Balanced Cellular and Humoral Immune Responses Targeting Multiple Antigens in Adults Receiving a Quadrivalent Inactivated Influenza Vaccine
Abstract
1. Introduction
2. Materials and Methods
2.1. Longitudinal Study Design
2.2. Design of Antigens Utilized in the Evaluation of Cellular Immunity
2.3. Functional T Cell Assay
2.4. Detection of Antibodies to Seasonal Influenza Viruses and Vaccine
2.5. Statistical Analysis
2.6. Study Approval
3. Results
3.1. Vaccine Study Design
3.2. Strategy to Measure Cellular Responses
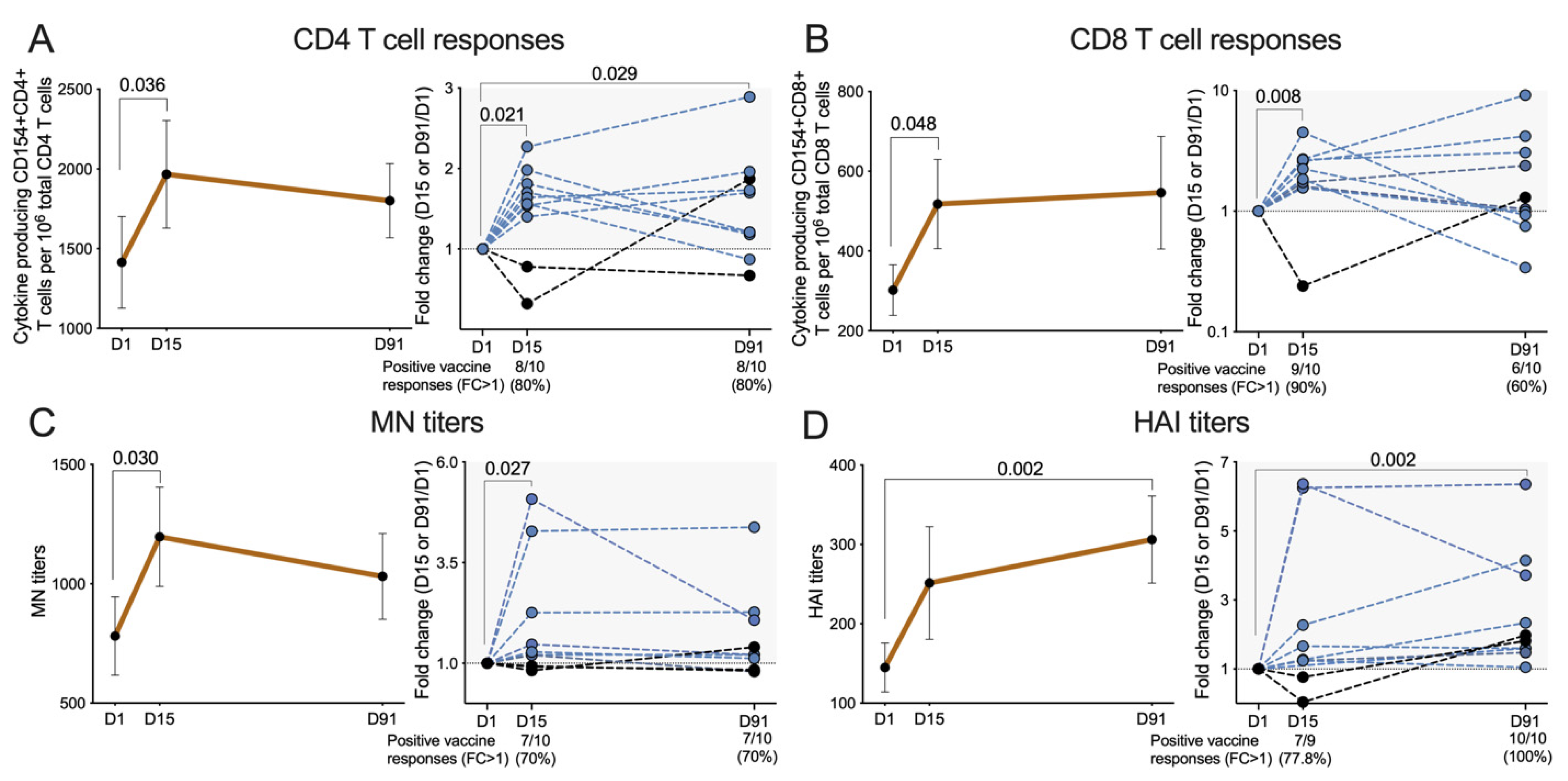
3.3. Overall CD4 T Cell Responses to the Flucelvax Quadrivalent Vaccine
3.4. Overall CD8 T Cell Responses to the Flucelvax Quadrivalent Vaccine
3.5. Overall Cellular Responses to PT and CMV Control Antigens
3.6. Serological Responses to Vaccination
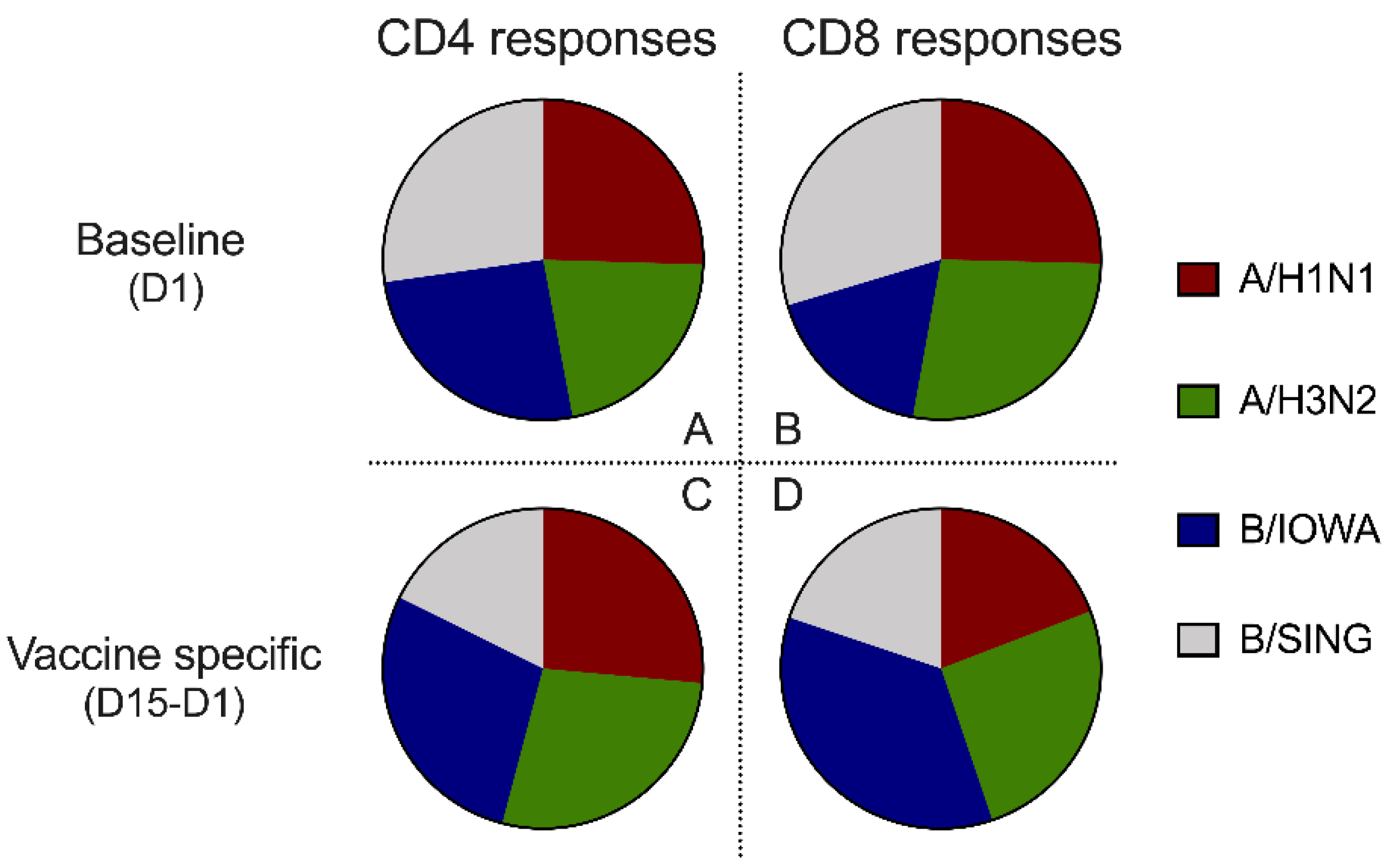
3.7. Balanced T Cell Reactivity at Baseline and Following Vaccination to the Four Different Strains
3.8. “Ceiling Effect” Observed in Both Cellular and Humoral Vaccine Responses
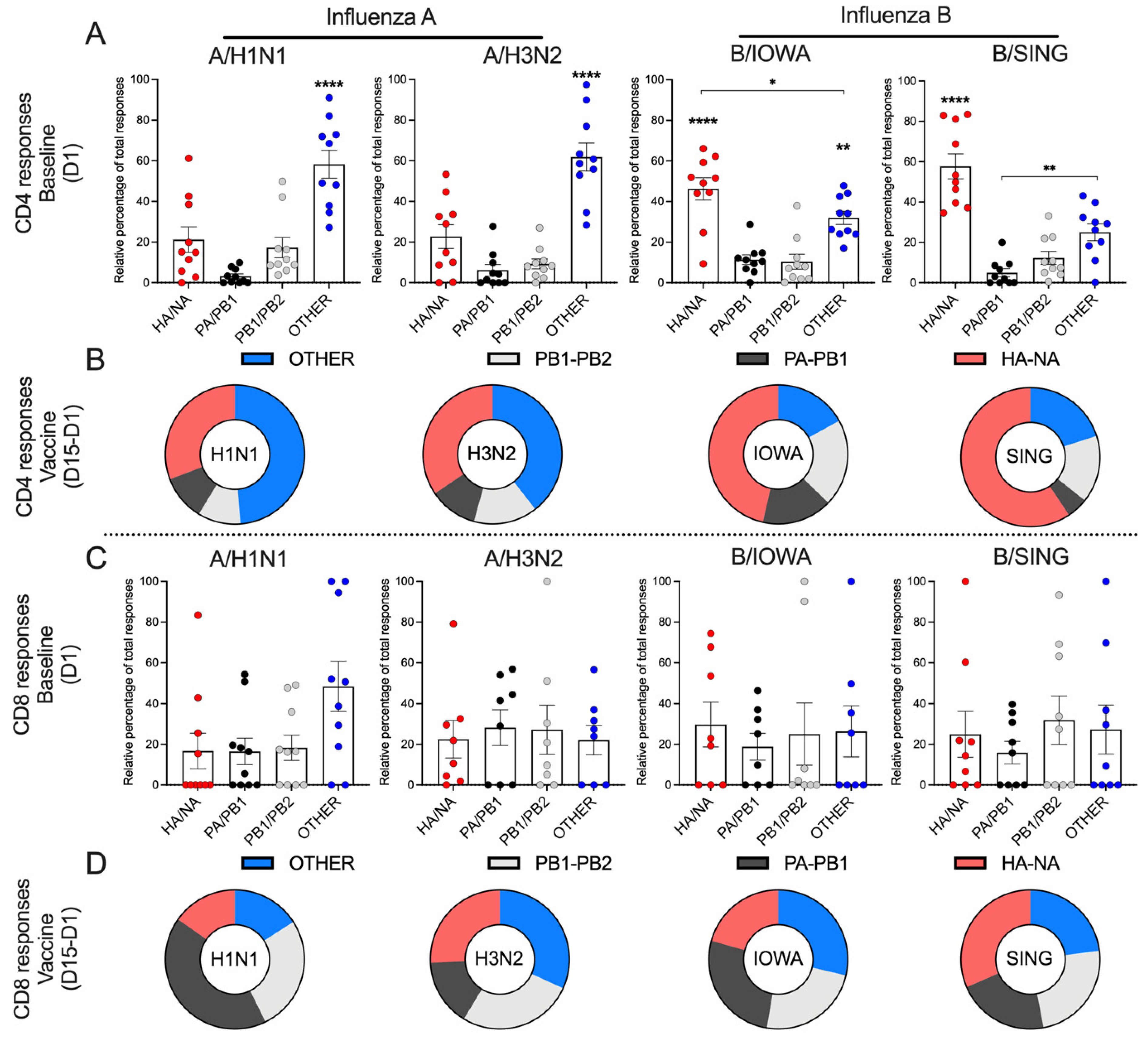
3.9. Type-Specific Immunodominance Patterns of CD4 and CD8 T Cell Responses
3.10. Patterns of Reactivity of CD4 and CD8 T Cell to Influenza A and B Viruses Following Vaccination
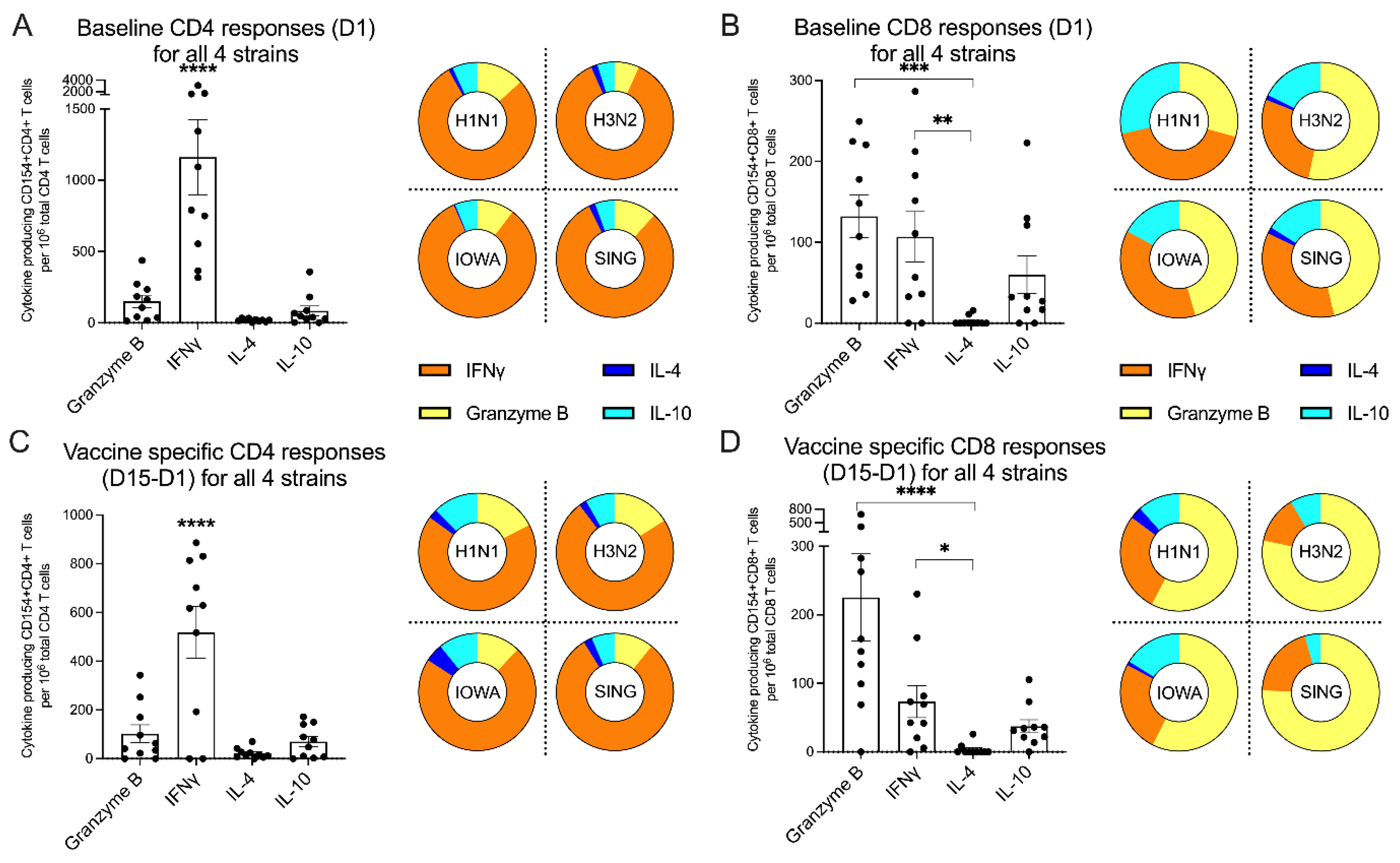
3.11. Differential Polarization of CD4 and CD8 Reactivity
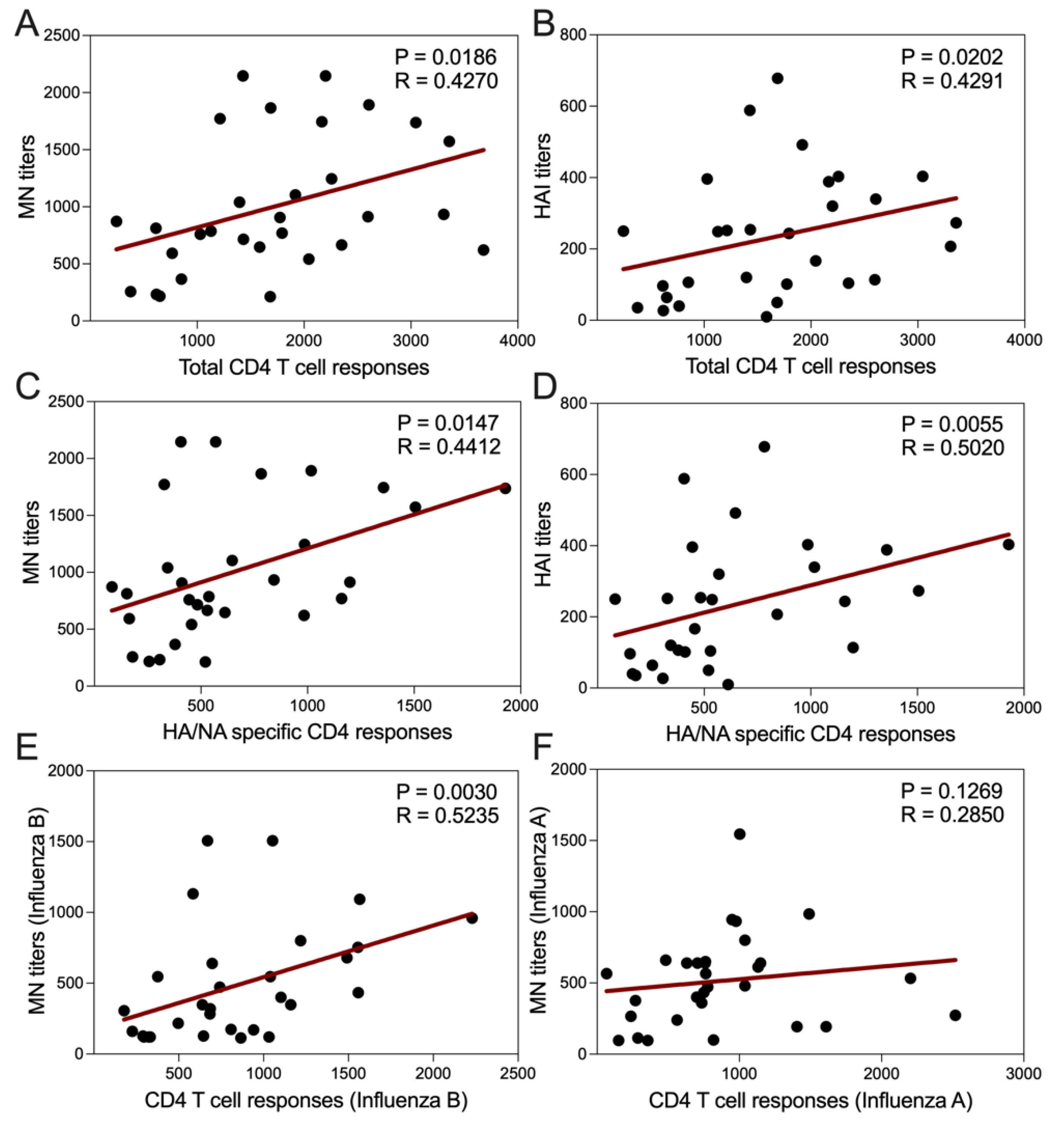
3.12. MN and HAI Titers Correlate with CD4 T Cell Reactivity
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lamb, R.A.; Choppin, P.W. The gene structure and replication of influenza virus. Annu. Rev. Biochem. 1983, 52, 467–506. [Google Scholar] [CrossRef]
- Kim, H.; Webster, R.G.; Webby, R.J. Influenza Virus: Dealing with a Drifting and Shifting Pathogen. Viral. Immunol. 2018, 31, 174–183. [Google Scholar] [CrossRef]
- World Health Organization. Recommended composition of Influenza Virus Vaccines for Use in the 2018–2019 Northern Hemisphere Influenza Season. 2018. Available online: https://www.who.int/influenza/vaccines/virus/recommendations/2018_19_north/en/ (accessed on 8 March 2021).
- European Medicines Agency. Flucelvax Tetra: Summary of Product Characteristics. 2018. Available online: http://www.ema.europa.eu/ (accessed on 8 March 2021).
- Flucelvax Quadrivalent (Influenza Vaccine) [Package Insert]; Seqirus, Inc.: Holly Springs, NC, USA, 2018.
- Seqirus Inc. Holly Springs (NC). Flucelvax Quadrivalent (Influenza Vaccine): Cell-based Vaccine for Prevention of Seasonal Influenza. 2018. Available online: https://flu.seqirus.com/FlucelvaxCategory/Flucelvax/p/FlucelvaxIn?seasonCategoryCode=Seqirus_InSeason (accessed on 8 March 2021).
- Richards, K.A.; Moritzky, S.; Shannon, I.; Fitzgerald, T.; Yang, H.; Branche, A.; Topham, D.J.; Treanor, J.J.; Nayak, J.; Sant, A.J. Recombinant HA-based vaccine outperforms split and subunit vaccines in elicitation of influenza-specific CD4 T cells and CD4 T cell-dependent antibody responses in humans. NPJ Vaccines 2020, 5, 77. [Google Scholar] [CrossRef] [PubMed]
- Lamb, Y.N. Cell-Based Quadrivalent Inactivated Influenza Virus Vaccine (Flucelvax((R)) Tetra/Flucelvax Quadrivalent((R))): A Review in the Prevention of Influenza. Drugs 2019, 79, 1337–1348. [Google Scholar] [CrossRef]
- Belongia, E.A.; Skowronski, D.M.; McLean, H.Q.; Chambers, C.; Sundaram, M.E.; De Serres, G. Repeated annual influenza vaccination and vaccine effectiveness: Review of evidence. Expert. Rev. Vaccines 2017, 16, 1–14. [Google Scholar] [CrossRef]
- Richards, K.A.; Shannon, I.; Treanor, J.J.; Yang, H.; Nayak, J.L.; Sant, A.J. Evidence That Blunted CD4 T-Cell Responses Underlie Deficient Protective Antibody Responses to Influenza Vaccines in Repeatedly Vaccinated Human Subjects. J. Infect. Dis. 2020, 222, 273–277. [Google Scholar] [CrossRef]
- Eichelberger, M.C.; Morens, D.M.; Taubenberger, J.K. Neuraminidase as an influenza vaccine antigen: A low hanging fruit, ready for picking to improve vaccine effectiveness. Curr. Opin. Immunol. 2018, 53, 38–44. [Google Scholar] [CrossRef] [PubMed]
- Greenshields-Watson, A.; Attaf, M.; MacLachlan, B.J.; Whalley, T.; Rius, C.; Wall, A.; Lloyd, A.; Hughes, H.; Strange, K.E.; Mason, G.H.; et al. CD4(+) T Cells Recognize Conserved Influenza A Epitopes through Shared Patterns of V-Gene Usage and Complementary Biochemical Features. Cell. Rep. 2020, 32, 107885. [Google Scholar] [CrossRef]
- Li, Z.T.; Zarnitsyna, V.I.; Lowen, A.C.; Weissman, D.; Koelle, K.; Kohlmeier, J.E.; Antia, R. Why Are CD8 T Cell Epitopes of Human Influenza A Virus Conserved? J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Wooden, S.L.; Koff, W.C. The Human Vaccines Project: Towards a comprehensive understanding of the human immune response to immunization. Hum. Vaccin. Immunother. 2018, 14, 2214–2216. [Google Scholar] [CrossRef] [PubMed]
- Carrasco Pro, S.; Sidney, J.; Paul, S.; Lindestam Arlehamn, C.; Weiskopf, D.; Peters, B.; Sette, A. Automatic Generation of Validated Specific Epitope Sets. J. Immunol. Res. 2015, 2015, 763461. [Google Scholar] [CrossRef]
- Weiskopf, D.; Cerpas, C.; Angelo, M.A.; Bangs, D.J.; Sidney, J.; Paul, S.; Peters, B.; Sanches, F.P.; Silvera, C.G.; Costa, P.R.; et al. Human CD8+ T-Cell Responses Against the 4 Dengue Virus Serotypes Are Associated With Distinct Patterns of Protein Targets. J. Infect. Dis. 2015, 212, 1743–1751. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Aevermann, B.D.; Anderson, T.K.; Burke, D.F.; Dauphin, G.; Gu, Z.; He, S.; Kumar, S.; Larsen, C.N.; Lee, A.J.; et al. Influenza Research Database: An integrated bioinformatics resource for influenza virus research. Nucleic Acids Res. 2017, 45, D466–D474. [Google Scholar] [CrossRef]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data—From vision to reality. Euro. Surveill. 2017, 22. [Google Scholar] [CrossRef]
- da Silva Antunes, R.; Quiambao, L.G.; Soldevila, F.; Sutherland, A.; Peters, B.; Sette, A. Lack of evidence supporting a role of IFN-beta and TGF-beta in differential polarization of Bordetella pertussis specific-T cell responses. Cytokine 2021, 137, 155313. [Google Scholar] [CrossRef]
- da Silva Antunes, R.; Quiambao, L.G.; Sutherland, A.; Soldevila, F.; Dhanda, S.K.; Armstrong, S.K.; Brickman, T.J.; Merkel, T.; Peters, B.; Sette, A. Development and Validation of a Bordetella pertussis Whole-Genome Screening Strategy. J. Immunol. Res. 2020, 2020, 8202067. [Google Scholar] [CrossRef]
- Mateus, J.; Grifoni, A.; Tarke, A.; Sidney, J.; Ramirez, S.I.; Dan, J.M.; Burger, Z.C.; Rawlings, S.A.; Smith, D.M.; Phillips, E.; et al. Selective and cross-reactive SARS-CoV-2 T cell epitopes in unexposed humans. Science 2020, 370, 89–94. [Google Scholar] [CrossRef] [PubMed]
- United States Census Bureau. Quick Facts: Tennessee. 2019. Available online: https://www.census.gov/quickfacts/TN (accessed on 8 March 2021).
- McMichael, A.J.; Gotch, F.; Cullen, P.; Askonas, B.; Webster, R.G. The human cytotoxic T cell response to influenza A vaccination. Clin. Exp. Immunol. 1981, 43, 276–284. [Google Scholar] [PubMed]
- Dan, J.M.; Lindestam Arlehamn, C.S.; Weiskopf, D.; da Silva Antunes, R.; Havenar-Daughton, C.; Reiss, S.M.; Brigger, M.; Bothwell, M.; Sette, A.; Crotty, S. A Cytokine-Independent Approach To Identify Antigen-Specific Human Germinal Center T Follicular Helper Cells and Rare Antigen-Specific CD4+ T Cells in Blood. J. Immunol. 2016, 197, 983–993. [Google Scholar] [CrossRef] [PubMed]
- Yam, K.K.; Gupta, J.; Brewer, A.; Scheifele, D.W.; Halperin, S.; Ward, B.J.; Public Health Agency of Canada/Canadian Institute of Health Research, I.R.N.R.T.S.I. Unusual patterns of IgG avidity in some young children following two doses of the adjuvanted pandemic H1N1 (2009) influenza virus vaccine. Clin. Vaccine Immunol. 2013, 20, 459–467. [Google Scholar] [CrossRef] [PubMed]
- Nayak, J.L.; Fitzgerald, T.F.; Richards, K.A.; Yang, H.; Treanor, J.J.; Sant, A.J. CD4+ T-cell expansion predicts neutralizing antibody responses to monovalent, inactivated 2009 pandemic influenza A(H1N1) virus subtype H1N1 vaccine. J. Infect. Dis. 2013, 207, 297–305. [Google Scholar] [CrossRef]
- Vidor, E. The nature and consequences of intra- and inter-vaccine interference. J. Comp. Pathol. 2007, 137 Suppl 1, S62–S66. [Google Scholar] [CrossRef]
- Dionne, B.; Brett, M.; Culbreath, K.; Mercier, R.C. Potential Ceiling Effect of Healthcare Worker Influenza Vaccination on the Incidence of Nosocomial Influenza Infection. Infect. Control Hosp. Epidemiol. 2016, 37, 840–844. [Google Scholar] [CrossRef]
- Sasaki, S.; He, X.S.; Holmes, T.H.; Dekker, C.L.; Kemble, G.W.; Arvin, A.M.; Greenberg, H.B. Influence of prior influenza vaccination on antibody and B-cell responses. PLoS ONE 2008, 3, e2975. [Google Scholar] [CrossRef]
- Bancroft, T.; Dillon, M.B.; da Silva Antunes, R.; Paul, S.; Peters, B.; Crotty, S.; Lindestam Arlehamn, C.S.; Sette, A. Th1 versus Th2 T cell polarization by whole-cell and acellular childhood pertussis vaccines persists upon re-immunization in adolescence and adulthood. Cell Immunol. 2016, 304–305, 35–43. [Google Scholar] [CrossRef] [PubMed]
- Frey, S.; Vesikari, T.; Szymczakiewicz-Multanowska, A.; Lattanzi, M.; Izu, A.; Groth, N.; Holmes, S. Clinical efficacy of cell culture-derived and egg-derived inactivated subunit influenza vaccines in healthy adults. Clin. Infect. Dis. 2010, 51, 997–1004. [Google Scholar] [CrossRef]
- Bart, S.; Cannon, K.; Herrington, D.; Mills, R.; Forleo-Neto, E.; Lindert, K.; Abdul Mateen, A. Immunogenicity and safety of a cell culture-based quadrivalent influenza vaccine in adults: A Phase III, double-blind, multicenter, randomized, non-inferiority study. Hum. Vaccin. Immunother. 2016, 12, 2278–2288. [Google Scholar] [CrossRef] [PubMed]
- Chung, J.R.; Flannery, B.; Thompson, M.G.; Gaglani, M.; Jackson, M.L.; Monto, A.S.; Nowalk, M.P.; Talbot, H.K.; Treanor, J.J.; Belongia, E.A.; et al. Seasonal Effectiveness of Live Attenuated and Inactivated Influenza Vaccine. Pediatrics 2016, 137, e20153279. [Google Scholar] [CrossRef]
- CDC (Centers for Disease Control and Prevention). ACIP Votes Down Use of LAIV for 2016–2017 Flu Season. 2016. Available online: https://www.cdc.gov/media/releases/2016/s0622-laiv-flu.html (accessed on 8 March 2021).
- Sridhar, S.; Begom, S.; Bermingham, A.; Hoschler, K.; Adamson, W.; Carman, W.; Bean, T.; Barclay, W.; Deeks, J.J.; Lalvani, A. Cellular immune correlates of protection against symptomatic pandemic influenza. Nat. Med. 2013, 19, 1305–1312. [Google Scholar] [CrossRef] [PubMed]
- He, X.S.; Holmes, T.H.; Zhang, C.; Mahmood, K.; Kemble, G.W.; Lewis, D.B.; Dekker, C.L.; Greenberg, H.B.; Arvin, A.M. Cellular immune responses in children and adults receiving inactivated or live attenuated influenza vaccines. J. Virol. 2006, 80, 11756–11766. [Google Scholar] [CrossRef]
- Schmidt, M.E.; Varga, S.M. The CD8 T Cell Response to Respiratory Virus Infections. Front. Immunol. 2018, 9, 678. [Google Scholar] [CrossRef]
- Awate, S.; Babiuk, L.A.; Mutwiri, G. Mechanisms of action of adjuvants. Front. Immunol. 2013, 4, 114. [Google Scholar] [CrossRef] [PubMed]
- Hayward, A.C.; Wang, L.; Goonetilleke, N.; Fragaszy, E.B.; Bermingham, A.; Copas, A.; Dukes, O.; Millett, E.R.; Nazareth, I.; Nguyen-Van-Tam, J.S.; et al. Natural T Cell-mediated Protection against Seasonal and Pandemic Influenza. Results of the Flu Watch Cohort Study. Am. J. Respir. Crit. Care Med. 2015, 191, 1422–1431. [Google Scholar] [CrossRef] [PubMed]
- CDC (Centers for Disease Control and Prevention). Update: Influenza Activity in the United States during the 2018–19 Season and Composition of the 2019–20 Influenza Vaccine. 2019. Available online: https://www.cdc.gov/mmwr/volumes/68/wr/mm6824a3.htm#:~:text=Using%20data%20available%20from%20October,deaths%20in%20the%20United%20States (accessed on 8 March 2021).
- Hensen, L.; Kedzierska, K.; Koutsakos, M. Innate and adaptive immunity toward influenza B viruses. Future Microbiol. 2020, 15, 1045–1058. [Google Scholar] [CrossRef] [PubMed]
| Participant ID | Age at Immunization | BMI | Gender | Race |
|---|---|---|---|---|
| 001 | 44 | 30.4 | Male | Caucasian |
| 002 | 47 | 24.5 | Female | Caucasian |
| 003 | 45 | 29.8 | Male | Caucasian |
| 004 | 23 | 25.8 | Female | Caucasian |
| 005 | 22 | 22.5 | Male | Caucasian |
| 006 | 33 | 22.1 | Female | Caucasian |
| 007 | 19 | 23.0 | Male | Caucasian |
| 008 | 33 | 21.6 | Female | Caucasian |
| 009 | 41 | 32.4 | Female | African American |
| 010 | 49 | 28.3 | Female | Caucasian |
| Total | 35.60 ± 3.54 | 26.04 ± 1.24 | 60% Female, 40% Male | 90% Caucasian, 10% African American |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yu, E.D.; Grifoni, A.; Sutherland, A.; Voic, H.; Wang, E.; Frazier, A.; Jimenez-Truque, N.; Yoder, S.; Welsh, S.; Wooden, S.; et al. Balanced Cellular and Humoral Immune Responses Targeting Multiple Antigens in Adults Receiving a Quadrivalent Inactivated Influenza Vaccine. Vaccines 2021, 9, 426. https://doi.org/10.3390/vaccines9050426
Yu ED, Grifoni A, Sutherland A, Voic H, Wang E, Frazier A, Jimenez-Truque N, Yoder S, Welsh S, Wooden S, et al. Balanced Cellular and Humoral Immune Responses Targeting Multiple Antigens in Adults Receiving a Quadrivalent Inactivated Influenza Vaccine. Vaccines. 2021; 9(5):426. https://doi.org/10.3390/vaccines9050426
Chicago/Turabian StyleYu, Esther Dawen, Alba Grifoni, Aaron Sutherland, Hannah Voic, Eric Wang, April Frazier, Natalia Jimenez-Truque, Sandra Yoder, Sabrina Welsh, Stacey Wooden, and et al. 2021. "Balanced Cellular and Humoral Immune Responses Targeting Multiple Antigens in Adults Receiving a Quadrivalent Inactivated Influenza Vaccine" Vaccines 9, no. 5: 426. https://doi.org/10.3390/vaccines9050426
APA StyleYu, E. D., Grifoni, A., Sutherland, A., Voic, H., Wang, E., Frazier, A., Jimenez-Truque, N., Yoder, S., Welsh, S., Wooden, S., Koff, W., Creech, B., Sette, A., & da Silva Antunes, R. (2021). Balanced Cellular and Humoral Immune Responses Targeting Multiple Antigens in Adults Receiving a Quadrivalent Inactivated Influenza Vaccine. Vaccines, 9(5), 426. https://doi.org/10.3390/vaccines9050426







