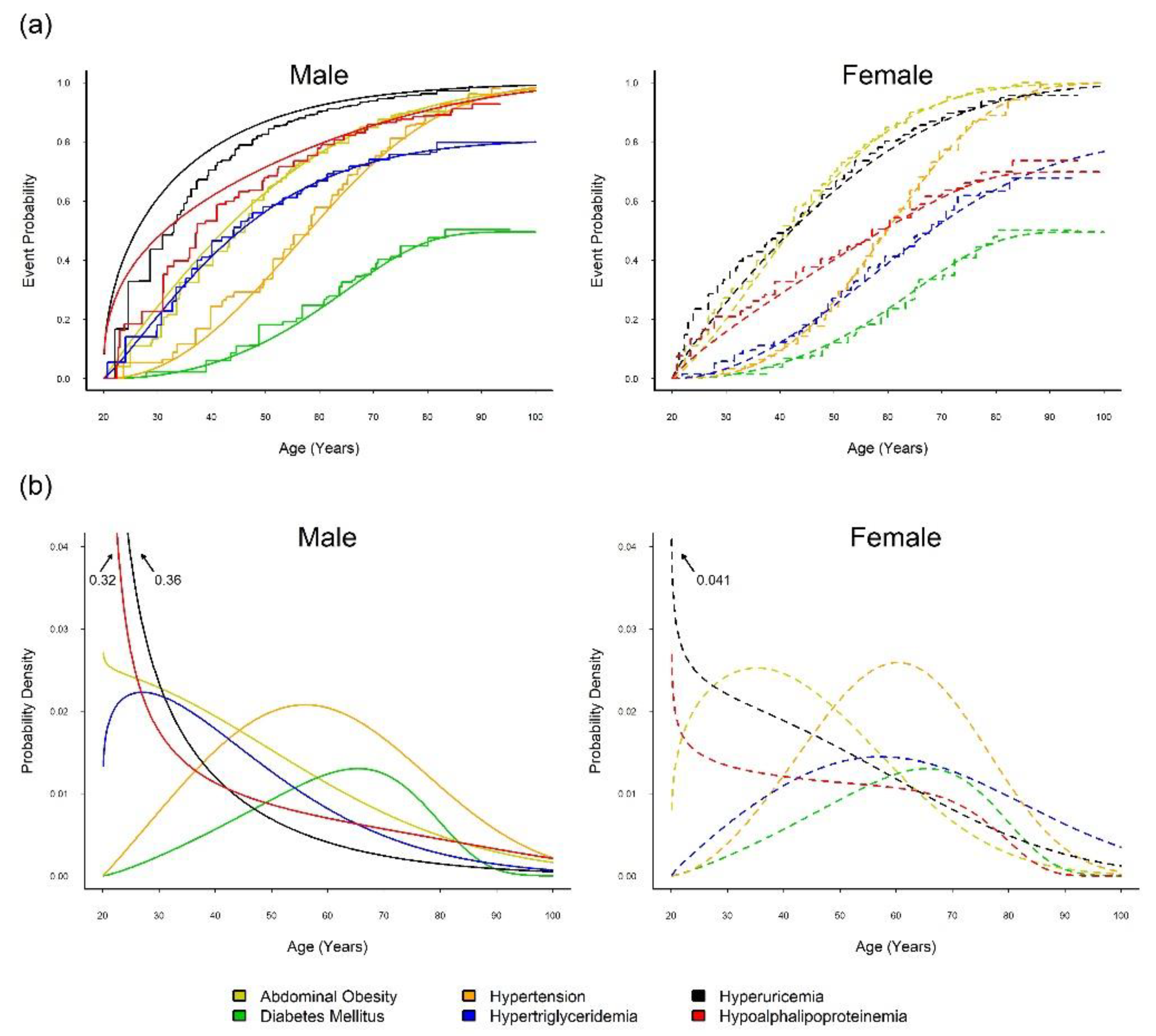
3.1. Onset Sequence Study
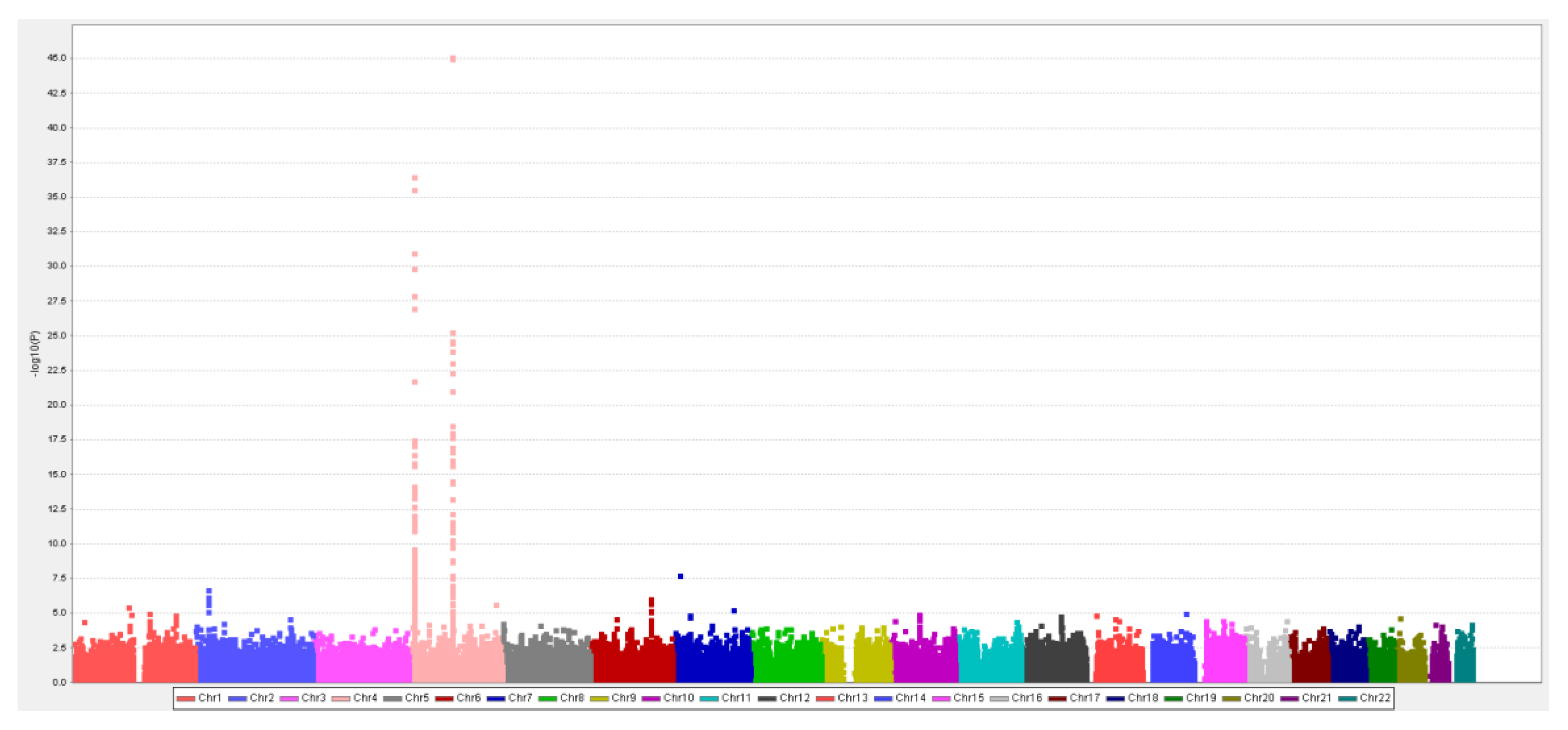
3.2. MR Study: SUA-SNP Discovery and Selection
3.3. MR Study: WGRS
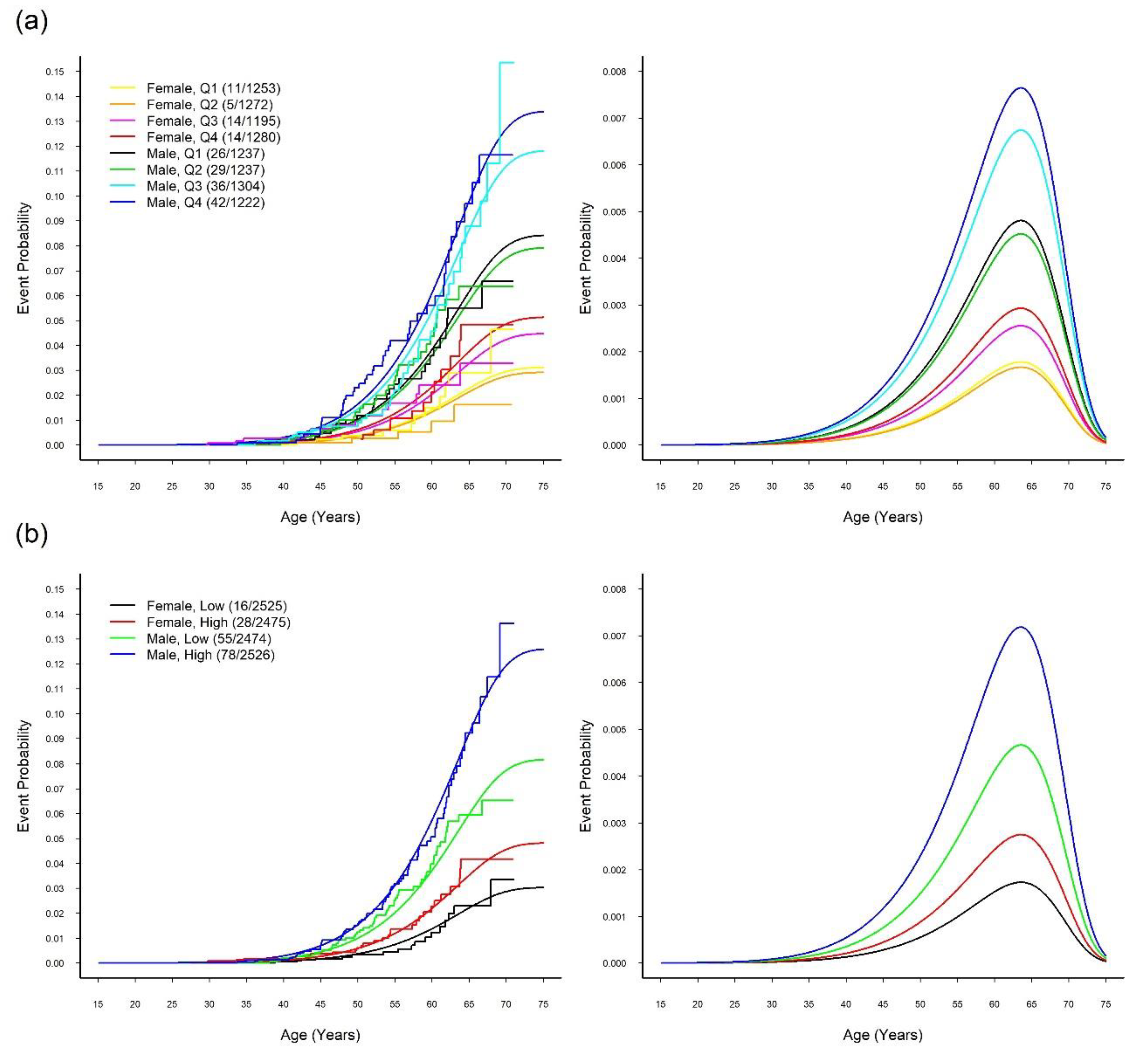
3.4. MR Study: Relationship between SUA and CVD
4. Discussions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Mozaffarian, D.; Benjamin, E.J.; Go, A.S.; Arnett, D.K.; Blaha, M.J.; Cushman, M.; De Ferranti, S.; Després, J.P.; Fullerton, H.J.; Howard, V.J.; et al. Heart disease and stroke statistics 2015 update: A report from the American Heart Association. Circulation 2015, 131, 434–441. [Google Scholar] [CrossRef]
- Keenan, T.; Zhao, W.; Rasheed, A.; Ho, W.K.; Malik, R.; Felix, J.F.; Young, R.; Shah, N.; Samuel, M.; Sheikh, N.; et al. Causal assessment of serum urate levels in cardiometabolic diseases through a Mendelian Randomization study. J. Am. Coll. Cardiol. 2016, 67, 407–416. [Google Scholar] [CrossRef] [PubMed]
- Zhao, G.; Huang, L.; Song, M.; Song, Y. Baseline serum uric acid level as a predictor of cardiovascular disease related mortality and all-cause mortality: A meta-analysis of prospective studies. Atherosclerosis 2013, 231, 61–68. [Google Scholar] [CrossRef] [PubMed]
- Feig, D.I.; Kang, D.H.; Johnson, R.J. Uric Acid and Cardiovascular Risk. N. Engl. J. Med. 2008, 359, 1811–1821. [Google Scholar] [CrossRef] [PubMed]
- Chuang, S.Y.; Chen, J.H.; Yeh, W.T.; Wu, C.C.; Pan, W.H. Hyperuricemia and increased risk of ischemic heart disease in a large Chinese cohort. Int. J. Cardiol. 2012, 154, 316–321. [Google Scholar] [CrossRef] [PubMed]
- Chu, N.F.; Wang, D.J.; Liou, S.H.; Shieh, S.M. Relationship between hyperuricemia and other cardiovascular disease risk factors among adult males in Taiwan. Eur. J. Epidemiol. 2000, 16, 13–17. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Iso, H.; Murakami, Y.; Miura, K.; Nagai, M.; Sugiyama, D.; Ueshima, H.; Okamura, T. Serum Uric Acid and Mortality Form Cardiovascular Disease: EPOCH-JAPAN Study. J. Atheroscler. Thromb. 2016, 23, 692–703. [Google Scholar] [CrossRef]
- Pan, W.H.; Flegal, K.M.; Chang, H.Y.; Yeh, W.T.; Yeh, C.J.; Lee, W.C. Body mass index and obesity-related metabolic disorders in Taiwanese and US whites and blacks: Implications for definitions of overweight and obesity for Asians. Am. J. Clin. Nutr. 2004, 79, 31–39. [Google Scholar] [CrossRef]
- Nejatinamini, S.; Ataie-Jafari, A.; Qorbani, M.; Nikoohemat, S.; Kelishadi, R.; Asayesh, H. Association between serum uric acid level and metabolic syndrome components. J. Diabetes Metab. Disord. 2015, 14, 70. [Google Scholar] [CrossRef]
- Tsay, Y.C.; Chen, C.H.; Pan, W.H. Ages at Onset of 5 Cardiometabolic Diseases Adjusting for Nonsusceptibility: Implications for the Pathogenesis of Metabolic Syndrome. Am. J. Epidemiol. 2016, 184, 366–377. [Google Scholar] [CrossRef]
- Smith, G.D.; Ebrahim, S. ‘Mendelian randomization’: Can genetic epidemiology contribute to understanding environmental determinants of disease? Int. J. Epidemiol. 2003, 32, 1–22. [Google Scholar] [CrossRef] [PubMed]
- Tyrrell, J.; Jones, S.; Beaumont, R.; Astley, C.M.; Lovell, R.; Yaghootkar, H.; Tuke, M.; Ruth, K.S.; Freathy, R.M.; Hirschhorn, J.N.; et al. Height, body mass index, and socioeconomic status: Mendelian randomisation study in UK Biobank. BMJ 2016, 352, i582. [Google Scholar] [CrossRef] [PubMed]
- Glymour, M.M. Alcohol and cardiovascular disease. BMJ 2014, 349, g4334. [Google Scholar] [CrossRef] [PubMed]
- Kleber, M.E.; Delgado, G.; Grammer, T.B.; Silbernagel, G.; Huang, J.; Krämer, B.K.; Ritz, E.; März, W. Uric Acid and Cardiovascular Events: A Mendelian Randomization Study. J. Am. Soc. Nephrol. 2015, 26, 2831–2838. [Google Scholar] [CrossRef] [PubMed]
- White, J.; Sofat, R.; Hemani, G.; Shah, T.; Engmann, J.; Dale, C.; Shah, S.; Kruger, F.A.; Giambartolomei, C.; Swerdlow, D.I.; et al. Plasma urate concentration and risk of coronary heart disease: A Mendelian randomisation analysis. Lancet Diabetes Endocrinol. 2016, 4, 327–336. [Google Scholar] [CrossRef]
- Stark, K.; Reinhard, W.; Grassl, M.; Erdmann, J.; Schunkert, H.; Illig, T.; Hengstenberg, C. Common Polymorphisms Influencing Serum Uric Acid Levels Contribute to Susceptibility to Gout, but Not to Coronary Artery Disease. PLoS ONE 2009, 4, e7729. [Google Scholar] [CrossRef] [PubMed]
- Palmer, T.M.; Nordestgaard, B.G.; Benn, M.; Tybjærg-Hansen, A.; Smith, G.D.; Lawlor, D.A.; Timpson, N.J. Association of plasma uric acid with ischaemic heart disease and blood pressure: Mendelian randomisation analysis of two large cohorts. BMJ 2013, 347, f4262. [Google Scholar] [CrossRef]
- Yang, B.; Mo, Z.; Wu, C.; Yang, H.; Yang, X.; He, Y.; Gui, L.; Zhou, L.; Guo, H.; Zhang, X.; et al. A genome-wide association study identifies common variants influencing serum uric acid concentrations in a Chinese population. BMC Med. Genomics 2014, 7, 10. [Google Scholar] [CrossRef]
- Chen, C.H.; Tsay, Y.C.; Wu, Y.C.; Horng, C.F. Logistic-AFT location-scale mixture regression models with nonsusceptibility for left-truncated and general interval-censored data. Stat. Med. 2013, 32, 4285–4305. [Google Scholar] [CrossRef]
- Chuang, S.Y.; Bai, C.H.; Chen, W.H.; Lien, L.M.; Pan, W.H. Fibrinogen Independently Predicts the Development of Ischemic Stroke in a Taiwanese Population: CVDFACTS Study. Stroke 2009, 40, 1578–1584. [Google Scholar] [CrossRef]
- Weng, L.C.; Yeh, W.T.; Bai, C.H.; Chen, H.J.; Chuang, S.Y.; Chang, H.Y.; Lin, B.F.; Chen, K.J.; Pan, W.H. Is Ischemic Stroke Risk Related to Folate Status or Other Nutrients Correlated With Folate Intake? Stroke 2008, 39, 3152–3158. [Google Scholar] [CrossRef]
- Frydman, H. A Note on Nonparametric Estimation of the Distribution Function from Interval-Censored and Truncated Observations. J. R. Stat. Soc. Ser. B 1994, 56, 71–74. [Google Scholar] [CrossRef]
- Turnbull, B.W. The Empirical Distribution Function with Arbitrarily Grouped, Censored and Truncated Data. J. R. Stat. Soc. Ser. B 1976, 38, 290–295. [Google Scholar] [CrossRef]
- Chen, C.H.; Yang, J.H.; Chiang, C.W.; Hsiung, C.N.; Wu, P.E.; Chang, L.C.; Chu, H.W.; Chang, J.; Song, I.W.; Yang, S.L.; et al. Population structure of Han Chinese in the modern Taiwanese population based on 10,000 participants in the Taiwan Biobank project. Hum. Mol. Genet. 2016, 25, 5321–5331. [Google Scholar] [CrossRef]
- Yang, H.C.; Lin, C.H.; Hsu, C.L.; Hung, S.I.; Wu, J.Y.; Pan, W.H.; Chen, Y.T.; Fann, C.S.J. A comparison of major histocompatibility complex SNPs in Han Chinese residing in Taiwan and Caucasians. J. Biomed. Sci. 2006, 13, 489–498. [Google Scholar] [CrossRef]
- Benjamini, Y.; Drai, D.; Elmer, G.; Kafkafi, N.; Golani, I. Controlling the false discovery rate in behavior genetics research. Behav. Brain Res. 2001, 125, 279–284. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.R.; Bender, D.; Maller, J.; Sklar, P.; De Bakker, P.I.W.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Cheng, Y.C.; Hsiao, F.C.; Yeh, E.C.; Lin, W.J.; Tang, C.Y.L.; Tseng, H.C.; Wu, H.T.; Liu, C.K.; Chen, C.C.; Chen, Y.T.; et al. VarioWatch: Providing large-scale and comprehensive annotations on human genomic variants in the next generation sequencing era. Nucleic Acids Res. 2012, 40, W76–W81. [Google Scholar] [CrossRef]
- Barrett, J.C.; Fry, B.; Maller, J.; Daly, M.J. Haploview: Analysis and visualization of LD and haplotype maps. Bioinformatics 2005, 21, 263–265. [Google Scholar] [CrossRef]
- Mokry, L.E.; Ross, S.; Timpson, N.J.; Sawcer, S.; Smith, G.D.; Richards, J.B. Obesity and Multiple Sclerosis: A Mendelian Randomization Study. PLoS Med. 2016, 13, e1002053. [Google Scholar] [CrossRef]
- Bowden, J.; Smith, G.D.; Burgess, S. Mendelian randomization with invalid instruments: Effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 2015, 44, 512–525. [Google Scholar] [CrossRef]
- Pierce, B.L.; Ahsan, H.; Vanderweele, T.J. Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int. J. Epidemiol. 2011, 40, 740–752. [Google Scholar] [CrossRef]
- Chen, J.H.; Chuang, S.Y.; Chen, H.J.; Yeh, W.T.; Pan, W.H. Serum uric acid level as an independent risk factor for all-cause, cardiovascular, and ischemic stroke mortality: A chinese cohort study. Arthritis Rheum. 2009, 61, 225–232. [Google Scholar] [CrossRef]
- Chen, J.H.; Lan, J.L.; Cheng, C.F.; Liang, W.M.; Lin, H.Y.; Tsay, G.J.; Yeh, W.T.; Pan, W.H. Effect of Urate-Lowering Therapy on All-Cause and Cardiovascular Mortality in Hyperuricemic Patients without Gout: A Case-Matched Cohort Study. PLoS ONE 2015, 10, e0145193. [Google Scholar] [CrossRef]
- Culleton, B.F.; Larson, M.G.; Kannel, W.B.; Levy, D. Serum uric acid and risk for cardiovascular disease and death: The Framingham Heart Study. Ann. Intern. Med. 1999, 131, 7–13. [Google Scholar] [CrossRef]
- Moriarity, J.T.; Folsom, A.R.; Iribarren, C.; Nieto, F.J.; Rosamond, W.D. Serum uric acid and risk of coronary heart disease: Atherosclerosis Risk in Communities (ARIC) Study. Ann. Epidemiol. 2000, 10, 136–143. [Google Scholar] [CrossRef]
- International Diabetes Federation. 2015 Diabetes Atlas; International Diabetes Federation: Brussels, Belgium, 2015. [Google Scholar]
- Chuang, S.Y.; Lee, S.C.; Hsieh, Y.T.; Pan, W.H. Trends in hyperuricemia and gout prevalence: Nutrition and Health Survey in Taiwan from 1993–1996 to 2005–2008. Asia Pac. J. Clin. Nutr. 2011, 20, 301–308. [Google Scholar]
- Annemans, L.; Spaepen, E.; Gaskin, M.; Bonnemaire, M.; Malier, V.; Gilbert, T.; Nuki, G. Gout in the UK and Germany: Prevalence, comorbidities and management in general practice 2000–2005. Ann. Rheum. Dis. 2008, 67, 960–966. [Google Scholar] [CrossRef]
- Feig, D.I.; Soletsky, B.; Johnson, R.J. Effect of Allopurinol on Blood Pressure of Adolescents with Newly Diagnosed Essential Hypertension: A Randomized Trial. JAMA 2008, 300, 924–932. [Google Scholar] [CrossRef]
- Robertson, A.J.; Struthers, A.D. A Randomized Controlled Trial of Allopurinol in Patients with Peripheral Arterial Disease. Can. J. Cardiol. 2016, 32, 190–196. [Google Scholar] [CrossRef]
- Gavin, A.D.; Struthers, A.D. Allopurinol reduces B-type natriuretic peptide concentrations and haemoglobin but does not alter exercise capacity in chronic heart failure. Heart 2005, 91, 749–753. [Google Scholar] [CrossRef]
- Mulay, S.R.; Evan, A.; Anders, H.J. Molecular mechanisms of crystal-related kidney inflammation and injury. Implications for cholesterol embolism, crystalline nephropathies and kidney stone disease. Nephrol. Dial. Transp. 2014, 29, 507–514. [Google Scholar] [CrossRef]
- Duncan, D.J.; Yang, Z.; Hopkins, P.M.; Steele, D.S.; Harrison, S.M. TNF-alpha and IL-1beta increase Ca2+ leak from the sarcoplasmic reticulum and susceptibility to arrhythmia in rat ventricular myocytes. Cell Calcium 2010, 47, 378–386. [Google Scholar] [CrossRef]
- Park, J.H.; Jin, Y.M.; Hwang, S.; Cho, D.H.; Kang, D.H.; Jo, I. Uric acid attenuates nitric oxide production by decreasing the interaction between endothelial nitric oxide synthase and calmodulin in human umbilical vein endothelial cells: A mechanism for uric acid-induced cardiovascular disease development. Nitric Oxide 2013, 32, 36–42. [Google Scholar] [CrossRef]
- Sautin, Y.Y.; Nakagawa, T.; Zharikov, S.; Johnson, R.J. Adverse effects of the classic antioxidant uric acid in adipocytes: NADPH oxidase-mediated oxidative/nitrosative stress. Am. J. Physiol. Cell Physiol. 2007, 293, C584–C596. [Google Scholar] [CrossRef]
- Johnson, R.J.; Bakris, G.L.; Borghi, C.; Chonchol, M.B.; Feldman, D.; Lanaspa, M.A.; Merriman, T.R.; Moe, O.W.; Mount, D.B.; Lozada, L.G.S.; et al. Hyperuricemia, Acute and Chronic Kidney Disease, Hypertension, and Cardiovascular Disease: Report of a Scientific Workshop Organized by the National Kidney Foundation. Am. J. Kidney Dis. 2018, 71, 851–865. [Google Scholar] [CrossRef]
- Sirker, A.; Zhang, M.; Shah, A.M. NADPH oxidases in cardiovascular disease: Insights from in vivo models and clinical studies. Basic Res. Cardiol. 2011, 106, 735–747. [Google Scholar] [CrossRef]
- Targher, G.; Byrne, C.D.; Lonardo, A.; Zoppini, G.; Barbui, C. Non-alcoholic fatty liver disease and risk of incident cardiovascular disease: A meta-analysis. J. Hepatol. 2016, 65, 589–600. [Google Scholar] [CrossRef]
- Lonardo, A.; Nascimbeni, F.; Mantovani, A.; Targher, G. Hypertension, diabetes, atherosclerosis and NASH: Cause or consequence? J. Hepatol. 2018, 68, 335–352. [Google Scholar] [CrossRef]
- Suzuki, A.; Angulo, P.; Lymp, J.; St Sauver, J.; Muto, A.; Okada, T.; Lindor, K. Chronological development of elevated aminotransferases in a nonalcoholic population. Hepatology 2005, 41, 64–71. [Google Scholar] [CrossRef]
- Vanni, E.; Bugianesi, E.; Kotronen, A.; De Minicis, S.; Yki-Jarvinen, H.; Svegliati-Baroni, G. From the metabolic syndrome to NAFLD or vice versa? Dig Liver Dis. 2010, 42, 320–330. [Google Scholar] [CrossRef]
- Yki-Järvinen, H. Non-alcoholic fatty liver disease as a cause and a consequence of metabolic syndrome. Lancet Diabetes Endocrinol. 2014, 2, 901–910. [Google Scholar] [CrossRef]
- Lonardo, A.; Ballestri, S.; Marchesini, G.; Angulo, P.; Loria, P. Nonalcoholic fatty liver disease: A precursor of the metabolic syndrome. Dig. Liver Dis. 2015, 47, 181–190. [Google Scholar] [CrossRef]
- Ballestri, S.; Zona, S.; Targher, G.; Romagnoli, D.; Baldelli, E.; Nascimbeni, F.; Roverato, A.; Guaraldi, G.; Lonardo, A. Nonalcoholic fatty liver disease is associated with an almost twofold increased risk of incident type 2 diabetes and metabolic syndrome. Evidence from a systematic review and meta-analysis. J. Gastroenterol. Hepatol. 2016, 31, 936–944. [Google Scholar] [CrossRef]
- Lonardo, A.; Nascimbeni, F.; Ballestri, S.; Fairweather, D.; Win, S.; Than, T.A.; Abdelmalek, M.F.; Suzuki, A. Sex Differences in NAFLD: State of the Art and Identification of Research Gaps. Hepatology 2019. [Google Scholar] [CrossRef]
- Yang, J.D.; Abdelmalek, M.F.; Guy, C.D.; Gill, R.M.; Lavine, J.E.; Yates, K.; Klair, J.; Terrault, N.A.; Clark, J.M.; Unalp-Arida, A.; et al. Patient Sex, Reproductive Status, and Synthetic Hormone Use Associate with Histologic Severity of Nonalcoholic Steatohepatitis. Clin. Gastroenterol. Hepatol. 2017, 15, 127–131. [Google Scholar] [CrossRef]
| 1st Stage (N = 7000) | 2nd Stage (N = 3000) | Combined (N = 10,000) | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Chr | SNP | BP | Gene | Alleles | Beta | P-Value | FDR | Beta | P-Value | Beta | P-Value | F | R2 |
| 4 | rs4148155 | 89054667 | ABCG2 | A/G | 0.31 | 9.35 × 10−46 | 2.86 × 10−40 | 0.32 | 5.60 × 10−22 | 0.31 | 4.34 × 10−66 | 37.7 | 0.022 |
| 4 | rs3733588 | 9997303 | SLC2A9 | A/G | −0.26 | 2.91 × 10−37 | 5.93 × 10−32 | −0.21 | 2.80 × 10−11 | −0.24 | 1.73 × 10−46 | 25.9 | 0.015 |
| 4 | rs2725211 | 88970375 | PKD2 | C/T | 0.25 | 1.86 × 10−25 | 1.14 × 10−20 | 0.30 | 1.50 × 10−17 | 0.27 | 3.18 × 10−41 | 24.2 | 0.014 |
| 4 | rs17013282 | 88765873 | MEPE | G/A | 0.19 | 1.03 × 10−11 | 1.15 × 10−7 | 0.21 | 1.38 × 10−6 | 0.20 | 4.95 × 10−17 | 11.1 | 0.007 |
| 4 | rs17013187 | 88733531 | IBSP | C/T | 0.16 | 1.49 × 10−9 | 1.35 × 10−5 | 0.14 | 6.63 × 10−4 | 0.15 | 2.85 × 10−12 | 8.8 | 0.005 |
| 4 | rs3756224 | 10105739 | WDR1 | T/C | 0.11 | 1.05 × 10−7 | 6.67 × 10−4 | 0.10 | 1.73 × 10−3 | 0.11 | 9.45 × 10−10 | 7.9 | 0.005 |
| 2 | rs1260326 | 27730940 | GCKR | C/T | 0.10 | 7.90 × 10−7 | 4.38 × 10−3 | 0.11 | 6.55 × 10−4 | 0.10 | 2.33 × 10−9 | 8.5 | 0.005 |
| 1 | rs4072037 | 155162067 | MUC1 | T/C | 0.11 | 8.18 × 10−6 | 3.60 × 10−2 | 0.11 | 4.56 × 10−3 | 0.11 | 1.43 × 10−7 | 6.8 | 0.004 |
| Characteristic | WGRS | P-Value † | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Q1 (N = 2490) | Q2 (N = 2509) | Q3 (N = 2499) | Q4 (N = 2502) | ||||||
| Mean | (SD) | Mean | (SD) | Mean | (SD) | Mean | (SD) | ||
| SUA (mg/dL) | 5.4 | (1.3) | 5.6 | (1.5) | 5.8 | (1.5) | 6.0 | (1.6) | <0.0001 * |
| Sex (Men%) | 49.7% | 49.3% | 52.2% | 48.8% | 0.080 | ||||
| Age (yr) | 49.1 | (11.4) | 48.9 | (11.0) | 48.6 | (11.0) | 48.8 | (11.1) | 0.140 |
| BMI (kg/m2) | 24.2 | (3.6) | 24.3 | (3.6) | 24.3 | (3.6) | 24.3 | (3.6) | 0.570 |
| FG (mg/dL) | 96.6 | (21.6) | 96.0 | (20.7) | 96.4 | (19.1) | 97.4 | (25.0) | 0.136 |
| T-CHO (mg/dL) | 193.4 | (35.9) | 191.8 | (35.1) | 191.9 | (34.1) | 194.1 | (36.6) | 0.060 |
| TG (mg/dL) | 118.0 | (104.0) | 113.8 | (83.0) | 117.9 | (81.9) | 124.7 | (118.3) | 0.002 * |
| HDL-C (mg/dL) | 53.7 | (13.4) | 53.3 | (13.2) | 52.8 | (12.8) | 52.8 | (13.1) | 0.050 |
| LDL-C (mg/dL) | 120.3 | (31.8) | 120.4 | (32.2) | 120.4 | (31.1) | 121.4 | (31.1) | 0.520 |
| eGFR (mL/min/1.73m2) | 103.4 | (24.4) | 102.7 | (24.6) | 102.2 | (24.7) | 102.9 | (25.1) | 0.41 |
| SGOT (U/L) | 24.6 | (16.6) | 24.6 | (12.5) | 24.7 | (13.1) | 24.2 | (10.4) | 0.582 |
| SGPT (U/L) | 24.5 | (18.8) | 25.2 | (21.6) | 25.8 | (23.5) | 24.8 | (19.4) | 0.110 |
| SBP (mmHg) | 115.7 | (16.9) | 116.0 | (17.1) | 116.7 | (17.3) | 116.9 | (18.0) | 0.060 |
| DBP (mmHg) | 71.6 | (10.8) | 72.0 | (10.9) | 72.7 | (11.3) | 72.8 | (11.3) | 0.010 * |
| Characteristic | WGRS | P-Value † | |||||||
|---|---|---|---|---|---|---|---|---|---|
| Q1 (N = 2490) | Q2 (N = 2509) | Q3 (N = 2499) | Q4 (N = 2502) | ||||||
| N | % | N | % | N | % | N | % | ||
| Sex | 0.08 | ||||||||
| Male | 1237 | 49.7 | 1237 | 49.3 | 1304 | 52.2 | 1222 | 48.8 | |
| Female | 1253 | 50.3 | 1272 | 50.7 | 1195 | 47.8 | 1280 | 51.2 | |
| Drinking habit | 0.38 | ||||||||
| Yes | 207 | 8.3 | 183 | 7.3 | 204 | 8.2 | 204 | 8.2 | |
| No | 2194 | 88.1 | 2239 | 89.2 | 2215 | 88.6 | 2232 | 89.2 | |
| Quit | 89 | 3.57 | 87 | 3.5 | 80 | 3.2 | 66 | 2.6 | |
| Smoking habit | 0.16 | ||||||||
| Yes | 263 | 10.6 | 289 | 11.5 | 312 | 12.5 | 295 | 11.8 | |
| Few | 214 | 8.6 | 206 | 8.2 | 207 | 8.3 | 213 | 8.5 | |
| No | 1748 | 70.2 | 1723 | 68.7 | 1656 | 66.3 | 1703 | 68.1 | |
| Quit | 265 | 10.6 | 291 | 11.6 | 324 | 13.0 | 291 | 11.6 | |
| Education | 0.071 | ||||||||
| Elementary School | 177 | 7.1 | 172 | 6.9 | 163 | 6.5 | 199 | 8.0 | |
| Junior-high/Senior-high | 1063 | 42.7 | 1045 | 41.7 | 984 | 39.4 | 1027 | 41.1 | |
| BS/MS/PhD | 1248 | 50.2 | 1289 | 51.4 | 1351 | 54.1 | 1276 | 51.0 | |
| Marriage | 0.66 | ||||||||
| Single | 300 | 12.1 | 279 | 11.1 | 298 | 11.9 | 274 | 11.0 | |
| Married | 1936 | 77.8 | 1989 | 79.4 | 1961 | 78.5 | 1963 | 78.6 | |
| Divorced/Widowed | 253 | 10.2 | 238 | 9.5 | 239 | 9.6 | 262 | 10.5 | |
| Regular Exercise | 0.94 | ||||||||
| No | 1453 | 58.4 | 1473 | 58.7 | 1472 | 58.9 | 1481 | 59.2 | |
| Yes | 1021 | 40.8 | 1027 | 41.1 | 1036 | 41.3 | 1037 | 41.7 | |
| Predictors | Logistic | Location | Scale | Shape | AIC | ||||
|---|---|---|---|---|---|---|---|---|---|
| OR (95% C) | P-Value | EST (95% CI) | P-Value | EST (95% CI) | P-Value | EST (95% CI) | P-Value | ||
| (a)continuousWGRS | |||||||||
| Intercept | 1 | Referent | 4.16 a (4.11, 4.21) | <0.001 | −2.37 a (−3.00, −1.73) | <0.001 | 1.42 a (0.35, 2.49) | 0.009 | 2594.09 |
| Male | 2.87 a (2.01, 4.10) | <0.001 | |||||||
| WGRS | 1.41 c (1.04, 1.91) | 0.029 | |||||||
| (b)four-group WGRS | |||||||||
| Intercept | 1 | Referent | 4.16 a (4.11, 4.21) | <0.001 | −2.37 a (−3.00, −1.73) | <0.001 | 1.42 b (0.35, 2.50) | 0.009 | 2593.36 |
| Male | 2.86 a (2.00, 4.10) | <0.001 | |||||||
| Q2 | 0.94 (0.57, 1.53) | 0.790 | |||||||
| Q3 | 1.46 (0.93, 2.29) | 0.103 | |||||||
| Q4 | 1.68 c (1.08, 2.62) | 0.022 | |||||||
| (c)two-group WGRS | |||||||||
| Intercept | 1 | Referent | 4.16 a (4.11, 4.21) | <0.001 | −2.37 a (−3.00, −1.73) | <0.001 | 1.42 b (0.35, 2.50) | 0.009 | 2589.89 |
| Male | 2.85 a (1.99, 4.07) | <0.001 | |||||||
| High | 1.62 b (1.17, 2.23) | 0.003 | |||||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chiang, K.-M.; Tsay, Y.-C.; Vincent Ng, T.-C.; Yang, H.-C.; Huang, Y.-T.; Chen, C.-H.; Pan, W.-H. Is Hyperuricemia, an Early-Onset Metabolic Disorder, Causally Associated with Cardiovascular Disease Events in Han Chinese? J. Clin. Med. 2019, 8, 1202. https://doi.org/10.3390/jcm8081202
Chiang K-M, Tsay Y-C, Vincent Ng T-C, Yang H-C, Huang Y-T, Chen C-H, Pan W-H. Is Hyperuricemia, an Early-Onset Metabolic Disorder, Causally Associated with Cardiovascular Disease Events in Han Chinese? Journal of Clinical Medicine. 2019; 8(8):1202. https://doi.org/10.3390/jcm8081202
Chicago/Turabian StyleChiang, Kuang-Mao, Yuh-Chyuan Tsay, Ta-Chou Vincent Ng, Hsin-Chou Yang, Yen-Tsung Huang, Chen-Hsin Chen, and Wen-Harn Pan. 2019. "Is Hyperuricemia, an Early-Onset Metabolic Disorder, Causally Associated with Cardiovascular Disease Events in Han Chinese?" Journal of Clinical Medicine 8, no. 8: 1202. https://doi.org/10.3390/jcm8081202
APA StyleChiang, K.-M., Tsay, Y.-C., Vincent Ng, T.-C., Yang, H.-C., Huang, Y.-T., Chen, C.-H., & Pan, W.-H. (2019). Is Hyperuricemia, an Early-Onset Metabolic Disorder, Causally Associated with Cardiovascular Disease Events in Han Chinese? Journal of Clinical Medicine, 8(8), 1202. https://doi.org/10.3390/jcm8081202





