Microbiome of the Skin and Gut in Atopic Dermatitis (AD): Understanding the Pathophysiology and Finding Novel Management Strategies
Abstract
1. Introduction
2. Skin Microbiota and AD
2.1. Skin Microbiota in AD
2.2. Epidermal Barrier Status Impacts the Composition of the Skin Microbiota
2.3. Staphylococcal Biofilms in AD
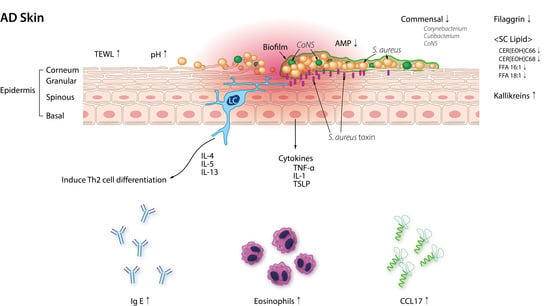
2.4. S. aureus Damages the Skin Barrier and Exacerbates AD Inflammation
2.5. Skin Dysbiosis and AD
2.6. Effect of Treatment on the Skin Microbiota in AD
3. Gut Microbiota and AD
3.1. Gut Microbiota in AD
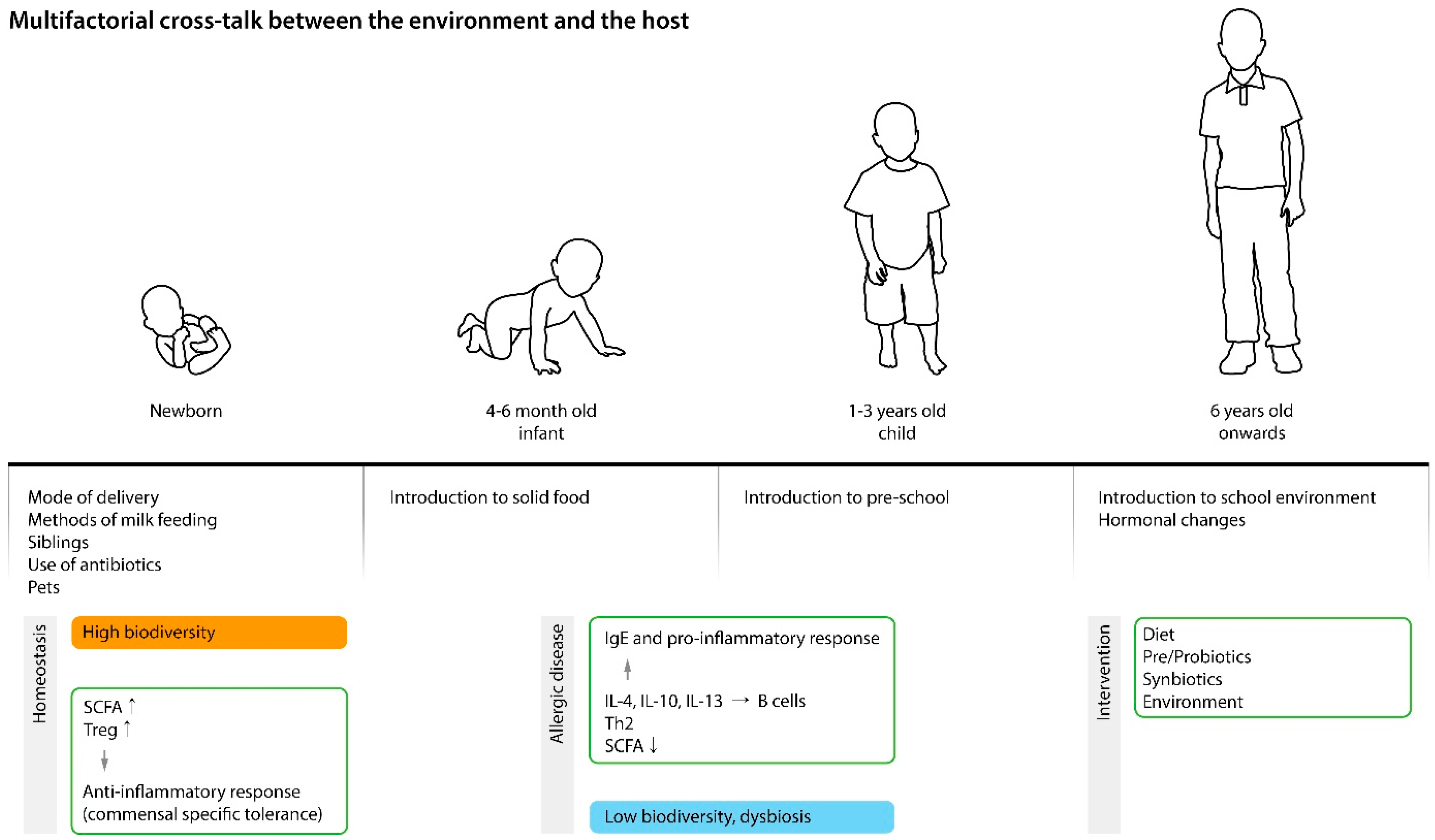
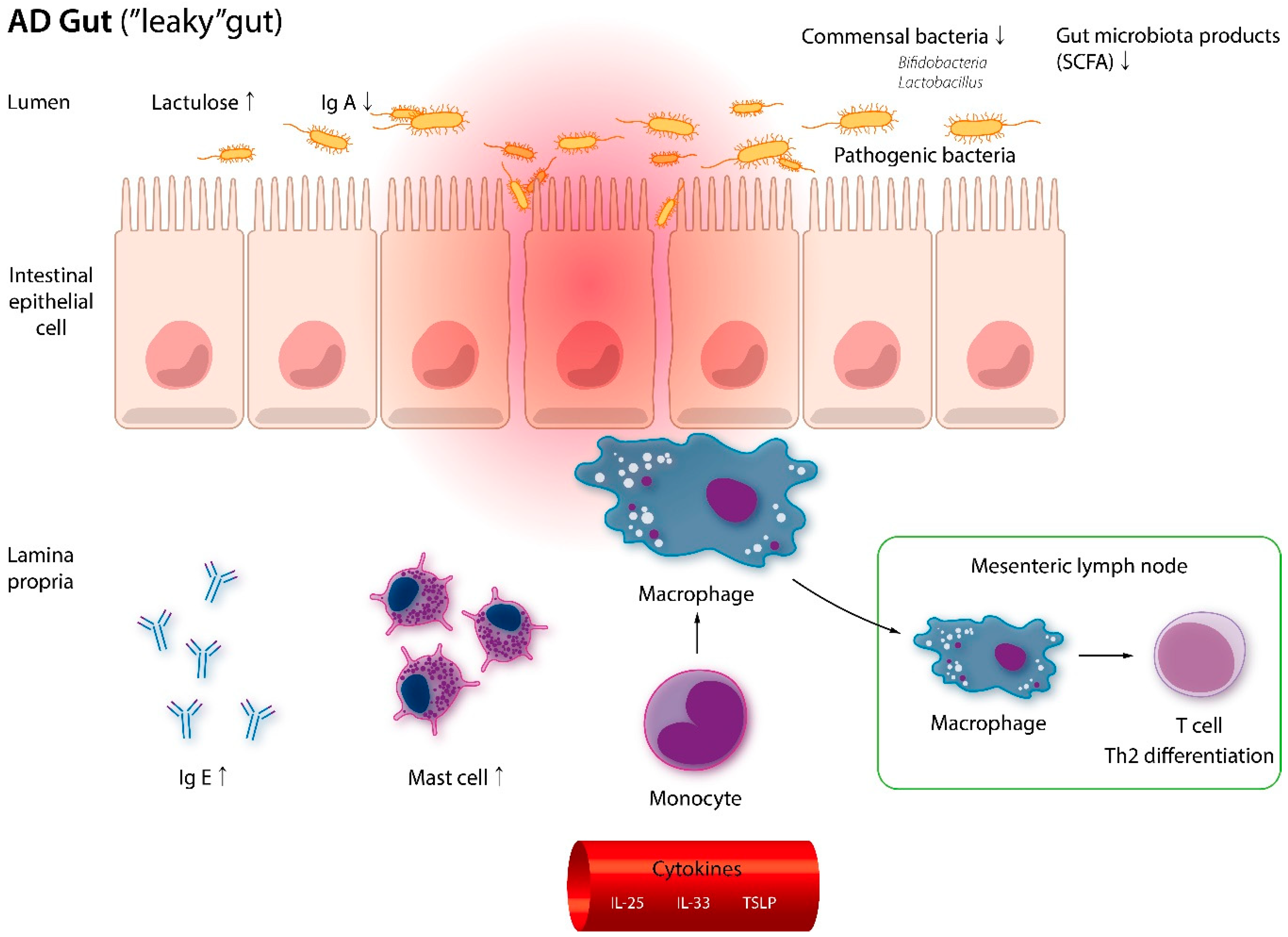
3.2. Association between Gut Dysbiosis and AD
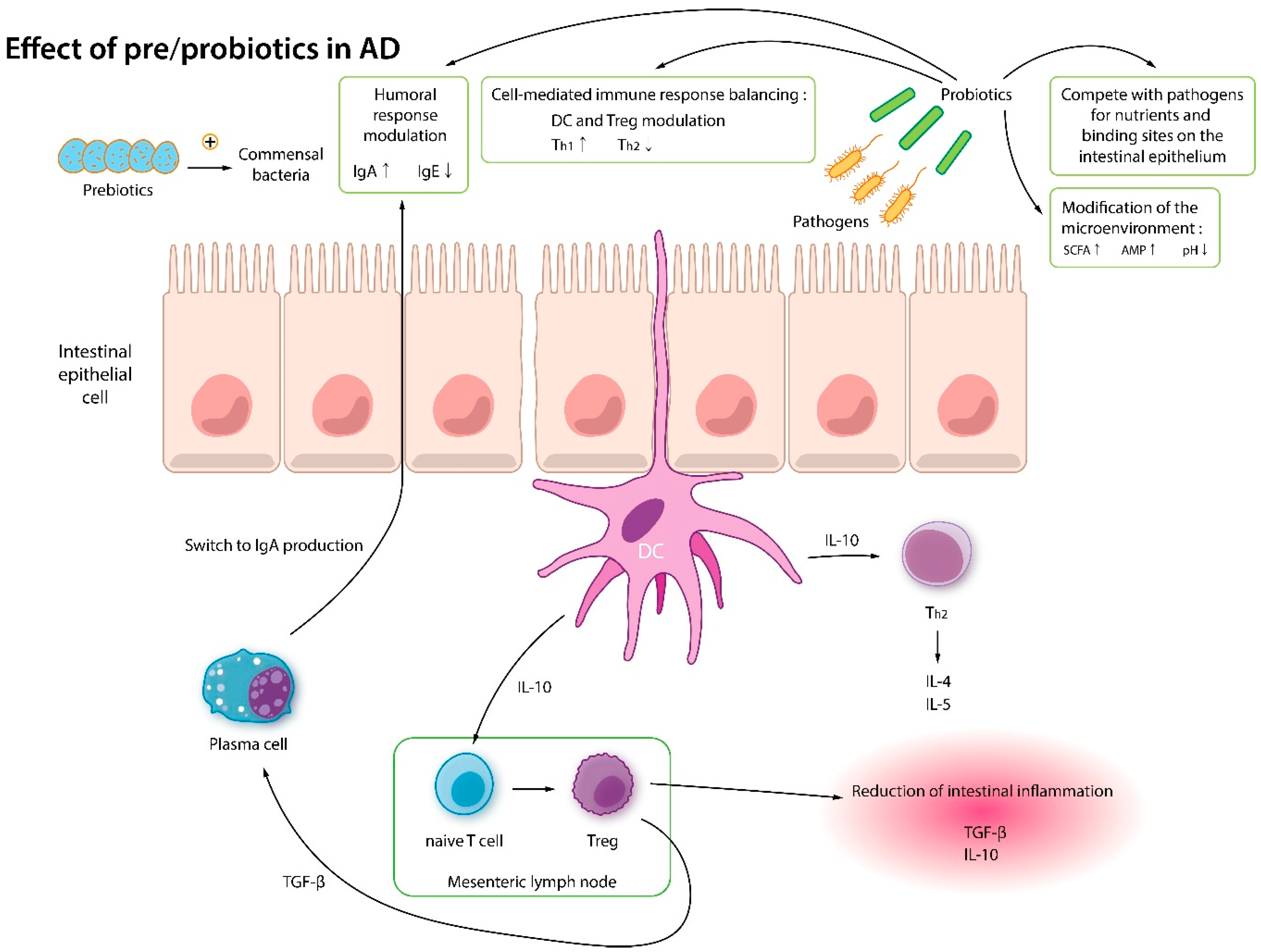
4. Impact of Probiotics/Prebiotics on AD
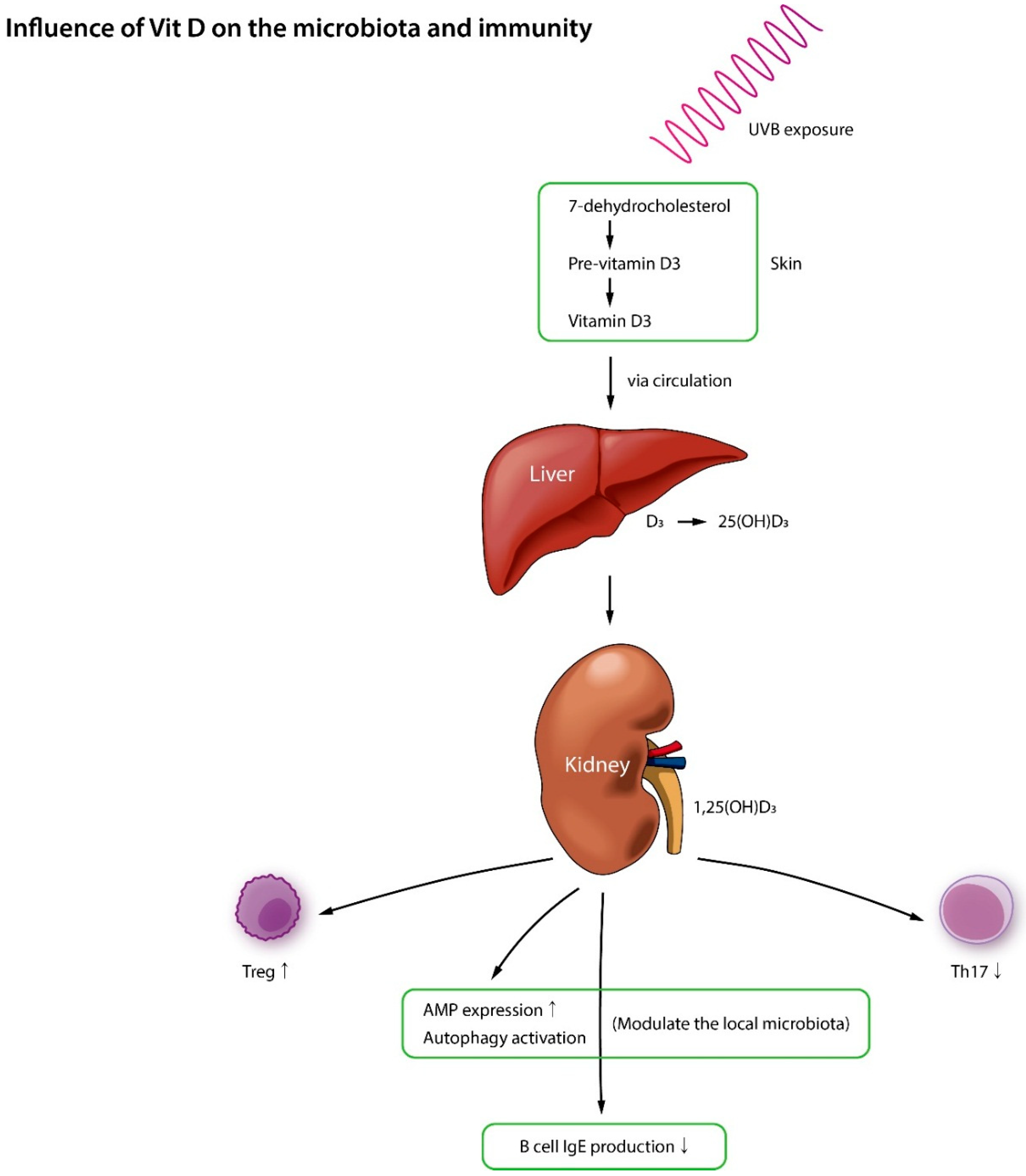
5. Vitamin D, Microbiota, and AD
6. Air Pollution, Gut Microbiota, and AD
7. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Grice, E.A.; Segre, J.A. The human microbiome: Our second genome. Ann. Rev. Genom. Hum. Genet. 2012, 13, 151–170. [Google Scholar] [CrossRef] [PubMed]
- Torres-Fuentes, C.; Schellekens, H.; Dinan, T.G.; Cryan, J.F. The microbiota-gut-brain axis in obesity. Lancet Gastroenterol. Hepatol. 2017, 2, 747–756. [Google Scholar] [CrossRef]
- Sheehan, D.; Shanahan, F. The Gut Microbiota in Inflammatory Bowel Disease. Gastroenterol. Clin. N. Am. 2017, 46, 143–154. [Google Scholar] [CrossRef] [PubMed]
- Blandino, G.; Inturri, R.; Lazzara, F.; Di Rosa, M.; Malaguarnera, L. Impact of gut microbiota on diabetes mellitus. Diabetes Metab. 2016, 42, 303–315. [Google Scholar] [CrossRef]
- Hyun, D.W.; Min, H.J.; Kim, M.S.; Whon, T.W.; Shin, N.R.; Kim, P.S.; Kim, H.S.; Lee, J.Y.; Kang, W.; Choi, A.M.K.; et al. Dysbiosis of Inferior Turbinate Microbiota Is Associated with High Total IgE Levels in Patients with Allergic Rhinitis. Infect. Immun. 2018, 86. [Google Scholar] [CrossRef] [PubMed]
- Yamazaki, Y.; Nakamura, Y.; Nunez, G. Role of the microbiota in skin immunity and atopic dermatitis. Allergol. Int. 2017, 66, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Rapin, A.; Pattaroni, C.; Marsland, B.J.; Harris, N.L. Microbiota Analysis Using an Illumina MiSeq Platform to Sequence 16S rRNA Genes. Curr. Protoc. Mouse Biol. 2017, 7, 100–129. [Google Scholar] [CrossRef]
- Liang, Y.; Chang, C.; Lu, Q. The Genetics and Epigenetics of Atopic Dermatitis-Filaggrin and Other Polymorphisms. Clin. Rev. Allergy Immunol. 2016, 51, 315–328. [Google Scholar] [CrossRef] [PubMed]
- Deckers, I.A.; McLean, S.; Linssen, S.; Mommers, M.; van Schayck, C.P.; Sheikh, A. Investigating international time trends in the incidence and prevalence of atopic eczema 1990-2010: A systematic review of epidemiological studies. PLoS ONE 2012, 7, e39803. [Google Scholar] [CrossRef]
- Wadonda-Kabondo, N.; Sterne, J.A.; Golding, J.; Kennedy, C.T.; Archer, C.B.; Dunnill, M.G. A prospective study of the prevalence and incidence of atopic dermatitis in children aged 0-42 months. Br. J. Dermatol. 2003, 149, 1023–1028. [Google Scholar] [CrossRef]
- Guo, Y.; Li, P.; Tang, J.; Han, X.; Zou, X.; Xu, G.; Xu, Z.; Wei, F.; Liu, Q.; Wang, M.; et al. Prevalence of Atopic Dermatitis in Chinese Children aged 1-7 ys. Sci. Rep. 2016, 6, 29751. [Google Scholar] [CrossRef] [PubMed]
- Asher, M.I.; Montefort, S.; Bjorksten, B.; Lai, C.K.; Strachan, D.P.; Weiland, S.K.; Williams, H. Worldwide time trends in the prevalence of symptoms of asthma, allergic rhinoconjunctivitis, and eczema in childhood: ISAAC Phases One and Three repeat multicountry cross-sectional surveys. Lancet (London England) 2006, 368, 733–743. [Google Scholar] [CrossRef]
- Flohr, C.; Pascoe, D.; Williams, H.C. Atopic dermatitis and the ’hygiene hypothesis’: too clean to be true? Br. J. Dermatol. 2005, 152, 202–216. [Google Scholar] [CrossRef]
- Grice, E.A.; Kong, H.H.; Renaud, G.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Wolfsberg, T.G.; Turner, M.L.; Segre, J.A. A diversity profile of the human skin microbiota. Genome Res. 2008, 18, 1043–1050. [Google Scholar] [CrossRef] [PubMed]
- Belkaid, Y.; Segre, J.A. Dialogue between skin microbiota and immunity. Science 2014, 346, 954–959. [Google Scholar] [CrossRef] [PubMed]
- Findley, K.; Oh, J.; Yang, J.; Conlan, S.; Deming, C.; Meyer, J.A.; Schoenfeld, D.; Nomicos, E.; Park, M.; Kong, H.H.; et al. Topographic diversity of fungal and bacterial communities in human skin. Nature 2013, 498, 367–370. [Google Scholar] [CrossRef]
- Cogen, A.L.; Nizet, V.; Gallo, R.L. Skin microbiota: A source of disease or defence? Br. J. Dermatol. 2008, 158, 442–455. [Google Scholar] [CrossRef]
- Orenstein, A.; Klein, D.; Kopolovic, J.; Winkler, E.; Malik, Z.; Keller, N.; Nitzan, Y. The use of porphyrins for eradication of Staphylococcus aureus in burn wound infections. FEMS Immunol. Med. Microbiol. 1997, 19, 307–314. [Google Scholar] [CrossRef] [PubMed]
- Shi, B.; Bangayan, N.J.; Curd, E.; Taylor, P.A.; Gallo, R.L.; Leung, D.Y.M.; Li, H. The skin microbiome is different in pediatric versus adult atopic dermatitis. J. Allergy Clin. Immunol. 2016, 138, 1233–1236. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.; Conlan, S.; Polley, E.C.; Segre, J.A.; Kong, H.H. Shifts in human skin and nares microbiota of healthy children and adults. Genome Med. 2012, 4, 77. [Google Scholar] [CrossRef]
- Shu, M.; Kuo, S.; Wang, Y.; Jiang, Y.; Liu, Y.T.; Gallo, R.L.; Huang, C.M. Porphyrin metabolisms in human skin commensal Propionibacterium acnes bacteria: potential application to monitor human radiation risk. Curr. Med. Chem. 2013, 20, 562–568. [Google Scholar]
- Shu, M.; Wang, Y.; Yu, J.; Kuo, S.; Coda, A.; Jiang, Y.; Gallo, R.L.; Huang, C.M. Fermentation of Propionibacterium acnes, a commensal bacterium in the human skin microbiome, as skin probiotics against methicillin-resistant Staphylococcus aureus. PLoS ONE 2013, 8, e55380. [Google Scholar] [CrossRef]
- Wang, Y.; Dai, A.; Huang, S.; Kuo, S.; Shu, M.; Tapia, C.P.; Yu, J.; Two, A.; Zhang, H.; Gallo, R.L.; et al. Propionic acid and its esterified derivative suppress the growth of methicillin-resistant Staphylococcus aureus USA300. Benef. Microbes 2014, 5, 161–168. [Google Scholar] [CrossRef]
- Paller, A.S.; Kong, H.H.; Seed, P.; Naik, S.; Scharschmidt, T.C.; Gallo, R.L.; Luger, T.; Irvine, A.D. The Microbiome in Atopic Dermatitis. J. Allergy Clin. Immunol. 2018. [Google Scholar] [CrossRef]
- Totte, J.E.; van der Feltz, W.T.; Hennekam, M.; van Belkum, A.; van Zuuren, E.J.; Pasmans, S.G. Prevalence and odds of Staphylococcus aureus carriage in atopic dermatitis: a systematic review and meta-analysis. Br. J. Dermatol. 2016, 175, 687–695. [Google Scholar] [CrossRef]
- Gonzalez, M.E.; Schaffer, J.V.; Orlow, S.J.; Gao, Z.; Li, H.; Alekseyenko, A.V.; Blaser, M.J. Cutaneous microbiome effects of fluticasone propionate cream and adjunctive bleach baths in childhood atopic dermatitis. J. Am. Acad. Dermatol. 2016, 75, 481–493.e488. [Google Scholar] [CrossRef]
- Kong, H.H.; Oh, J.; Deming, C.; Conlan, S.; Grice, E.A.; Beatson, M.A.; Nomicos, E.; Polley, E.C.; Komarow, H.D.; Murray, P.R.; et al. Temporal shifts in the skin microbiome associated with disease flares and treatment in children with atopic dermatitis. Genome Res. 2012, 22, 850–859. [Google Scholar] [CrossRef]
- Oh, J.; Freeman, A.F.; Park, M.; Sokolic, R.; Candotti, F.; Holland, S.M.; Segre, J.A.; Kong, H.H. The altered landscape of the human skin microbiome in patients with primary immunodeficiencies. Genome Res. 2013, 23, 2103–2114. [Google Scholar] [CrossRef]
- Seite, S.; Flores, G.E.; Henley, J.B.; Martin, R.; Zelenkova, H.; Aguilar, L.; Fierer, N. Microbiome of affected and unaffected skin of patients with atopic dermatitis before and after emollient treatment. JDD 2014, 13, 1365–1372. [Google Scholar] [PubMed]
- Francuzik, W.; Franke, K.; Schumann, R.R.; Heine, G.; Worm, M. Propionibacterium acnes Abundance Correlates Inversely with Staphylococcus aureus: Data from Atopic Dermatitis Skin Microbiome. Acta Derm. Venereol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Bradley, C.W.; Morris, D.O.; Rankin, S.C.; Cain, C.L.; Misic, A.M.; Houser, T.; Mauldin, E.A.; Grice, E.A. Longitudinal Evaluation of the Skin Microbiome and Association with Microenvironment and Treatment in Canine Atopic Dermatitis. J. Investig. Dermatol. 2016, 136, 1182–1190. [Google Scholar] [CrossRef] [PubMed]
- Byrd, A.L.; Deming, C.; Cassidy, S.K.B.; Harrison, O.J.; Ng, W.I.; Conlan, S.; Belkaid, Y.; Segre, J.A.; Kong, H.H. Staphylococcus aureus and Staphylococcus epidermidis strain diversity underlying pediatric atopic dermatitis. Sci. Trans. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Fleury, O.M.; McAleer, M.A.; Feuillie, C.; Formosa-Dague, C.; Sansevere, E.; Bennett, D.E.; Towell, A.M.; McLean, W.H.I.; Kezic, S.; Robinson, D.A.; et al. Clumping Factor B Promotes Adherence of Staphylococcus aureus to Corneocytes in Atopic Dermatitis. Infect. Immun. 2017, 85. [Google Scholar] [CrossRef]
- Simpson, E.L.; Villarreal, M.; Jepson, B.; Rafaels, N.; David, G.; Hanifin, J.; Taylor, P.; Boguniewicz, M.; Yoshida, T.; De Benedetto, A.; et al. Patients with Atopic Dermatitis Colonized with Staphylococcus aureus Have a Distinct Phenotype and Endotype. J. Investig. Dermatol. 2018, 138, 2224–2233. [Google Scholar] [CrossRef] [PubMed]
- Shi, B.; Leung, D.Y.M.; Taylor, P.A.; Li, H. Methicillin-Resistant Staphylococcus aureus Colonization Is Associated with Decreased Skin Commensal Bacteria in Atopic Dermatitis. J. Investig. Dermatol. 2018, 138, 1668–1671. [Google Scholar] [CrossRef]
- Zhang, E.; Tanaka, T.; Tajima, M.; Tsuboi, R.; Nishikawa, A.; Sugita, T. Characterization of the skin fungal microbiota in patients with atopic dermatitis and in healthy subjects. Microb. Immunol. 2011, 55, 625–632. [Google Scholar] [CrossRef] [PubMed]
- Bjerre, R.D.; Bandier, J.; Skov, L.; Engstrand, L.; Johansen, J.D. The role of the skin microbiome in atopic dermatitis: a systematic review. Br. J. Dermatol. 2017, 177, 1272–1278. [Google Scholar] [CrossRef] [PubMed]
- Baurecht, H.; Ruhlemann, M.C.; Rodriguez, E.; Thielking, F.; Harder, I.; Erkens, A.S.; Stolzl, D.; Ellinghaus, E.; Hotze, M.; Lieb, W.; et al. Epidermal lipid composition, barrier integrity and eczematous inflammation are associated with skin microbiome configuration. J. Allergy Clin. Immunol. 2018. [Google Scholar] [CrossRef]
- Li, S.; Villarreal, M.; Stewart, S.; Choi, J.; Ganguli-Indra, G.; Babineau, D.C.; Philpot, C.; David, G.; Yoshida, T.; Boguniewicz, M.; et al. Altered composition of epidermal lipids correlates with Staphylococcus aureus colonization status in atopic dermatitis. Br. J. Dermatol. 2017, 177, e125–e127. [Google Scholar] [CrossRef] [PubMed]
- Parsons, J.B.; Yao, J.; Frank, M.W.; Jackson, P.; Rock, C.O. Membrane disruption by antimicrobial fatty acids releases low-molecular-weight proteins from Staphylococcus aureus. J. Bacteriol. 2012, 194, 5294–5304. [Google Scholar] [CrossRef]
- Cabanillas, B.; Novak, N. Atopic dermatitis and filaggrin. Curr. Opin. Immunol. 2016, 42, 1–8. [Google Scholar] [CrossRef]
- McAleer, M.A.; Irvine, A.D. The multifunctional role of filaggrin in allergic skin disease. J. Allergy Clin. Immunol. 2013, 131, 280–291. [Google Scholar] [CrossRef]
- Miajlovic, H.; Fallon, P.G.; Irvine, A.D.; Foster, T.J. Effect of filaggrin breakdown products on growth of and protein expression by Staphylococcus aureus. J. Allergy Clin. Immunol. 2010, 126, 1184–1190.e1183. [Google Scholar] [CrossRef]
- Komatsu, N.; Saijoh, K.; Kuk, C.; Liu, A.C.; Khan, S.; Shirasaki, F.; Takehara, K.; Diamandis, E.P. Human tissue kallikrein expression in the stratum corneum and serum of atopic dermatitis patients. Exp. Dermatol. 2007, 16, 513–519. [Google Scholar] [CrossRef] [PubMed]
- Voegeli, R.; Rawlings, A.V.; Breternitz, M.; Doppler, S.; Schreier, T.; Fluhr, J.W. Increased stratum corneum serine protease activity in acute eczematous atopic skin. Br. J. Dermatol. 2009, 161, 70–77. [Google Scholar] [CrossRef]
- Borgono, C.A.; Michael, I.P.; Komatsu, N.; Jayakumar, A.; Kapadia, R.; Clayman, G.L.; Sotiropoulou, G.; Diamandis, E.P. A potential role for multiple tissue kallikrein serine proteases in epidermal desquamation. J. Biol. Chem. 2007, 282, 3640–3652. [Google Scholar] [CrossRef]
- Sakabe, J.; Yamamoto, M.; Hirakawa, S.; Motoyama, A.; Ohta, I.; Tatsuno, K.; Ito, T.; Kabashima, K.; Hibino, T.; Tokura, Y. Kallikrein-related peptidase 5 functions in proteolytic processing of profilaggrin in cultured human keratinocytes. J. Biol. Chem. 2013, 288, 17179–17189. [Google Scholar] [CrossRef]
- Stefansson, K.; Brattsand, M.; Roosterman, D.; Kempkes, C.; Bocheva, G.; Steinhoff, M.; Egelrud, T. Activation of proteinase-activated receptor-2 by human kallikrein-related peptidases. J. Investig. Dermatol. 2008, 128, 18–25. [Google Scholar] [CrossRef]
- Yamasaki, K.; Schauber, J.; Coda, A.; Lin, H.; Dorschner, R.A.; Schechter, N.M.; Bonnart, C.; Descargues, P.; Hovnanian, A.; Gallo, R.L. Kallikrein-mediated proteolysis regulates the antimicrobial effects of cathelicidins in skin. FASEB J. 2006, 20, 2068–2080. [Google Scholar] [CrossRef] [PubMed]
- Hata, T.R.; Gallo, R.L. Antimicrobial peptides, skin infections, and atopic dermatitis. Semin. Cutan. Med. Surg. 2008, 27, 144–150. [Google Scholar] [CrossRef] [PubMed]
- Nakatsuji, T.; Chen, T.H.; Narala, S.; Chun, K.A.; Two, A.M.; Yun, T.; Shafiq, F.; Kotol, P.F.; Bouslimani, A.; Melnik, A.V.; et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Trans. Med. 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Di Domenico, E.G.; Cavallo, I.; Bordignon, V.; Prignano, G.; Sperduti, I.; Gurtner, A.; Trento, E.; Toma, L.; Pimpinelli, F.; Capitanio, B.; et al. Inflammatory cytokines and biofilm production sustain Staphylococcus aureus outgrowth and persistence: a pivotal interplay in the pathogenesis of Atopic Dermatitis. Sci. Rep. 2018, 8, 9573. [Google Scholar] [CrossRef]
- Allen, H.B.; Vaze, N.D.; Choi, C.; Hailu, T.; Tulbert, B.H.; Cusack, C.A.; Joshi, S.G. The presence and impact of biofilm-producing staphylococci in atopic dermatitis. JAMA Dermatol. 2014, 150, 260–265. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, T.; Biagini Myers, J.M.; Herr, A.B.; Khurana Hershey, G.K. Staphylococcal Biofilms in Atopic Dermatitis. Curr. Allergy Asthma Rep. 2017, 17, 81. [Google Scholar] [CrossRef] [PubMed]
- Scherr, T.D.; Heim, C.E.; Morrison, J.M.; Kielian, T. Hiding in Plain Sight: Interplay between Staphylococcal Biofilms and Host Immunity. Front. Immunol. 2014, 5, 37. [Google Scholar] [CrossRef] [PubMed]
- Scherr, T.D.; Hanke, M.L.; Huang, O.; James, D.B.; Horswill, A.R.; Bayles, K.W.; Fey, P.D.; Torres, V.J.; Kielian, T. Staphylococcus aureus Biofilms Induce Macrophage Dysfunction Through Leukocidin AB and Alpha-Toxin. mBio 2015, 6. [Google Scholar] [CrossRef] [PubMed]
- Thurlow, L.R.; Hanke, M.L.; Fritz, T.; Angle, A.; Aldrich, A.; Williams, S.H.; Engebretsen, I.L.; Bayles, K.W.; Horswill, A.R.; Kielian, T. Staphylococcus aureus biofilms prevent macrophage phagocytosis and attenuate inflammation in vivo. J. Immunol. 2011, 186, 6585–6596. [Google Scholar] [CrossRef]
- Tankersley, A.; Frank, M.B.; Bebak, M.; Brennan, R. Early effects of Staphylococcus aureus biofilm secreted products on inflammatory responses of human epithelial keratinocytes. J. Inflamm. 2014, 11, 17. [Google Scholar] [CrossRef] [PubMed]
- Takai, T. TSLP expression: Cellular sources, triggers, and regulatory mechanisms. Allergol. Int. 2012, 61, 3–17. [Google Scholar] [CrossRef] [PubMed]
- Wilson, S.R.; The, L.; Batia, L.M.; Beattie, K.; Katibah, G.E.; McClain, S.P.; Pellegrino, M.; Estandian, D.M.; Bautista, D.M. The epithelial cell-derived atopic dermatitis cytokine TSLP activates neurons to induce itch. Cell 2013, 155, 285–295. [Google Scholar] [CrossRef]
- Ong, P.Y.; Leung, D.Y. Bacterial and Viral Infections in Atopic Dermatitis: A Comprehensive Review. Clin. Rev. Allergy Immunol. 2016, 51, 329–337. [Google Scholar] [CrossRef] [PubMed]
- Son, E.D.; Kim, H.J.; Park, T.; Shin, K.; Bae, I.H.; Lim, K.M.; Cho, E.G.; Lee, T.R. Staphylococcus aureus inhibits terminal differentiation of normal human keratinocytes by stimulating interleukin-6 secretion. J. Dermatol. Sci. 2014, 74, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Den Reijer, P.M.; Haisma, E.M.; Lemmens-den Toom, N.A.; Willemse, J.; Koning, R.I.; Demmers, J.A.; Dekkers, D.H.; Rijkers, E.; El Ghalbzouri, A.; Nibbering, P.H.; et al. Detection of Alpha-Toxin and Other Virulence Factors in Biofilms of Staphylococcus aureus on Polystyrene and a Human Epidermal Model. PLoS ONE 2016, 11, e0145722. [Google Scholar] [CrossRef]
- Sonesson, A.; Przybyszewska, K.; Eriksson, S.; Morgelin, M.; Kjellstrom, S.; Davies, J.; Potempa, J.; Schmidtchen, A. Identification of bacterial biofilm and the Staphylococcus aureus derived protease, staphopain, on the skin surface of patients with atopic dermatitis. Sci. Rep. 2017, 7, 8689. [Google Scholar] [CrossRef] [PubMed]
- Song, L.; Hobaugh, M.R.; Shustak, C.; Cheley, S.; Bayley, H.; Gouaux, J.E. Structure of staphylococcal alpha-hemolysin, a heptameric transmembrane pore. Science 1996, 274, 1859–1866. [Google Scholar] [CrossRef]
- Berube, B.J.; Bubeck Wardenburg, J. Staphylococcus aureus alpha-toxin: nearly a century of intrigue. Toxins 2013, 5, 1140–1166. [Google Scholar] [CrossRef] [PubMed]
- Nakatsuji, T.; Chen, T.H.; Two, A.M.; Chun, K.A.; Narala, S.; Geha, R.S.; Hata, T.R.; Gallo, R.L. Staphylococcus aureus Exploits Epidermal Barrier Defects in Atopic Dermatitis to Trigger Cytokine Expression. J. Investig. Dermatol. 2016, 136, 2192–2200. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.R.; Nakatsuji, T.; Sanford, J.A.; Vrbanac, A.F.; Gallo, R.L. Staphylococcus aureus Induces Increased Serine Protease Activity in Keratinocytes. J. Investig. Dermatol. 2017, 137, 377–384. [Google Scholar] [CrossRef] [PubMed]
- Fischer, J.; Meyer-Hoffert, U. Regulation of kallikrein-related peptidases in the skin-from physiology to diseases to therapeutic options. Thromb. Haemost. 2013, 110, 442–449. [Google Scholar] [CrossRef]
- Brauweiler, A.M.; Goleva, E.; Leung, D.Y.M. Staphylococcus aureus Lipoteichoic Acid Damages the Skin Barrier through an IL-1 mediated Pathway. J. Investig. Dermatol. 2019. [Google Scholar] [CrossRef]
- Williams, M.R.; Nakatsuji, T.; Gallo, R.L. Staphylococcus aureus: Master Manipulator of the Skin. Cell Host Microbe 2017, 22, 579–581. [Google Scholar] [CrossRef]
- Spaulding, A.R.; Salgado-Pabon, W.; Kohler, P.L.; Horswill, A.R.; Leung, D.Y.; Schlievert, P.M. Staphylococcal and streptococcal superantigen exotoxins. Clin. Microbiol. Rev. 2013, 26, 422–447. [Google Scholar] [CrossRef] [PubMed]
- Vu, A.T.; Baba, T.; Chen, X.; Le, T.A.; Kinoshita, H.; Xie, Y.; Kamijo, S.; Hiramatsu, K.; Ikeda, S.; Ogawa, H.; et al. Staphylococcus aureus membrane and diacylated lipopeptide induce thymic stromal lymphopoietin in keratinocytes through the Toll-like receptor 2-Toll-like receptor 6 pathway. J. Allergy Clin. Immunol. 2010, 126, 985–993. [Google Scholar] [CrossRef] [PubMed]
- Nakamura, Y.; Oscherwitz, J.; Cease, K.B.; Chan, S.M.; Munoz-Planillo, R.; Hasegawa, M.; Villaruz, A.E.; Cheung, G.Y.; McGavin, M.J.; Travers, J.B.; et al. Staphylococcus delta-toxin induces allergic skin disease by activating mast cells. Nature 2013, 503, 397–401. [Google Scholar] [CrossRef] [PubMed]
- Naik, S.; Bouladoux, N.; Wilhelm, C.; Molloy, M.J.; Salcedo, R.; Kastenmuller, W.; Deming, C.; Quinones, M.; Koo, L.; Conlan, S.; et al. Compartmentalized control of skin immunity by resident commensals. Science 2012, 337, 1115–1119. [Google Scholar] [CrossRef]
- Scharschmidt, T.C.; Vasquez, K.S.; Truong, H.A.; Gearty, S.V.; Pauli, M.L.; Nosbaum, A.; Gratz, I.K.; Otto, M.; Moon, J.J.; Liese, J.; et al. A Wave of Regulatory T Cells into Neonatal Skin Mediates Tolerance to Commensal Microbes. Immunity 2015, 43, 1011–1021. [Google Scholar] [CrossRef]
- Kobayashi, T.; Glatz, M.; Horiuchi, K.; Kawasaki, H.; Akiyama, H.; Kaplan, D.H.; Kong, H.H.; Amagai, M.; Nagao, K. Dysbiosis and Staphylococcus aureus Colonization Drives Inflammation in Atopic Dermatitis. Immunity 2015, 42, 756–766.e745. [Google Scholar] [CrossRef] [PubMed]
- Kennedy, E.A.; Connolly, J.; Hourihane, J.O.; Fallon, P.G.; McLean, W.H.I.; Murray, D.; Jo, J.H.; Segre, J.A.; Kong, H.H.; Irvine, A.D. Skin microbiome before development of atopic dermatitis: Early colonization with commensal staphylococci at 2 months is associated with a lower risk of atopic dermatitis at 1 year. J. Allergy Clin. Immunol. 2017, 139, 166–172. [Google Scholar] [CrossRef]
- Meylan, P.; Lang, C.; Mermoud, S.; Johannsen, A.; Norrenberg, S.; Hohl, D.; Vial, Y.; Prod’hom, G.; Greub, G.; Kypriotou, M.; et al. Skin Colonization by Staphylococcus aureus Precedes the Clinical Diagnosis of Atopic Dermatitis in Infancy. J. Investig. Dermatol. 2017, 137, 2497–2504. [Google Scholar] [CrossRef]
- Bunikowski, R.; Mielke, M.E.; Skarabis, H.; Worm, M.; Anagnostopoulos, I.; Kolde, G.; Wahn, U.; Renz, H. Evidence for a disease-promoting effect of Staphylococcus aureus-derived exotoxins in atopic dermatitis. J. Allergy Clin. Immunol. 2000, 105, 814–819. [Google Scholar] [CrossRef]
- Kollmann, T.R.; Levy, O.; Montgomery, R.R.; Goriely, S. Innate immune function by Toll-like receptors: distinct responses in newborns and the elderly. Immunity 2012, 37, 771–783. [Google Scholar] [CrossRef]
- McGovern, N.; Shin, A.; Low, G.; Low, D.; Duan, K.; Yao, L.J.; Msallam, R.; Low, I.; Shadan, N.B.; Sumatoh, H.R.; et al. Human fetal dendritic cells promote prenatal T-cell immune suppression through arginase-2. Nature 2017, 546, 662–666. [Google Scholar] [CrossRef]
- Yang, S.; Fujikado, N.; Kolodin, D.; Benoist, C.; Mathis, D. Immune tolerance. Regulatory T cells generated early in life play a distinct role in maintaining self-tolerance. Science 2015, 348, 589–594. [Google Scholar] [CrossRef] [PubMed]
- Thome, J.J.; Bickham, K.L.; Ohmura, Y.; Kubota, M.; Matsuoka, N.; Gordon, C.; Granot, T.; Griesemer, A.; Lerner, H.; Kato, T.; et al. Early-life compartmentalization of human T cell differentiation and regulatory function in mucosal and lymphoid tissues. Nat. Med. 2016, 22, 72–77. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.; Choi, J.Y.; Shin, J.W.; Huh, C.H.; Park, K.C.; Du, M.H.; Yoon, S.; Na, J.I. Changes in Lesional and Non-lesional Skin Microbiome During Treatment of Atopic Dermatitis. Acta Derm. Venereol. 2018. [Google Scholar] [CrossRef] [PubMed]
- Sidbury, R.; Davis, D.M.; Cohen, D.E.; Cordoro, K.M.; Berger, T.G.; Bergman, J.N.; Chamlin, S.L.; Cooper, K.D.; Feldman, S.R.; Hanifin, J.M.; et al. Guidelines of care for the management of atopic dermatitis: Section 3. Management and treatment with phototherapy and systemic agents. J. Am. Acad. Dermatol. 2014, 71, 327–349. [Google Scholar] [CrossRef]
- Darne, S.; Leech, S.N.; Taylor, A.E. Narrowband ultraviolet B phototherapy in children with moderate-to-severe eczema: A comparative cohort study. Br. J. Dermatol. 2014, 170, 150–156. [Google Scholar] [CrossRef]
- Yoshimura, M.; Namura, S.; Akamatsu, H.; Horio, T. Antimicrobial effects of phototherapy and photochemotherapy in vivo and in vitro. Br. J. Dermatol. 1996, 135, 528–532. [Google Scholar] [CrossRef] [PubMed]
- Silva, S.H.; Guedes, A.C.; Gontijo, B.; Ramos, A.M.; Carmo, L.S.; Farias, L.M.; Nicoli, J.R. Influence of narrow-band UVB phototherapy on cutaneous microbiota of children with atopic dermatitis. JEADV 2006, 20, 1114–1120. [Google Scholar] [CrossRef] [PubMed]
- Felton, S.; Navid, F.; Schwarz, A.; Schwarz, T.; Glaser, R.; Rhodes, L.E. Ultraviolet radiation-induced upregulation of antimicrobial proteins in health and disease. Photochem. Photobiol. Sci. 2013, 12, 29–36. [Google Scholar] [CrossRef]
- Wu, G.D.; Lewis, J.D. Analysis of the human gut microbiome and association with disease. Clin. Gastroenterol. Hepatol. 2013, 11, 774–777. [Google Scholar] [CrossRef] [PubMed]
- Arrieta, M.C.; Stiemsma, L.T.; Amenyogbe, N.; Brown, E.M.; Finlay, B. The intestinal microbiome in early life: health and disease. Front. Immunol. 2014, 5, 427. [Google Scholar] [CrossRef]
- Fujimura, K.E.; Slusher, N.A.; Cabana, M.D.; Lynch, S.V. Role of the gut microbiota in defining human health. Exp. Rev. Anti-Infect. Ther. 2010, 8, 435–454. [Google Scholar] [CrossRef]
- Tannock, G.W.; Fuller, R.; Smith, S.L.; Hall, M.A. Plasmid profiling of members of the family Enterobacteriaceae, lactobacilli, and bifidobacteria to study the transmission of bacteria from mother to infant. J. Clin. Microbiol. 1990, 28, 1225–1228. [Google Scholar] [PubMed]
- Macdonald, T.T.; Monteleone, G. Immunity, inflammation, and allergy in the gut. Science 2005, 307, 1920–1925. [Google Scholar] [CrossRef] [PubMed]
- Kelly, D.; Campbell, J.I.; King, T.P.; Grant, G.; Jansson, E.A.; Coutts, A.G.; Pettersson, S.; Conway, S. Commensal anaerobic gut bacteria attenuate inflammation by regulating nuclear-cytoplasmic shuttling of PPAR-gamma and RelA. Nat. Immunol. 2004, 5, 104–112. [Google Scholar] [CrossRef]
- Corr, S.C.; Hill, C.; Gahan, C.G. Understanding the mechanisms by which probiotics inhibit gastrointestinal pathogens. Adv. Food Nutr. Res. 2009, 56, 1–15. [Google Scholar] [CrossRef]
- Wagner, C.L.; Taylor, S.N.; Johnson, D. Host factors in amniotic fluid and breast milk that contribute to gut maturation. Clin. Rev. Allergy Immunol. 2008, 34, 191–204. [Google Scholar] [CrossRef] [PubMed]
- Penders, J.; Thijs, C.; Vink, C.; Stelma, F.F.; Snijders, B.; Kummeling, I.; van den Brandt, P.A.; Stobberingh, E.E. Factors influencing the composition of the intestinal microbiota in early infancy. Pediatrics 2006, 118, 511–521. [Google Scholar] [CrossRef]
- Penders, J.; Thijs, C.; van den Brandt, P.A.; Kummeling, I.; Snijders, B.; Stelma, F.; Adams, H.; van Ree, R.; Stobberingh, E.E. Gut microbiota composition and development of atopic manifestations in infancy: The KOALA Birth Cohort Study. Gut 2007, 56, 661–667. [Google Scholar] [CrossRef] [PubMed]
- Klaassens, E.S.; Boesten, R.J.; Haarman, M.; Knol, J.; Schuren, F.H.; Vaughan, E.E.; de Vos, W.M. Mixed-species genomic microarray analysis of fecal samples reveals differential transcriptional responses of bifidobacteria in breast- and formula-fed infants. App. Environ. Microbiol. 2009, 75, 2668–2676. [Google Scholar] [CrossRef] [PubMed]
- Valles, Y.; Artacho, A.; Pascual-Garcia, A.; Ferrus, M.L.; Gosalbes, M.J.; Abellan, J.J.; Francino, M.P. Microbial succession in the gut: Directional trends of taxonomic and functional change in a birth cohort of Spanish infants. PLoS Genet. 2014, 10, e1004406. [Google Scholar] [CrossRef] [PubMed]
- Palmer, C.; Bik, E.M.; DiGiulio, D.B.; Relman, D.A.; Brown, P.O. Development of the human infant intestinal microbiota. PLoS Biol. 2007, 5, e177. [Google Scholar] [CrossRef]
- Fallani, M.; Amarri, S.; Uusijarvi, A.; Adam, R.; Khanna, S.; Aguilera, M.; Gil, A.; Vieites, J.M.; Norin, E.; Young, D.; et al. Determinants of the human infant intestinal microbiota after the introduction of first complementary foods in infant samples from five European centres. Microbiology (Reading England) 2011, 157, 1385–1392. [Google Scholar] [CrossRef]
- Tanaka, M.; Nakayama, J. Development of the gut microbiota in infancy and its impact on health in later life. Allergol. Int. 2017, 66, 515–522. [Google Scholar] [CrossRef]
- Purchiaroni, F.; Tortora, A.; Gabrielli, M.; Bertucci, F.; Gigante, G.; Ianiro, G.; Ojetti, V.; Scarpellini, E.; Gasbarrini, A. The role of intestinal microbiota and the immune system. Eur. Rev. Med. Pharmacol. Sci. 2013, 17, 323–333. [Google Scholar]
- Macpherson, A.J.; Harris, N.L. Interactions between commensal intestinal bacteria and the immune system. Nat. Rev. Immunol. 2004, 4, 478–485. [Google Scholar] [CrossRef]
- Atarashi, K.; Nishimura, J.; Shima, T.; Umesaki, Y.; Yamamoto, M.; Onoue, M.; Yagita, H.; Ishii, N.; Evans, R.; Honda, K.; et al. ATP drives lamina propria T(H)17 cell differentiation. Nature 2008, 455, 808–812. [Google Scholar] [CrossRef]
- Nguyen, Q.N.; Himes, J.E.; Martinez, D.R.; Permar, S.R. The Impact of the Gut Microbiota on Humoral Immunity to Pathogens and Vaccination in Early Infancy. PLoS Pathog. 2016, 12, e1005997. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, S.; Narisawa, Y.; Arase, S.; Okamatsu, H.; Ikenaga, T.; Tajiri, Y.; Kumemura, M. Differences in fecal microflora between patients with atopic dermatitis and healthy control subjects. J. Allergy Clin. Immunol. 2003, 111, 587–591. [Google Scholar] [CrossRef]
- Song, H.; Yoo, Y.; Hwang, J.; Na, Y.C.; Kim, H.S. Faecalibacterium prausnitzii subspecies-level dysbiosis in the human gut microbiome underlying atopic dermatitis. J. Allergy Clin. Immunol. 2016, 137, 852–860. [Google Scholar] [CrossRef] [PubMed]
- Fieten, K.B.; Totte, J.E.E.; Levin, E.; Reyman, M.; Meijer, Y.; Knulst, A.; Schuren, F.; Pasmans, S. Fecal Microbiome and Food Allergy in Pediatric Atopic Dermatitis: A Cross-Sectional Pilot Study. Int. Arch. Allergy Immunol. 2018, 175, 77–84. [Google Scholar] [CrossRef] [PubMed]
- Gensollen, T.; Blumberg, R.S. Correlation between early-life regulation of the immune system by microbiota and allergy development. J. Allergy Clin. Immunol. 2017, 139, 1084–1091. [Google Scholar] [CrossRef]
- Lee, S.Y.; Lee, E.; Park, Y.M.; Hong, S.J. Microbiome in the Gut-Skin Axis in Atopic Dermatitis. Allergy Asthma Immunol. Res. 2018, 10, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Abrahamsson, T.R.; Jakobsson, H.E.; Andersson, A.F.; Bjorksten, B.; Engstrand, L.; Jenmalm, M.C. Low diversity of the gut microbiota in infants with atopic eczema. J. Allergy Clin. Immunol. 2012, 129, 434–440. [Google Scholar] [CrossRef]
- Penders, J.; Stobberingh, E.E.; Thijs, C.; Adams, H.; Vink, C.; van Ree, R.; van den Brandt, P.A. Molecular fingerprinting of the intestinal microbiota of infants in whom atopic eczema was or was not developing. Clin. Exp. Allergy 2006, 36, 1602–1608. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.; Lee, S.Y.; Kang, M.J.; Kim, K.; Won, S.; Kim, B.J.; Choi, K.Y.; Kim, B.S.; Cho, H.J.; Kim, Y.; et al. Clostridia in the gut and onset of atopic dermatitis via eosinophilic inflammation. Ann. Allergy Asthma Immunol. 2016, 117, 91–92.e91. [Google Scholar] [CrossRef]
- Kirjavainen, P.V.; Arvola, T.; Salminen, S.J.; Isolauri, E. Aberrant composition of gut microbiota of allergic infants: A target of bifidobacterial therapy at weaning? Gut 2002, 51, 51–55. [Google Scholar] [CrossRef]
- Nylund, L.; Nermes, M.; Isolauri, E.; Salminen, S.; de Vos, W.M.; Satokari, R. Severity of atopic disease inversely correlates with intestinal microbiota diversity and butyrate-producing bacteria. Allergy 2015, 70, 241–244. [Google Scholar] [CrossRef]
- Fujimura, K.E.; Sitarik, A.R.; Havstad, S.; Lin, D.L.; Levan, S.; Fadrosh, D.; Panzer, A.R.; LaMere, B.; Rackaityte, E.; Lukacs, N.W.; et al. Neonatal gut microbiota associates with childhood multisensitized atopy and T cell differentiation. Nat. Med. 2016, 22, 1187–1191. [Google Scholar] [CrossRef]
- Kalliomaki, M.; Kirjavainen, P.; Eerola, E.; Kero, P.; Salminen, S.; Isolauri, E. Distinct patterns of neonatal gut microflora in infants in whom atopy was and was not developing. J. Allergy Clin. Immunol. 2001, 107, 129–134. [Google Scholar] [CrossRef] [PubMed]
- Ismail, I.H.; Oppedisano, F.; Joseph, S.J.; Boyle, R.J.; Licciardi, P.V.; Robins-Browne, R.M.; Tang, M.L. Reduced gut microbial diversity in early life is associated with later development of eczema but not atopy in high-risk infants. Pediatr. Allergy Immunol. 2012, 23, 674–681. [Google Scholar] [CrossRef] [PubMed]
- He, F.; Ouwehand, A.C.; Isolauri, E.; Hashimoto, H.; Benno, Y.; Salminen, S. Comparison of mucosal adhesion and species identification of bifidobacteria isolated from healthy and allergic infants. FEMS Immunol. Med. Microbiol. 2001, 30, 43–47. [Google Scholar] [CrossRef] [PubMed]
- Ouwehand, A.C.; Isolauri, E.; He, F.; Hashimoto, H.; Benno, Y.; Salminen, S. Differences in Bifidobacterium flora composition in allergic and healthy infants. J. Allergy Clin. Immunol. 2001, 108, 144–145. [Google Scholar] [CrossRef]
- Suzuki, S.; Shimojo, N.; Tajiri, Y.; Kumemura, M.; Kohno, Y. Differences in the composition of intestinal Bifidobacterium species and the development of allergic diseases in infants in rural Japan. Clin. Exp. Allergy 2007, 37, 506–511. [Google Scholar] [CrossRef]
- Francino, M.P. Early development of the gut microbiota and immune health. Pathogens (Basel Switzerland) 2014, 3, 769–790. [Google Scholar] [CrossRef]
- Belkaid, Y.; Hand, T.W. Role of the microbiota in immunity and inflammation. Cell 2014, 157, 121–141. [Google Scholar] [CrossRef]
- Platt, A.M.; Mowat, A.M. Mucosal macrophages and the regulation of immune responses in the intestine. Immunol. Lett. 2008, 119, 22–31. [Google Scholar] [CrossRef]
- Romagnani, S. Regulation of the T cell response. Clin. Exp. Allergy 2006, 36, 1357–1366. [Google Scholar] [CrossRef]
- Groux, H.; O’Garra, A.; Bigler, M.; Rouleau, M.; Antonenko, S.; de Vries, J.E.; Roncarolo, M.G. A CD4+ T-cell subset inhibits antigen-specific T-cell responses and prevents colitis. Nature 1997, 389, 737–742. [Google Scholar] [CrossRef] [PubMed]
- Akdis, M. Healthy immune response to allergens: T regulatory cells and more. Curr. Opin. Immunol. 2006, 18, 738–744. [Google Scholar] [CrossRef] [PubMed]
- Geuking, M.B.; Cahenzli, J.; Lawson, M.A.; Ng, D.C.; Slack, E.; Hapfelmeier, S.; McCoy, K.D.; Macpherson, A.J. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 2011, 34, 794–806. [Google Scholar] [CrossRef]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Josefowicz, S.Z.; Niec, R.E.; Kim, H.Y.; Treuting, P.; Chinen, T.; Zheng, Y.; Umetsu, D.T.; Rudensky, A.Y. Extrathymically generated regulatory T cells control mucosal TH2 inflammation. Nature 2012, 482, 395–399. [Google Scholar] [CrossRef] [PubMed]
- Round, J.L.; Mazmanian, S.K. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc. Natl. Acad. Sci. USA 2010, 107, 12204–12209. [Google Scholar] [CrossRef]
- Salloum, N.; Hussein, H.M.; Jammaz, R.; Jiche, S.; Uthman, I.W.; Abdelnoor, A.M.; Rahal, E.A. Epstein-Barr virus DNA modulates regulatory T-cell programming in addition to enhancing interleukin-17A production via Toll-like receptor 9. PLoS ONE 2018, 13, e0200546. [Google Scholar] [CrossRef]
- Kim, H.J.; Kim, H.Y.; Lee, S.Y.; Seo, J.H.; Lee, E.; Hong, S.J. Clinical efficacy and mechanism of probiotics in allergic diseases. Korean J. Pediatr. 2013, 56, 369–376. [Google Scholar] [CrossRef]
- Seite, S.; Bieber, T. Barrier function and microbiotic dysbiosis in atopic dermatitis. Clin. Cosmet. Investig. Dermatol. 2015, 8, 479–483. [Google Scholar] [CrossRef]
- Johnson, C.C.; Ownby, D.R. The infant gut bacterial microbiota and risk of pediatric asthma and allergic diseases. Trans. Res. 2017, 179, 60–70. [Google Scholar] [CrossRef]
- Salem, I.; Ramser, A.; Isham, N.; Ghannoum, M.A. The Gut Microbiome as a Major Regulator of the Gut-Skin Axis. Front. Microbiol. 2018, 9, 1459. [Google Scholar] [CrossRef]
- Samuelson, D.R.; Welsh, D.A.; Shellito, J.E. Regulation of lung immunity and host defense by the intestinal microbiota. Front. Microbiol. 2015, 6, 1085. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, A.; Bruhs, A.; Schwarz, T. The Short-Chain Fatty Acid Sodium Butyrate Functions as a Regulator of the Skin Immune System. J. Investig. Dermatol. 2017, 137, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Penders, J.; Gerhold, K.; Stobberingh, E.E.; Thijs, C.; Zimmermann, K.; Lau, S.; Hamelmann, E. Establishment of the intestinal microbiota and its role for atopic dermatitis in early childhood. J. Allergy Clin. Immunol. 2013, 132, 601–607.e608. [Google Scholar] [CrossRef] [PubMed]
- Hill, C.; Guarner, F.; Reid, G.; Gibson, G.R.; Merenstein, D.J.; Pot, B.; Morelli, L.; Canani, R.B.; Flint, H.J.; Salminen, S.; et al. Expert consensus document. The International Scientific Association for Probiotics and Prebiotics consensus statement on the scope and appropriate use of the term probiotic. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 506–514. [Google Scholar] [CrossRef] [PubMed]
- Hutkins, R.W.; Krumbeck, J.A.; Bindels, L.B.; Cani, P.D.; Fahey, G., Jr.; Goh, Y.J.; Hamaker, B.; Martens, E.C.; Mills, D.A.; Rastal, R.A.; et al. Prebiotics: why definitions matter. Curr. Opin. Biotechnol. 2016, 37, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ceapa, C.; Wopereis, H.; Rezaiki, L.; Kleerebezem, M.; Knol, J.; Oozeer, R. Influence of fermented milk products, prebiotics and probiotics on microbiota composition and health. Best Pract. Res. Clin. Gastroenterol. 2013, 27, 139–155. [Google Scholar] [CrossRef] [PubMed]
- Kwon, H.K.; Lee, C.G.; So, J.S.; Chae, C.S.; Hwang, J.S.; Sahoo, A.; Nam, J.H.; Rhee, J.H.; Hwang, K.C.; Im, S.H. Generation of regulatory dendritic cells and CD4+Foxp3+ T cells by probiotics administration suppresses immune disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 2159–2164. [Google Scholar] [CrossRef] [PubMed]
- Sawada, J.; Morita, H.; Tanaka, A.; Salminen, S.; He, F.; Matsuda, H. Ingestion of heat-treated Lactobacillus rhamnosus GG prevents development of atopic dermatitis in NC/Nga mice. Clin. Exp. Allergy 2007, 37, 296–303. [Google Scholar] [CrossRef] [PubMed]
- Shida, K.; Takahashi, R.; Iwadate, E.; Takamizawa, K.; Yasui, H.; Sato, T.; Habu, S.; Hachimura, S.; Kaminogawa, S. Lactobacillus casei strain Shirota suppresses serum immunoglobulin E and immunoglobulin G1 responses and systemic anaphylaxis in a food allergy model. Clin. Exp. Allergy 2002, 32, 563–570. [Google Scholar] [CrossRef]
- Gueniche, A.; Benyacoub, J.; Philippe, D.; Bastien, P.; Kusy, N.; Breton, L.; Blum, S.; Castiel-Higounenc, I. Lactobacillus paracasei CNCM I-2116 (ST11) inhibits substance P-induced skin inflammation and accelerates skin barrier function recovery in vitro. EJD 2010, 20, 731–737. [Google Scholar] [CrossRef]
- Matsumoto, M.; Ebata, T.; Hirooka, J.; Hosoya, R.; Inoue, N.; Itami, S.; Tsuji, K.; Yaginuma, T.; Muramatsu, K.; Nakamura, A.; et al. Antipruritic effects of the probiotic strain LKM512 in adults with atopic dermatitis. Ann. Allergy Asthma Immunol. 2014, 113, 209–216.e207. [Google Scholar] [CrossRef] [PubMed]
- Nagpal, R.; Wang, S.; Ahmadi, S.; Hayes, J.; Gagliano, J.; Subashchandrabose, S.; Kitzman, D.W.; Becton, T.; Read, R.; Yadav, H. Human-origin probiotic cocktail increases short-chain fatty acid production via modulation of mice and human gut microbiome. Sci. Rep. 2018, 8, 12649. [Google Scholar] [CrossRef] [PubMed]
- Kalliomaki, M.; Salminen, S.; Arvilommi, H.; Kero, P.; Koskinen, P.; Isolauri, E. Probiotics in primary prevention of atopic disease: a randomised placebo-controlled trial. Lancet (London England) 2001, 357, 1076–1079. [Google Scholar] [CrossRef]
- Rautava, S.; Kainonen, E.; Salminen, S.; Isolauri, E. Maternal probiotic supplementation during pregnancy and breast-feeding reduces the risk of eczema in the infant. J. Allergy Clin. Immunol. 2012, 130, 1355–1360. [Google Scholar] [CrossRef]
- Makrgeorgou, A.; Leonardi-Bee, J.; Bath-Hextall, F.J.; Murrell, D.F.; Tang, M.L.; Roberts, A.; Boyle, R.J. Probiotics for treating eczema. Cochrane Database Syst. Rev. 2018, 11, Cd006135. [Google Scholar] [CrossRef]
- Gueniche, A.; Knaudt, B.; Schuck, E.; Volz, T.; Bastien, P.; Martin, R.; Rocken, M.; Breton, L.; Biedermann, T. Effects of nonpathogenic Gram-negative bacterium Vitreoscilla filiformis lysate on atopic dermatitis: a prospective, randomized, double-blind, placebo-controlled clinical study. Br. J. Dermatol. 2008, 159, 1357–1363. [Google Scholar] [CrossRef]
- Paharik, A.E.; Parlet, C.P.; Chung, N.; Todd, D.A.; Rodriguez, E.I.; Van Dyke, M.J.; Cech, N.B.; Horswill, A.R. Coagulase-Negative Staphylococcal Strain Prevents Staphylococcus aureus Colonization and Skin Infection by Blocking Quorum Sensing. Cell Host Microbe 2017, 22, 746–756. [Google Scholar] [CrossRef]
- Myles, I.A.; Earland, N.J.; Anderson, E.D.; Moore, I.N.; Kieh, M.D.; Williams, K.W.; Saleem, A.; Fontecilla, N.M.; Welch, P.A.; Darnell, D.A.; et al. First-in-human topical microbiome transplantation with Roseomonas mucosa for atopic dermatitis. JCI Insight 2018, 3. [Google Scholar] [CrossRef] [PubMed]
- Gueniche, A.; Philippe, D.; Bastien, P.; Reuteler, G.; Blum, S.; Castiel-Higounenc, I.; Breton, L.; Benyacoub, J. Randomised double-blind placebo-controlled study of the effect of Lactobacillus paracasei NCC 2461 on skin reactivity. Benef. Microbes 2014, 5, 137–145. [Google Scholar] [CrossRef]
- Mutgi, K.; Koo, J. Update on the role of systemic vitamin D in atopic dermatitis. Pediatr. Dermatol. 2013, 30, 303–307. [Google Scholar] [CrossRef] [PubMed]
- Heimbeck, I.; Wjst, M.; Apfelbacher, C.J. Low vitamin D serum level is inversely associated with eczema in children and adolescents in Germany. Allergy 2013, 68, 906–910. [Google Scholar] [CrossRef]
- Del Pinto, R.; Ferri, C.; Cominelli, F. Vitamin D Axis in Inflammatory Bowel Diseases: Role, Current Uses and Future Perspectives. Int. J. Mol. Sci. 2017, 18, 2360. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.T.; Stenger, S.; Li, H.; Wenzel, L.; Tan, B.H.; Krutzik, S.R.; Ochoa, M.T.; Schauber, J.; Wu, K.; Meinken, C.; et al. Toll-like receptor triggering of a vitamin D-mediated human antimicrobial response. Science 2006, 311, 1770–1773. [Google Scholar] [CrossRef] [PubMed]
- Amon, U.; Baier, L.; Yaguboglu, R.; Ennis, M.; Holick, M.F.; Amon, J. Serum 25-hydroxyvitamin D levels in patients with skin diseases including psoriasis, infections, and atopic dermatitis. Dermato-endocrinology 2018, 10, e1442159. [Google Scholar] [CrossRef] [PubMed]
- Talsness, C.E.; Penders, J.; Jansen, E.; Damoiseaux, J.; Thijs, C.; Mommers, M. Influence of vitamin D on key bacterial taxa in infant microbiota in the KOALA Birth Cohort Study. PLoS ONE 2017, 12, e0188011. [Google Scholar] [CrossRef]
- Salim, S.Y.; Kaplan, G.G.; Madsen, K.L. Air pollution effects on the gut microbiota: a link between exposure and inflammatory disease. Gut Microbes 2014, 5, 215–219. [Google Scholar] [CrossRef]
- Valles, Y.; Francino, M.P. Air Pollution, Early Life Microbiome, and Development. Curr. Environ. Health Rep. 2018, 5, 512–521. [Google Scholar] [CrossRef] [PubMed]
- Hassoun, Y.; James, C.; Bernstein, D.I. The Effects of Air Pollution on the Development of Atopic Disease. Clin. Rev. Allergy Immunol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.L.; Su, H.J.; Sheu, H.M.; Yu, H.S.; Guo, Y.L. Traffic-related air pollution, climate, and prevalence of eczema in Taiwanese school children. J. Investig. Dermatol. 2008, 128, 2412–2420. [Google Scholar] [CrossRef]
- Penard-Morand, C.; Raherison, C.; Charpin, D.; Kopferschmitt, C.; Lavaud, F.; Caillaud, D.; Annesi-Maesano, I. Long-term exposure to close-proximity air pollution and asthma and allergies in urban children. Eur. Respir. J. 2010, 36, 33–40. [Google Scholar] [CrossRef]
- Mutlu, E.A.; Engen, P.A.; Soberanes, S.; Urich, D.; Forsyth, C.B.; Nigdelioglu, R.; Chiarella, S.E.; Radigan, K.A.; Gonzalez, A.; Jakate, S.; et al. Particulate matter air pollution causes oxidant-mediated increase in gut permeability in mice. Part. Fibre Toxicol. 2011, 8, 19. [Google Scholar] [CrossRef] [PubMed]
- Nicholson, J.K.; Holmes, E.; Kinross, J.; Burcelin, R.; Gibson, G.; Jia, W.; Pettersson, S. Host-gut microbiota metabolic interactions. Science 2012, 336, 1262–1267. [Google Scholar] [CrossRef] [PubMed]
| Gut | Skin | |
|---|---|---|
| Density | 1012/g of intestinal matter | 106/cm2 |
| Diversity | Bacteria dominant - 7–8 Phyla of bacteria (~100 species/individual) Fungi and virus rare | Bacteria dominant - 7–8 Phyla of bacteria (~40 species/individual) Up to 10% fungi and 40% viral/phage colonization |
| Niche | Mucus Epithelial surfaces Crypts | Stratum corneum (surface) Appendages (e.g., Hair follicles, sebaceous glands) |
| Community establishment | Early life | Early life Puberty |
| Nutrients | Rich | Poor |
| Effect on the immune system | Control the development of gut-associated lymphoid structures Innate immunity activation Control the induction, function, homeostasis of the regulatory immune network Colonization resistance | Control of innate immunity - Produce AMPs (e.g., Cathelicidin, β-defensin) - Increase expression of the complement system and IL-1 Control of adaptive immunity - Increase IL-17A and IFN-γ production by dermal T cells - Control the regulatory immune network Colonization resistance - Bacteriocin, serine protease Esp, and phenol-soluble modulin (PSMs) production by S. epidermidis - Short-chain fatty acid and porphyrin production by Cutibacterium |
| Range of effect | Local Systemic | Local Systemic (possibly) |
| AD Skin (Non-Affected Areas) | AD Skin (Lesions) |
|---|---|
| Actinobacteria (phylum) Corynebacterium (genus) Cutibacterium (genus) Rothia (genus) Actinomyces (genus) | Decreased relative abundance |
| Bacteroides (phylum) Prvotella (genus) | Decreased relative abundance |
| Proteobacteria (phylum) Acinetobacter (genus) | Decreased relative abundance |
| Firmicutes (phylum) Streptococcus (genus) Staphylococcus (genus) Granulicatella (genus) | Decreased relative abundance of Streptococcus/Granulicatella Increased absolute and relative abundance of Staphylococcus |
| Virulence Factors | Mechanisms for Increased AD Severity |
|---|---|
| α-toxin | Directly forms pores in keratinocytes, eroding the integrity of the epidermal barrier |
| Protease | Facilitate dissolution of the stratum corneum |
| Staphylococcal superantigens (SEA, SEB, SEC, TSST-1) | Trigger B cell expansion and cytokine release from keratinocytes Non-specific APC-mediated T cell activation |
| Protein A | Triggers inflammatory response from keratinocytes through the tumor necrosis factor receptor 1 (TNFR1) |
| PSM α | Stimulates keratinocyte production of IL-36, and Th17 inflammation |
| PSM γ (δ-toxin) | Stimulates dermal mast cells and induces skin inflammation |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kim, J.E.; Kim, H.S. Microbiome of the Skin and Gut in Atopic Dermatitis (AD): Understanding the Pathophysiology and Finding Novel Management Strategies. J. Clin. Med. 2019, 8, 444. https://doi.org/10.3390/jcm8040444
Kim JE, Kim HS. Microbiome of the Skin and Gut in Atopic Dermatitis (AD): Understanding the Pathophysiology and Finding Novel Management Strategies. Journal of Clinical Medicine. 2019; 8(4):444. https://doi.org/10.3390/jcm8040444
Chicago/Turabian StyleKim, Jung Eun, and Hei Sung Kim. 2019. "Microbiome of the Skin and Gut in Atopic Dermatitis (AD): Understanding the Pathophysiology and Finding Novel Management Strategies" Journal of Clinical Medicine 8, no. 4: 444. https://doi.org/10.3390/jcm8040444
APA StyleKim, J. E., & Kim, H. S. (2019). Microbiome of the Skin and Gut in Atopic Dermatitis (AD): Understanding the Pathophysiology and Finding Novel Management Strategies. Journal of Clinical Medicine, 8(4), 444. https://doi.org/10.3390/jcm8040444