Advances in the Relationship between Respiratory Viruses and Asthma
Abstract
:1. Viral Infections as a Risk Factor for Asthma
2. Genetic Factors, Viral Infections, and Asthma
3. Respiratory Viruses
3.1. Picornaviridae Family
3.2. Pneumoviridae Family
3.3. Metapneumovirus:
3.4. Orthopneumovirus:
3.5. Orthomyxoviridae Family
3.6. Paramyxoviridae Family
3.7. Adenoviridae Family
3.8. Coronaviridae Family
4. Pathogenic Mechanisms between the Relation of Infections by Respiratory Viruses and Asthma
4.1. Respiratory Syncytial Virus
4.2. Rhinovirus
4.3. Metapneumovirus
4.4. Bocavirus
4.5. Influenza
4.6. Coronavirus
5. Viral Infections and Asthma Exacerbations
6. Current Therapeutics for Viral Infection in Asthma
6.1. Inhaled Corticosteroids (ICS)
6.2. Omalizumab
6.3. Mepolizumab
6.4. Dupilumab
6.5. Benralizumab
6.6. Anti-Thymic Stromal Lymphopoietin mAb (Tezepelumab)
7. Directions for Future Research
8. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| RV | Rhinovirus |
| RSV | Respiratory syncytial virus |
| Th2 | T helper type 2 |
| ILC2 | Innate lymphoid cells type 2 |
| IgE | Immunoglobulin E |
| TSLP | Thymic stromal lymphopoietin |
| NOD2 | Nucleotide-binding oligomerization domain containing 2 |
| TLR | Toll-like receptor |
| TNF-α | Tumor necrosis factor-alpha |
| IFN | Interferon |
| hMPV | Human metapneumovirus |
References
- Global Initiative for Asthma, Global Strategy for Asthma Management and Prevention. 2023. Available online: http://www.ginasthma.org (accessed on 3 April 2023).
- Asher, M.I.; Rutter, C.E.; Bissell, K.; Chiang, C.-Y.; El Sony, A.; Ellwood, E.; Ellwood, P.; García-Marcos, L.; Marks, G.B.; Morales, E.; et al. Worldwide trends in the burden of asthma symptoms in school-aged children: Global Asthma Network Phase I cross-sectional study. Lancet 2021, 398, 1569–1580. [Google Scholar] [CrossRef] [PubMed]
- Kuruvilla, M.E.; Lee, F.E.-H.; Lee, G.B. Understanding Asthma Phenotypes, Endotypes, and Mechanisms of Disease. Clin. Rev. Allergy Immunol. 2019, 56, 219–233. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.R. Viral infection and airway epithelial immunity in asthma. Int. J. Mol. Sci. 2022, 23, 9914. [Google Scholar] [CrossRef] [PubMed]
- Jartti, T.; Gern, J.E. Role of viral infections in the development and exacerbation of asthma in children. J. Allergy Clin. Immunol. 2017, 140, 895–906. [Google Scholar] [CrossRef]
- Priante, E.; Cavicchiolo, M.E.; Baraldi, E. RSV infection and respiratory sequelae. Minerva Pediatr. 2018, 70, 623–633. [Google Scholar] [CrossRef] [PubMed]
- Alsayed, A.R.; Abed, A.; Abu-Samak, M.; Alshammari, F.; Alshammari, B. Etiologies of Acute Bronchiolitis in Children at Risk for Asthma, with Emphasis on the Human Rhinovirus Genotyping Protocol. J. Clin. Med. 2023, 12, 3909. [Google Scholar] [CrossRef]
- Holt, P.G.; Mok, D.; Panda, D.; Renn, L.; Fabozzi, G.; deKlerk, N.H.; Kusel, M.M.H.; Serralha, M.; Hollams, E.M.; Holt, B.J.; et al. Developmental regulation of type 1 and type 3 interferon production and risk for infant infections and asthma development. J. Allergy Clin. Immunol. 2019, 143, 1176–1182.e5. [Google Scholar] [CrossRef]
- Makrinioti, H.; Hasegawa, K.; Lakoumentas, J.; Xepapadaki, P.; Tsolia, M.; Castro-Rodriguez, J.A.; Feleszko, W.; Jartti, T.; Johnston, S.L.; Bush, A.; et al. The role of respiratory syncytial virus- and rhinovirus-induced bronchiolitis in recurrent wheeze and asthma—A systematic review and meta-analysis. Pediatr. Allergy Immunol. 2022, 33, e13741. [Google Scholar] [CrossRef]
- García-García, M.; Calvo, C.; Casas, I.; Bracamonte, T.; Rellán, A.; Gozalo, F.; Tenorio, T.; Pérez-Breña, P. Human metapneumovirus bronchiolitis in infancy is an important risk factor for asthma at age 5. Pediatr. Pulmonol. 2007, 42, 458–464. [Google Scholar] [CrossRef]
- Soto, J.A.; Gálvez, N.M.; Benavente, F.M.; Pizarro-Ortega, M.S.; Lay, M.K.; Riedel, C.; Bueno, S.M.; Gonzalez, P.A.; Kalergis, A.M. Human metapneumovirus: Mechanisms and molecular targets used by the virus to avoid the inmune system. Front. Immunol. 2018, 9, 2466. [Google Scholar] [CrossRef]
- Jartti, T.; Bønnelykke, K.; Elenius, V.; Feleszko, W. Role of viruses in asthma. Semin. Immunopathol. 2020, 42, 61–74. [Google Scholar] [CrossRef] [PubMed]
- Pividori, M.; Schoettler, N.; Nicolae, D.L.; Ober, C.; Im, H.K. Shared and distinct genetic risk factors for childhood-onset and adult-onset asthma: Genome-wide and transcriptome-wide studies. Lancet Respir. Med. 2019, 7, 509–522. [Google Scholar] [CrossRef] [PubMed]
- Anderson, J.; Do, L.A.H.; Wurzel, D.; Toh, Z.Q.; Mulholland, K.; Pellicci, D.G.; Licciardi, P.V. Severe respiratory syncytial virus disease in preterm infants: A case of innate immaturity. Thorax 2021, 76, 942–950. [Google Scholar] [CrossRef] [PubMed]
- Reese, S.E.; Xu, C.-J.; Dekker, H.T.D.; Lee, M.K.; Sikdar, S.; Ruiz-Arenas, C.; Merid, S.K.; Rezwan, F.I.; Page, C.M.; Ullemar, V.; et al. Epigenome-wide meta-analysis of DNA methylation and childhood asthma. J. Allergy Clin. Immunol. 2019, 143, 2062–2074. [Google Scholar] [CrossRef]
- Stikker, B.S.; Hendriks, R.W.; Stadhouders, R. Decoding the genetic and epigenetic basis of asthma. Allergy 2023, 78, 940–956. [Google Scholar] [CrossRef]
- Bonnelykke, K.; Coleman, A.T.; Evans, M.D.; Thorsen, J.; Waage, J.; Vissing, N.H.; Carlsson, C.J.; Stokholm, J.; Chawes, B.L.; Jessen, L.E.; et al. Cadherin-related family member 3 genetics and rhinovirus C respiratory illnesses. Am. J. Respir. Crit. Care Med. 2018, 197, 589–594. [Google Scholar] [CrossRef]
- Bønnelykke, K.; Sleiman, P.; Nielsen, K.; Kreiner-Møller, E.; Mercader, J.M.; Belgrave, D.; Dekker, H.T.D.; Husby, A.; Sevelsted, A.; Faura-Tellez, G.; et al. A genome-wide association study identifies CDHR3 as a susceptibility locus for early childhood asthma with severe exacerbations. Nat. Genet. 2014, 46, 51–55. [Google Scholar] [CrossRef]
- Ortega, H.; Nickle, D.; Carter, L. Rhinovirus and asthma: Challenges and opportunities. Rev. Med. Virol. 2021, 31, e2193. [Google Scholar] [CrossRef]
- Esteban, I.; Stein, R.T.; Polack, F.P. A durable relationship: Respiratory syncytial virus bronchiolitis and asthma past their golden anniversary. Vaccines 2020, 8, 201. [Google Scholar] [CrossRef]
- Zhou, Y.; Tong, L.; Li, M.; Wang, Y.; Li, L.; Yang, D.; Zhang, Y.; Chen, Z. Recurrent Wheezing and Asthma after Respiratory Syncytial Virus Bronchiolitis. Front. Pediatr. 2021, 9, 649003. [Google Scholar] [CrossRef]
- Wang, X.; Li, Y.; Nair, H.; Campbell, H.; Reeves, R.M.; van Wijhe, M.; Fischer, T.K.; Simonsen, L.; Trebbien, R.; Tong, S.; et al. Time-Varying Association Between Severe Respiratory Syncytial Virus Infections and Subsequent Severe Asthma and Wheeze and Influences of Age at the Infection. J. Infect. Dis. 2022, 226 (Suppl. 1), S38–S44. [Google Scholar] [CrossRef] [PubMed]
- Díaz-Chiguer, D.L.; Tirado-Mendoza, R.; Márquez-Navarro, A.; Ambrosio-Hernández, J.R.; Ruiz-Fraga, I.; Aguilar-Vargas, R.E.; Lira-Martínez, J.M.; López-Valdés, J.C. Detección y caracterización molecular de virus respiratorios causantes de infección respiratoria aguda en población adulta. Gac. Medica Mex. 2019, 155, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Pío, A.; Leowski, J.; Luelmo, F. Programa de la Organización Mundial de la Salud de infecciones respiratorias agudas en la infancia. Boletín Oficina Sanit. Panam. (OSP) 1984, 96, 283–293. Available online: https://iris.paho.org/handle/10665.2/17017 (accessed on 19 February 2023).
- Wylie, K.M. The Virome of the Human Respiratory Tract. Clin. Chest Med. 2017, 38, 11–19. [Google Scholar] [CrossRef] [PubMed]
- Willner, D.; Haynes, M.R.; Furlan, M.; Hanson, N.; Kirby, B.; Lim, Y.W.; Rainey, P.B.; Schmieder, R.; Youle, M.; Conrad, D.; et al. Case Studies of the Spatial Heterogeneity of DNA Viruses in the Cystic Fibrosis Lung. Am. J. Respir. Cell Mol. Biol. 2012, 46, 127–131. [Google Scholar] [CrossRef] [PubMed]
- Shao, L.; Shen, W.; Wang, S.; Qiu, J. Recent Advances in Molecular Biology of Human Bocavirus 1 and Its Applications. Front. Microbiol 2021, 12, 696604. [Google Scholar] [CrossRef]
- Cox, R.G.; Mainou, B.A.; Johnson, M.; Hastings, A.K.; Schuster, J.E.; Dermody, T.S.; Williams, J.V. Human metapneumovirus is capable of entering cells by fusion with endosomal membranes. PLoS Pathog. 2015, 11, e1005303. [Google Scholar] [CrossRef]
- Yuan, S.; Balaji, S.; Lomakin, I.V.; Xiong, Y. Coronavirus Nsp1: Immune Response Supression and Protein Expression Inhibition. Front Microbiol 2021, 12, 752214. [Google Scholar] [CrossRef]
- Whitney, S.; Manal, O. StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Zell, R. Picornaviridae—The ever-growing virus family. Arch. Virol. 2018, 163, 299–317. [Google Scholar] [CrossRef]
- Ambros, V.; Baltimore, D. Protein is linked to the 5′ end of poliovirus RNA by a phosphodiester linkage to tyrosine. J. Biol. Chem. 1978, 253, 5263–5266. [Google Scholar] [CrossRef]
- Zell, R.; Knowles, N.J.; Simmonds, P. A proposed division of the family Picornaviridae into subfamilies based on phylogenetic relationships and functional genomic organization. Arch. Virol. 2021, 166, 2927–2935. [Google Scholar] [CrossRef] [PubMed]
- Bakker, S.E.; Duquerroy, S.; Galloux, M.; Loney, C.; Conner, E.; Eléouët, J.-F.; Rey, F.A.; Bhella, D. The respiratory syncytial virus nucleoprotein–RNA complex forms a left-handed helical nucleocapsid. J. Gen. Virol. 2013, 94 Pt 8, 1734–1738. [Google Scholar] [CrossRef] [PubMed]
- Rima, B.; Collins, P.; Easton, A.; Fouchier, R.; Kurath, G.; Lamb, R.A.; Lee, B.; Maisner, A.; Rota, P.; Wang, L.; et al. ICTV Virus Taxonomy Profile: Pneumoviridae. J. Gen. Virol. 2017, 98, 2912–2913. [Google Scholar] [CrossRef] [PubMed]
- Hause, B.M.; Padmanabhan, A.; Pedersen, K.; Gidlewski, T. Feral swine virome is dominated by single-stranded DNA viruses and contains a novel Orthopneumovirus which circulates both in feral and domestic swine. J. Gen. Virol. 2016, 97, 2090–2095. [Google Scholar] [CrossRef]
- Hause, B.M.; Collin, E.A.; Liu, R.; Huang, B.; Sheng, Z.; Lu, W.; Wang, D.; Nelson, E.A.; Li, F. Characterization of a novel influenza virus in cattle and Swine: Proposal for a new genus in the Orthomyxoviridae family. mBio 2014, 5, e00031-14. [Google Scholar] [CrossRef]
- Lvov, D.K.; Shchelkanov, M.Y.; Alkhovsky, S.V.; Deryabin, P.G. Single-Stranded RNA Viruses. In Zoonotic Viruses in Northern Eurasia; Academic Press: Cambridge, MA, USA, 2015; pp. 135–392. [Google Scholar] [CrossRef]
- Salomon, R.; Webster, R.G. The influenza virus enigma. Cell 2009, 136, 402–410. [Google Scholar] [CrossRef]
- Rima, B.; Collins, P.; Easton, A.; Fouchier, R.; Kurath, G.; Lamb, R.A.; Lee, B.; Maisner, A.; Rota, P.; Wang, L.-F. Problems of classification in the family Paramyxoviridae. Arch. Virol. 2018, 163, 1395–1404. [Google Scholar] [CrossRef]
- Rima, B.; Balkema-Buschmann, A.; Dundon, W.G.; Duprex, P.; Easton, A.; Fouchier, R.; Kurath, G.; Lamb, R.; Lee, B.; Rota, P.; et al. ICTV Virus Taxonomy Profile: Paramyxoviridae. J. Gen. Virol. 2019, 100, 1593–1594. [Google Scholar] [CrossRef]
- Orqueda, A.S.; Lucion, M.F.; Juárez MD, V.; Barquez, R.; Stach, P.; Nievas, A.; Gentile, Á. Vigilancia de virus sincicial respiratorio e influenza en niños escolarizados asistidos en un hospital pediátrico durante 2 meses del segundo semestre de 2021. Arch. Argent. Pediatr. 2022, 120, 269–273. [Google Scholar] [CrossRef]
- Li, Y.; Reeves, R.M.; Wang, X.; Bassat, Q.; Brooks, W.A.; Cohen, C.; Moore, D.P.; Nunes, M.; Rath, B.; Campbell, H.; et al. Global patterns in monthly activity of influenza virus, respiratory syncytial virus, parainfluenza virus, and metapneumovirus: A systematic analysis. Lancet Glob. Health 2019, 7, e1031–e1045. [Google Scholar] [CrossRef]
- Kojaoghlanian, T.; Flomenberg, P.; Horwitz, M.S. The impact of adenovirus infection on the immunocompromised host. Rev. Med. Virol. 2003, 13, 155–171. [Google Scholar] [CrossRef] [PubMed]
- Lenaerts, L.; De Clercq, E.; Naesens, L. Clinical features and treatment of adenovirus infections. Rev. Med. Virol. 2008, 18, 357–374. [Google Scholar] [CrossRef] [PubMed]
- De Groot, R.J.; Baker, S.C.; Baric, R.S.; Brown, C.S.; Drosten, C.; Enjuanes, L.; Fouchier, R.A.M.; Galiano, M.; Gorbalenya, A.E.; Memish, Z.A.; et al. Middle East respiratory syndrome coronavirus (MERS-CoV): Announcement of the Coronavirus Study Group. J. Virol. 2013, 87, 7790–7792. [Google Scholar] [CrossRef] [PubMed]
- Bárcena, M.; Oostergetel, G.T.; Bartelink, W.; Faas, F.G.A.; Verkleij, A.; Rottier, P.J.M.; Koster, A.J.; Bosch, B.J. Cryo-electron tomography of mouse hepatitis virus: Insights into the structure of the coronavirion. Proc. Natl. Acad. Sci. USA 2009, 106, 582–587. [Google Scholar] [CrossRef]
- Novak, N.; Cabanillas, B. Viruses and asthma: The role of common respiratory viruses in asthma and its potential meaning for SARS-CoV-2. Immunology 2020, 161, 83–93. [Google Scholar] [CrossRef]
- Hayashi, Y.; Sada, M.; Shirai, T.; Okayama, K.; Kimura, R.; Kondo, M.; Okodo, M.; Tsugawa, T.; Ryo, A.; Kimura, H. Rhinovirus Infection and Virus-Induced Asthma. Viruses 2022, 14, 2616. [Google Scholar] [CrossRef]
- Kimura, H.; Yoshizumi, M.; Ishii, H.; Oishi, K.; Ryo, A. Cytokine production and signaling pathways in respiratory virus infection. Front. Microbiol. 2013, 4, 276. [Google Scholar] [CrossRef]
- Price, A.S.; Kennedy, J.L. T-helper 2 mechanisms involved in human rhinovirus infections and asthma. Ann. Allergy Asthma Immunol. 2022, 129, 681–691. [Google Scholar] [CrossRef]
- Whetstone, C.E.; Ranjbar, M.; Omer, H.; Cusack, R.P.; Gauvreau, G.M. The Role of Airway Epithelial Cell Alarmins in Asthma. Cells 2022, 11, 1105. [Google Scholar] [CrossRef]
- Kumar, K.; Singanayagam, A.; Johnston, S.L. Respiratory Virus Infections in Asthma: Research Developments and Therapeutic Advances. Acta Med. Acad. 2020, 49, 130–143. [Google Scholar] [CrossRef]
- Xiao, C.; Puddicombe, S.M.; Field, S.; Haywood, J.; Broughton-Head, V.; Puxeddu, I.; Haitchi, H.M.; Vernon-Wilson, E.; Sammut, D.; Bedke, N.; et al. Defective epithelial barrier function in asthma. J. Allergy Clin. Immunol. 2011, 128, 549–556.e12. [Google Scholar] [CrossRef] [PubMed]
- Veerati, P.C.; Troy, N.M.; Reid, A.T.; Li, N.F.; Nichol, K.S.; Kaur, P.; Maltby, S.; Wark, P.A.B.; Knight, D.A.; Bosco, A.; et al. Airway Epithelial Cell Immunity Is Delayed during Rhinovirus Infection in Asthma and COPD. Front. Immunol. 2020, 11, 974. [Google Scholar] [CrossRef] [PubMed]
- Iwata, A.; Toda, Y.; Furuya, H.; Nakajima, H. Group 2 innate lymphoid cells in human asthma. Allergol. Int. 2023, 72, 194–200. [Google Scholar] [CrossRef] [PubMed]
- Ricciardolo, F.L.M.; Sprio, A.E.; Baroso, A.; Gallo, F.; Riccardi, E.; Bertolini, F.; Carriero, V.; Arrigo, E.; Ciprandi, G. Characterization of T2-Low and T2-High Asthma Phenotypes in Real-Life. Biomedicines 2021, 9, 1684. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez-Rodriguez, N.; Gogoi, M.; McKenzie, A.N. Group 2 innate lymphoid cells: Team players in regulating asthma. Annu. Rev. Immunol. 2021, 39, 167–198. [Google Scholar] [CrossRef]
- Manti, S.; Piedimonte, G. An overview on the RSV-mediated mechanisms in the onset of non-allergic asthma. Front. Pediatr. 2022, 10, 998296. [Google Scholar] [CrossRef]
- Bozic, C.R.; Lu, B.; Höpken, U.E.; Gerard, C.; Gerard, N.P. Neurogenic amplification of immune complex inflammation. Science 1996, 273, 1722–1725. [Google Scholar] [CrossRef]
- Binns, E.; Tuckerman, J.; Licciardi, P.V.; Wurzel, D. Respiratory syncytial virus, recurrent wheeze and asthma: A narrative review of pathophysiology, prevention and future directions. J. Paediatr. Child Health 2022, 58, 1741–1746. [Google Scholar] [CrossRef]
- Liu, D.; Tang, Z.; Bajinka, O.; Dai, P.; Wu, G.; Qin, L.; Tan, Y. miR-34b/c-5p/CXCL10 Axis Induced by RSV Infection Mediates a Mechanism of Airway Hyperresponsive Diseases. Biology 2023, 12, 317. [Google Scholar] [CrossRef]
- Luz Garcia-Garcia, M.; Calvo Rey, C.; del Rosal Rabes, T. Asma y virus en el nino. Arch. Bronconeumol. 2016, 52, 269–273. [Google Scholar] [CrossRef]
- Basnet, S.; Palmenberg, A.C.; Gern, J.E. Rhinoviruses and Their Receptors. Chest 2019, 155, 1018–1025. [Google Scholar] [CrossRef] [PubMed]
- Yuan, X.-H.; Pang, L.-L.; Yang, J.; Jin, Y. Comparison of immune response to human rhinovirus C and respiratory syncytial virus in highly differentiated human airway epithelial cells. Virol. J. 2022, 19, 81. [Google Scholar] [CrossRef] [PubMed]
- Roh, D.E.; Park, S.-H.; Choi, H.J.; Kim, Y.H. Comparison of cytokine expression profiles in infants with a rhinovirus induced lower respiratory tract infection with or without wheezing: A comparison with respiratory syncytial virus. Korean J. Pediatr. 2017, 60, 296–301. [Google Scholar] [CrossRef]
- Nakagome, K.; Bochkov, Y.A.; Ashraf, S.; Brockman-Schneider, R.A.; Evans, M.D.; Pasic, T.R.; Gern, J.E. Effects of rhinovirus species on viral replication and cytokine production. J. Allergy Clin. Immunol. 2014, 134, 332–341.e10. [Google Scholar] [CrossRef] [PubMed]
- Uddin, S.; Thomas, M. Human Metapneumovirus. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Kolli, D.; Gupta, M.R.; Sbrana, E.; Velayutham, T.S.; Chao, H.; Casola, A.; Garofalo, R.P. Alveolar macrophages contribute to the pathogenesis of human metapneumovirus infection while protecting against respiratory syncytial virus infection. Am. J. Respir. Cell Mol. Biol. 2014, 51, 502–515. [Google Scholar] [CrossRef]
- Chauhan, A.; Singh, M.; Agarwal, A.; Paul, N. Correlation of TSLP, IL-33, and CD4 + CD25 + FOXP3 + T regulatory (Treg) in pediatric asthma. J. Asthma 2015, 52, 868–872. [Google Scholar] [CrossRef]
- Christensen, A.; Nordbø, S.A.; Krokstad, S.; Rognlien, A.G.W.; Døllner, H. Human bocavirus in children: Mono-detection, high viral load and viraemia are associated with respiratory tract infection. J. Clin. Virol. 2010, 49, 158–162. [Google Scholar] [CrossRef]
- Lukkarinen, H.; Söderlund-Venermo, M.; Vuorinen, T.; Allander, T.; Hedman, K.; Simell, O.; Ruuskanen, O.; Jartti, T. Human bocavirus 1 may suppress rhinovirus-associated immune response in wheezing children. J. Allergy Clin. Immunol. 2014, 133, 256–258.e4. [Google Scholar] [CrossRef]
- Coverstone, A.M.; Wang, L.; Sumino, K. Beyond Respiratory Syncytial Virus and Rhinovirus in the Pathogenesis and Exacerbation of Asthma. Immunol. Allergy Clin. N. Am. 2019, 39, 391–401. [Google Scholar] [CrossRef]
- Bhat, R.; Almajhdi, F.N. Induction of Immune Responses and Immune Evasion by Human Bocavirus. Int. Arch. Allergy Immunol. 2021, 182, 728–735. [Google Scholar] [CrossRef]
- Kusel, M.; Kebadze, T.; Johnston, S.; Holt, P.; Sly, P. Febrile respiratory illnesses in infancy and atopy are risk factors for persistent asthma and wheeze. Eur. Respir. J. 2012, 39, 876–882. [Google Scholar] [CrossRef] [PubMed]
- Lund, S.; Walford, H.H.; Doherty, T.A. Type 2 innate lymphoid cells in allergic disease. Curr. Immunol. Rev. 2013, 9, 214–221. [Google Scholar] [CrossRef] [PubMed]
- Ishikawa, H.; Sasaki, H.; Fukui, T.; Fujita, K.; Kutsukake, E.; Matsumoto, T. Mice with asthma are more resistant to influenza virus infection and NK cells activated by the induction of asthma have potentially protective effects. J. Clin. Immunol. 2012, 32, 256–267. [Google Scholar] [CrossRef]
- Fehr, A.R.; Perlman, S. Coronaviruses: An overview of their replication and pathogenesis. Methods Mol. Biol. 2015, 1282, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Diamond, M.S.; Kanneganti, T.-D. Innate immunity: The first line of defense against SARS-CoV-2. Nat. Immunol. 2022, 23, 165–176. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Hadjadj, J.; Yatim, N.; Barnabei, L.; Corneau, A.; Boussier, J.; Smith, N.; Péré, H.; Charbit, B.; Bondet, V.; Chenevier-Gobeaux, C.; et al. Impaired type I interferon activity and inflammatory responses in severe COVID-19 patients. Science 2020, 369, 718–724. [Google Scholar] [CrossRef]
- Hou, Y.J.; Okuda, K.; Edwards, C.E.; Martinez, D.R.; Asakura, T.; Dinnon, K.H., III; Kato, T.; Lee, R.E.; Yount, B.L.; Mascenik, T.M.; et al. SARS-CoV-2 reverse genetics reveals a variable infection gradient in the respiratory tract. Cell 2020, 182, 429–446. [Google Scholar] [CrossRef]
- Jackson, D.J.; Busse, W.W.; Bacharier, L.B.; Kattan, M.; O’Connor, G.T.; Wood, R.A.; Visness, C.M.; Durham, S.R.; Larson, D.; Esnault, S.; et al. Association of respiratory allergy, asthma, and expression of the SARS-CoV-2 receptor ACE2. J. Allergy Clin. Immunol. 2020, 146, 203–206.e3. [Google Scholar] [CrossRef]
- Jayavelu, N.D.; Altman, M.C.; Benson, B.; Dufort, M.J.; Vanderwall, E.R.; Rich, L.M.; White, M.P.; Becker, P.M.; Togias, A.; Jackson, D.J.; et al. Type 2 inflammation reduces SARS-CoV-2 replication in the airway epithelium in allergic asthma through functional alteration of ciliated epithelial cells. J. Allergy Clin. Immunol. 2023, 152, 56–67. [Google Scholar] [CrossRef]
- Kimura, H.; Francisco, D.; Conway, M.; Martinez, F.D.; Vercelli, D.; Polverino, F.; Billheimer, D.; Kraft, M. Type 2 inflammation modulates ACE2 and TMPRSS2 in airway epithelial cells. J. Allergy Clin. Immunol. 2020, 146, 80–88.e8. [Google Scholar] [CrossRef] [PubMed]
- Skevaki, C.; Karsonova, A.; Karaulov, A.; Fomina, D.; Xie, M.; Chinthrajah, S.; Nadeau, K.C.; Renz, H. SARS-CoV-2 infection and COVID-19 in asthmatics: A complex relationship. Nat. Rev. Immunol. 2021, 21, 202–203. [Google Scholar] [CrossRef]
- Morais-Almeida, M.; Bousquet, J. COVID-19 and asthma: To have or not to have T2 inflammation makes a difference? Pulmonology 2020, 26, 261–263. [Google Scholar] [CrossRef] [PubMed]
- Di Vincenzo, S.; Ferrante, G.; Ferraro, M.; Cascio, C.; Malizia, V.; Licari, A.; La Grutta, S.; Pace, E. Oxidative Stress, Environmental Pollution, and Lifestyle as Determinants of Asthma in Children. Biology 2023, 12, 133. [Google Scholar] [CrossRef] [PubMed]
- Nakagome, K.; Nagata, M. Innate Immune Responses by Respiratory Viruses, Including Rhinovirus, during Asthma Exacerbation. Front. Immunol. 2022, 13, 865973. [Google Scholar] [CrossRef]
- Radzikowska, U.; Eljaszewicz, A.; Tan, G.; Stocker, N.; Heider, A.; Westermann, P.; Steiner, S.; Dreher, A.; Wawrzyniak, P.; Rückert, B.; et al. Rhinovirus-induced epithelial RIG-I inflammasome suppresses antiviral immunity and promotes inflammation in asthma and COVID-19. Nat. Commun. 2023, 14, 2329. [Google Scholar] [CrossRef]
- Dinwiddie, D.L.; Kaukis, N.; Pham, S.; Hardin, O.; Stoner, A.N.; Kincaid, J.C.; Caid, K.; Kirkpatrick, C.; Pomeroy, K.; Putt, C.; et al. Viral infection and allergy status impact severity of asthma symptoms in children with asthma exacerbations. Ann. Allergy Asthma Immunol. 2022, 129, 319–326.e3. [Google Scholar] [CrossRef]
- Bagnasco, D.; Testino, E.; Nicola, S.; Melissari, L.; Russo, M.; Canevari, R.F.; Brussino, L.; Passalacqua, G. Specific Therapy for T2 Asthma. J. Pers. Med. 2022, 12, 593. [Google Scholar] [CrossRef]
- Ramphul, M. Increased inhaled corticosteroids for treating acute asthma exacerbations. Clin. Exp. Allergy 2023, 53, 388–391. [Google Scholar] [CrossRef]
- McDowell, P.J.; Busby, J.; Heaney, L.G. Asthma Exacerbations in Severe Asthma: Why Systemic Corticosteroids May not Always Be the Best Treatment Option. Curr. Treat. Options Allergy 2023, 10, 53–63. [Google Scholar] [CrossRef]
- Esquivel, A.; Busse, W.W.; Calatroni, A.; Togias, A.G.; Grindle, K.G.; Bochkov, Y.A.; Gruchalla, R.S.; Kattan, M.; Kercsmar, C.M.; Khurana Hershey, G.; et al. Effects of Omalizumab on Rhinovirus Infections, Illnesses, and Exacerbations of Asthma. Am. J. Respir. Crit. Care Med. 2017, 196, 985–992. [Google Scholar] [CrossRef]
- Teach, S.J.; Gill, M.A.; Togias, A.; Sorkness, C.A.; Arbes, S.J., Jr.; Calatroni, A.; Wildfire, J.J.; Gergen, P.J.; Cohen, R.; Pongracic, J.A.; et al. Preseasonal treatment with either omalizumab or an inhaled corticosteroid boost to prevent fall asthma exacerbations. J. Allergy Clin. Immunol. 2015, 136, 1476–1485. [Google Scholar] [CrossRef] [PubMed]
- Fu, Z.; Xu, Y.; Cai, C. Efficacy and safety of omalizumab in children with moderate-to-severe asthma: A meta-analysis. J. Asthma 2021, 58, 1350–1358. [Google Scholar] [CrossRef] [PubMed]
- Gupta, A.; Ikeda, M.; Geng, B.; Azmi, J.; Price, R.G.; Bradford, E.S.; Yancey, S.W.; Steinfeld, J. Long-term safety and pharmacodynamics of mepolizumab in children with severe asthma with an eosinophilic phenotype. J. Allergy Clin. Immunol. 2019, 144, 1336–1342.e7. [Google Scholar] [CrossRef]
- Lugogo, N.L.; Bogart, M.; Corbridge, T.; Packnett, E.R.; Wu, J.; Hahn, B. Impact of mepolizumab in patients with high-burden severe asthma within a managed care population. J. Asthma 2023, 60, 811–823. [Google Scholar] [CrossRef] [PubMed]
- Bacharier, L.B.; Jackson, D.J. Biologics in the treatment of asthma in children and adolescents. J. Allergy Clin. Immunol. 2023, 151, 581–589. [Google Scholar] [CrossRef]
- Zaazouee, M.S.; Alwarraqi, A.G.; Mohammed, Y.A.; Badheeb, M.A.; Farhat, A.M.; Eleyan, M.; Morad, A.; Zeid, M.A.-A.; Mohamed, A.S.; AbuEl-Enien, H.; et al. Dupilumab efficacy and safety in patients with moderate to severe asthma: A systematic review and meta-analysis. Front. Pharmacol. 2022, 13, 992731. [Google Scholar] [CrossRef]
- Laviolette, M.; Gossage, D.L.; Gauvreau, G.; Leigh, R.; Olivenstein, R.; Katial, R.; Busse, W.W.; Wenzel, S.; Wu, Y.; Datta, V.; et al. Effects of benralizumab on airway eosinophils in asthmatic patients with sputum eosinophilia. J. Allergy Clin. Immunol. 2013, 132, 1086–1096.e5. [Google Scholar] [CrossRef]
- Nakagome, K.; Nagata, M. Involvement and Possible Role of Eosinophils in Asthma Exacerbation. Front. Immunol. 2018, 9, 2220. [Google Scholar] [CrossRef]
- Hossain, F.M.A.; Choi, J.Y.; Uyangaa, E.; Park, S.O.; Eo, S.K. The Interplay between Host Immunity and Respiratory Viral Infection in Asthma Exacerbation. Immune Netw. 2019, 19, e31. [Google Scholar] [CrossRef]
- Radzikowska, U.; Baerenfaller, K.; Cornejo-Garcia, J.A.; Karaaslan, C.; Barletta, E.; Sarac, B.E.; Zhakparov, D.; Villaseñor, A.; Eguiluz-Gracia, I.; Mayorga, C.; et al. Omics technologies in allergy and asthma research: An EAACI position paper. Allergy 2022, 77, 2888–2908. [Google Scholar] [CrossRef] [PubMed]
- Kampmann, B.; Madhi, S.A.; Munjal, I.; Simões, E.A.; Pahud, B.A.; Llapur, C.; Baker, J.; Marc, G.P.; Radley, D.; Shittu, E.; et al. Bivalent Prefusion F Vaccine in Pregnancy to Prevent RSV Illness in Infants. N. Engl. J. Med. 2023, 388, 1451–1464. [Google Scholar] [CrossRef] [PubMed]
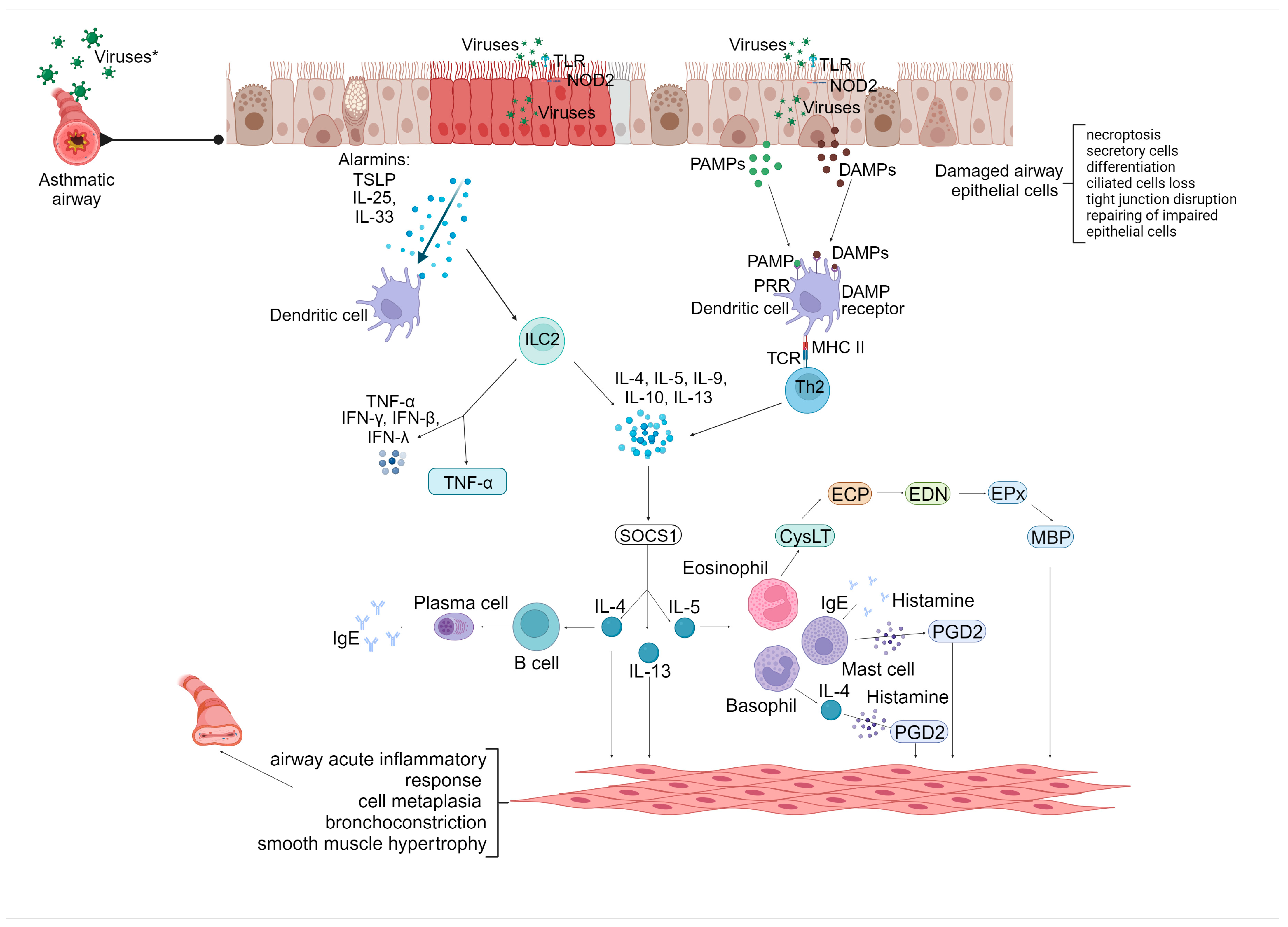
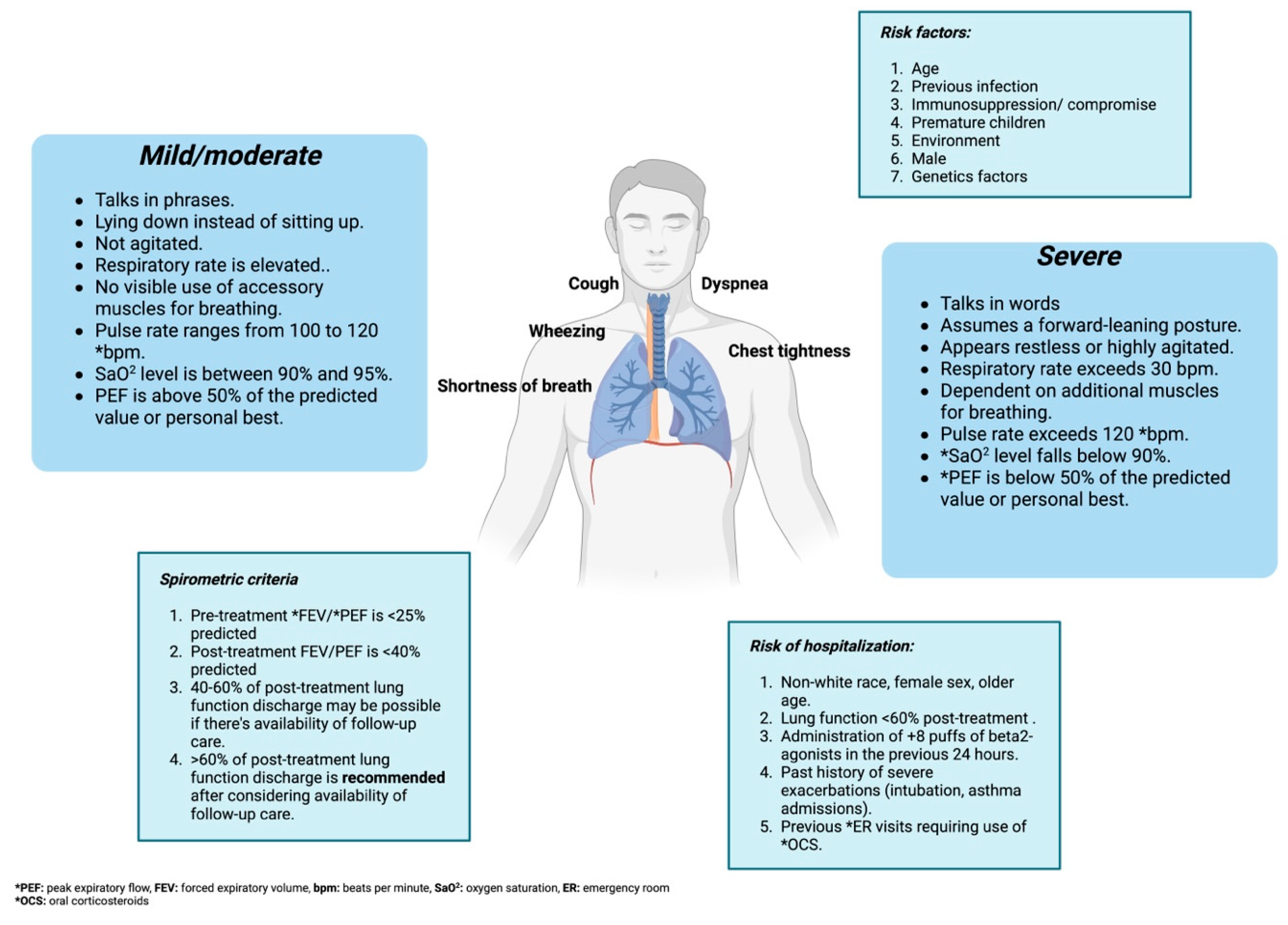

| Species | Family | Subfamily | Genus | Type | Subgroup | Gene | Molecular Characterization | Protein Expression–Receptor | Capsid |
|---|---|---|---|---|---|---|---|---|---|
| Rhinovirus | Picornaviridae | Enterovirus | RNA | HRV-A, HRV-B, HRV-C | IFIH1 | The surface of the rhinovirus has shaped-like canyons that provide an *attachment site for its receptors on the surface of susceptible target cells. * Attachment site: ICAM-1. | VP1—binding site function to the receptors of the surface. VP1, VP2, VP3—antigenic diversity. VP4—the ssRNA is anchored to its core capsid. Receptor * LDRL—attachment and internalization. * ICAM-1RV-A, RV-B * CDHR3-RV-C. | Its viral capsid encircles the RNA genome, composed of 60 identical copies of its structural proteins. [27]. | |
| Respiratory syncytial virus | Pneumoviridae | Pneumovirinae | Orthopneumovirus/Pneumovirus | RNA | A and B | M, M1, M2, M2-1 NS1, NS2 P | RSV virions consist of the structure of a nucleocapsid packaged in a lipid envelope matter that is derived from the host cell plasma membrane. These spherical particles are 100-350 nm in diameter. | F, G glycoproteins—viral neutralization antigens. Signals a cleaved. N—terminal pattern. F—viral penetration and syncytium emergence pathway. Receptor CX3CR1, nucleolin, EGFR, IFG1R, HSPGs. | Made from a polymerase protein, nucleoprotein, and a phosphoprotein. |
| Metapneumovirus | Pneumoviridae | Pneumovirinae | Metapneumovirus | RNA | A (A1, A2) and B (B1, B2) | N, P, M, F, M2, SH, G, L | The size of its virions can go from 150–600 nm with a spike-like envelope. Its *-ssRNA genome is suspected to be the starring cause of the main lower respiratory tract illness among children. | Glycoproteins. F—fusion protein. * SH protein—small hydrophobic protein. G—attachment protein. Receptor RGD-binding integrins. αvβ1 integrin. | The metapneumovirus forms a left-handed helical nucleocapsid to shelter the RNA genome as well as protect its RNA from nucleases. [28]. |
| Human bocavirus | Parvoviridae | Parvovirinae | Bocavirus | DNA | HBoV1, HBoV2, HBoV3, HBoV4 | RNA Pol III | Bocaparvoviruses have both a proximal and a distal *polyadenylation site within their promoters. This transcribes exclusively the (pre-) mRNA precursor. The NP1 expression in bocavirus has an open reading action that is in its genome center. *Polyadenylation sites: (pA)p and (pA)d. | Non-structural proteins NS1, NS1-70, NS2, NS3, NS4, and NP1. Structural proteins VP1, VP2, and VP3. | The HBoV capsid has a protective coat function for the genome, thus logging on to start the infection process. |
| Adenovirus | Adenoviridae | Mastadenovirus | DNA | HAdV A, HAdV B HAdV C HAdV D, HAdV E, HAdV F, HAdV G | E1, E3 dl309 (mutant) | There are 49 serological types of adenoviruses that have the capacity to infect humans. This commonly causes upper and lower (acute) respiratory infections. These infections can lead to a severe or fatal outcome in immunocompetent/immunocompromised patients. Different HAdVs groups own slight changes to their tropisms, correlating with other clinical manifestations. | Protein V—viral DNA to the cell nucleus pathway. Protein IX—transcriptional activator. p32K—unique structural protein. | Built out of an icosahedral capsid made up of 252 capsomeres. A double-stranded DNA virus of 70 to 90 in size. | |
| Coronavirus | Coronaviridae | Coronavirinae | Alphacoronavirus, Betacoronavirus, Gammacoronavirus, Deltacoronavirus | RNA | Betacoronavirus—A, B, C, D | ORF1ab N gene | This virus has a unique way of operating its genome expression. The replication merely stands on the frameshifting of its ribosome during the translation process of the genome and the progeny virions assembly. | * Nsp1 | The coronavirus forms helical capsids, which are resistant to RNase because of the binding properties of the N protein. [29]. |
| Parainfluenza 1,3 | Paramyxoviridae | Paramyxovirinae | Respirovirus | RNA | HPIV-4a, HPIV-4b | Parainfluenza virions have pleomorphic properties, in which the diameter ranges from 150 to 200 μm. They have been grouped into four serotypes, where they are seemingly divided into respirovirus, which includes HPIV-1 and HPIV-3, and rubulavirus, which HPIV-2 and HPIV-4 are a part. | N—Nucleocapsid protein. P—Phosphoprotein. F—Fusion glycoprotein. M—Matrix protein. HN—Hemagglutinin neuraminidase glycoprotein. L—RNA polymerase. Receptor α2-3-linked * SAs with sulfated sialyl-Lewis motif α2-8-linked * SAs. | The PIV5-N nucleocapsid ring encapsulates a nuclease-resistant 78-nt RNA strand. | |
| Parainfluenza 2,4 | Paramyxoviridae | Paramyxovirinae | Rubulavirus | ||||||
| Influenza | Orthomyxovirus | Orthomyxovirus | RNA | A, B, C | M gene | RNA genomes with a negative segmented RNA strand are one of the principles of the formation of the virion. It requires the possession of an RNA-dependent RNA polymerase native from a viral origin for replication. The drift and shift of the virus has antigenic properties that are being enabled by the genome structure of the influenza virus. [30]. | PB1, PB2, PA. Receptor. A2,3- and α2,6 receptors. | The capsid is surrounded by the M1 matrix protein made out of a lipid bilayer where hemagglutinin, neuraminidase, and M2 proteins are tailored. | |
| Enterovirus | Picornaviridae | Enterovirus | RNA | Coxsackievirus A 1-22, 24 Rhinovirus Poliovirus Echovirus 1–7, 9, 11–27, 29–34 | The non-enveloped virus, such as enterovirus, has approximately 7500 nucleotides, containing a single open reading frame that encodes polyproteins that are then processed to yield the structural proteins. | Precursor proteins (polyproteins). P1 - VP1, VP2, VP3—capsid surface. - VP4—within the capsid. P2, P3. | They have a non-enveloped icosahedral * +ssRNA capsid. Their replication takes place in the cytoplasm. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Romero-Tapia, S.d.J.; Guzmán Priego, C.G.; Del-Río-Navarro, B.E.; Sánchez-Solis, M. Advances in the Relationship between Respiratory Viruses and Asthma. J. Clin. Med. 2023, 12, 5501. https://doi.org/10.3390/jcm12175501
Romero-Tapia SdJ, Guzmán Priego CG, Del-Río-Navarro BE, Sánchez-Solis M. Advances in the Relationship between Respiratory Viruses and Asthma. Journal of Clinical Medicine. 2023; 12(17):5501. https://doi.org/10.3390/jcm12175501
Chicago/Turabian StyleRomero-Tapia, Sergio de Jesús, Crystell Guadalupe Guzmán Priego, Blanca E. Del-Río-Navarro, and Manuel Sánchez-Solis. 2023. "Advances in the Relationship between Respiratory Viruses and Asthma" Journal of Clinical Medicine 12, no. 17: 5501. https://doi.org/10.3390/jcm12175501
APA StyleRomero-Tapia, S. d. J., Guzmán Priego, C. G., Del-Río-Navarro, B. E., & Sánchez-Solis, M. (2023). Advances in the Relationship between Respiratory Viruses and Asthma. Journal of Clinical Medicine, 12(17), 5501. https://doi.org/10.3390/jcm12175501







