An Overview of Influenza Viruses and Vaccines
Abstract
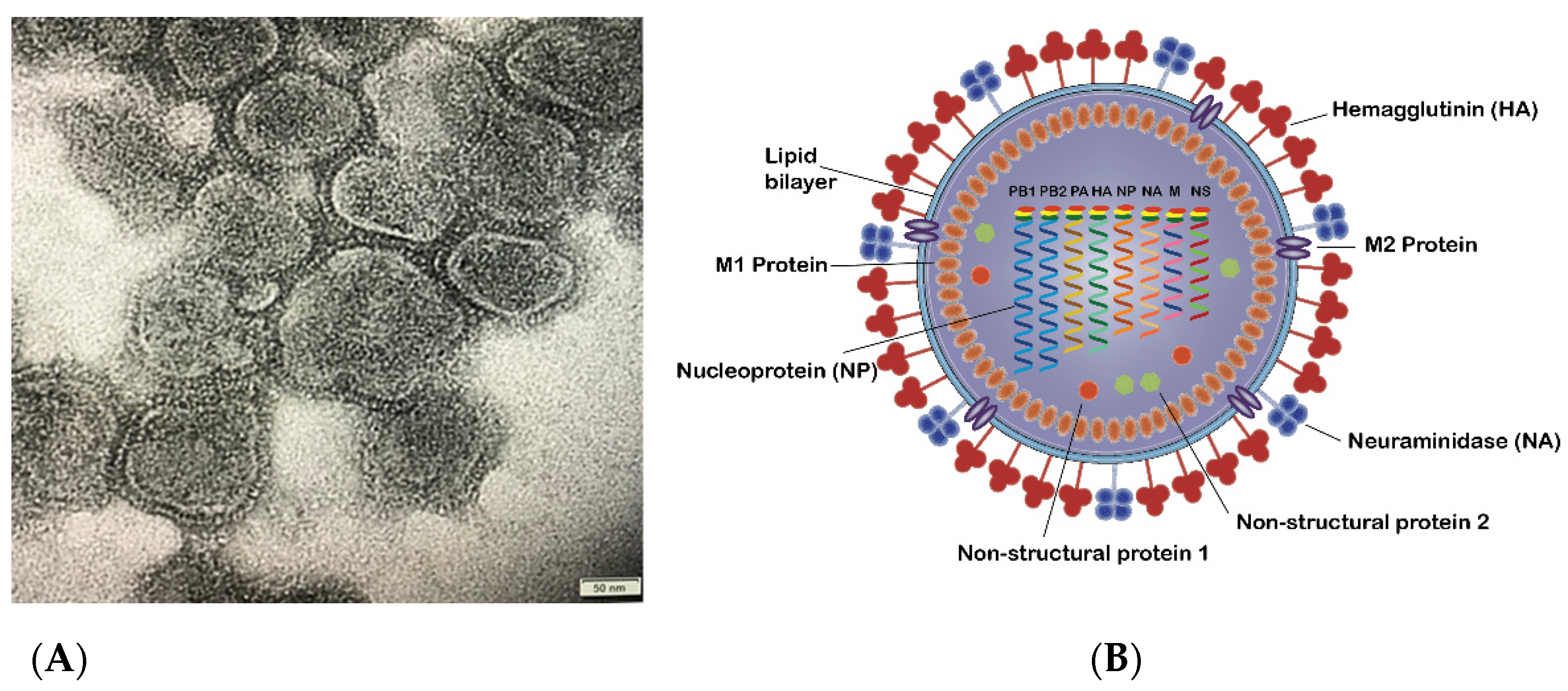
1. The Influenza Virus and Subtypes
2. Influenza Virus Life Cycle
- (1)
- Virus attachment to the sialic acid receptor: the first stage of viral infection is the attachment of HA to the sialic acids located at the host cell membrane’s surface. Sialic acids are connected to the carbohydrates of HA via glycosidic linkage. There are two linkages which are essential for the specificity of HA; (i) α (2, 3) linkage which is abundant in the digestive tract in avian or in the bronchial tissue in human, monkeys, horses, as well as upper respiratory tract of the lung epithelium in swine, and (ii) α (2, 6) which is found on the cell surface of the human upper respiratory tract as well as the trachea of bats and swine [4,18].
- (2)
- Entry of the virus into the host cell: receptor-mediated endocytosis occurs upon virus binding, and the virus enters the host cell in an endosome.
- (3)
- Fusion and uncoating of the virus particles: The acidic pH of the endosome (pH 5–6) causes the fusion of the viral and endosomal membranes and opens up the M2 ion channel protein and acidifies the nucleus, thus allowing vRNP to be released from M1 to enter the host cell’s cytoplasm and then to the nucleus.
- (4)
- vRNPs entry into the nucleus followed by transcription and replication: the viral proteins that constitute the vRNP (NP, PA, PB1, and PB2) detect the nuclear localization signals which can attach to the cellular nuclear import machinery and consequently, enter the nucleus to undergo the transcription and replication processes. The negative-sense RNA is first transformed into a positive-sense RNA and serves as a template for the generation of viral RNA, followed by the internal RNA synthesis initiated by the viral RNA-dependent RNA polymerase (RdRp). Through its C-terminal domain, the RdRp associates with the large subunit of RNA Polymerase II (Pol II), which continue the transcription to produce mature mRNA. vRNPs are then exported out from the viral core through the nuclear pores [19,20].
- (5)
- The assembly of virus components and budding: the virus’ protein components, i.e., HA, NA, and M2, are transported to the membrane’s apical region where the virions bud from the polarized epithelial cells.
- (6)
- The release of new virions from the host cells: after forming viral particles, the sialic acid residues from glycoproteins and glycolipids are cleaved by NA, enabling the newly synthesized viral particles to be released from the host membrane and spread to the nearby cells [21].
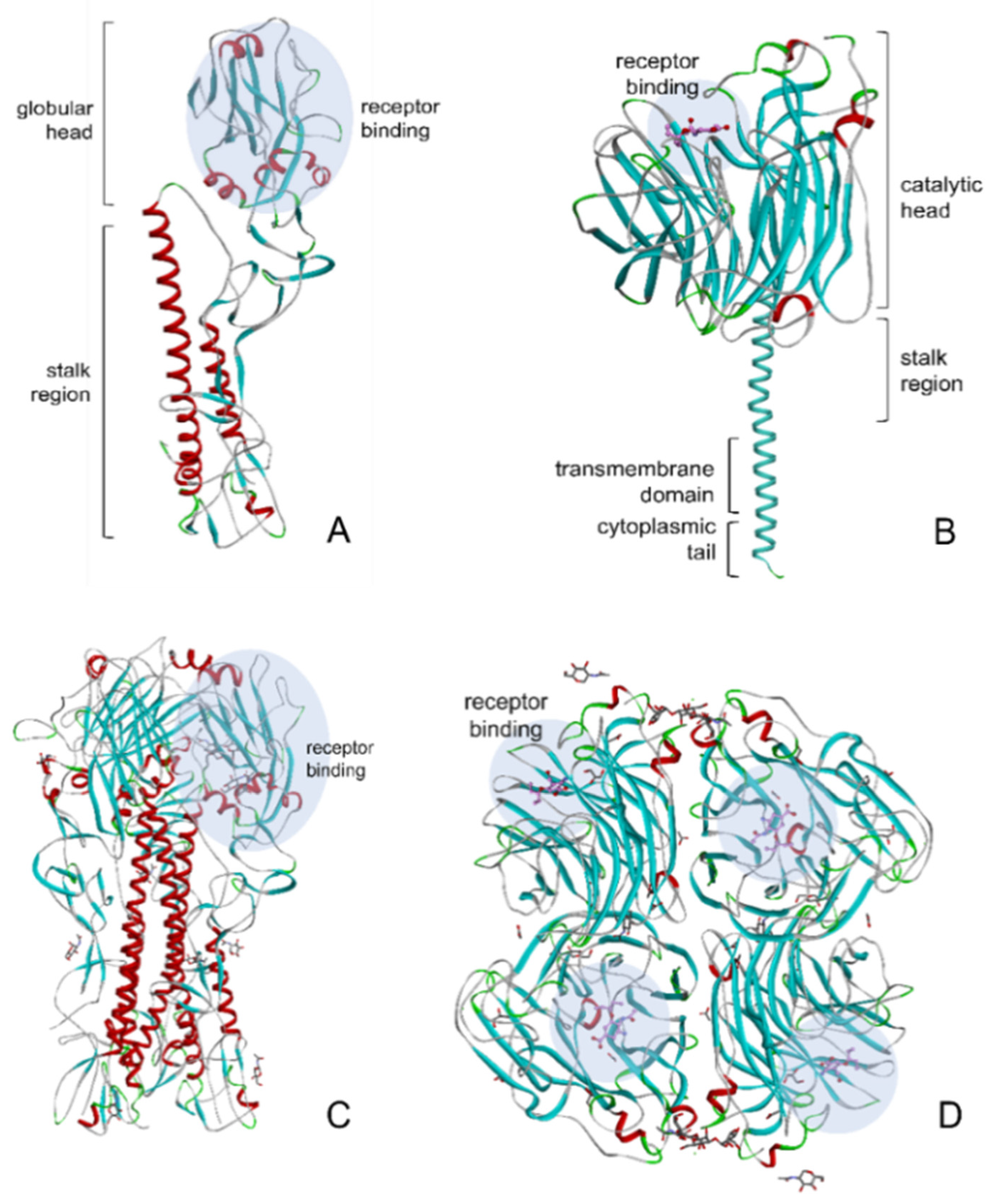
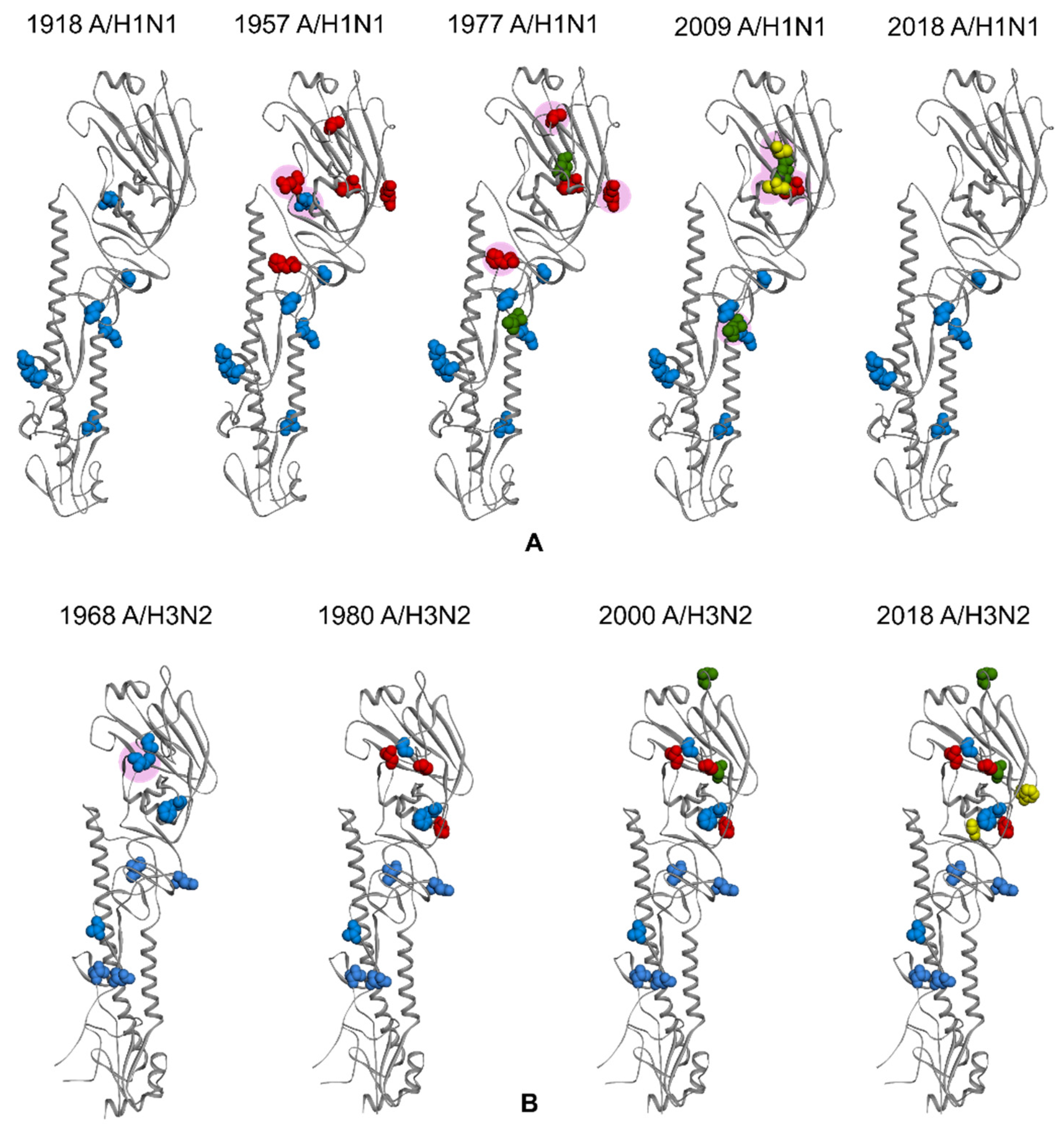
3. HA and NA of Influenza Virus
4. Glycosylation of HA and NA
5. The Influenza Epidemic vs. Pandemic
6. Vaccines against Influenza Epidemic and Pandemic
6.1. Inactivated Influenza Vaccine (IIV)
6.1.1. Whole-Virus Inactivated Vaccines (WIV)
6.1.2. Split-Virus Inactivated Vaccines
6.1.3. Subunit Inactivated Vaccines
6.2. Live Attenuated Influenza Vaccines (LAIV)
6.3. Recombinant HA Vaccine
7. Influenza Vaccine Manufacturing Processes
7.1. Egg-Based Vaccines
7.2. Cell-Based Vaccines
8. Influenza Vaccine Formulation and Ingredients
8.1. Active Components
8.2. Adjuvants
8.3. Stabilizers
8.4. Preservatives
8.5. Trace Components
9. Novel Influenza Vaccine Platforms
9.1. Virus-Like Particle (VLP) Vaccines
9.2. Antigen-Presenting Cell (APC) Inducible Vaccines
9.3. Nanoparticle-Based Influenza Vaccines
9.4. Universal Influenza Vaccines
10. Characterization of Vaccine Products
10.1. Purity
10.2. Quantity
10.3. Homogeneity
10.4. Stability
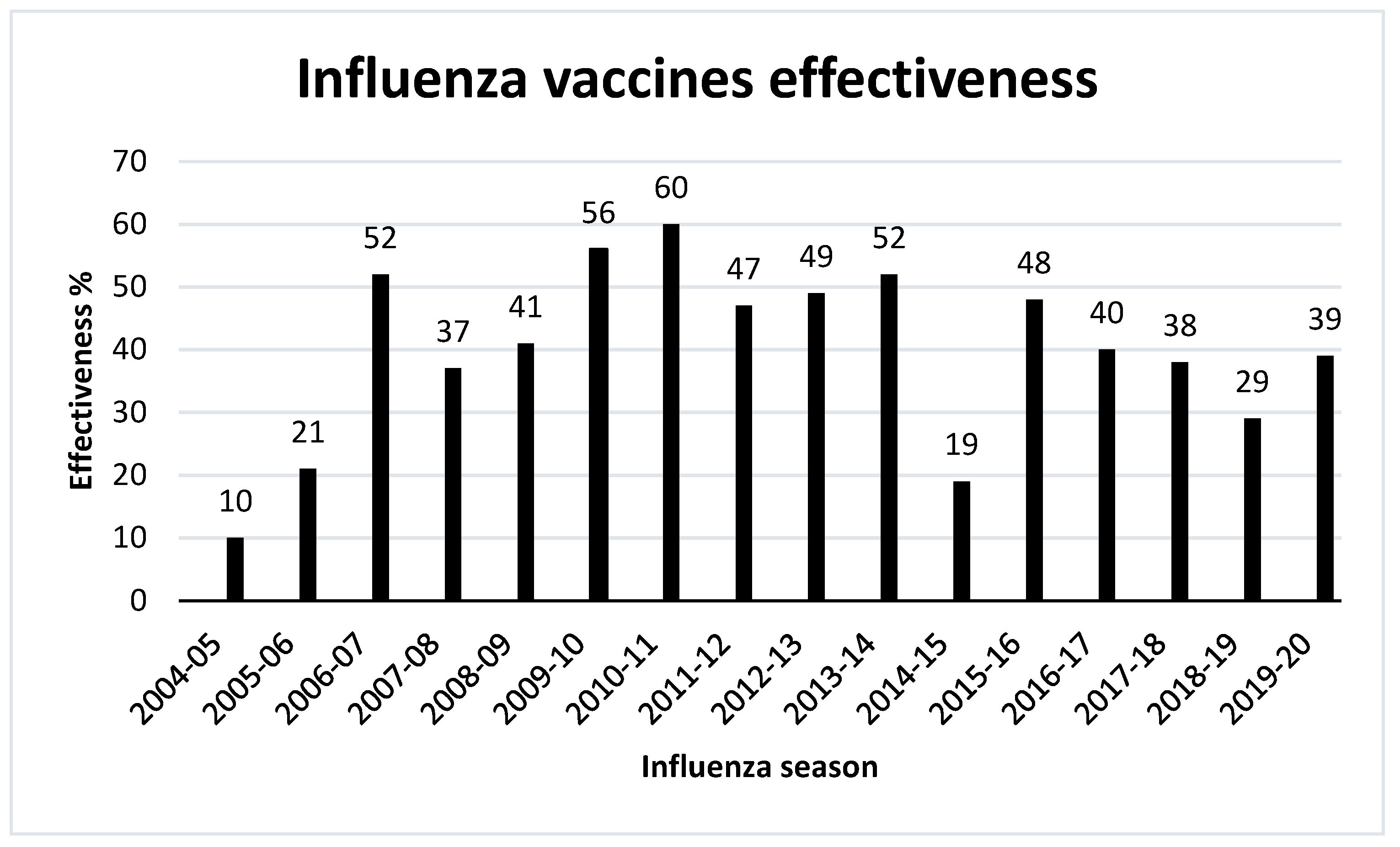
11. Effectiveness of Influenza Vaccines
12. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
References
- Klimov, A.I.; Garten, R.; Russell, C.; Barr, I.G.; Besselaar, T.G.; Daniels, R.; Engelhardt, O.G.; Grohmann, G.; Itamura, S.; Kelso, A.; et al. WHO recommendations for the viruses to be used in the 2012 Southern Hemisphere Influenza Vaccine: Epidemiology, antigenic and genetic characteristics of influenza A(H1N1)pdm09, A(H3N2) and B influenza viruses collected from February to September 2011. Vaccine 2012, 30, 6461–6471. [Google Scholar] [CrossRef] [PubMed]
- CDC. Types of Influenza Viruses. Available online: https://www.cdc.gov/flu/about/viruses/types.htm (accessed on 22 January 2020).
- Asha, K.; Kumar, B. Emerging Influenza D Virus Threat: What We Know So Far! J. Clin. Med. 2019, 8, 192. [Google Scholar] [CrossRef]
- Dawson, W.K.; Lazniewski, M.; Plewczynski, D. RNA structure interactions and ribonucleoprotein processes of the influenza A virus. Brief. Funct. Genom. 2017, 17, 402–414. [Google Scholar] [CrossRef]
- Saunders-Hastings, P.R.; Krewski, D. Reviewing the History of Pandemic Influenza: Understanding Patterns of Emergence and Transmission. Pathogens 2016, 5, 66. [Google Scholar] [CrossRef] [PubMed]
- Seladi-Schulman, J.; Steel, J.; Lowen, A.C. Spherical influenza viruses have a fitness advantage in embryonated eggs, while filament-producing strains are selected in vivo. J. Virol. 2013, 87, 13343–13353. [Google Scholar] [CrossRef]
- Gaymard, A.; Le Briand, N.; Frobert, E.; Lina, B.; Escuret, V. Functional balance between neuraminidase and haemagglutinin in influenza viruses. Clin. Microbiol. Infect. 2016, 22, 975–983. [Google Scholar] [CrossRef] [PubMed]
- Harris, A.; Cardone, G.; Winkler, D.C.; Heymann, J.B.; Brecher, M.; White, J.M.; Steven, A.C. Influenza virus pleiomorphy characterized by cryoelectron tomography. Proc. Natl. Acad. Sci. USA 2006, 103, 19123. [Google Scholar] [CrossRef] [PubMed]
- Moulès, V.; Terrier, O.; Yver, M.; Riteau, B.; Moriscot, C.; Ferraris, O.; Julien, T.; Giudice, E.; Rolland, J.-P.; Erny, A.; et al. Importance of viral genomic composition in modulating glycoprotein content on the surface of influenza virus particles. Virology 2011, 414, 51–62. [Google Scholar] [CrossRef]
- Tong, S.; Li, Y.; Rivailler, P.; Conrardy, C.; Castillo, D.A.; Chen, L.M.; Recuenco, S.; Ellison, J.A.; Davis, C.T.; York, I.A.; et al. A distinct lineage of influenza A virus from bats. Proc. Natl. Acad. Sci. USA 2012, 109, 4269–4274. [Google Scholar] [CrossRef]
- De Vries, R.D.; Herfst, S.; Richard, M. Avian Influenza A Virus Pandemic Preparedness and Vaccine Development. Vaccines 2018, 6, 46. [Google Scholar] [CrossRef]
- Asaduzzaman, S.M.; Ma, J.; van den Driessche, P. The coexistence or replacement of two subtypes of influenza. Math. Biosci. 2015, 270, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Petric, M.; Comanor, L.; Petti, C.A. Role of the Laboratory in Diagnosis of Influenza during Seasonal Epidemics and Potential Pandemics. J. Infect. Dis. 2006, 194, S98–S110. [Google Scholar] [CrossRef] [PubMed]
- Zebedee, S.L.; Lamb, R.A. Influenza A virus M2 protein: Monoclonal antibody restriction of virus growth and detection of M2 in virions. J. Virol. 1988, 62, 2762–2772. [Google Scholar] [CrossRef]
- Shaw, M.; Palese, P. Orthomyxoviridae: The viruses and their replication. In Fields Virol, 5th ed.; Knipe, D.M., Howley, P.M., Eds.; Lippincott Williams & Wilkins: Philadelphia, PA, USA, 2007; pp. 1647–1689. [Google Scholar]
- Bouvier, N.M.; Palese, P. The biology of influenza viruses. Vaccine 2008, 26 (Suppl. 4), D49–D53. [Google Scholar] [CrossRef]
- Wolff, T.; Veit, M. Influenza B, C and D Viruses (Orthomyxoviridae). In Encyclopedia of Virology; Academic Press: Cambridge, MA, USA, 2021; pp. 561–574. [Google Scholar] [CrossRef]
- Kuchipudi, S.V.; Nelli, R.K.; Gontu, A.; Satyakumar, R.; Surendran Nair, M.; Subbiah, M. Sialic Acid Receptors: The Key to Solving the Enigma of Zoonotic Virus Spillover. Viruses 2021, 13, 262. [Google Scholar] [CrossRef]
- Samji, T. Influenza A: Understanding the viral life cycle. Yale J. Biol. Med. 2009, 82, 153–159. [Google Scholar] [PubMed]
- Engelhardt, O.G.; Smith, M.; Fodor, E. Association of the influenza A virus RNA-dependent RNA polymerase with cellular RNA polymerase II. J. Virol. 2005, 79, 5812–5818. [Google Scholar] [CrossRef] [PubMed]
- Nayak, D.P.; Balogun, R.A.; Yamada, H.; Zhou, Z.H.; Barman, S. Influenza virus morphogenesis and budding. Virus Res. 2009, 143, 147–161. [Google Scholar] [CrossRef]
- Gamblin, S.J.; Skehel, J.J. Influenza hemagglutinin and neuraminidase membrane glycoproteins. J. Biol. Chem. 2010, 285, 28403–28409. [Google Scholar] [CrossRef]
- Hai, R.; Krammer, F.; Tan, G.S.; Pica, N.; Eggink, D.; Maamary, J.; Margine, I.; Albrecht, R.A.; Palese, P. Influenza viruses expressing chimeric hemagglutinins: Globular head and stalk domains derived from different subtypes. J. Virol. 2012, 86, 5774–5781. [Google Scholar] [CrossRef]
- Velkov, T.; Ong, C.; Baker, M.A.; Kim, H.; Li, J.; Nation, R.L.; Huang, J.X.; Cooper, M.A.; Rockman, S. The antigenic architecture of the hemagglutinin of influenza H5N1 viruses. Mol. Immunol. 2013, 56, 705–719. [Google Scholar] [CrossRef] [PubMed]
- Wohlbold, T.J.; Krammer, F. In the shadow of hemagglutinin: A growing interest in influenza viral neuraminidase and its role as a vaccine antigen. Viruses 2014, 6, 2465–2494. [Google Scholar] [CrossRef]
- Von Itzstein, M.; Thomson, R. Anti-influenza drugs: The development of sialidase inhibitors. Handb. Exp. Pharm. 2009, 189, 111–154. [Google Scholar] [CrossRef]
- Byrd-Leotis, L.; Cummings, R.D.; Steinhauer, D.A. The Interplay between the Host Receptor and Influenza Virus Hemagglutinin and Neuraminidase. Int. J. Mol. Sci. 2017, 18, 1541. [Google Scholar] [CrossRef] [PubMed]
- York, I.A.; Stevens, J.; Alymova, I.V. Influenza virus N-linked glycosylation and innate immunity. Biosci. Rep. 2019, 39, BSR20171505. [Google Scholar] [CrossRef] [PubMed]
- Vigerust, D.J.; Shepherd, V.L. Virus glycosylation: Role in virulence and immune interactions. Trends Microbiol. 2007, 15, 211–218. [Google Scholar] [CrossRef]
- Kim, P.; Jang, Y.H.; Kwon, S.B.; Lee, C.M.; Han, G.; Seong, B.L. Glycosylation of Hemagglutinin and Neuraminidase of Influenza A Virus as Signature for Ecological Spillover and Adaptation among Influenza Reservoirs. Viruses 2018, 10, 183. [Google Scholar] [CrossRef]
- Job, E.R.; Deng, Y.M.; Barfod, K.K.; Tate, M.D.; Caldwell, N.; Reddiex, S.; Maurer-Stroh, S.; Brooks, A.G.; Reading, P.C. Addition of glycosylation to influenza A virus hemagglutinin modulates antibody-mediated recognition of H1N1 2009 pandemic viruses. J. Immunol. 2013, 190, 2169–2177. [Google Scholar] [CrossRef]
- Wanzeck, K.; Boyd, K.L.; McCullers, J.A. Glycan shielding of the influenza virus hemagglutinin contributes to immunopathology in mice. Am. J. Respir. Crit. Care Med. 2011, 183, 767–773. [Google Scholar] [CrossRef]
- Kobayashi, Y.; Suzuki, Y. Evidence for N-glycan shielding of antigenic sites during evolution of human influenza A virus hemagglutinin. J. Virol. 2012, 86, 3446–3451. [Google Scholar] [CrossRef]
- Cruz, E.; Cain, J.; Crossett, B.; Kayser, V. Site-specific glycosylation profile of influenza A (H1N1) hemagglutinin through tandem mass spectrometry. Hum. Vaccines Immunother. 2018, 14, 508–517. [Google Scholar] [CrossRef]
- Fiore, A.E.; Uyeki, T.M.; Broder, K.; Finelli, L.; Euler, G.L.; Singleton, J.A.; Iskander, J.K.; Wortley, P.M.; Shay, D.K.; Bresee, J.S.; et al. Prevention and control of influenza with vaccines: Recommendations of the Advisory Committee on Immunization Practices (ACIP), 2010. MMWR Recomm. Rep. 2010, 59, 1–62. [Google Scholar]
- Barr, I.G.; McCauley, J.; Cox, N.; Daniels, R.; Engelhardt, O.G.; Fukuda, K.; Grohmann, G.; Hay, A.; Kelso, A.; Klimov, A.; et al. Epidemiological, antigenic and genetic characteristics of seasonal influenza A(H1N1), A(H3N2) and B influenza viruses: Basis for the WHO recommendation on the composition of influenza vaccines for use in the 2009–2010 Northern Hemisphere season. Vaccine 2010, 28, 1156–1167. [Google Scholar] [CrossRef]
- Chang, D.; Zaia, J. Why Glycosylation Matters in Building a Better Flu Vaccine. Mol. Cell Proteom. 2019, 18, 2348–2358. [Google Scholar] [CrossRef]
- Saito, T.; Taylor, G.; Webster, R.G. Steps in maturation of influenza A virus neuraminidase. J. Virol. 1995, 69, 5011–5017. [Google Scholar] [CrossRef]
- Wang, N.; Glidden, E.J.; Murphy, S.R.; Pearse, B.R.; Hebert, D.N. The cotranslational maturation program for the type II membrane glycoprotein influenza neuraminidase. J. Biol. Chem. 2008, 283, 33826–33837. [Google Scholar] [CrossRef] [PubMed]
- Bao, D.; Xue, R.; Zhang, M.; Lu, C.; Ma, T.; Ren, C.; Zhang, T.; Yang, J.; Teng, Q.; Li, X.; et al. N-Linked Glycosylation Plays an Important Role in Budding of Neuraminidase Protein and Virulence of Influenza Viruses. J. Virol. 2021, 95, e02042-20. [Google Scholar] [CrossRef]
- Iuliano, A.D.; Roguski, K.M.; Chang, H.H.; Muscatello, D.J.; Palekar, R.; Tempia, S.; Cohen, C.; Gran, J.M.; Schanzer, D.; Cowling, B.J.; et al. Estimates of global seasonal influenza-associated respiratory mortality: A modelling study. Lancet 2018, 391, 1285–1300. [Google Scholar] [CrossRef]
- WHO. Influenza (Seasonal). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(seasonal) (accessed on 2 December 2019).
- Fleming, D.M.; Zambon, M.; Bartelds, A.I.M.; De Jong, J.C. The duration and magnitude of influenza epidemics: A study of surveillance data from sentinel general practices in England, Wales and the Netherlands. Eur. J. Epidemiol. 1999, 15, 467–473. [Google Scholar] [CrossRef] [PubMed]
- Boni, M.F. Vaccination and antigenic drift in influenza. Vaccine 2008, 26, C8–C14. [Google Scholar] [CrossRef] [PubMed]
- McCaughey, C. Influenza: A virus of our times. Ulst. Med. J. 2010, 79, 46–51. [Google Scholar]
- Potter, C.W. A history of influenza. J. Appl. Microbiol. 2001, 91, 572–579. [Google Scholar] [CrossRef]
- Nguyen-Van-Tam, J.S.; Hampson, A.W. The epidemiology and clinical impact of pandemic influenza. Vaccine 2003, 21, 1762–1768. [Google Scholar] [CrossRef]
- Scholtissek, C.; Rohde, W.; Von Hoyningen, V.; Rott, R. On the origin of the human influenza virus subtypes H2N2 and H3N2. Virology 1978, 87, 13–20. [Google Scholar] [CrossRef]
- De Wit, E.; Fouchier, R.A. Emerging influenza. J. Clin. Virol. 2008, 41, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Jester, B.; Uyeki, T.M.; Jernigan, D.B.; Tumpey, T.M. Historical and clinical aspects of the 1918 H1N1 pandemic in the United States. Virology 2019, 527, 32–37. [Google Scholar] [CrossRef]
- Johnson, N.P.; Mueller, J. Updating the accounts: Global mortality of the 1918–1920 “Spanish” influenza pandemic. Bull. Hist. Med. 2002, 76, 105–115. [Google Scholar] [CrossRef] [PubMed]
- Kayser, V.; Ramzan, I. Vaccines and Vaccination: History and Emerging Issues. Hum. Vaccines Immunother. 2021, 17. [Google Scholar] [CrossRef]
- Spreeuwenberg, P.; Kroneman, M.; Paget, J. Reassessing the Global Mortality Burden of the 1918 Influenza Pandemic. Am. J. Epidemiol. 2018, 187, 2561–2567. [Google Scholar] [CrossRef]
- Taubenberger, J.K.; Kash, J.C.; Morens, D.M. The 1918 influenza pandemic: 100 years of questions answered and unanswered. Sci. Transl. Med. 2019, 11, eaau5485. [Google Scholar] [CrossRef]
- Viboud, C.; Simonsen, L.; Fuentes, R.; Flores, J.; Miller, M.A.; Chowell, G. Global Mortality Impact of the 1957-1959 Influenza Pandemic. J. Infect. Dis. 2016, 213, 738–745. [Google Scholar] [CrossRef]
- Oxford, J.S. Influenza A pandemics of the 20th century with special reference to 1918: Virology, pathology and epidemiology. Rev. Med. Virol. 2000, 10, 119–133. [Google Scholar] [CrossRef]
- CDC. 1968 Pandemic (H3N2 Virus). Available online: https://www.cdc.gov/flu/pandemic-resources/1968-pandemic.html (accessed on 5 December 2019).
- Mena, I.; Nelson, M.I.; Quezada-Monroy, F.; Dutta, J.; Cortes-Fernández, R.; Lara-Puente, J.H.; Castro-Peralta, F.; Cunha, L.F.; Trovão, N.S.; Lozano-Dubernard, B.; et al. Origins of the 2009 H1N1 influenza pandemic in swine in Mexico. eLife 2016, 5, e16777. [Google Scholar] [CrossRef] [PubMed]
- WHO. What Is the Pandemic (H1N1) 2009 Virus? Available online: https://www.who.int/csr/disease/swineflu/frequently_asked_questions/about_disease/en/ (accessed on 29 November 2020).
- Taubenberger, J.K.; Morens, D.M. Influenza: The once and future pandemic. Public Health Rep. 2010, 125 (Suppl. 3), 16–26. [Google Scholar] [CrossRef]
- Paules, C.; Subbarao, K. Influenza. Lancet 2017, 390, 697–708. [Google Scholar] [CrossRef]
- Peiris, J.S.M.; Yu, W.C.; Leung, C.W.; Cheung, C.Y.; Ng, W.F.; Nicholls, J.M.; Ng, T.K.; Chan, K.H.; Lai, S.T.; Lim, W.L.; et al. Re-emergence of fatal human influenza A subtype H5N1 disease. Lancet 2004, 363, 617–619. [Google Scholar] [CrossRef]
- Chowdhury, S.; Hossain, M.E.; Ghosh, P.K.; Ghosh, S.; Hossain, M.B.; Beard, C.; Rahman, M.; Rahman, M.Z. The Pattern of Highly Pathogenic Avian Influenza H5N1 Outbreaks in South Asia. Trop. Med. Infect. Dis. 2019, 4, 138. [Google Scholar] [CrossRef]
- WHO. Influenza (Avian and Other Zoonotic). Available online: https://www.who.int/news-room/fact-sheets/detail/influenza-(avian-and-other-zoonotic) (accessed on 15 January 2020).
- CDC. Highly Pathogenic Asian Avian Influenza A(H5N1) in People. Available online: https://www.cdc.gov/flu/avianflu/h5n1-people.htm (accessed on 10 September 2021).
- WHO. Avian Influenza Weekly Update Number 716. Available online: https://www.who.int/docs/default-source/wpro---documents/emergency/surveillance/avian-influenza/ai-20191122.pdf?sfvrsn=30d65594_42 (accessed on 4 December 2019).
- Young, B.; Sadarangani, S.; Jiang, L.; Wilder-Smith, A.; Chen, M.I. Duration of Influenza Vaccine Effectiveness: A Systematic Review, Meta-analysis, and Meta-regression of Test-Negative Design Case-Control Studies. J. Infect. Dis. 2018, 217, 731–741. [Google Scholar] [CrossRef] [PubMed]
- Weir, J.P.; Gruber, M.F. An overview of the regulation of influenza vaccines in the United States. Influenza Other Respir. Viruses 2016, 10, 354–360. [Google Scholar] [CrossRef]
- Gutierrez, A.F.; El Sahly, H. Recombinant hemagglutinin protein vaccine: A new option in immunization against influenza. Future Virol. 2015, 10, 1057–1067. [Google Scholar] [CrossRef]
- Yamayoshi, S.; Kawaoka, Y. Current and future influenza vaccines. Nat. Med. 2019, 25, 212–220. [Google Scholar] [CrossRef]
- Xie, H.; Wan, X.F.; Ye, Z.P.; Plant, E.P.; Zhao, Y.Q.; Xu, Y.F.; Li, X.; Finch, C.; Zhao, N.; Kawano, T.; et al. H3N2 Mismatch of 2014-15 Northern Hemisphere Influenza Vaccines and Head-to-head Comparison between Human and Ferret Antisera derived Antigenic Maps. Sci. Rep. 2015, 5, 15279. [Google Scholar] [CrossRef]
- Statler, V.A.; Albano, F.R.; Airey, J.; Sawlwin, D.C.; Graves Jones, A.; Matassa, V.; Heijnen, E.; Edelman, J.; Marshall, G.S. Immunogenicity and safety of a quadrivalent inactivated influenza vaccine in children 6–59 months of age: A phase 3, randomized, noninferiority study. Vaccine 2019, 37, 343–351. [Google Scholar] [CrossRef] [PubMed]
- McKeage, K. Inactivated quadrivalent split-virus seasonal influenza vaccine (Fluarix® quadrivalent): A review of its use in the prevention of disease caused by influenza A and B. Drugs 2013, 73, 1587–1594. [Google Scholar] [CrossRef]
- Bekkat-Berkani, R.; Ray, R.; Jain, V.K.; Chandrasekaran, V.; Innis, B.L. Evidence update: GlaxoSmithKline’s inactivated quadrivalent influenza vaccines. Expert Rev. Vaccines 2016, 15, 201–214. [Google Scholar] [CrossRef]
- Robertson, C.A.; DiazGranados, C.A.; Decker, M.D.; Chit, A.; Mercer, M.; Greenberg, D.P. Fluzone® High-Dose Influenza Vaccine. Expert Rev. Vaccines 2016, 15, 1495–1505. [Google Scholar] [CrossRef]
- Suryadevara, M.; Domachowske, J.B. Quadrivalent influenza vaccine in the United States. Hum. Vaccines Immunother. 2014, 10, 596–599. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sullivan, S.G.; Arriola, C.S.; Bocacao, J.; Burgos, P.; Bustos, P.; Carville, K.S.; Cheng, A.C.; Chilver, M.B.; Cohen, C.; Deng, Y.-M.; et al. Heterogeneity in influenza seasonality and vaccine effectiveness in Australia, Chile, New Zealand and South Africa: Early estimates of the 2019 influenza season. Eurosurveillance 2019, 24, 1900645. [Google Scholar] [CrossRef] [PubMed]
- Montomoli, E.; Torelli, A.; Manini, I.; Gianchecchi, E. Immunogenicity and Safety of the New Inactivated Quadrivalent Influenza Vaccine Vaxigrip Tetra: Preliminary Results in Children ≥6 Months and Older Adults. Vaccines 2018, 6, 14. [Google Scholar] [CrossRef]
- Zhao, L.; Young, K.; Gemmill, I. Summary of the NACI Seasonal Influenza Vaccine Statement for 2019-2020. Can. Commun. Dis. Rep. 2019, 45, 149–155. [Google Scholar] [CrossRef]
- Tsai, T.F. Fluad®-MF59®-Adjuvanted Influenza Vaccine in Older Adults. Infect. Chemother. 2013, 45, 159–174. [Google Scholar] [CrossRef]
- Essink, B.; Fierro, C.; Rosen, J.; Figueroa, A.L.; Zhang, B.; Verhoeven, C.; Edelman, J.; Smolenov, I. Immunogenicity and safety of MF59-adjuvanted quadrivalent influenza vaccine versus standard and alternate B strain MF59-adjuvanted trivalent influenza vaccines in older adults. Vaccine 2020, 38, 242–250. [Google Scholar] [CrossRef] [PubMed]
- Dunkle, L.; Izikson, R.; Post, P.; Cox, M. Introducing Modern Recombinant Technology to the Realm of Seasonal Influenza Vaccine: Flublok® For Prevention of Influenza in Adults. EC Microbiol. 2015, 2, 224–234. [Google Scholar]
- Lamb, Y.N. Cell-Based Quadrivalent Inactivated Influenza Virus Vaccine (Flucelvax® Tetra/Flucelvax Quadrivalent®): A Review in the Prevention of Influenza. Drugs 2019, 79, 1337–1348. [Google Scholar] [CrossRef] [PubMed]
- Baxter, R.; Eaton, A.; Hansen, J.; Aukes, L.; Caspard, H.; Ambrose, C.S. Safety of quadrivalent live attenuated influenza vaccine in subjects aged 2–49years. Vaccine 2017, 35, 1254–1258. [Google Scholar] [CrossRef] [PubMed]
- Hoft, D.F.; Lottenbach, K.R.; Blazevic, A.; Turan, A.; Blevins, T.P.; Pacatte, T.P.; Yu, Y.; Mitchell, M.C.; Hoft, S.G.; Belshe, R.B. Comparisons of the Humoral and Cellular Immune Responses Induced by Live Attenuated Influenza Vaccine and Inactivated Influenza Vaccine in Adults. Clin. Vaccine Immunol. 2017, 24, e00414-16. [Google Scholar] [CrossRef]
- Al-Mazrou, A.; Scheifele, D.W.; Soong, T.; Bjornson, G. Comparison of adverse reactions to whole-virion and split-virion influenza vaccines in hospital personnel. CMAJ 1991, 145, 213–218. [Google Scholar]
- Soema, P.C.; Kompier, R.; Amorij, J.P.; Kersten, G.F. Current and next generation influenza vaccines: Formulation and production strategies. Eur. J. Pharm. Biopharm. 2015, 94, 251–263. [Google Scholar] [CrossRef]
- Sabbaghi, A.; Miri, S.M.; Keshavarz, M.; Zargar, M.; Ghaemi, A. Inactivation methods for whole influenza vaccine production. Rev. Med. Virol. 2019, 29, e2074. [Google Scholar] [CrossRef]
- Lee, K.K.; Sahin, Y.Z.; Neeleman, R.; Trout, B.L.; Kayser, V. Quantitative determination of the surfactant-induced split ratio of influenza virus by fluorescence spectroscopy. Hum. Vaccines Immunother. 2016, 12, 1757–1765. [Google Scholar] [CrossRef]
- Squarcione, S.; Sgricia, S.; Biasio, L.R.; Perinetti, E. Comparison of the reactogenicity and immunogenicity of a split and a subunit-adjuvanted influenza vaccine in elderly subjects. Vaccine 2003, 21, 1268–1274. [Google Scholar] [CrossRef]
- Khalaj-Hedayati, A.; Chua, C.L.L.; Smooker, P.; Lee, K.W. Nanoparticles in influenza subunit vaccine development: Immunogenicity enhancement. Influenza Other Respir. Viruses 2019, 14, 92–101. [Google Scholar] [CrossRef] [PubMed]
- Maassab, H.F. Adaptation and growth characteristics of influenza virus at 25 degrees C. Nature 1967, 213, 612–614. [Google Scholar] [CrossRef]
- Beyer, W.E.P.; Palache, A.M.; de Jong, J.C.; Osterhaus, A.D.M.E. Cold-adapted live influenza vaccine versus inactivated vaccine: Systemic vaccine reactions, local and systemic antibody response, and vaccine efficacy: A meta-analysis. Vaccine 2002, 20, 1340–1353. [Google Scholar] [CrossRef]
- Suzuki, T.; Kawaguchi, A.; Ainai, A.; Tamura, S.-i.; Ito, R.; Multihartina, P.; Setiawaty, V.; Pangesti, K.N.A.; Odagiri, T.; Tashiro, M.; et al. Relationship of the quaternary structure of human secretory IgA to neutralization of influenza virus. Proc. Natl. Acad. Sci. USA 2015, 112, 7809–7814. [Google Scholar] [CrossRef] [PubMed]
- King, J.C., Jr.; Treanor, J.; Fast, P.E.; Wolff, M.; Yan, L.; Iacuzio, D.; Readmond, B.; O’Brien, D.; Mallon, K.; Highsmith, W.E.; et al. Comparison of the safety, vaccine virus shedding, and immunogenicity of influenza virus vaccine, trivalent, types A and B, live cold-adapted, administered to human immunodeficiency virus (HIV)-infected and non-HIV-infected adults. J. Infect. Dis. 2000, 181, 725–728. [Google Scholar] [CrossRef]
- Belshe, R.B.; Mendelman, P.M.; Treanor, J.; King, J.; Gruber, W.C.; Piedra, P.; Bernstein, D.I.; Hayden, F.G.; Kotloff, K.; Zangwill, K.; et al. The efficacy of live attenuated, cold-adapted, trivalent, intranasal influenzavirus vaccine in children. N. Engl. J. Med. 1998, 338, 1405–1412. [Google Scholar] [CrossRef]
- Keitel, W.A.; Couch, R.B.; Cate, T.R.; Maassab, H.F. Variability in infectivity of cold-adapted recombinant influenza virus vaccines in humans. J. Infect. Dis. 1994, 169, 477. [Google Scholar] [CrossRef]
- Cox, M.M.; Hollister, J.R. FluBlok, a next generation influenza vaccine manufactured in insect cells. Biologicals 2009, 37, 182–189. [Google Scholar] [CrossRef]
- Geisler, C.; Jarvis, D.L. Adventitious viruses in insect cell lines used for recombinant protein expression. Protein Expr. Purif. 2018, 144, 25–32. [Google Scholar] [CrossRef]
- Grohskopf, L.A.; Sokolow, L.Z.; Broder, K.R.; Walter, E.B.; Fry, A.M.; Jernigan, D.B. Prevention and Control of Seasonal Influenza with Vaccines: Recommendations of the Advisory Committee on Immunization Practices-United States, 2018–2019 Influenza Season. MMWR Recomm Rep. 2018, 67, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Cox, M.M.; Patriarca, P.A.; Treanor, J. FluBlok, a recombinant hemagglutinin influenza vaccine. Influenza Other Respir. Viruses 2008, 2, 211–219. [Google Scholar] [CrossRef] [PubMed]
- Treanor, J.J.; Wilkinson, B.E.; Masseoud, F.; Hu-Primmer, J.; Battaglia, R.; O’Brien, D.; Wolff, M.; Rabinovich, G.; Blackwelder, W.; Katz, J.M. Safety and immunogenicity of a recombinant hemagglutinin vaccine for H5 influenza in humans. Vaccine 2001, 19, 1732–1737. [Google Scholar] [CrossRef]
- Wong, S.S.; Webby, R.J. Traditional and new influenza vaccines. Clin. Microbiol. Rev. 2013, 26, 476–492. [Google Scholar] [CrossRef]
- DeMarcus, L.; Shoubaki, L.; Federinko, S. Comparing influenza vaccine effectiveness between cell-derived and egg-derived vaccines, 2017–2018 influenza season. Vaccine 2019, 37, 4015–4021. [Google Scholar] [CrossRef]
- Milian, E.; Kamen, A.A. Current and emerging cell culture manufacturing technologies for influenza vaccines. Biomed Res. Int. 2015, 2015, 504831. [Google Scholar] [CrossRef] [PubMed]
- Rajao, D.S.; Perez, D.R. Universal Vaccines and Vaccine Platforms to Protect against Influenza Viruses in Humans and Agriculture. Front. Microbiol. 2018, 9, 123. [Google Scholar] [CrossRef]
- CDC. Cell-Based Flu Vaccines. Available online: https://www.cdc.gov/flu/protect/vaccine/cell-based.htm (accessed on 24 November 2020).
- Couch, R.B. Seasonal inactivated influenza virus vaccines. Vaccine 2008, 26 (Suppl. 4), D5–D9. [Google Scholar] [CrossRef]
- Tate, M.D.; Job, E.R.; Deng, Y.M.; Gunalan, V.; Maurer-Stroh, S.; Reading, P.C. Playing hide and seek: How glycosylation of the influenza virus hemagglutinin can modulate the immune response to infection. Viruses 2014, 6, 1294–1316. [Google Scholar] [CrossRef] [PubMed]
- Krause, J.C.; Crowe, J.E., Jr. Committing the Oldest Sins in the Newest Kind of Ways-Antibodies Targeting the Influenza Virus Type A Hemagglutinin Globular Head. Microbiol. Spectr. 2014, 2. [Google Scholar] [CrossRef]
- Ping, J.; Lopes, T.J.S.; Nidom, C.A.; Ghedin, E.; Macken, C.A.; Fitch, A.; Imai, M.; Maher, E.A.; Neumann, G.; Kawaoka, Y. Development of high-yield influenza A virus vaccine viruses. Nat. Commun. 2015, 6, 8148. [Google Scholar] [CrossRef]
- Stohr, K.; Bucher, D.; Colgate, T.; Wood, J. Influenza virus surveillance, vaccine strain selection, and manufacture. Methods Mol. Biol. 2012, 865, 147–162. [Google Scholar] [CrossRef]
- Skowronski, D.M.; Janjua, N.Z.; De Serres, G.; Sabaiduc, S.; Eshaghi, A.; Dickinson, J.A.; Fonseca, K.; Winter, A.L.; Gubbay, J.B.; Krajden, M.; et al. Low 2012-13 influenza vaccine effectiveness associated with mutation in the egg-adapted H3N2 vaccine strain not antigenic drift in circulating viruses. PLoS ONE 2014, 9, e92153. [Google Scholar] [CrossRef]
- Mochalova, L.; Gambaryan, A.; Romanova, J.; Tuzikov, A.; Chinarev, A.; Katinger, D.; Katinger, H.; Egorov, A.; Bovin, N. Receptor-binding properties of modern human influenza viruses primarily isolated in Vero and MDCK cells and chicken embryonated eggs. Virology 2003, 313, 473–480. [Google Scholar] [CrossRef][Green Version]
- Zost, S.J.; Parkhouse, K.; Gumina, M.E.; Kim, K.; Diaz Perez, S.; Wilson, P.C.; Treanor, J.J.; Sant, A.J.; Cobey, S.; Hensley, S.E. Contemporary H3N2 influenza viruses have a glycosylation site that alters binding of antibodies elicited by egg-adapted vaccine strains. Proc. Natl. Acad. Sci. USA 2017, 114, 12578. [Google Scholar] [CrossRef]
- Hegde, N.R. Cell culture-based influenza vaccines: A necessary and indispensable investment for the future. Hum. Vaccines Immunother. 2015, 11, 1223–1234. [Google Scholar] [CrossRef] [PubMed]
- An, Y.; Rininger, J.A.; Jarvis, D.L.; Jing, X.; Ye, Z.; Aumiller, J.J.; Eichelberger, M.; Cipollo, J.F. Comparative glycomics analysis of influenza Hemagglutinin (H5N1) produced in vaccine relevant cell platforms. J. Proteome Res. 2013, 12, 3707–3720. [Google Scholar] [CrossRef] [PubMed]
- Hutter, J.; Rodig, J.V.; Hoper, D.; Seeberger, P.H.; Reichl, U.; Rapp, E.; Lepenies, B. Toward animal cell culture-based influenza vaccine design: Viral hemagglutinin N-glycosylation markedly impacts immunogenicity. J. Immunol. 2013, 190, 220–230. [Google Scholar] [CrossRef] [PubMed]
- Paules, C.I.; Sullivan, S.G.; Subbarao, K.; Fauci, A.S. Chasing Seasonal Influenza—The Need for a Universal Influenza Vaccine. N. Engl. J. Med. 2018, 378, 7–9. [Google Scholar] [CrossRef]
- Lin, Y.; Wharton, S.A.; Whittaker, L.; Dai, M.; Ermetal, B.; Lo, J.; Pontoriero, A.; Baumeister, E.; Daniels, R.S.; McCauley, J.W. The characteristics and antigenic properties of recently emerged subclade 3C.3a and 3C.2a human influenza A(H3N2) viruses passaged in MDCK cells. Influenza Other Respir. Viruses 2017, 11, 263–274. [Google Scholar] [CrossRef]
- Tregoning, J.S.; Russell, R.F.; Kinnear, E. Adjuvanted influenza vaccines. Hum. Vaccines Immunother. 2018, 14, 550–564. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, T. [Influenza vaccine and adjuvant]. Yakugaku Zasshi 2011, 131, 1723–1731. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Pelliccia, M.; Andreozzi, P.; Paulose, J.; D’Alicarnasso, M.; Cagno, V.; Donalisio, M.; Civra, A.; Broeckel, R.M.; Haese, N.; Jacob Silva, P.; et al. Additives for vaccine storage to improve thermal stability of adenoviruses from hours to months. Nat. Commun. 2016, 7, 13520. [Google Scholar] [CrossRef] [PubMed]
- Chung, E.H. Vaccine allergies. Clin. Exp. Vaccine Res. 2014, 3, 50–57. [Google Scholar] [CrossRef]
- Grgacic, E.V.; Anderson, D.A. Virus-like particles: Passport to immune recognition. Methods 2006, 40, 60–65. [Google Scholar] [CrossRef]
- Zhao, Q.; Li, S.; Yu, H.; Xia, N.; Modis, Y. Virus-like particle-based human vaccines: Quality assessment based on structural and functional properties. Trends Biotechnol. 2013, 31, 654–663. [Google Scholar] [CrossRef]
- Gao, X.; Wang, W.; Li, Y.; Zhang, S.; Duan, Y.; Xing, L.; Zhao, Z.; Zhang, P.; Li, Z.; Li, R.; et al. Enhanced Influenza VLP vaccines comprising matrix-2 ectodomain and nucleoprotein epitopes protects mice from lethal challenge. Antivir. Res. 2013, 98, 4–11. [Google Scholar] [CrossRef]
- Giurgea, L.T.; Morens, D.M.; Taubenberger, J.K.; Memoli, M.J. Influenza Neuraminidase: A Neglected Protein and Its Potential for a Better Influenza Vaccine. Vaccines 2020, 8, 409. [Google Scholar] [CrossRef]
- Kumar, A.; Meldgaard, T.S.; Bertholet, S. Novel Platforms for the Development of a Universal Influenza Vaccine. Front. Immunol. 2018, 9, 600. [Google Scholar] [CrossRef]
- Low, J.G.; Lee, L.S.; Ooi, E.E.; Ethirajulu, K.; Yeo, P.; Matter, A.; Connolly, J.E.; Skibinski, D.A.; Saudan, P.; Bachmann, M.; et al. Safety and immunogenicity of a virus-like particle pandemic influenza A (H1N1) 2009 vaccine: Results from a double-blinded, randomized Phase I clinical trial in healthy Asian volunteers. Vaccine 2014, 32, 5041–5048. [Google Scholar] [CrossRef]
- Pillet, S.; Couillard, J.; Trépanier, S.; Poulin, J.-F.; Yassine-Diab, B.; Guy, B.; Ward, B.J.; Landry, N. Immunogenicity and safety of a quadrivalent plant-derived virus like particle influenza vaccine candidate—Two randomized Phase II clinical trials in 18 to 49 and ≥50 years old adults. PLoS ONE 2019, 14, e0216533. [Google Scholar] [CrossRef]
- Ren, Z.; Zhao, Y.; Liu, J.; Ji, X.; Meng, L.; Wang, T.; Sun, W.; Zhang, K.; Sang, X.; Yu, Z.; et al. Intramuscular and intranasal immunization with an H7N9 influenza virus-like particle vaccine protects mice against lethal influenza virus challenge. Int. Immunopharmacol. 2018, 58, 109–116. [Google Scholar] [CrossRef]
- Ramirez, A.; Morris, S.; Maucourant, S.; D’Ascanio, I.; Crescente, V.; Lu, I.N.; Farinelle, S.; Muller, C.P.; Whelan, M.; Rosenberg, W. A virus-like particle vaccine candidate for influenza A virus based on multiple conserved antigens presented on hepatitis B tandem core particles. Vaccine 2018, 36, 873–880. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Mohan, T.; Zhu, W.; Wang, C.; Deng, L.; Wang, B.Z. Sequential Immunizations with heterosubtypic virus-like particles elicit cross protection against divergent influenza A viruses in mice. Sci. Rep. 2018, 8, 4577. [Google Scholar] [CrossRef] [PubMed]
- López-Macías, C.; Ferat-Osorio, E.; Tenorio-Calvo, A.; Isibasi, A.; Talavera, J.; Arteaga-Ruiz, O.; Arriaga-Pizano, L.; Hickman, S.P.; Allende, M.; Lenhard, K.; et al. Safety and immunogenicity of a virus-like particle pandemic influenza A (H1N1) 2009 vaccine in a blinded, randomized, placebo-controlled trial of adults in Mexico. Vaccine 2011, 29, 7826–7834. [Google Scholar] [CrossRef] [PubMed]
- Kang, H.-J.; Chu, K.-B.; Lee, D.-H.; Lee, S.-H.; Park, B.R.; Kim, M.-C.; Kang, S.-M.; Quan, F.-S. Influenza M2 virus-like particle vaccination enhances protection in combination with avian influenza HA VLPs. PLoS ONE 2019, 14, e0216871. [Google Scholar] [CrossRef]
- Pillet, S.; Aubin, É.; Trépanier, S.; Bussière, D.; Dargis, M.; Poulin, J.-F.; Yassine-Diab, B.; Ward, B.J.; Landry, N. A plant-derived quadrivalent virus like particle influenza vaccine induces cross-reactive antibody and T cell response in healthy adults. Clin. Immunol. 2016, 168, 72–87. [Google Scholar] [CrossRef]
- Banchereau, J.; Steinman, R.M. Dendritic cells and the control of immunity. Nature 1998, 392, 245–252. [Google Scholar] [CrossRef]
- Fonteneau, J.F.; Gilliet, M.; Larsson, M.; Dasilva, I.; Munz, C.; Liu, Y.J.; Bhardwaj, N. Activation of influenza virus-specific CD4+ and CD8+ T cells: A new role for plasmacytoid dendritic cells in adaptive immunity. Blood 2003, 101, 3520–3526. [Google Scholar] [CrossRef]
- Abdel-Motal, U.M.; Guay, H.M.; Wigglesworth, K.; Welsh, R.M.; Galili, U. Immunogenicity of influenza virus vaccine is increased by anti-gal-mediated targeting to antigen-presenting cells. J. Virol. 2007, 81, 9131–9141. [Google Scholar] [CrossRef]
- Grodeland, G.; Mjaaland, S.; Tunheim, G.; Fredriksen, A.B.; Bogen, B. The specificity of targeted vaccines for APC surface molecules influences the immune response phenotype. PLoS ONE 2013, 8, e80008. [Google Scholar] [CrossRef]
- Muszkat, M.; Greenbaum, E.; Ben-Yehuda, A.; Oster, M.; Yeu’l, E.; Heimann, S.; Levy, R.; Friedman, G.; Zakay-Rones, Z. Local and systemic immune response in nursing-home elderly following intranasal or intramuscular immunization with inactivated influenza vaccine. Vaccine 2003, 21, 1180–1186. [Google Scholar] [CrossRef]
- Su, F.; Patel, G.B.; Hu, S.; Chen, W. Induction of mucosal immunity through systemic immunization: Phantom or reality? Hum. Vaccines Immunother. 2016, 12, 1070–1079. [Google Scholar] [CrossRef] [PubMed]
- Al-Halifa, S.; Gauthier, L.; Arpin, D.; Bourgault, S.; Archambault, D. Nanoparticle-Based Vaccines against Respiratory Viruses. Front. Immunol. 2019, 10, 22. [Google Scholar] [CrossRef]
- Rioux, G.; Mathieu, C.; Russell, A.; Bolduc, M.; Laliberte-Gagne, M.E.; Savard, P.; Leclerc, D. PapMV nanoparticles improve mucosal immune responses to the trivalent inactivated flu vaccine. J. Nanobiotechnol. 2014, 12, 19. [Google Scholar] [CrossRef]
- Hiremath, J.; Kang, K.I.; Xia, M.; Elaish, M.; Binjawadagi, B.; Ouyang, K.; Dhakal, S.; Arcos, J.; Torrelles, J.B.; Jiang, X.; et al. Entrapment of H1N1 Influenza Virus Derived Conserved Peptides in PLGA Nanoparticles Enhances T Cell Response and Vaccine Efficacy in Pigs. PLoS ONE 2016, 11, e0151922. [Google Scholar] [CrossRef]
- Karch, C.P.; Li, J.; Kulangara, C.; Paulillo, S.M.; Raman, S.K.; Emadi, S.; Tan, A.; Helal, Z.H.; Fan, Q.; Khan, M.I.; et al. Vaccination with self-adjuvanted protein nanoparticles provides protection against lethal influenza challenge. Nanomedicine 2017, 13, 241–251. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.; Webster, R.G.; Webby, R.J. Influenza Virus: Dealing with a Drifting and Shifting Pathogen. Viral Immunol. 2018, 31, 174–183. [Google Scholar] [CrossRef]
- Ohmit, S.E.; Petrie, J.G.; Cross, R.T.; Johnson, E.; Monto, A.S. Influenza hemagglutination-inhibition antibody titer as a correlate of vaccine-induced protection. J. Infect. Dis. 2011, 204, 1879–1885. [Google Scholar] [CrossRef]
- Corti, D.; Voss, J.; Gamblin, S.J.; Codoni, G.; Macagno, A.; Jarrossay, D.; Vachieri, S.G.; Pinna, D.; Minola, A.; Vanzetta, F.; et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 2011, 333, 850–856. [Google Scholar] [CrossRef]
- Nachbagauer, R.; Feser, J.; Naficy, A.; Bernstein, D.I.; Guptill, J.; Walter, E.B.; Berlanda-Scorza, F.; Stadlbauer, D.; Wilson, P.C.; Aydillo, T.; et al. A chimeric hemagglutinin-based universal influenza virus vaccine approach induces broad and long-lasting immunity in a randomized, placebo-controlled phase I trial. Nat. Med. 2021, 27, 106–114. [Google Scholar] [CrossRef] [PubMed]
- Sautto, G.A.; Kirchenbaum, G.A.; Abreu, R.B.; Ecker, J.W.; Pierce, S.R.; Kleanthous, H.; Ross, T.M. A Computationally Optimized Broadly Reactive Antigen Subtype-Specific Influenza Vaccine Strategy Elicits Unique Potent Broadly Neutralizing Antibodies against Hemagglutinin. J. Immunol. 2020, 204, 375–385. [Google Scholar] [CrossRef]
- Van Doorn, E.; Liu, H.; Ben-Yedidia, T.; Hassin, S.; Visontai, I.; Norley, S.; Frijlink, H.W.; Hak, E. Evaluating the immunogenicity and safety of a BiondVax-developed universal influenza vaccine (Multimeric-001) either as a standalone vaccine or as a primer to H5N1 influenza vaccine: Phase IIb study protocol. Medicine 2017, 96, e6339. [Google Scholar] [CrossRef] [PubMed]
- Atsmon, J.; Kate-Ilovitz, E.; Shaikevich, D.; Singer, Y.; Volokhov, I.; Haim, K.Y.; Ben-Yedidia, T. Safety and immunogenicity of multimeric-001—A novel universal influenza vaccine. J. Clin. Immunol. 2012, 32, 595–603. [Google Scholar] [CrossRef] [PubMed]
- Preiss, S.; Garçon, N.; Cunningham, A.L.; Strugnell, R.; Friedland, L.R. Vaccine provision: Delivering sustained & widespread use. Vaccine 2016, 34, 6665–6671. [Google Scholar] [CrossRef] [PubMed]
- Dey, A.K.; Malyala, P.; Singh, M. Physicochemical and functional characterization of vaccine antigens and adjuvants. Expert Rev. Vaccines 2014, 13, 671–685. [Google Scholar] [CrossRef]
- Kon, T.C.; Onu, A.; Berbecila, L.; Lupulescu, E.; Ghiorgisor, A.; Kersten, G.F.; Cui, Y.-Q.; Amorij, J.-P.; Van der Pol, L. Influenza Vaccine Manufacturing: Effect of Inactivation, Splitting and Site of Manufacturing. Comparison of Influenza Vaccine Production Processes. PLoS ONE 2016, 11, e0150700. [Google Scholar] [CrossRef]
- Pedersen, J.C. Hemagglutination-inhibition assay for influenza virus subtype identification and the detection and quantitation of serum antibodies to influenza virus. Methods Mol. Biol. 2014, 1161, 11–25. [Google Scholar] [CrossRef]
- Kaufmann, L.; Syedbasha, M.; Vogt, D.; Hollenstein, Y.; Hartmann, J.; Linnik, J.E.; Egli, A. An Optimized Hemagglutination Inhibition (HI) Assay to Quantify Influenza-specific Antibody Titers. J. Vis. Exp. 2017, 55833. [Google Scholar] [CrossRef]
- Defang, G.N.; Martin, N.J.; Burgess, T.H.; Millar, E.V.; Pecenka, L.A.; Danko, J.R.; Arnold, J.C.; Kochel, T.J.; Luke, T.C. Comparative Analysis of Hemagglutination Inhibition Titers Generated Using Temporally Matched Serum and Plasma Samples. PLoS ONE 2012, 7, e48229. [Google Scholar] [CrossRef]
- Wood, J.M.; Weir, J.P. Standardisation of inactivated influenza vaccines-Learning from history. Influenza Other Respir. Viruses 2018, 12, 195–201. [Google Scholar] [CrossRef]
- Minor, P.D. Assaying the Potency of Influenza Vaccines. Vaccines 2015, 3, 90–104. [Google Scholar] [CrossRef]
- Engelhardt, O.G.; Edge, C.; Dunleavy, U.; Guilfoyle, K.; Harvey, R.; Major, D.; Newman, R.; Penn, R.; Skeldon, S.; Storey, C.; et al. Comparison of single radial immunodiffusion, SDS-PAGE and HPLC potency assays for inactivated influenza vaccines shows differences in ability to predict immunogenicity of haemagglutinin antigen. Vaccine 2018, 36, 4339–4345. [Google Scholar] [CrossRef]
- Cole, J.L.; Lary, J.W.; P Moody, T.; Laue, T.M. Analytical ultracentrifugation: Sedimentation velocity and sedimentation equilibrium. Methods Cell Biol. 2008, 84, 143–179. [Google Scholar] [CrossRef]
- Weigel, T.; Soliman, R.; Wolff, M.W.; Reichl, U. Hydrophobic-interaction chromatography for purification of influenza A and B virus. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 2019, 1117, 103–117. [Google Scholar] [CrossRef]
- Shytuhina, A.; Pristatsky, P.; He, J.; Casimiro, D.R.; Schwartz, R.M.; Hoang, V.M.; Ha, S. Development and application of a reversed-phase high-performance liquid chromatographic method for quantitation and characterization of a Chikungunya virus-like particle vaccine. J. Chromatogr. A 2014, 1364, 192–197. [Google Scholar] [CrossRef]
- Rustandi, R.R.; Wang, F.; Lancaster, C.; Kristopeit, A.; Thiriot, D.S.; Heinrichs, J.H. Ion-Exchange Chromatography to Analyze Components of a Clostridium difficile Vaccine. Methods Mol. Biol. 2016, 1476, 269–277. [Google Scholar] [CrossRef] [PubMed]
- Lancaster, C.; Rustandi, R.R.; Pannizzo, P.; Ha, S. A Size-Exclusion Chromatography Method for Analysis of Clostridium difficile Vaccine Toxins. Methods Mol. Biol. 2016, 1476, 279–287. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Vandersluis, M.; Stout, J.; Haupts, U.; Sanders, M.; Jacquemart, R. Affinity chromatography for vaccines manufacturing: Finally ready for prime time? Vaccine 2019, 37, 5491–5503. [Google Scholar] [CrossRef]
- Vajda, J.; Weber, D.; Brekel, D.; Hundt, B.; Müller, E. Size distribution analysis of influenza virus particles using size exclusion chromatography. J. Chromatogr. A 2016, 1465, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Tay, T.; Agius, C.; Hamilton, R.; Bodle, J.; Rockman, S. Investigation into alternative testing methodologies for characterization of influenza virus vaccine. Hum. Vaccines Immunother. 2015, 11, 1673–1684. [Google Scholar] [CrossRef][Green Version]
- Sahin, Z.; Neeleman, R.; Haines, J.; Kayser, V. Preparation-free method can enable rapid surfactant screening during industrial processing of influenza vaccines. Vaccine 2019, 37, 1073–1079. [Google Scholar] [CrossRef] [PubMed]
- Sahin, Z.; Akkoc, S.; Neeleman, R.; Haines, J.; Kayser, V. Nile Red fluorescence spectrum decomposition enables rapid screening of large protein aggregates in complex biopharmaceutical formulations like influenza vaccines. Vaccine 2017, 35, 3026–3032. [Google Scholar] [CrossRef]
- Downard, K.M.; Morrissey, B.; Schwahn, A.B. Mass spectrometry analysis of the influenza virus. Mass Spectrom. Rev. 2009, 28, 35–49. [Google Scholar] [CrossRef] [PubMed]
- Frahm, G.E.; Pochopsky, A.W.T.; Clarke, T.M.; Johnston, M.J.W. Evaluation of Microflow Digital Imaging Particle Analysis for Sub-Visible Particles Formulated with an Opaque Vaccine Adjuvant. PLoS ONE 2016, 11, e0150229. [Google Scholar] [CrossRef]
- Noble, J.E.; Bailey, M.J. Quantitation of protein. Methods Enzym. 2009, 463, 73–95. [Google Scholar] [CrossRef]
- Rhodes, D.G.; Bossio, R.E.; Laue, T.M. Determination of size, molecular weight, and presence of subunits. Methods Enzym. 2009, 463, 691–723. [Google Scholar] [CrossRef]
- Liu, J.; Andya, J.D.; Shire, S.J. A critical review of analytical ultracentrifugation and field flow fractionation methods for measuring protein aggregation. AAPS J. 2006, 8, E580–E589. [Google Scholar] [CrossRef]
- Baldwin, M.A.; Medzihradszky, K.F.; Lock, C.M.; Fisher, B.; Settineri, T.A.; Burlingame, A.L. Matrix-assisted laser desorption/ionization coupled with quadrupole/orthogonal acceleration time-of-flight mass spectrometry for protein discovery, identification, and structural analysis. Anal. Chem. 2001, 73, 1707–1720. [Google Scholar] [CrossRef]
- Tarasov, M.; Shanko, A.; Kordyukova, L.; Katlinski, A. Characterization of Inactivated Influenza Vaccines Used in the Russian National Immunization Program. Vaccines 2020, 8, 488. [Google Scholar] [CrossRef]
- Filipe, V.; Hawe, A.; Carpenter, J.F.; Jiskoot, W. Analytical approaches to assess the degradation of therapeutic proteins. TrAC Trends Anal. Chem. 2013, 49, 118–125. [Google Scholar] [CrossRef]
- Lewnard, J.A.; Cobey, S. Immune History and Influenza Vaccine Effectiveness. Vaccines 2018, 6, 28. [Google Scholar] [CrossRef] [PubMed]
- CDC. CDC Seasonal Flu Vaccine Effectiveness Studies. Available online: https://www.cdc.gov/flu/vaccines-work/effectiveness-studies.htm (accessed on 12 September 2021).
- Sullivan, S.G.; Price, O.H.; Regan, A.K. Burden, effectiveness and safety of influenza vaccines in elderly, paediatric and pregnant populations. Ther. Adv. Vaccines Immunother. 2019, 7. [Google Scholar] [CrossRef]
- Dhakal, S.; Klein, S.L. Host Factors Impact Vaccine Efficacy: Implications for Seasonal and Universal Influenza Vaccine Programs. J. Virol. 2019, 93, e00797-19. [Google Scholar] [CrossRef]
- DiazGranados, C.A.; Dunning, A.J.; Kimmel, M.; Kirby, D.; Treanor, J.; Collins, A.; Pollak, R.; Christoff, J.; Earl, J.; Landolfi, V.; et al. Efficacy of High-Dose versus Standard-Dose Influenza Vaccine in Older Adults. N. Engl. J. Med. 2014, 371, 635–645. [Google Scholar] [CrossRef]
- Arnou, R.; Icardi, G.; De Decker, M.; Ambrozaitis, A.; Kazek, M.P.; Weber, F.; Van Damme, P. Intradermal influenza vaccine for older adults: A randomized controlled multicenter phase III study. Vaccine 2009, 27, 7304–7312. [Google Scholar] [CrossRef]

| No | Tradename | Manufacturer | Age Group | Country | References |
|---|---|---|---|---|---|
| Inactivated Influenza Vaccine, Split Virion, Egg-Based | |||||
| 1 | Afluria Quadrivalent | Seqirus Pty. Ltd. | Six months and over | US | [72] |
| 2 | Fluarix Quadrivalent | GlaxoSmithKline Biologicals | Six months and over | US | [73] |
| 3 | FluLaval Quadrivalent | ID Biomedical Corporation of Quebec | Six months and over | US | [74] |
| 4 | Fluzone Highdose Quadrivalent | Sanofi Pasteur, Inc. | 65 years and over | US | [75] |
| 5 | Fluzone Quadrivalent | Sanofi Pasteur, Inc. | Six months and over | US | [76] |
| 6 | FluQuadri | Sanofi-Aventis Australia | Six months and over | Australia | [77] |
| 7 | Vaxigrip Tetra | Sanofi-Aventis Australia | Six months and over | Australia | [78] |
| 8 | Fluarix Tetra | GlaxoSmithKline Biologicals | Six months and over | Australia | [74] |
| 9 | Afluria Quadrivalent | Seqirus Pty. Ltd. | Five years and over | Australia | [79] |
| 10 | Influvac Tetra | Mylan Health | Three years and over | Australia | [79] |
| Inactivated Influenza Vaccine, Surface Antigen, Adjuvanted, Egg-Based | |||||
| 11 | Fluad | Seqirus, Inc. | 65 years and over | US | [80] |
| 12 | Fluad Quadrivalent | Seqirus, Inc. | 65 years and over | US | [81] |
| 13 | Fluad Quad | Seqirus, Inc. | 65 years and over | Australia | [80] |
| Recombinant Vaccine | |||||
| 14 | Flublok Quadrivalent | Sanofi Pasteur, Inc. | 18 years and over | US | [82] |
| Inactivated Subunit Influenza Vaccine, Cell Culture-Based | |||||
| 15 | Flucelvax Quadrivalent | Seqirus, Inc. | Four years and over | US | [83] |
| Live Attenuated Influenza Vaccine | |||||
| 16 | FluMist Quadrivalent | MedImmune, LLC | Two years through 49 years | US | [84] |
| VLP Vaccine Candidate | Type of Response | Cross-Protection | Clinical Phase | Type of Species | Sponsor | References |
|---|---|---|---|---|---|---|
| BV VLP-HA-NA-M1 | Humoral and cellular | N/A | Preclinical | Mice | - | [132] |
| HBc VLP-M2e-HA2 (Tandiflu1) | Cellular | Yes | Preclinical | Mice | - | [133] |
| HBc VLP-M2e-NP | Humoral and cellular | Yes | Preclinical | Mice | - | [127] |
| Influenza VLP-HA (H1, H8, H13, H3, H4, H10) | Humoral | Yes | Preclinical | Mice | - | [134] |
| Recombinant A (H1N1) 2009 influenza VLP vaccine (HA) | Humoral | N/A | Phase II | Human | Novavax | [135] |
| HA and M2e5x | Humoral and cellular | N/A | Preclinical | Mice | - | [136] |
| Plant-based QVLP (HA) | Humoral and cellular | Yes | Phase I &II | Human | Medicago | [131,137] |
| Vaccine Candidate | Vaccine Type | Manufacturer | Clinical Phase | Participants | Mechanism of Action |
|---|---|---|---|---|---|
| Chimeric HA proteins [151] | Hemagglutinin based | Glaxo-SmithKline | Phase I | 66 | Ag-specific cellular response Broadly cross-reactive Abs |
| Computationally optimized broadly reactive antigens (COBRA) [152] | Computationally optimized antigens | Sanofi-Pasteur | Preclinical | - | Elicitation of a unique broad cross-reactive and cross-neutralizing Ab against HA. |
| NP, M1 and HA peptides (Multimeric-001) [153] | Recombinant proteins | BiondVax Pharmaceuticals Ltd./NIAID | Phase III | 12463 | Cellular (B- and T-cell) immune response. |
| Assay or Technique | Use | References |
|---|---|---|
| Identity | ||
| Hemagglutinin inhibition assay (HI Test) | Selection of the candidate virus vaccine CVV and stability | [158,159,160] |
| Single radial immunodiffusion assay SRID | Vaccine potency | [161,162,163] |
| Enzyme-linked immunosorbent assay ELISA | ||
| Sedimentation | ||
| Velocity | Mass and size | [164] |
| Equilibrium | Mass, association | [164] |
| Chromatography | ||
| Hydrophobic interaction | Purification | [165] |
| Reverse phase | Purification and stability | [166] |
| Ion exchange | Charge | [167] |
| Size exclusion | Size | [168] |
| Affinity | Specific interaction | [169] |
| Light scattering | ||
| Dynamic light scattering DLS | Size measurements and aggregation study | [170,171] |
| Static light scattering | ||
| UV-Visible absorbance spectroscopy | Agglomeration assessment and stability | [171,172,173] |
| Mass spectrometry | Protein quantification, vaccine potency | [34,174] |
| Fluorescence spectroscopy | Protein stability, agglomeration assessment | [89,172] |
| Microscopy | ||
| Transmission electron microscopy TEM | Structure-guided information and aggregation study | [89,171,172] |
| Flow imaging microscopy | Size | [175] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nuwarda, R.F.; Alharbi, A.A.; Kayser, V. An Overview of Influenza Viruses and Vaccines. Vaccines 2021, 9, 1032. https://doi.org/10.3390/vaccines9091032
Nuwarda RF, Alharbi AA, Kayser V. An Overview of Influenza Viruses and Vaccines. Vaccines. 2021; 9(9):1032. https://doi.org/10.3390/vaccines9091032
Chicago/Turabian StyleNuwarda, Rina Fajri, Abdulsalam Abdullah Alharbi, and Veysel Kayser. 2021. "An Overview of Influenza Viruses and Vaccines" Vaccines 9, no. 9: 1032. https://doi.org/10.3390/vaccines9091032
APA StyleNuwarda, R. F., Alharbi, A. A., & Kayser, V. (2021). An Overview of Influenza Viruses and Vaccines. Vaccines, 9(9), 1032. https://doi.org/10.3390/vaccines9091032






