Distinct Profiles of Cell-Free MicroRNAs in Plasma of Veterans with Post-Traumatic Stress Disorder
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Subjects
2.2. EV Isolation and RNA Extraction from Plasma
2.3. Sequencing Library Construction
2.4. Library Sequencing
2.5. Small RNA Sequencing Data Analysis
2.6. qRT-PCR
2.7. Functional Association of miRNA Target Genes
2.8. Ethics
3. Results
3.1. Demographic and Clinical Characteristics of Study Subjects
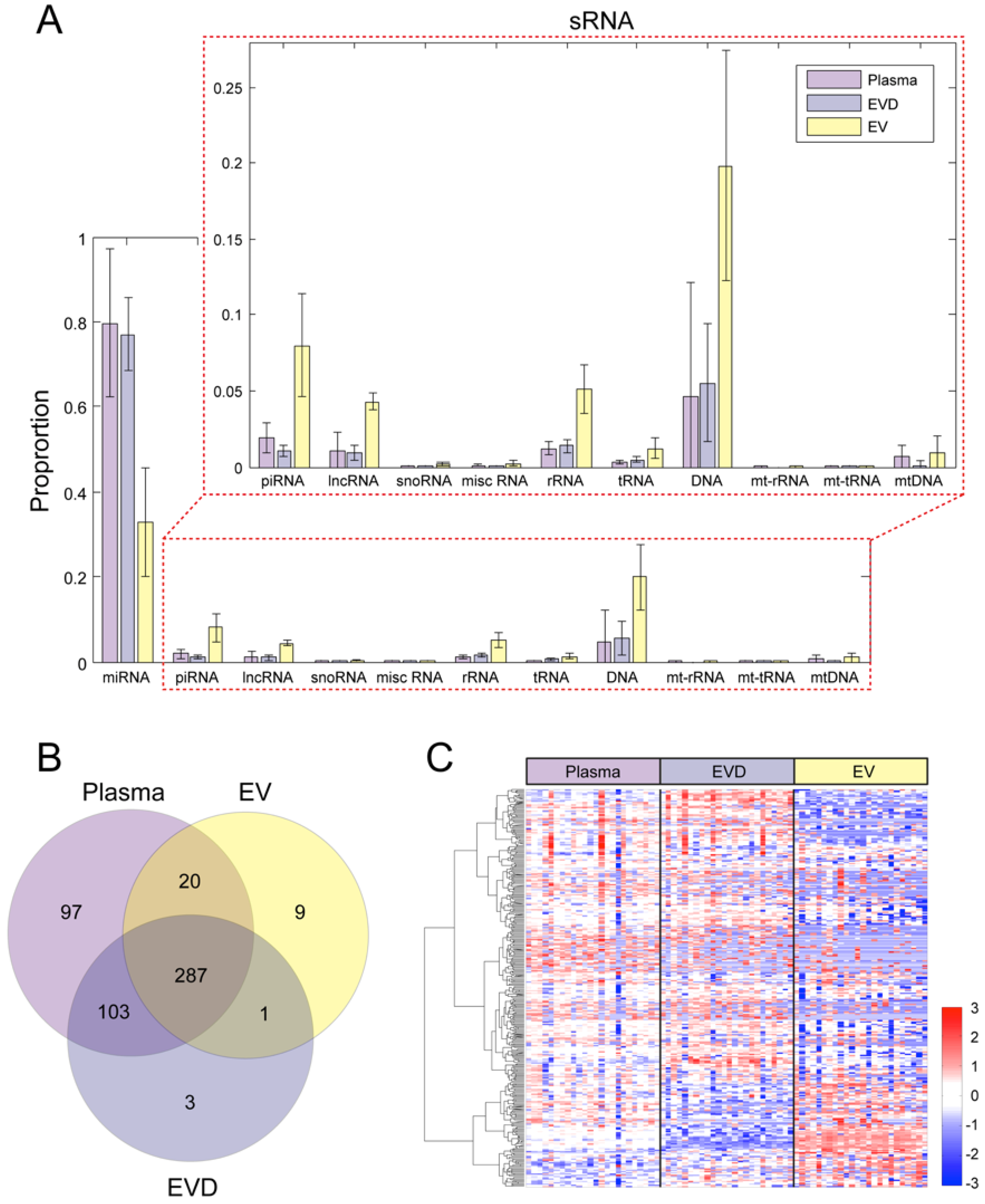
3.2. The Distribution of Small RNA in Circulation
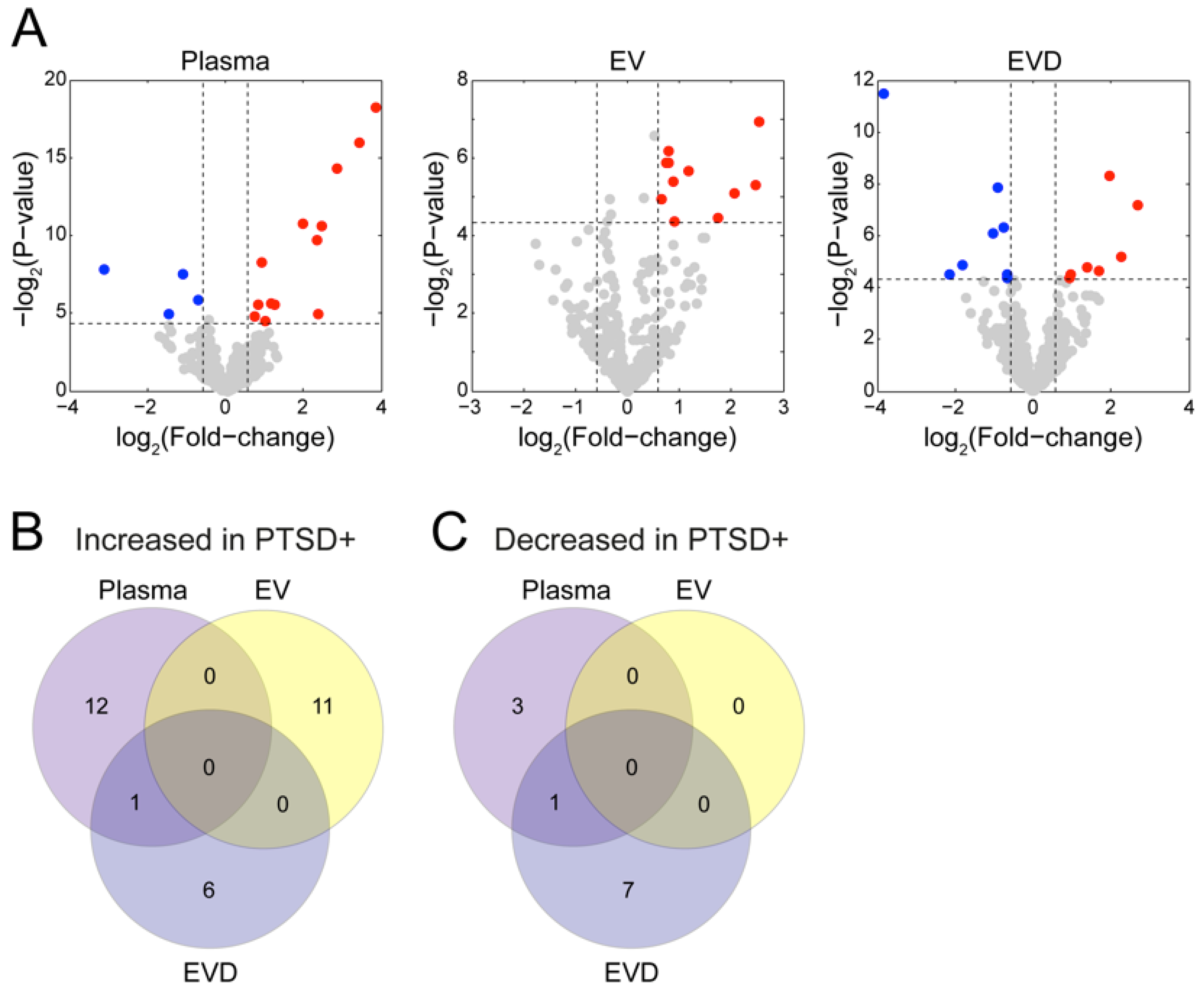
3.3. Circulating miRNAs Affected by PTSD
3.4. Other Types of Small RNAs Affected by PTSD
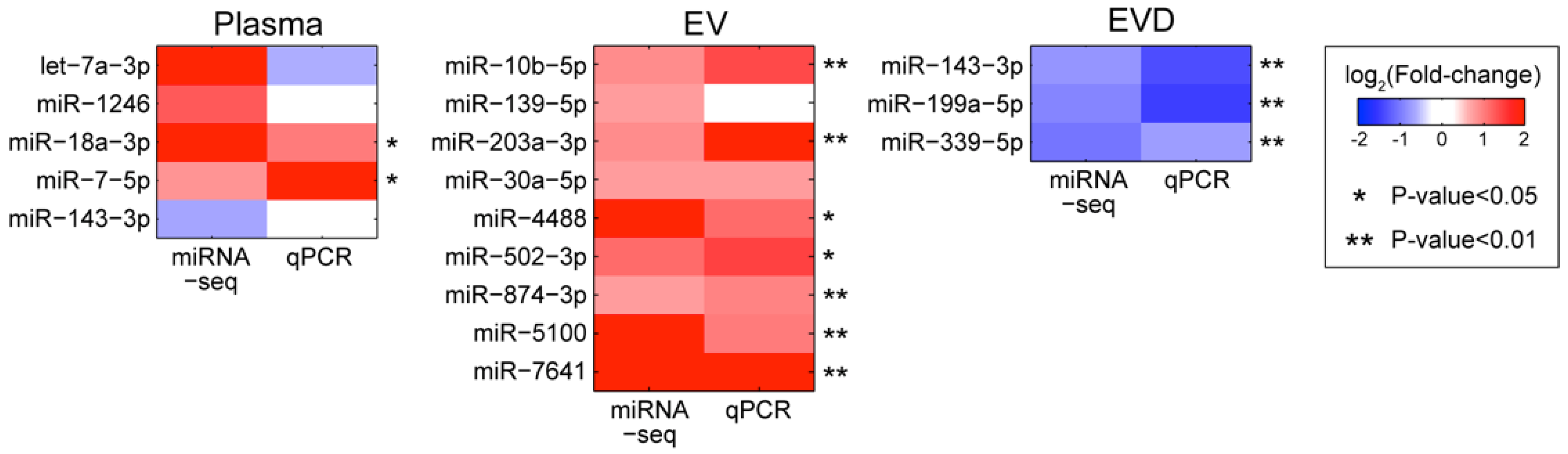
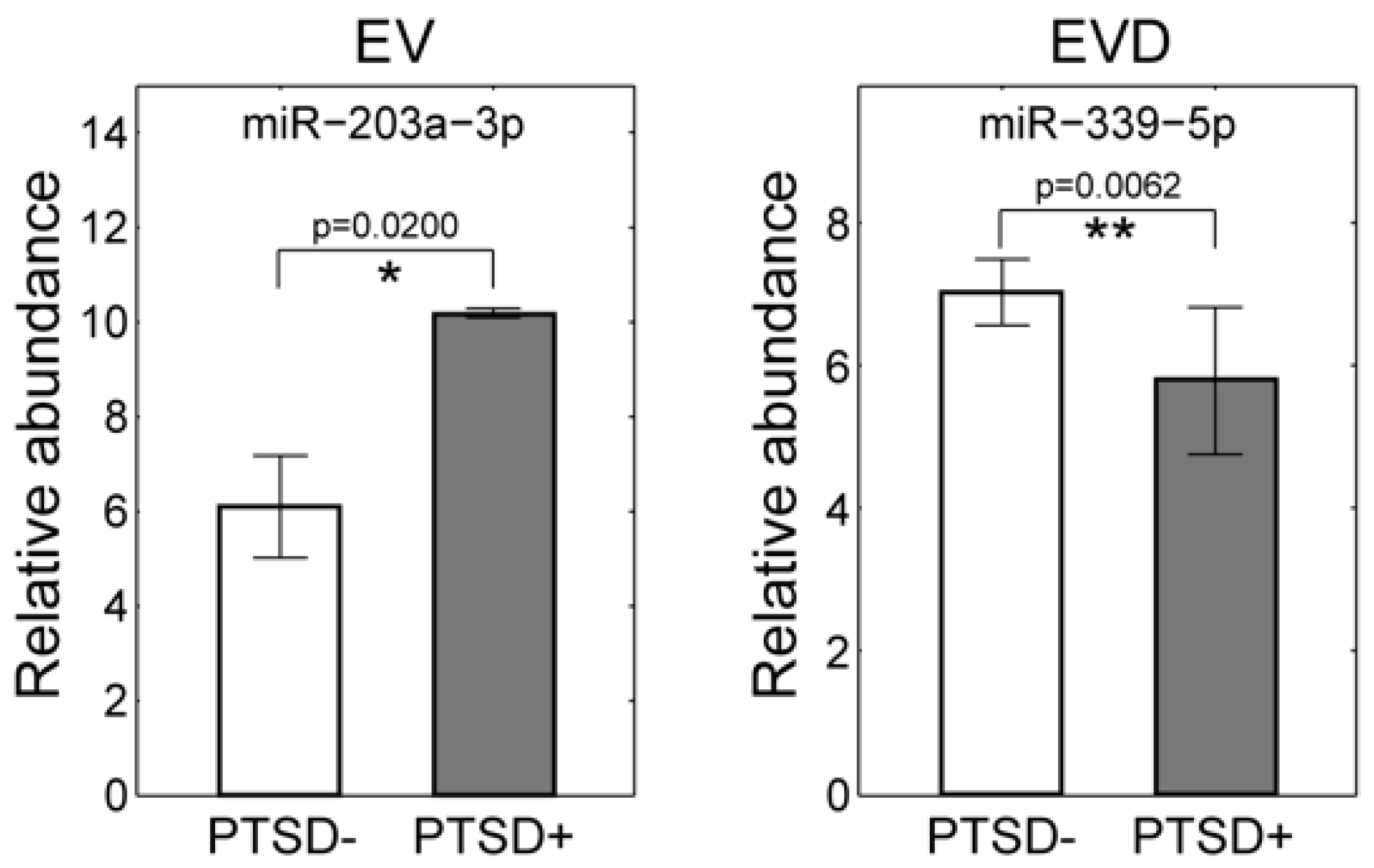
3.5. Validation of the miRNA Associated with PTSD Using qRT-PCR
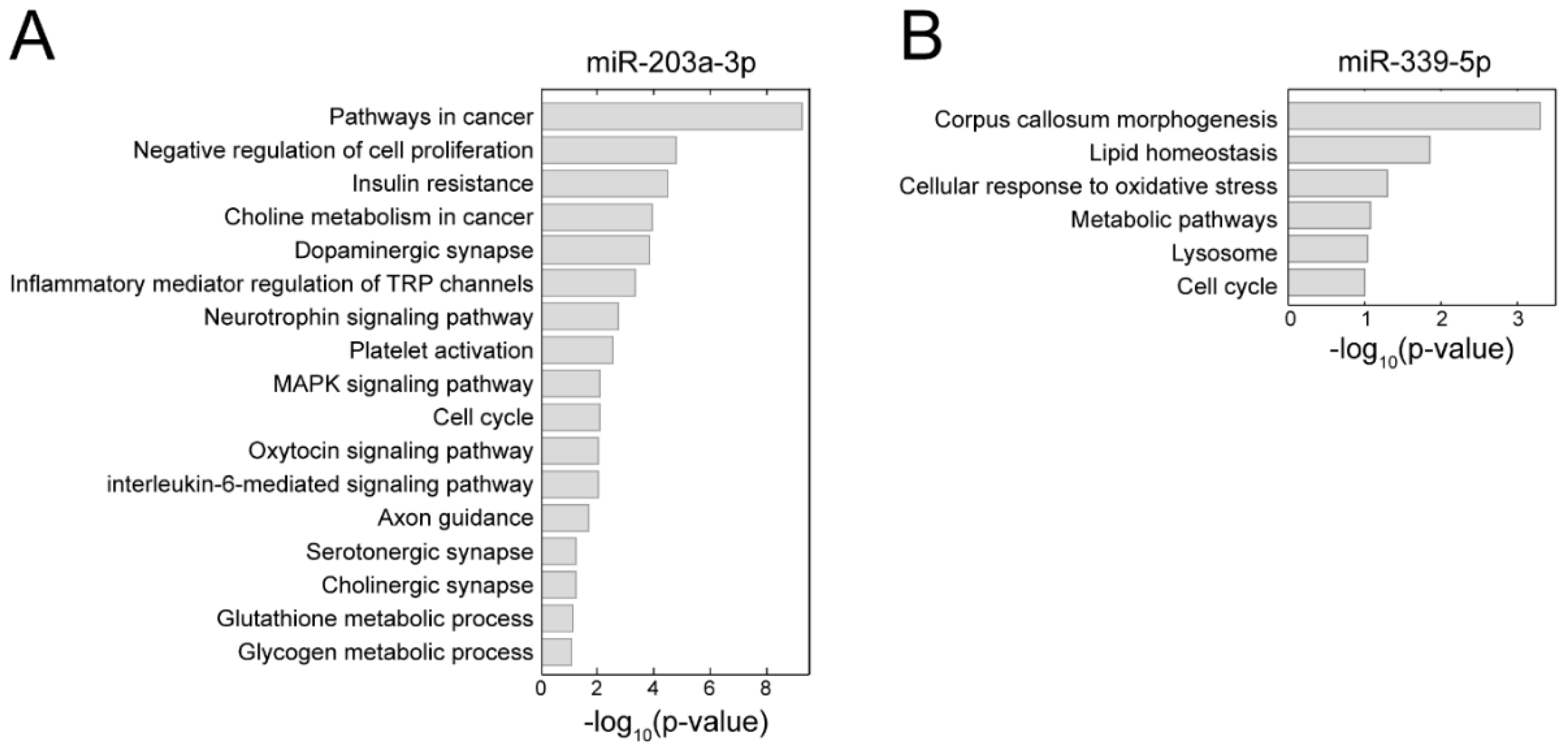
3.6. Biological Processes May Be Affected by PTSD-Associated miRNA
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Yehuda, R.; Hoge, C.W.; McFarlane, A.C.; Vermetten, E.; Lanius, R.A.; Nievergelt, C.M.; Hobfoll, S.E.; Koenen, K.C.; Neylan, T.C.; Hyman, S.E. Post-traumatic stress disorder. Nat. Rev. Dis. Primers 2015, 1, 15057. [Google Scholar] [CrossRef] [PubMed]
- Shalev, A.; Liberzon, I.; Marmar, C. Post-Traumatic Stress Disorder. N. Engl. J. Med. 2017, 376, 2459–2469. [Google Scholar] [CrossRef] [PubMed]
- Kulka, R.A.; Schlenger, W.E.; Fairbank, J.A.; Hough, R.L.; Jordan, B.K.; Marmar, C.R.; Weiss, D.S. Trauma and the Vietnam War Generation: Report of Findings from the National Vietnam Veterans Readjustment Study; Brunner/Mazel: Philadelphia, PA, USA, 1990. [Google Scholar]
- Kang, H.K.; Natelson, B.H.; Mahan, C.M.; Lee, K.Y.; Murphy, F.M. Post-traumatic stress disorder and chronic fatigue syndrome-like illness among Gulf War veterans: A population-based survey of 30,000 veterans. Am. J. Epidemiol. 2003, 157, 141–148. [Google Scholar] [CrossRef] [PubMed]
- Marmar, C.R.; Schlenger, W.; Henn-Haase, C.; Qian, M.; Purchia, E.; Li, M.; Corry, N.; Williams, C.S.; Ho, C.L.; Horesh, D.; et al. Course of Posttraumatic Stress Disorder 40 Years After the Vietnam War: Findings From the National Vietnam Veterans Longitudinal Study. JAMA Psychiatry 2015, 72, 875–881. [Google Scholar] [CrossRef] [PubMed]
- Vyas, K.J.; Fesperman, S.F.; Nebeker, B.J.; Gerard, S.K.; Boyd, N.D.; Delaney, E.M.; Webb-Murphy, J.A.; Johnston, S.L. Preventing PTSD and Depression and Reducing Health Care Costs in the Military: A Call for Building Resilience Among Service Members. Mil. Med. 2016, 181, 1240–1247. [Google Scholar] [CrossRef] [PubMed]
- Xue, C.; Ge, Y.; Tang, B.; Liu, Y.; Kang, P.; Wang, M.; Zhang, L. A meta-analysis of risk factors for combat-related PTSD among military personnel and veterans. PLoS ONE 2015, 10, e0120270. [Google Scholar] [CrossRef] [PubMed]
- Blacker, C.J.; Frye, M.A.; Morava, E.; Kozicz, T.; Veldic, M. A Review of Epigenetics of PTSD in Comorbid Psychiatric Conditions. Genes (Basel) 2019, 10, 140. [Google Scholar] [CrossRef] [PubMed]
- Gros, D.F.; Price, M.; Magruder, K.M.; Frueh, B.C. Symptom overlap in posttraumatic stress disorder and major depression. Psychiatry Res. 2012, 196, 267–270. [Google Scholar] [CrossRef]
- Bryant, R. Post-traumatic stress disorder vs traumatic brain injury. Dialogues Clin. Neurosci. 2011, 13, 251–262. [Google Scholar]
- DiMauro, J.; Carter, S.; Folk, J.B.; Kashdan, T.B. A historical review of trauma-related diagnoses to reconsider the heterogeneity of PTSD. J. Anxiety Disord. 2014, 28, 774–786. [Google Scholar] [CrossRef]
- Tufekci, K.U.; Meuwissen, R.L.; Genc, S. The role of microRNAs in biological processes. Methods Mol. Biol. 2014, 1107, 15–31. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P.S.; Parkin, R.K.; Kroh, E.M.; Fritz, B.R.; Wyman, S.K.; Pogosova-Agadjanyan, E.L.; Peterson, A.; Noteboom, J.; O’Briant, K.C.; Allen, A.; et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc. Natl. Acad. Sci. USA. 2008, 105, 10513–10518. [Google Scholar] [CrossRef] [PubMed]
- Park, N.J.; Zhou, H.; Elashoff, D.; Henson, B.S.; Kastratovic, D.A.; Abemayor, E.; Wong, D.T. Salivary microRNA: Discovery, characterization, and clinical utility for oral cancer detection. Clin. Cancer Res. 2009, 15, 5473–5477. [Google Scholar] [CrossRef] [PubMed]
- Hanke, M.; Hoefig, K.; Merz, H.; Feller, A.C.; Kausch, I.; Jocham, D.; Warnecke, J.M.; Sczakiel, G. A robust methodology to study urine microRNA as tumor marker: microRNA-126 and microRNA-182 are related to urinary bladder cancer. Urol. Oncol. 2010, 28, 655–661. [Google Scholar] [CrossRef] [PubMed]
- Matin, F.; Jeet, V.; Moya, L.; Selth, L.A.; Chambers, S.; Australian Prostate Cancer, B.; Clements, J.A.; Batra, J. A Plasma Biomarker Panel of Four MicroRNAs for the Diagnosis of Prostate Cancer. Sci. Rep. 2018, 8, 6653. [Google Scholar] [CrossRef] [PubMed]
- Leng, Q.; Lin, Y.; Jiang, F.; Lee, C.J.; Zhan, M.; Fang, H.; Wang, Y.; Jiang, F. A plasma miRNA signature for lung cancer early detection. Oncotarget 2017, 8, 111902–111911. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Xu, Y.; Jin, X.; Wang, Z.; Wu, Y.; Zhao, D.; Chen, G.; Li, D.; Wang, X.; Cao, H.; et al. A circulating miRNA signature as a diagnostic biomarker for non-invasive early detection of breast cancer. Breast Cancer Res. Treat. 2015, 154, 423–434. [Google Scholar] [CrossRef] [PubMed]
- Nagaraj, S.; Laskowska-Kaszub, K.; Debski, K.J.; Wojsiat, J.; Dabrowski, M.; Gabryelewicz, T.; Kuznicki, J.; Wojda, U. Profile of 6 microRNA in blood plasma distinguish early stage Alzheimer’s disease patients from non-demented subjects. Oncotarget 2017, 8, 16122–16143. [Google Scholar] [CrossRef] [PubMed]
- Khoo, S.K.; Petillo, D.; Kang, U.J.; Resau, J.H.; Berryhill, B.; Linder, J.; Forsgren, L.; Neuman, L.A.; Tan, A.C. Plasma-based circulating MicroRNA biomarkers for Parkinson’s disease. J. Parkinson’s Dis. 2012, 2, 321–331. [Google Scholar] [CrossRef]
- Mundalil Vasu, M.; Anitha, A.; Thanseem, I.; Suzuki, K.; Yamada, K.; Takahashi, T.; Wakuda, T.; Iwata, K.; Tsujii, M.; Sugiyama, T.; et al. Serum microRNA profiles in children with autism. Mol. Autism 2014, 5, 40. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.Y.; Lu, J.; Zhang, L.; Song, H.T.; Zhao, L.; Fan, H.M.; Zhong, A.F.; Niu, W.; Guo, Z.M.; Dai, Y.H.; et al. Aberrant microRNA expression in peripheral plasma and mononuclear cells as specific blood-based biomarkers in schizophrenia patients. J. Clin. Neurosci. Off. J. Neurosurg. Soc. Australas. 2015, 22, 570–574. [Google Scholar] [CrossRef] [PubMed]
- Endzelins, E.; Berger, A.; Melne, V.; Bajo-Santos, C.; Sobolevska, K.; Abols, A.; Rodriguez, M.; Santare, D.; Rudnickiha, A.; Lietuvietis, V.; et al. Detection of circulating miRNAs: Comparative analysis of extracellular vesicle-incorporated miRNAs and cell-free miRNAs in whole plasma of prostate cancer patients. BMC Cancer 2017, 17, 730. [Google Scholar] [CrossRef] [PubMed]
- Alexander, M.; Hu, R.; Runtsch, M.C.; Kagele, D.A.; Mosbruger, T.L.; Tolmachova, T.; Seabra, M.C.; Round, J.L.; Ward, D.M.; O’Connell, R.M. Exosome-delivered microRNAs modulate the inflammatory response to endotoxin. Nat. Commun. 2015, 6, 7321. [Google Scholar] [CrossRef]
- Chivet, M.; Javalet, C.; Laulagnier, K.; Blot, B.; Hemming, F.J.; Sadoul, R. Exosomes secreted by cortical neurons upon glutamatergic synapse activation specifically interact with neurons. J. Extracell. Vesicles 2014, 3, 24722. [Google Scholar] [CrossRef] [PubMed]
- Thompson, A.G.; Gray, E.; Heman-Ackah, S.M.; Mager, I.; Talbot, K.; Andaloussi, S.E.; Wood, M.J.; Turner, M.R. Extracellular vesicles in neurodegenerative disease - pathogenesis to biomarkers. Nat. Rev. Neurol. 2016, 12, 346–357. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Hochberg, F.H.; Jones, P.S. Extracellular vesicles: The growth as diagnostics and therapeutics; a survey. J. Extracell. Vesicles 2018, 7, 1438720. [Google Scholar] [CrossRef]
- Fallen, S.; Baxter, D.; Wu, X.; Kim, T.K.; Shynlova, O.; Lee, M.Y.; Scherler, K.; Lye, S.; Hood, L.; Wang, K. Extracellular vesicle RNAs reflect placenta dysfunction and are a biomarker source for preterm labour. J. Cell. Mol. Med. 2018, 22, 2760–2773. [Google Scholar] [CrossRef]
- Ghai, V.; Wu, X.; Bheda-Malge, A.; Argyropoulos, C.P.; Bernardo, J.F.; Orchard, T.; Galas, D.; Wang, K. Genome-wide Profiling of Urinary Extracellular Vesicle microRNAs Associated With Diabetic Nephropathy in Type 1 Diabetes. Kidney Int. Rep. 2018, 3, 555–572. [Google Scholar] [CrossRef]
- Balakathiresan, N.S.; Chandran, R.; Bhomia, M.; Jia, M.; Li, H.; Maheshwari, R.K. Serum and amygdala microRNA signatures of posttraumatic stress: Fear correlation and biomarker potential. J. Psychiatr Res. 2014, 57, 65–73. [Google Scholar] [CrossRef]
- Cho, J.H.; Lee, I.; Hammamieh, R.; Wang, K.; Baxter, D.; Scherler, K.; Etheridge, A.; Kulchenko, A.; Gautam, A.; Muhie, S.; et al. Molecular evidence of stress-induced acute heart injury in a mouse model simulating posttraumatic stress disorder. Proc. Natl. Acad. Sci. USA 2014, 111, 3188–3193. [Google Scholar] [CrossRef]
- Zhou, J.; Nagarkatti, P.; Zhong, Y.; Ginsberg, J.P.; Singh, N.P.; Zhang, J.; Nagarkatti, M. Dysregulation in microRNA expression is associated with alterations in immune functions in combat veterans with post-traumatic stress disorder. PLoS ONE 2014, 9, e94075. [Google Scholar] [CrossRef] [PubMed]
- Martin, C.G.; Kim, H.; Yun, S.; Livingston, W.; Fetta, J.; Mysliwiec, V.; Baxter, T.; Gill, J.M. Circulating miRNA associated with posttraumatic stress disorder in a cohort of military combat veterans. Psychiatry Res. 2017, 251, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Wingo, A.P.; Almli, L.M.; Stevens, J.S.; Klengel, T.; Uddin, M.; Li, Y.; Bustamante, A.C.; Lori, A.; Koen, N.; Stein, D.J.; et al. DICER1 and microRNA regulation in post-traumatic stress disorder with comorbid depression. Nat. Commun. 2015, 6, 10106. [Google Scholar] [CrossRef] [PubMed]
- Hammamieh, R.; Chakraborty, N.; Gautam, A.; Muhie, S.; Yang, R.; Donohue, D.; Kumar, R.; Daigle, B.J., Jr.; Zhang, Y.; Amara, D.A.; et al. Whole-genome DNA methylation status associated with clinical PTSD measures of OIF/OEF veterans. Transl. Psychiatry 2017, 7, e1169. [Google Scholar] [CrossRef] [PubMed]
- Lindqvist, D.; Fernstrom, J.; Grudet, C.; Ljunggren, L.; Traskman-Bendz, L.; Ohlsson, L.; Westrin, A. Increased plasma levels of circulating cell-free mitochondrial DNA in suicide attempters: Associations with HPA-axis hyperactivity. Transl. Psychiatry 2016, 6, e971. [Google Scholar] [CrossRef] [PubMed]
- Lindqvist, D.; Wolkowitz, O.M.; Mellon, S.; Yehuda, R.; Flory, J.D.; Henn-Haase, C.; Bierer, L.M.; Abu-Amara, D.; Coy, M.; Neylan, T.C.; et al. Proinflammatory milieu in combat-related PTSD is independent of depression and early life stress. Brain Behav. Immun. 2014, 42, 81–88. [Google Scholar] [CrossRef] [PubMed]
- Etheridge, A.; Wang, K.; Baxter, D.; Galas, D. Preparation of Small RNA NGS Libraries from Biofluids. Methods Mol. Biol. 2018, 1740, 163–175. [Google Scholar] [CrossRef]
- Wu, X.; Kim, T.K.; Baxter, D.; Scherler, K.; Gordon, A.; Fong, O.; Etheridge, A.; Galas, D.J.; Wang, K. sRNAnalyzer-a flexible and customizable small RNA sequencing data analysis pipeline. Nucleic Acids Res. 2017, 45, 12140–12151. [Google Scholar] [CrossRef]
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. Embnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef]
- Griffiths-Jones, S.; Grocock, R.J.; van Dongen, S.; Bateman, A.; Enright, A.J. MiRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 2006, 34, D140–D144. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Si, X.; Skogerbo, G.; Wang, J.; Cui, D.; Li, Y.; Sun, X.; Liu, L.; Sun, B.; Chen, R.; et al. piRBase: A web resource assisting piRNA functional study. Database 2014, 2014, bau110. [Google Scholar] [CrossRef] [PubMed]
- Lestrade, L.; Weber, M.J. snoRNA-LBME-db, a comprehensive database of human H/ACA and C/D box snoRNAs. Nucleic Acids Res. 2006, 34, D158–D162. [Google Scholar] [CrossRef] [PubMed]
- Volders, P.J.; Helsens, K.; Wang, X.; Menten, B.; Martens, L.; Gevaert, K.; Vandesompele, J.; Mestdagh, P. LNCipedia: A database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res. 2013, 41, D246–D251. [Google Scholar] [CrossRef] [PubMed]
- Bao, W.; Kojima, K.K.; Kohany, O. Repbase Update, a database of repetitive elements in eukaryotic genomes. Mob. DNA 2015, 6, 11. [Google Scholar] [CrossRef] [PubMed]
- Flicek, P.; Amode, M.R.; Barrell, D.; Beal, K.; Brent, S.; Carvalho-Silva, D.; Clapham, P.; Coates, G.; Fairley, S.; Fitzgerald, S.; et al. Ensembl 2012. Nucleic Acids Res. 2012, 40, D84–D90. [Google Scholar] [CrossRef]
- Friedlander, M.R.; Mackowiak, S.D.; Li, N.; Chen, W.; Rajewsky, N. miRDeep2 accurately identifies known and hundreds of novel microRNA genes in seven animal clades. Nucleic Acids Res. 2012, 40, 37–52. [Google Scholar] [CrossRef]
- Robinson, M.D.; Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 2010, 11, R25. [Google Scholar] [CrossRef]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2010, 26, 139–140. [Google Scholar] [CrossRef]
- Chou, C.H.; Chang, N.W.; Shrestha, S.; Hsu, S.D.; Lin, Y.L.; Lee, W.H.; Yang, C.D.; Hong, H.C.; Wei, T.Y.; Tu, S.J.; et al. miRTarBase 2016: Updates to the experimentally validated miRNA-target interactions database. Nucleic Acids Res. 2016, 44, D239–D247. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Santangelo, L.; Giurato, G.; Cicchini, C.; Montaldo, C.; Mancone, C.; Tarallo, R.; Battistelli, C.; Alonzi, T.; Weisz, A.; Tripodi, M. The RNA-Binding Protein SYNCRIP Is a Component of the Hepatocyte Exosomal Machinery Controlling MicroRNA Sorting. Cell Rep. 2016, 17, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Hui, Y.; Li, X.; Jia, Q. Silencing of lncRNA-CCDC26 Restrains the Growth and Migration of Glioma Cells In Vitro and In Vivo Via Targeting miR-203. Oncol. Res. 2017. [Google Scholar] [CrossRef]
- Kaur, P.; Tan, J.R.; Karolina, D.S.; Sepramaniam, S.; Armugam, A.; Wong, P.T.; Jeyaseelan, K. A long non-coding RNA, BC048612 and a microRNA, miR-203 coordinate the gene expression of neuronal growth regulator 1 (NEGR1) adhesion protein. Biochim. Biophys. Acta. 2016, 1863, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.T.; Jeon, D.; Chu, K.; Jung, K.H.; Moon, J.; Sunwoo, J.; Park, D.K.; Yang, H.; Park, J.H.; Kim, M.; et al. Inhibition of miR-203 Reduces Spontaneous Recurrent Seizures in Mice. Mol. Neurobiol. 2017, 54, 3300–3308. [Google Scholar] [CrossRef] [PubMed]
- Brogaard, L.; Heegaard, P.M.; Larsen, L.E.; Mortensen, S.; Schlegel, M.; Durrwald, R.; Skovgaard, K. Late regulation of immune genes and microRNAs in circulating leukocytes in a pig model of influenza A (H1N2) infection. Sci. Rep. 2016, 6, 21812. [Google Scholar] [CrossRef]
- Ferreira, R.B.; Ferreira, R.; Albuquerque, D.M.; Costa, F.F.; Franco-Penteado, C.F. miRNA-146a, miRNA-203a, and miRNA-223 Modulate Inflammatory Response in LPS- Acute Lung Injury in Sickle Cell Transgenic Mice. Blood 2015, 126, 3390. [Google Scholar]
- Wang, Y.; Dong, Q.; Gu, Y.; Groome, L.J. Up-regulation of miR-203 expression induces endothelial inflammatory response: Potential role in preeclampsia. Am. J. Reprod. Immunol. 2016, 76, 482–490. [Google Scholar] [CrossRef]
- de Oliveira, J.F.; Wiener, C.D.; Jansen, K.; Portela, L.V.; Lara, D.R.; Souza, L.D.M.; da Silva, R.A.; Moreira, F.P.; Oses, J.P. Serum levels of interleukins IL-6 and IL-10 in individuals with posttraumatic stress disorder in a population-based sample. Psychiatry Res. 2017, 260, 111–115. [Google Scholar] [CrossRef]
- Kaufer, D.; Friedman, A.; Seidman, S.; Soreq, H. Acute stress facilitates long-lasting changes in cholinergic gene expression. Nature 1998, 393, 373–377. [Google Scholar] [CrossRef]
- Nees, F.; Witt, S.H.; Flor, H. Neurogenetic Approaches to Stress and Fear in Humans as Pathophysiological Mechanisms for Posttraumatic Stress Disorder. Biol. Psychiatry 2018, 83, 810–820. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Kim, C.Y.; Koo, H.; Heo, Y.; Cheon, K. A Novel Animal Model Simulating the Beginning of Combat Exposure. Neuroimmunomodulation 2017, 24, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Blessing, E.M.; Reus, V.; Mellon, S.H.; Wolkowitz, O.M.; Flory, J.D.; Bierer, L.; Lindqvist, D.; Dhabhar, F.; Li, M.; Qian, M.; et al. Biological predictors of insulin resistance associated with posttraumatic stress disorder in young military veterans. Psychoneuroendocrinology 2017, 82, 91–97. [Google Scholar] [CrossRef] [PubMed]
- Vidovic, A.; Grubisic-Ilic, M.; Kozaric-Kovacic, D.; Gotovac, K.; Rakos, I.; Markotic, A.; Rabatic, S.; Dekaris, D.; Sabioncello, A. Exaggerated platelet reactivity to physiological agonists in war veterans with posttraumatic stress disorder. Psychoneuroendocrinology 2011, 36, 161–172. [Google Scholar] [CrossRef] [PubMed]
- Swartzman, S.; Booth, J.N.; Munro, A.; Sani, F. Posttraumatic stress disorder after cancer diagnosis in adults: A meta-analysis. Depress. Anxiety 2017, 34, 327–339. [Google Scholar] [CrossRef] [PubMed]
- Yaffe, K.; Vittinghoff, E.; Lindquist, K.; Barnes, D.; Covinsky, K.E.; Neylan, T.; Kluse, M.; Marmar, C. Posttraumatic stress disorder and risk of dementia among US veterans. Arch. Gen. Psychiatry 2010, 67, 608–613. [Google Scholar] [CrossRef] [PubMed]
- Long, J.M.; Ray, B.; Lahiri, D.K. MicroRNA-339-5p down-regulates protein expression of beta-site amyloid precursor protein-cleaving enzyme 1 (BACE1) in human primary brain cultures and is reduced in brain tissue specimens of Alzheimer disease subjects. J. Bio.l Chem. 2014, 289, 5184–5198. [Google Scholar] [CrossRef]
- Ren, R.J.; Zhang, Y.F.; Dammer, E.B.; Zhou, Y.; Wang, L.L.; Liu, X.H.; Feng, B.L.; Jiang, G.X.; Chen, S.D.; Wang, G.; et al. Peripheral Blood MicroRNA Expression Profiles in Alzheimer’s Disease: Screening, Validation, Association with Clinical Phenotype and Implications for Molecular Mechanism. Mol. Neurobiol. 2016, 53, 5772–5781. [Google Scholar] [CrossRef]
- Zhang, Y.; Wei, G.; Di, Z.; Zhao, Q. miR-339-5p inhibits alcohol-induced brain inflammation through regulating NF-kappaB pathway. Biochem. Biophys. Res. Commun. 2014, 452, 450–456. [Google Scholar] [CrossRef]
- Villarreal, G.; Hamilton, D.A.; Graham, D.P.; Driscoll, I.; Qualls, C.; Petropoulos, H.; Brooks, W.M. Reduced area of the corpus callosum in posttraumatic stress disorder. Psychiatry Res. 2004, 131, 227–235. [Google Scholar] [CrossRef]
- Kitayama, N.; Brummer, M.; Hertz, L.; Quinn, S.; Kim, Y.; Bremner, J.D. Morphologic alterations in the corpus callosum in abuse-related posttraumatic stress disorder: A preliminary study. J. Nerv. Ment. Dis. 2007, 195, 1027–1029. [Google Scholar] [CrossRef] [PubMed]
- Saar-Ashkenazy, R.; Cohen, J.E.; Guez, J.; Gasho, C.; Shelef, I.; Friedman, A.; Shalev, H. Reduced corpus-callosum volume in posttraumatic stress disorder highlights the importance of interhemispheric connectivity for associative memory. J. Trauma. Stress 2014, 27, 18–26. [Google Scholar] [CrossRef] [PubMed]
- Nedic Erjavec, G.; Konjevod, M.; Nikolac Perkovic, M.; Svob Strac, D.; Tudor, L.; Barbas, C.; Grune, T.; Zarkovic, N.; Pivac, N. Short overview on metabolomic approach and redox changes in psychiatric disorders. Redox Biol. 2018, 14, 178–186. [Google Scholar] [CrossRef] [PubMed]
- Freedman, J.E.; Gerstein, M.; Mick, E.; Rozowsky, J.; Levy, D.; Kitchen, R.; Das, S.; Shah, R.; Danielson, K.; Beaulieu, L.; et al. Diverse human extracellular RNAs are widely detected in human plasma. Nat. Commun. 2016, 7, 11106. [Google Scholar] [CrossRef] [PubMed]
- Bahn, J.H.; Zhang, Q.; Li, F.; Chan, T.M.; Lin, X.; Kim, Y.; Wong, D.T.; Xiao, X. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 2015, 61, 221–230. [Google Scholar] [CrossRef]
- Chandran, R.; Mehta, S.L.; Vemuganti, R. Non-coding RNAs and neuroprotection after acute CNS injuries. Neurochem. Int. 2017, 111, 12–22. [Google Scholar] [CrossRef] [PubMed]
- Gong, J.; Li, Y.; Liu, C.J.; Xiang, Y.; Li, C.; Ye, Y.; Zhang, Z.; Hawke, D.H.; Park, P.K.; Diao, L.; et al. A Pan-cancer Analysis of the Expression and Clinical Relevance of Small Nucleolar RNAs in Human Cancer. Cell Rep. 2017, 21, 1968–1981. [Google Scholar] [CrossRef]
- Qiu, W.; Guo, X.; Lin, X.; Yang, Q.; Zhang, W.; Zhang, Y.; Zuo, L.; Zhu, Y.; Li, C.R.; Ma, C.; et al. Transcriptome-wide piRNA profiling in human brains of Alzheimer’s disease. Neurobiol. Aging 2017, 57, 170–177. [Google Scholar] [CrossRef]
- Hong, Y.; Wang, C.; Fu, Z.; Liang, H.; Zhang, S.; Lu, M.; Sun, W.; Ye, C.; Zhang, C.Y.; Zen, K.; et al. Systematic characterization of seminal plasma piRNAs as molecular biomarkers for male infertility. Sci. Rep. 2016, 6, 24229. [Google Scholar] [CrossRef]
- Baraniskin, A.; Zaslavska, E.; Nopel-Dunnebacke, S.; Ahle, G.; Seidel, S.; Schlegel, U.; Schmiegel, W.; Hahn, S.; Schroers, R. Circulating U2 small nuclear RNA fragments as a novel diagnostic biomarker for primary central nervous system lymphoma. Neuro-Oncol. 2016, 18, 361–367. [Google Scholar] [CrossRef]
- Amorim, M.G.; Valieris, R.; Drummond, R.D.; Pizzi, M.P.; Freitas, V.M.; Sinigaglia-Coimbra, R.; Calin, G.A.; Pasqualini, R.; Arap, W.; Silva, I.T.; et al. A total transcriptome profiling method for plasma-derived extracellular vesicles: Applications for liquid biopsies. Sci. Rep. 2017, 7, 14395. [Google Scholar] [CrossRef] [PubMed]
- Blandford, S.N.; Galloway, D.A.; Moore, C.S. The roles of extracellular vesicle microRNAs in the central nervous system. Glia 2018, 66, 2267–2278. [Google Scholar] [CrossRef] [PubMed]
- Yanez-Mo, M.; Siljander, P.R.; Andreu, Z.; Zavec, A.B.; Borras, F.E.; Buzas, E.I.; Buzas, K.; Casal, E.; Cappello, F.; Carvalho, J.; et al. Biological properties of extracellular vesicles and their physiological functions. J. Extracell. Vesicles 2015, 4, 27066. [Google Scholar] [CrossRef] [PubMed]
- T, L.R.; Sanchez-Abarca, L.I.; Muntion, S.; Preciado, S.; Puig, N.; Lopez-Ruano, G.; Hernandez-Hernandez, A.; Redondo, A.; Ortega, R.; Rodriguez, C.; et al. MSC surface markers (CD44, CD73, and CD90) can identify human MSC-derived extracellular vesicles by conventional flow cytometry. Cell Commun. Signal. CCS 2016, 14, 2. [Google Scholar] [CrossRef]
- Whitham, M.; Parker, B.L.; Friedrichsen, M.; Hingst, J.R.; Hjorth, M.; Hughes, W.E.; Egan, C.L.; Cron, L.; Watt, K.I.; Kuchel, R.P.; et al. Extracellular Vesicles Provide a Means for Tissue Crosstalk during Exercise. Cell Metab. 2018, 27, 237–251.e4. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, S.A.; Perez-Gonzalez, R.; Sharma, A.; Huang, F.K.; Alldred, M.J.; Pawlik, M.; Kaur, G.; Ginsberg, S.D.; Neubert, T.A.; Levy, E. Enhanced exosome secretion in Down syndrome brain—A protective mechanism to alleviate neuronal endosomal abnormalities. Acta Neuropathol. Commun. 2017, 5, 65. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Guo, Y.; Yang, W.; Chen, L.; Ren, D.; Wu, C.; He, B.; Zheng, P.; Tong, W. Phosphorylation of connexin 43 induced by traumatic brain injury promotes exosome release. J. Neurophysiol. 2018, 119, 305–311. [Google Scholar] [CrossRef]
- Lee, L.J.; Yang, Z.; Rahman, M.; Ma, J.; Kwak, K.J.; McElroy, J.; Shilo, K.; Goparaju, C.; Yu, L.; Rom, W.; et al. Extracellular mRNA Detected by Tethered Lipoplex Nanoparticle Biochip for Lung Adenocarcinoma Detection. Am. J. Respir. Crit. Care Med. 2016, 193, 1431–1433. [Google Scholar] [CrossRef]
- Notarangelo, M.; Zucal, C.; Modelska, A.; Pesce, I.; Scarduelli, G.; Potrich, C.; Lunelli, L.; Pederzolli, C.; Pavan, P.; la Marca, G.; et al. Ultrasensitive detection of cancer biomarkers by nickel-based isolation of polydisperse extracellular vesicles from blood. EBioMedicine 2019, 43, 114–126. [Google Scholar] [CrossRef]
- Wang, X.; Kwak, K.J.; Yang, Z.; Zhang, A.; Zhang, X.; Sullivan, R.; Lin, D.; Lee, R.L.; Castro, C.; Ghoshal, K.; et al. Extracellular mRNA detected by molecular beacons in tethered lipoplex nanoparticles for diagnosis of human hepatocellular carcinoma. PLoS ONE 2018, 13, e0198552. [Google Scholar] [CrossRef]
- Frye, M.A.; McElroy, S.L.; Fuentes, M.; Sutor, B.; Schak, K.M.; Galardy, C.W.; Palmer, B.A.; Prieto, M.L.; Kung, S.; Sola, C.L.; et al. Development of a bipolar disorder biobank: Differential phenotyping for subsequent biomarker analyses. Int. J. Bipolar. Disord. 2015, 3, 30. [Google Scholar] [CrossRef] [PubMed]
- Frye, M.A.; Nassan, M.; Jenkins, G.D.; Kung, S.; Veldic, M.; Palmer, B.A.; Feeder, S.E.; Tye, S.J.; Choi, D.S.; Biernacka, J.M. Feasibility of investigating differential proteomic expression in depression: Implications for biomarker development in mood disorders. Transl. Psychiatry 2015, 5, e689. [Google Scholar] [CrossRef] [PubMed]
- Kim, E.Y.; Lee, M.Y.; Kim, S.H.; Ha, K.; Kim, K.P.; Ahn, Y.M. Diagnosis of major depressive disorder by combining multimodal information from heart rate dynamics and serum proteomics using machine-learning algorithm. Prog. Neuropsychopharmacol. Biol. Psychiatry 2017, 76, 65–71. [Google Scholar] [CrossRef] [PubMed]
| Cohort | Discovery Set | Validation Set | ||||||
|---|---|---|---|---|---|---|---|---|
| PTSD Status | PTSD- | PTSD+ | PTSD- | PTSD+ | ||||
| Mean | SD | Mean | SD | Mean | SD | Mean | SD | |
| Age (years) | 34.08 | 10.03 | 30.50 | 3.55 | 31.3 | 6.15 | 31.1 | 2.85 |
| BMI (kg/m2) | 26.13 | 2.45 | 27.13 | 3.24 | 26.7 | 3.66 | 28.03 | 4.40 |
| Ethnicity N (%) of Non-Hispanic White | 4 (33.3) | 6 (50.0) | 4 (40.0) | 5 (50.0) | ||||
| Education | 3.58 | 1.16 | 3.25 | 0.87 | 4.00 | 0.67 | 3.60 | 1.07 |
| CAPS Total Score life time | 0.42 | 1.44 | 81.17 | 12.26 | 4.6 | 5.87 | 91.2 | 14.54 |
| SCL90 Depression | 0.27 | 0.67 | 1.82 | 0.62 | 0.16 | 0.22 | 2.11 | 0.95 |
| Beck Depression Inventory II | 3.25 | 6.37 | 26.17 | 8.02 | 1.33 | 2.00 | 27 | 9.13 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, M.Y.; Baxter, D.; Scherler, K.; Kim, T.-K.; Wu, X.; Abu-Amara, D.; Flory, J.; Yehuda, R.; Marmar, C.; Jett, M.; et al. Distinct Profiles of Cell-Free MicroRNAs in Plasma of Veterans with Post-Traumatic Stress Disorder. J. Clin. Med. 2019, 8, 963. https://doi.org/10.3390/jcm8070963
Lee MY, Baxter D, Scherler K, Kim T-K, Wu X, Abu-Amara D, Flory J, Yehuda R, Marmar C, Jett M, et al. Distinct Profiles of Cell-Free MicroRNAs in Plasma of Veterans with Post-Traumatic Stress Disorder. Journal of Clinical Medicine. 2019; 8(7):963. https://doi.org/10.3390/jcm8070963
Chicago/Turabian StyleLee, Min Young, David Baxter, Kelsey Scherler, Taek-Kyun Kim, Xiaogang Wu, Duna Abu-Amara, Janine Flory, Rachel Yehuda, Charles Marmar, Marti Jett, and et al. 2019. "Distinct Profiles of Cell-Free MicroRNAs in Plasma of Veterans with Post-Traumatic Stress Disorder" Journal of Clinical Medicine 8, no. 7: 963. https://doi.org/10.3390/jcm8070963
APA StyleLee, M. Y., Baxter, D., Scherler, K., Kim, T.-K., Wu, X., Abu-Amara, D., Flory, J., Yehuda, R., Marmar, C., Jett, M., Lee, I., Wang, K., & Hood, L. (2019). Distinct Profiles of Cell-Free MicroRNAs in Plasma of Veterans with Post-Traumatic Stress Disorder. Journal of Clinical Medicine, 8(7), 963. https://doi.org/10.3390/jcm8070963






