Emerging SARS-CoV-2 Variants: A Review of Its Mutations, Its Implications and Vaccine Efficacy
Abstract
:1. Introduction
2. Nomenclature of SARS-CoV-2 Variants
- Exhibits one or more shared nucleotide differences from the ancestral lineage
- Comprises at least five genomes with >95% of the genome sequenced
- Exhibits at least one shared nucleotide change among genomes within the lineage
- Demonstrate a bootstrap value >70% for the lineage-defining node.
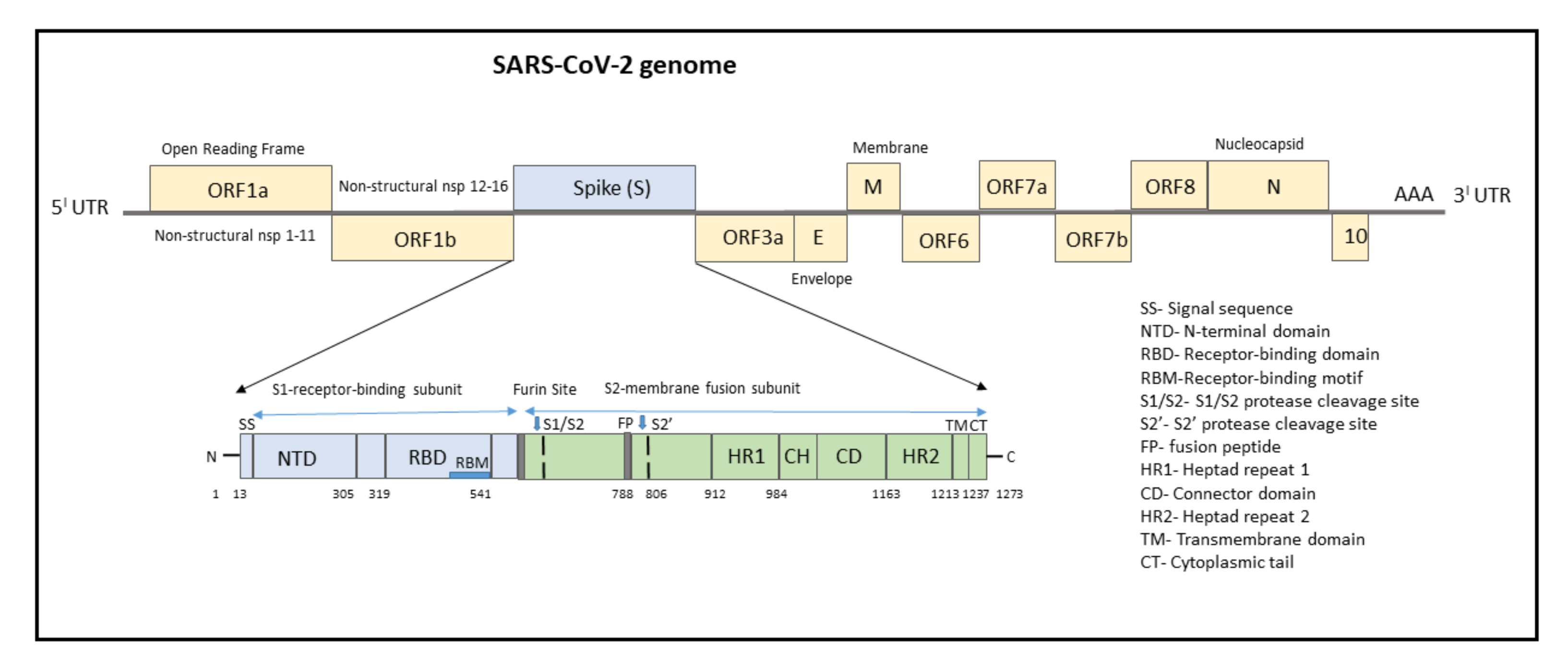
3. SARS-CoV-2 Genome
4. Mutations Associated with SARS-CoV-2 Genome
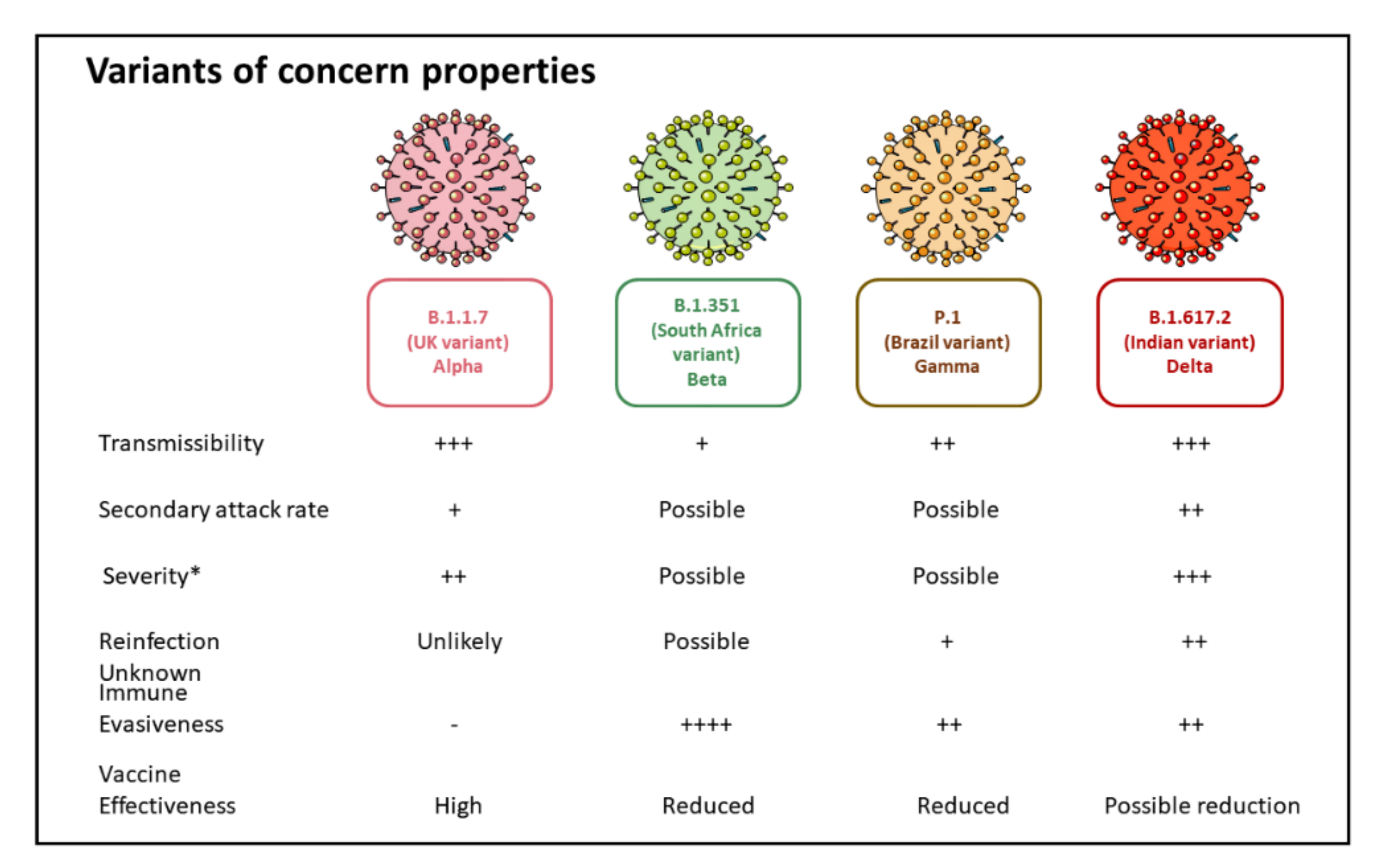
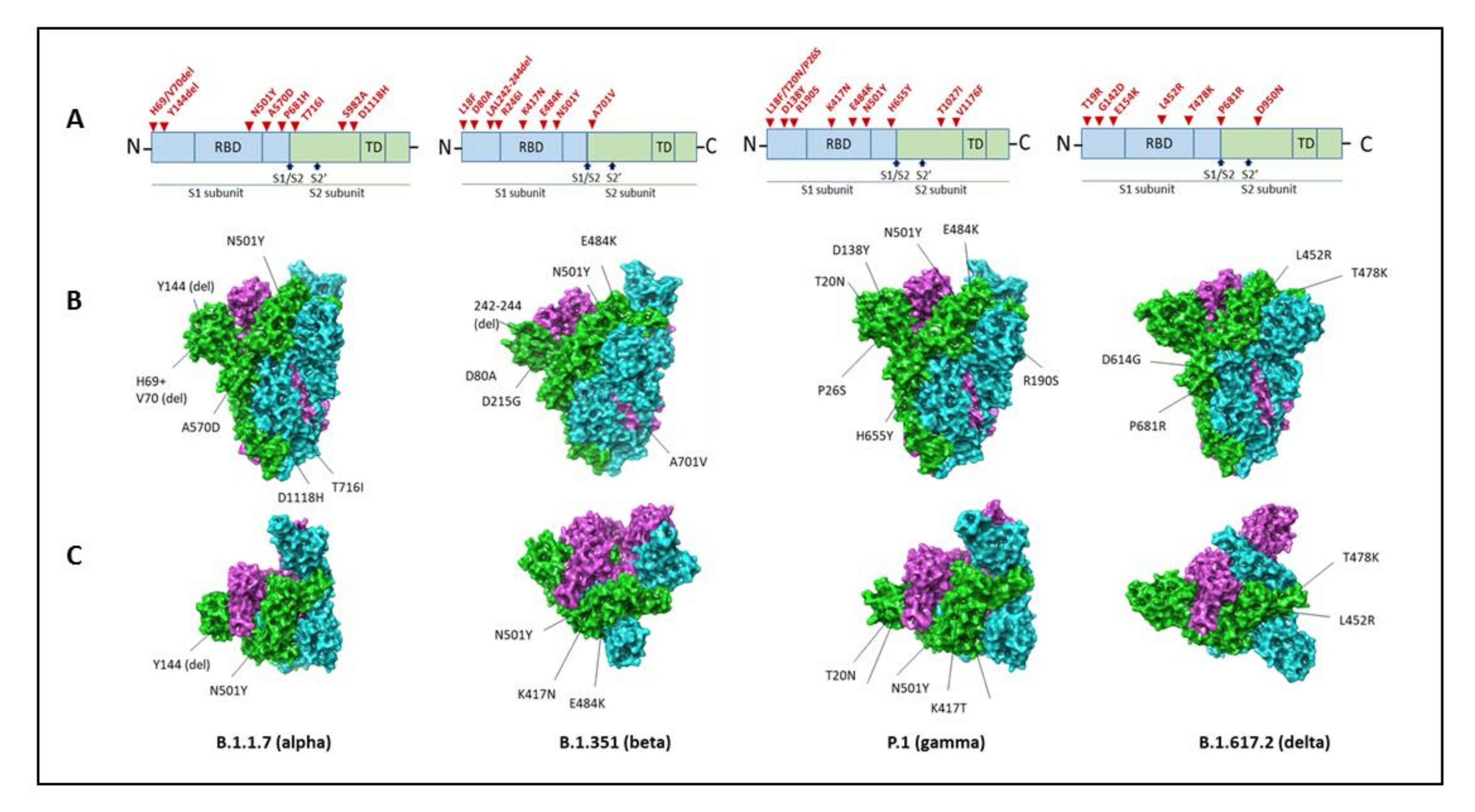
5. SARS-CoV-2 Variants of Concern
5.1. B.1.1.7 Variant
5.2. B.1.351 (20H/501Y.V2) Variant
5.3. P.1 (B.1.1.28.1) Variant
5.4. B.1.617 Variant (B.1.617.2 Variant)
| Pango Lineage/First Detected/ WHO Classification | Spike Protein Substitution | Other Mutations | Key Features |
|---|---|---|---|
| B.1.1.7 United Kingdom ALPHA (WHO) | Receptor-binding domain: N501Y Furin cleavage site: H69del, V70del, P681H Other mutations including N-terminal: A570D, T716I, S982A, D1118H Antibody supersite epitope: _Y144del, +/−E484K | NON- SYNONYMOUS Open reading frame (ORF)1ab: T1001I, A1708D, and I2230T. ORF8: Q27stop, R52I, and Y73C Nucleocapsid (N) protein: D3L and S235F SYNONYMOUS ORF1ab: C913T, C5986T, C14676T, C15279T, and T16176C Membrane (M) gene: T26801C DELETIONS ORF1ab: SGF 3675–3677del |
|
| B.1.351 South Africa BETA (WHO) | Receptor-binding domain: K417N, E484K, N501Y. N-terminal domain: L18F, D80A, D215G, R246I, D614G deletion at 242–244. S2 subunit: A701V. | NON- SYNONYMOUS ORF1ab: _T265I, _K1655N, H2799Y, S2900L, K3353R, D4527Y, T5912I ORF3a: Q57H, S71L Envelope (E): _P71L N viral protein: _T205I | |
| P.1 Brazil GAMMA (WHO) | Receptor-binding domain: K417N, E484K, N501Y N-terminal domain: L18F, T20N, P26S, D138Y, R190S, Carboxy(C)-terminal region of the S1 domain: D614G S1 and S2 site: _H655Y, T1027I, and V1176F | NON- SYNONYMOUS Open reading frame (ORF)1ab: S1188L, K1795Q, and E5665D ORF8: E92K Nucleocapsid (N) protein: P80K SYNONYMOUS ORF1ab: C913T, C5986T, C14676T, C15279T, and T16176C Membrane (M) gene: T26801C DELETIONS ORF1ab: _SGF 3675–3677del | |
| B.1.617.2 DELTA (WHO) | Receptor-binding domain: L452R, T478K Carboxy(C)-terminal region of the S1 domain: D614G Furin cleavage site: P681R | Spike protein: D950N, G142D, T19R, 156del, 157del, R185G |
6. SARS-CoV-2 Variants of Interest
7. Vaccines against SARS-CoV-2 Variants of Concern
7.1. Vaccine-Induced Immune Responses against SARS-CoV-2 Variants of Concern
7.1.1. Humoral Antibody Responses
| Pango Lineage/First Detected/WHO Classification | Spike Protein Substitution | Total Mutations | Key Features |
|---|---|---|---|
| B.1.525 UK/Nigeria ETA (WHO) | A67V, del69, del70, del144, E484K, D614G, Q677H, F888L | L5F, T95I, D253G, S477N, A701V ORF1a-nsp2: T85I ORF1a-nsp4: L438P ORF1a-nsp6: 9bp deletion, delta106–108. ORF1b-nsp12: P323L ORF1b-nsp13: Q88H ORF3a: Q57H N: P199L, M234I |
|
| B.1.526 New York, United States IOTA (WHO) | Receptor-binding domain: E484K, S477N Spike: L5F, T95I NTD: D253G Carboxy(C)-terminal region of the S1 domain: D614G S2 subunit: A701V. | A16500C, A22320G, T9867C | |
| B.1.617.1 India KAPPA (WHO) | T95I, G142D, E154K, L452R, E484Q, D614G, P681R, Q1071H. |
| |
| B.1.617.3 India | T19R, G142D, L452R, E484Q, D614G, P681R, D950N |
|
7.1.2. Cellular Antibody Responses
7.2. Efficacy of COVID-19 Vaccines against Variants of Concern
7.2.1. Vaccine Efficacy against B.1.1.7 Variant
7.2.2. Vaccine Efficacy against B.1.351 Variant
7.2.3. Vaccine Efficacy against the P.1 Variant
7.2.4. Vaccine Efficacy against the B.1.617.2 Variant
| Vaccine Name | Non-Variant and Variants of Concern | |||||
|---|---|---|---|---|---|---|
| Non-Variant | B.1.1.7 (Alpha) | B.1.351 (Beta) | P.1 (Gamma) | B.1.617.2 (Delta) | ||
| Pfizer Comirnaty (BNT162b2) | Overall efficacy: | 95% [165,166] | 90–93% 90% [167] Similar to non-variant [118,121] | 75% [165] | No evidence of reduced protection | 75–82% [139] Single dose: ~88% [168] |
| Asymptomatic disease: | N/A | Israel: 90.7–92.2% [159] | N/A | N/A | N/A | |
| Symptomatic disease: | Trial: 89–93.2% Israel CLALIT: 87–98% [169] England (80 + years): 85–93% [170] | Israel: 96.7–97.2% Canada: 89% [159] | Canada: 84% two doses [159] | Canada: 84% two doses [159] | Single dose: 33% Two doses: 88% [139] ~88% [164] Canada: 87% [159] | |
| Severe disease: | UK Siren: 95.3–100% [171] Israel CLALIT:75–100% [169] England (80 + years): 37–62% lower risk of mortality 80 + years [172] Israel (85 + years): 97% 7 days after 2nd-dose decreased mortality [173] | Qatar: 81.7–100% [165] | Qatar: 92.2–99.5% [165] | N/A | ||
| Reduced hospitalization: | England (80 + years): 33–52% in 80 + years [172] Israel (85 + years): 96.9% in 85 + years [173] | Canada: 95% [159] | Canada: 95% (two doses) [159] | Canada: 95% (two doses) [159] | Single dose: 46–99% Two doses: 86–99% [164] Scotland: 92% [140] Canada: 78% (single dose) [159] | |
| Decreased infection: | UK Siren: 76–97% [171] Israel CLALIT: 88–95% [169] Israel: 94.1% in 85 + years [173] | Qatar: 85.9–92.3% [165] | Qatar: 70.5–78.9% [165] | N/A | ||
| Moderna COVID-19 vaccine (mRNA 1273) | Overall efficacy: | 94.1% [174] 89.3–96.8% [174] >90% [175] >65 years: 61.4–95.2% Adolescents: 100% [176] | N/A | 6.5-fold reduction in neutralizing antibodies in laboratory study [160] | 2.6-fold reduction in neutralizing antibodies in laboratory study [160] | |
| Asymptomatic disease: | Single dose: 80% after 14 days Two doses: 90% after 14 days [125] | N/A | N/A | N/A | N/A | |
| Symptomatic disease: | 89.3–96.8% (Trial) Single dose: 80% after 14 days Two doses: 90% after 14 days [177] | Canada: 92% [159] | Canada: One dose 77% [159] | Canada: One dose 77% [159] | Canada: One dose 72% [159] | |
| Severe disease: | 100% [174] >95% [175] | N/A | N/A | N/A | N/A | |
| Reduced hospitalization: | N/A | Canada: 94% (two doses) [159] | Canada: 89% (single dose) [159] | Canada: 89% (single dose) [159] | Canada: 96% (single dose) [159] | |
| AstraZeneca/Oxford COVID-19 Vaccine AstraZeneca (AZD1222); | Overall efficacy: | 55–81% [178] 58–95% in 65 + years (US Study) 27–90% after 35 + days in 70 + years [172] 60–81% in 80 + years [170] | ~75% [107,178] | ≤10% [130] | Antibody titers reduced by 2.9-fold [179] 40.7–69.7% (Trial, non-variant specific) | 53–66% [140] Single dose: 60–71% [164] |
| Symptomatic disease: | 70.7–91.4% [107] 68–82% [180] 55–81% [178] | 43.6–84.5% [107,181] Two doses: 66% [172] | -76.8–54.8% [130] Canada: One dose 48% [159] | Canada: One dose 48% [159] | Single dose: 33% Two doses: 60% [139] ~67% [164] Canada: One dose 67% [159] | |
| Severe disease: | 100% [180] | N/A | N/A | |||
| Reduced hospitalization: | 100% [180] 60–81% in >80years [170] 74–89% in 70–79 years [170] | Canada: Single dose 85% prevention [159] | Canada: Single dose 83% prevention [159] | Canada: Single dose 83% prevention [159] | Single dose: 51–83% Two doses: 75–97% [164] 92% [182] Scotland: 73 [140] Canada: 88% (single dose) [159] | |
| Decreased infection: | 42–60.1% [107] Single dose: 60–70% [183] | N/A | N/A | N/A | N/A | |
| Johnson and Johnson Ad26. CoV2-S | Overall efficacy: | 66% [184] | 30.3–95.3% [185] 70% [133] | 72% (USA) 66% (Latin America) 57% (South Africa) [133] | 68% [133] | 60% [185] |
| Symptomatic disease: | 58.2–81.7% [133] Moderate-severe: 59–73.4% [133] | N/A | 41.2–78.7% [132] | 48.8–80.7% [132] | ||
| Severe disease: | 9.4–99.7% [133] Severe-critical: 54.6–89.1% [133] 85.4% [184] | N/A | 46.2–95.4% [133] | 7.8–99.7% [133] | N/A | |
| Novavax NVX-CoV2373 | Overall efficacy: | 89% [186] | ~85.6% [187] ~86% [187,188] | ~49% [186] 60% [187] | No evidence yet | No evidence yet |
| Symptomatic disease: | 73.8–99.5% [188] 100% against severe disease and hospitalization [187,189] | 71.3–93.5% [188] | 28.4–63.1% [133] 6.1–72.8% [137] 60% [137] | |||
| Sputnik V (also known as Gam- Covid-Vac) | Overall efficacy: | ~91% [137] | No evidence yet | No evidence yet | ||
| Symptomatic disease: | 85.6–95.2% [190] | No evidence yet | No evidence yet | |||
| Severe disease: | 94.4–100% [191] | |||||
8. Countermeasures to Treat Anticipated Continued Viral Mutations
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- WHO Coronavirus Disease (COVID-19) Dashboard with Vaccination Data. Available online: https://covid19.who.int/region/amro/country/us (accessed on 24 September 2021).
- Cascella, M.; Rajnik, M.; Aleem, A.; Dulebohn, S.C.; Di Napoli, R. Features, Evaluation, and Treatment of Coronavirus (COVID-19). In StatPearls; StatPearls Publishing LLC: Treasure Island, FL, USA, 2021. [Google Scholar]
- Astrezeneca. The Natural Evolution of SARS-CoV-2: How Science Responds to These Challenges. Available online: https://www.astrazeneca.com/what-science-can-do/topics/disease-understanding/the-natural-evolution-of-sars-cov-2.html (accessed on 24 September 2021).
- Ju, B.; Zhang, Q.; Ge, J.; Wang, R.; Sun, J.; Ge, X.; Yu, J.; Shan, S.; Zhou, B.; Song, S.; et al. Human neutralizing antibodies elicited by SARS-CoV-2 infection. Nature 2020, 584, 115–119. [Google Scholar] [CrossRef]
- McCallum, M.; De Marco, A.; Lempp, F.A.; Tortorici, M.A.; Pinto, D.; Walls, A.C.; Beltramello, M.; Chen, A.; Liu, Z.; Zatta, F.; et al. N-terminal domain antigenic mapping reveals a site of vulnerability for SARS-CoV-2. Cell 2021, 184, 2332–2347.e16. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Jackson, C.B.; Mou, H.; Ojha, A.; Peng, H.; Quinlan, B.D.; Rangarajan, E.S.; Pan, A.; Vanderheiden, A.; Suthar, M.S.; et al. SARS-CoV-2 spike-protein D614G mutation increases virion spike density and infectivity. Nat. Commun. 2020, 11, 6013. [Google Scholar] [CrossRef]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking Changes in SARS-CoV-2 Spike: Evidence that D614G Increases Infectivity of the COVID-19 Virus. Cell 2020, 182, 812–827.e19. [Google Scholar] [CrossRef]
- Burki, T. Understanding variants of SARS-CoV-2. Lancet 2021, 397, 462. [Google Scholar] [CrossRef]
- CDC. SARS-CoV-2 Variant Classifications and Definitions. Available online: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant-info.html (accessed on 24 September 2021).
- Rambaut, A.; Holmes, E.C.; O’Toole, Á.; Hill, V.; McCrone, J.T.; Ruis, C.; du Plessis, L.; Pybus, O.G. A dynamic nomenclature proposal for SARS-CoV-2 lineages to assist genomic epidemiology. Nat. Microbiol. 2020, 5, 1403–1407. [Google Scholar] [CrossRef]
- CDC. About Variants of the Virus that Causes COVID-19. Available online: https://www.cdc.gov/coronavirus/2019-ncov/variants/variant.html (accessed on 24 September 2021).
- Snijder, E.J.; Bredenbeek, P.J.; Dobbe, J.C.; Thiel, V.; Ziebuhr, J.; Poon, L.L.; Guan, Y.; Rozanov, M.; Spaan, W.J.; Gorbalenya, A.E. Unique and conserved features of genome and proteome of SARS-coronavirus, an early split-off from the coronavirus group 2 lineage. J. Mol. Biol. 2003, 331, 991–1004. [Google Scholar] [CrossRef]
- Lehmann, K.C.; Gulyaeva, A.; Zevenhoven-Dobbe, J.C.; Janssen, G.M.; Ruben, M.; Overkleeft, H.S.; van Veelen, P.A.; Samborskiy, D.V.; Kravchenko, A.A.; Leontovich, A.M.; et al. Discovery of an essential nucleotidylating activity associated with a newly delineated conserved domain in the RNA polymerase-containing protein of all nidoviruses. Nucleic Acids Res. 2015, 43, 8416–8434. [Google Scholar] [CrossRef] [Green Version]
- Yang, Y.; Du, L. SARS-CoV-2 spike protein: A key target for eliciting persistent neutralizing antibiodies. Signal Transduct. Target. Ther. 2021, 26, 1–3. [Google Scholar] [CrossRef]
- Huang, Y.; Yang, C.; Xu, X.-F.; Liu, S.-W. Structural and functional properties of SARS-CoV-2 spike protein: Potential antivirus drug development for COVID-19. Acta Pharmacol. Sin. 2020, 41, 1141–1149. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Pöhlmann, S. A Multibasic Cleavage Site in the Spike Protein of SARS-CoV-2 Is Essential for Infection of Human Lung Cells. Mol. Cell 2020, 78, 779–784.e5. [Google Scholar] [CrossRef] [PubMed]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Broer, R.; Boson, B.; Spaan, W.; Cosset, F.L.; Corver, J. Important role for the transmembrane domain of severe acute respiratory syndrome coronavirus spike protein during entry. J. Virol. 2006, 80, 1302–1310. [Google Scholar] [CrossRef] [Green Version]
- Millet, J.K.; Whittaker, G.R. Physiological and molecular triggers for SARS-CoV membrane fusion and entry into host cells. Virology 2018, 517, 3–8. [Google Scholar] [CrossRef]
- Xia, S.; Zhu, Y.; Liu, M.; Lan, Q.; Xu, W.; Wu, Y.; Ying, T.; Liu, S.; Shi, Z.; Jiang, S.; et al. Fusion mechanism of 2019-nCoV and fusion inhibitors targeting HR1 domain in spike protein. Cell. Mol. Immunol. 2020, 17, 765–767. [Google Scholar] [CrossRef]
- Teoh, K.T.; Siu, Y.L.; Chan, W.L.; Schlüter, M.A.; Liu, C.J.; Peiris, J.S.; Bruzzone, R.; Margolis, B.; Nal, B. The SARS coronavirus E protein interacts with PALS1 and alters tight junction formation and epithelial morphogenesis. Mol. Biol. Cell 2010, 21, 3838–3852. [Google Scholar] [CrossRef] [Green Version]
- Schoeman, D.; Fielding, B.C. Is There a Link Between the Pathogenic Human Coronavirus Envelope Protein and Immunopathology? A Review of the Literature. Front. Microbiol. 2020, 11, 2086. [Google Scholar] [CrossRef]
- Astuti, I.; Ysrafil. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab. Syndr. 2020, 14, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Del Vecchio, C.; Brancaccio, G.; Brazzale, A.R.; Lavezzo, E.; Onelia, F.; Franchin, E.; Manuto, L.; Bianca, F.; Cianci, V.; Cattelan, A.; et al. Emergence of N antigen SARS-CoV-2 genetic variants escaping detection of antigenic tests. medRxiv 2021. [Google Scholar] [CrossRef]
- Ren, Y.; Shu, T.; Wu, D.; Mu, J.; Wang, C.; Huang, M.; Han, Y.; Zhang, X.-Y.; Zhou, W.; Qiu, Y.; et al. The ORF3a protein of SARS-CoV-2 induces apoptosis in cells. Cell. Mol. Immunol. 2020, 17, 881–883. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.S.; Choudhury, P.P.; Basu, P.; Jana, S.S. Molecular conservation and differential mutation on ORF3a gene in Indian SARS-CoV2 genomes. Genomics 2020, 112, 3226–3237. [Google Scholar] [CrossRef]
- Shah, A. Novel Coronavirus-Induced NLRP3 Inflammasome Activation: A Potential Drug Target in the Treatment of COVID-19. Front. Immunol. 2020, 11, 1021. [Google Scholar] [CrossRef]
- Yuen, C.K.; Lam, J.Y.; Wong, W.M.; Mak, L.F.; Wang, X.; Chu, H.; Cai, J.P.; Jin, D.Y.; To, K.K.; Chan, J.F.; et al. SARS-CoV-2 nsp13, nsp14, nsp15 and orf6 function as potent interferon antagonists. Emerg. Microbes Infect. 2020, 9, 1418–1428. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Chen, Y.; Li, Y.; Huang, F.; Luo, B.; Yuan, Y.; Xia, B.; Ma, X.; Yang, T.; Yu, F.; et al. The ORF8 protein of SARS-CoV-2 mediates immune evasion through down-regulating MHC-Ι. Proc. Natl. Acad. Sci. USA 2021, 118, e2024202118. [Google Scholar] [CrossRef]
- Jiang, H.W.; Zhang, H.N.; Meng, Q.F.; Xie, J.; Li, Y.; Chen, H.; Zheng, Y.X.; Wang, X.N.; Qi, H.; Zhang, J.; et al. SARS-CoV-2 Orf9b suppresses type I interferon responses by targeting TOM70. Cell. Mol. Immunol. 2020, 17, 998–1000. [Google Scholar] [CrossRef] [PubMed]
- Hassan, S.S.; Lundstrom, K.; Serrano-Aroca, Á.; Adadi, P.; Aljabali, A.; Redwan, E.; Lal, A.; Kandimalla, R.; El-Aziz, T.; Choudhury, P.; et al. Emergence of Unique SARS-CoV-2 ORF10 Variants and Their Impact on Protein Structure and Function. Preprints 2021, 2021070554. [Google Scholar] [CrossRef]
- Li, J.Y.; Liao, C.H.; Wang, Q.; Tan, Y.J.; Luo, R.; Qiu, Y.; Ge, X.Y. The ORF6, ORF8 and nucleocapsid proteins of SARS-CoV-2 inhibit type I interferon signaling pathway. Virus Res. 2020, 286, 198074. [Google Scholar] [CrossRef]
- Miorin, L.; Kehrer, T.; Sanchez-Aparicio, M.T.; Zhang, K.; Cohen, P.; Patel, R.S.; Cupic, A.; Makio, T.; Mei, M.; Moreno, E.; et al. SARS-CoV-2 Orf6 hijacks Nup98 to block STAT nuclear import and antagonize interferon signaling. Proc. Natl. Acad. Sci. USA 2020, 117, 28344–28354. [Google Scholar] [CrossRef]
- Li, T.; Zhang, Y.; Fu, L.; Yu, C.; Li, X.; Li, Y.; Zhang, X.; Rong, Z.; Wang, Y.; Ning, H.; et al. siRNA targeting the leader sequence of SARS-CoV inhibits virus replication. Gene 2005, 12, 751–761. [Google Scholar] [CrossRef] [Green Version]
- Kim, D.; Lee, J.Y.; Yang, J.S.; Kim, J.W.; Kim, V.N.; Chang, H. The Architecture of SARS-CoV-2 Transcriptome. Cell 2020, 181, 914–921.e10. [Google Scholar] [CrossRef]
- Hossain, M.U.; Bhattacharjee, A.; Emon, M.T.H.; Chowdhury, Z.M.; Ahammad, I.; Mosaib, M.G.; Moniruzzaman, M.; Rahman, M.H.; Islam, M.N.; Ahmed, I.; et al. Novel mutations in NSP-1 and PLPro of SARS-CoV-2 NIB-1 genome mount for effective therapeutics. J. Genet. Eng. Biotechnol. 2021, 19, 52. [Google Scholar] [CrossRef]
- Benedetti, F.; Snyder, G.A.; Giovanetti, M.; Angeletti, S.; Gallo, R.C.; Ciccozzi, M.; Zella, D. Emerging of a SARS-CoV-2 viral strain with a deletion in nsp1. J. Transl. Med. 2020, 18, 329. [Google Scholar] [CrossRef]
- Lin, J.-W.; Tang, C.; Wei, H.-C.; Du, B.; Chen, C.; Wang, M.; Zhou, Y.; Yu, M.-X.; Cheng, L.; Kuivanen, S.; et al. Genomic monitoring of SARS-CoV-2 uncovers an Nsp1 deletion variant that modulates type I interferon response. Cell Host Microbe 2021, 29, 489–502.e8. [Google Scholar] [CrossRef]
- Laha, S.; Chakraborty, J.; Das, S.; Manna, S.K.; Biswas, S.; Chatterjee, R. Characterizations of SARS-CoV-2 mutational profile, spike protein stability and viral transmission. Infect. Genet. Evol. 2020, 85, 104445. [Google Scholar] [CrossRef] [PubMed]
- Tomaszewski, T.; DeVries, R.S.; Dong, M.; Bhatia, G.; Norsworthy, M.D.; Zheng, X.; Caetano-Anollés, G. New pathways of mutational change in SARS-CoV-2 proteomes involve regions of intrinsic disorder important for virus replication and release. Evol. Bioinform. 2020, 16, 1176934320965149. [Google Scholar] [CrossRef]
- Nagy, Á.; Pongor, S.; Győrffy, B. Different mutations in SARS-CoV-2 associate with severe and mild outcome. Int. J. Antimicrob. Agents 2021, 57, 106272. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Yan, L.; Huang, Y.; Liu, F.; Zhao, Y.; Cao, L.; Wang, T.; Sun, Q.; Ming, Z.; Zhang, L. Structure of the RNA-dependent RNA polymerase from COVID-19 virus. Science 2020, 368, 779–782. [Google Scholar] [CrossRef] [Green Version]
- Wang, R.; Chen, J.; Gao, K.; Hozumi, Y.; Yin, C.; Wei, G. Characterizing SARS-CoV-2 mutations in the United States. Res. Sq. 2020. [Google Scholar] [CrossRef]
- Wang, R.; Chen, J.; Gao, K.; Hozumi, Y.; Yin, C.; Wei, G.-W. Analysis of SARS-CoV-2 mutations in the United States suggests presence of four substrains and novel variants. Commun. Biol. 2021, 4, 228. [Google Scholar] [CrossRef]
- Zeng, W.; Liu, G.; Ma, H.; Zhao, D.; Yang, Y.; Liu, M.; Mohammed, A.; Zhao, C.; Yang, Y.; Xie, J. Biochemical characterization of SARS-CoV-2 nucleocapsid protein. Biochem. Biophys. Res. Commun. 2020, 527, 618–623. [Google Scholar] [CrossRef] [PubMed]
- Shen, L.; Bard, J.D.; Triche, T.J.; Judkins, A.R.; Biegel, J.A.; Gai, X. Emerging variants of concern in SARS-CoV-2 membrane protein: A highly conserved target with potential pathological and therapeutic implications. Emerg. Microbes Infect. 2021, 10, 885–893. [Google Scholar] [CrossRef] [PubMed]
- Mou, K.; Abdalla, M.; Wei, D.Q.; Khan, M.T.; Lodhi, M.S.; Darwish, D.B.; Sharaf, M.; Tu, X. Emerging mutations in envelope protein of SARS-CoV-2 and their effect on thermodynamic properties. Inform. Med. Unlocked 2021, 25, 100675. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.S.; Hoque, M.N.; Islam, M.R.; Islam, I.; Mishu, I.D.; Rahaman, M.M.; Sultana, M.; Hossain, M.A. Mutational insights into the envelope protein of SARS-CoV-2. Gene Rep. 2021, 22, 100997. [Google Scholar]
- Ramanathan, M.; Ferguson, I.D.; Miao, W.; Khavari, P.A. SARS-CoV-2 B.1.1.7 and B.1.351 Spike variants bind human ACE2 with increased affinity. Lancet Infect. Dis. 2021, 21, 1070. [Google Scholar] [CrossRef]
- Greaney, A.J.; Loes, A.N.; Crawford, K.H.D.; Starr, T.N.; Malone, K.D.; Chu, H.Y.; Bloom, J.D. Comprehensive mapping of mutations in the SARS-CoV-2 receptor-binding domain that affect recognition by polyclonal human plasma antibodies. Cell Host Microbe 2021, 29, 463–476.e6. [Google Scholar] [CrossRef] [PubMed]
- Nelson, G.; Buzko, O.; Spilman, P.; Niazi, K.; Rabizadeh, S.; Soon-Shiong, P. Molecular dynamic simulation reveals E484K mutation enhances spike RBD-ACE2 affinity and the combination of E484K, K417N and N501Y mutations (501Y.V2 variant) induces conformational change greater than N501Y mutant alone, potentially resulting in an escape mutant. bioRxiv 2021. [Google Scholar] [CrossRef]
- Starr, T.N.; Greaney, A.J.; Dingens, A.S.; Bloom, J.D. Complete map of SARS-CoV-2 RBD mutations that escape the monoclonal antibody LY-CoV555 and its cocktail with LY-CoV016. Cell Rep. Med. 2021, 2, 100255. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beltran, W.F.; Lam, E.C.; St Denis, K.; Nitido, A.D.; Garcia, Z.H.; Hauser, B.M.; Feldman, J.; Pavlovic, M.N.; Gregory, D.J.; Poznansky, M.C.; et al. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 2021, 184, 2372–2383.e9. [Google Scholar] [CrossRef]
- Zhang, W.; Davis, B.D.; Chen, S.S.; Sincuir Martinez, J.M.; Plummer, J.T.; Vail, E. Emergence of a Novel SARS-CoV-2 Variant in Southern California. JAMA 2021, 325, 1324–1326. [Google Scholar] [CrossRef]
- Gaebler, C.; Wang, Z.; Lorenzi, J.C.C.; Muecksch, F.; Finkin, S.; Tokuyama, M.; Cho, A.; Jankovic, M.; Schaefer-Babajew, D.; Oliveira, T.Y.; et al. Evolution of antibody immunity to SARS-CoV-2. Nature 2021, 591, 639–644. [Google Scholar] [CrossRef]
- Kemp, S.; Datir, R.; Collier, D.; Ferreira, I.; Carabelli, A.; Harvey, W.; Robertson, D.; Gupta, R. Recurrent emergence and transmission of a SARS-CoV-2 Spike deletion ΔH69/ΔV70. bioRxiv 2020. [Google Scholar] [CrossRef]
- McCarthy, K.R.; Rennick, L.J.; Nambulli, S.; Robinson-McCarthy, L.R.; Bain, W.G.; Haidar, G.; Duprex, W.P. Recurrent deletions in the SARS-CoV-2 spike glycoprotein drive antibody escape. Science 2021, 371, 1139. [Google Scholar] [CrossRef]
- Tran, K.N.; Miller, C.; Waldman, S. The SARS-CoV-2 Variant and its Impact on Diagnostic Testing. Available online: https://health.ucdavis.edu/blog/lab-best-practice/the-sars-cov-2-variant-and-its-impact-on-diagnostic-testing/2021/01 (accessed on 13 October 2021).
- McCallum, M.; Bassi, J.; Marco, A.D.; Chen, A.; Walls, A.C.; Iulio, J.D.; Tortorici, M.A.; Navarro, M.-J.; Silacci-Fregni, C.; Saliba, C.; et al. SARS-CoV-2 immune evasion by the B.1.427/B.1.429 variant of concern. Science 2021, 373, 648–654. [Google Scholar] [CrossRef]
- Zhang, L.; Jackson, C.B.; Mou, H.; Ojha, A.; Rangarajan, E.S.; Izard, T.; Farzan, M.; Choe, H. The D614G mutation in the SARS-CoV-2 spike protein reduces S1 shedding and increases infectivity. bioRxiv 2020. [Google Scholar] [CrossRef]
- Braun, K.M.; Moreno, G.K.; Halfmann, P.J.; Hodcroft, E.B.; Baker, D.A.; Boehm, E.C.; Weiler, A.M.; Haj, A.K.; Hatta, M.; Chiba, S.; et al. Transmission of SARS-CoV-2 in domestic cats imposes a narrow bottleneck. PLoS Pathog. 2021, 17, e1009373. [Google Scholar] [CrossRef]
- Wu, K.; Werner, A.P.; Moliva, J.I.; Koch, M.; Choi, A.; Stewart-Jones, G.B.E.; Bennett, H.; Boyoglu-Barnum, S.; Shi, W.; Graham, B.S.; et al. mRNA-1273 vaccine induces neutralizing antibodies against spike mutants from global SARS-CoV-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- Galloway, S.E.; Paul, P.; MacCannell, D.R.; Johansson, M.A.; Brooks, J.T.; MacNeil, A.; Slayton, R.B.; Tong, S.; Silk, B.J.; Armstrong, G.L.; et al. Emergence of SARS-CoV-2 B.1.1.7 Lineage—United States, 29 December 2020–12 January 2021. MMWR Morb. Mortal. Wkly Rep. 2021, 70, 95–99. [Google Scholar] [CrossRef]
- Volz, E.; Mishra, S.; Chand, M.; Barrett, J.C.; Johnson, R.; Geidelberg, L.; Hinsley, W.R.; Laydon, D.J.; Dabrera, G.; O’Toole, Á.; et al. Assessing transmissibility of SARS-CoV-2 lineage B.1.1.7 in England. Nature 2021, 593, 266–269. [Google Scholar] [CrossRef] [PubMed]
- Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/en/activities/tracking-SARS-CoV-2-variants/ (accessed on 13 October 2021).
- Grint, D.J.; Wing, K.; Williamson, E.; McDonald, H.I.; Bhaskaran, K.; Evans, D.; Evans, S.J.; Walker, A.J.; Hickman, G.; Nightingale, E.; et al. Case fatality risk of the SARS-CoV-2 variant of concern B.1.1.7 in England, 16 November to 5 February. Eurosurveillance 2021, 26, 2100256. [Google Scholar] [CrossRef] [PubMed]
- Davies, N.G.; Jarvis, C.I.; van Zandvoort, K.; Clifford, S.; Sun, F.Y.; Funk, S.; Medley, G.; Jafari, Y.; Meakin, S.R.; Lowe, R.; et al. Increased mortality in community-tested cases of SARS-CoV-2 lineage B.1.1.7. Nature 2021, 593, 270–274. [Google Scholar] [CrossRef]
- Davies, N.G.; Barnard, R.C.; Jarvis, C.I.; Kucharski, A.J.; Munday, J.; Pearson, C.A.B.; Russell, T.W.; Tully, D.C.; Abbott, S.; Gimma, A.; et al. Estimated transmissibility and severity of novel SARS-CoV-2 Variant of Concern 202012/01 in England. medRxiv 2020. [Google Scholar] [CrossRef]
- Aleem, A.; Akbar Samad, A.B.; Slenker, A.K. Emerging Variants of SARS-CoV-2 And Novel Therapeutics against Coronavirus (COVID-19). In StatPearls; StatPearls Publishing LLC: Treasure Island, FL, USA, 2021. [Google Scholar]
- Buchan, S.A.; Tibebu, S.; Daneman, N.; Whelan, M.; Vanniyasingam, T.; Murti, M.; Brown, K.A. Increased household secondary attacks rates with Variant of Concern SARS-CoV-2 index cases. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef]
- Golubchik, T.; Lythgoe, K.A.; Hall, M.; Ferretti, L.; Fryer, H.R.; MacIntyre-Cockett, G.; de Cesare, M.; Trebes, A.; Piazza, P.; Buck, D.; et al. Early analysis of a potential link between viral load and the N501Y mutation in the SARS-COV-2 spike protein. medRxiv 2021, 1. [Google Scholar] [CrossRef]
- Gómez, C.E.; Perdiguero, B.; Esteban, M. Emerging SARS-CoV-2 Variants and Impact in Global Vaccination Programs against SARS-CoV-2/COVID-19. Vaccines 2021, 9, 243. [Google Scholar] [CrossRef] [PubMed]
- Gu, H.; Chen, Q.; Yang, G.; He, L.; Fan, H.; Deng, Y.Q.; Wang, Y.; Teng, Y.; Zhao, Z.; Cui, Y.; et al. Adaptation of SARS-CoV-2 in BALB/c mice for testing vaccine efficacy. Science 2020, 369, 1603–1607. [Google Scholar] [CrossRef] [PubMed]
- Meng, T.; Cao, H.; Zhang, H.; Kang, Z.; Xu, D.; Gong, H.; Wang, J.; Li, Z.; Cui, X.; Xu, H.; et al. The transmembrane serine protease inhibitors are potential antiviral drugs for 2019-nCoV targeting the insertion sequence-induced viral infectivity enhancement. bioRxiv 2020. [Google Scholar] [CrossRef]
- Thomson, E.C.; Rosen, L.E.; Shepherd, J.G.; Spreafico, R.; da Silva Filipe, A.; Wojcechowskyj, J.A.; Davis, C.; Piccoli, L.; Pascall, D.J.; Dillen, J.; et al. Circulating SARS-CoV-2 spike N439K variants maintain fitness while evading antibody-mediated immunity. Cell 2021, 184, 1171–1187.e20. [Google Scholar] [CrossRef] [PubMed]
- Graham, M.S.; Sudre, C.H.; May, A.; Antonelli, M.; Murray, B.; Varsavsky, T.; Kläser, K.; Canas, L.S.; Molteni, E.; Modat, M.; et al. Changes in symptomatology, reinfection, and transmissibility associated with the SARS-CoV-2 variant B.1.1.7: An ecological study. Lancet Public Health 2021, 6, e335–e345. [Google Scholar] [CrossRef]
- Faulkner, N.; Ng, K.W.; Wu, M.; Harvey, R.; Margaritis, M.; Paraskevopoulou, S.; Houlihan, C.F.; Hussain, S.; Greco, M.; Bolland, W.; et al. Reduced antibody cross-reactivity following infection with B.1.1.7 than with parental SARS-CoV-2 strains. Elife 2021, 10, e69317. [Google Scholar] [CrossRef] [PubMed]
- O’Toole, Á.; Hill, V.; Pybus, O.G.; Watts, A.; Bogoch, I.I.; Khan, K.; Messina, J.P.; Tegally, H.; Lessells, R.R.; Giandhari, J.; et al. Tracking the international spread of SARS-CoV-2 lineages B.1.1.7 and B.1.351/501Y-V2. Wellcome Open Res. 2021, 6, 121. [Google Scholar] [CrossRef]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Emergence and rapid spread of a new severe acute respiratory syndrome-related coronavirus 2 (SARS-CoV-2) lineage with multiple spike mutations in South Africa. medRxiv 2020, 10. [Google Scholar] [CrossRef]
- Tegally, H.; Wilkinson, E.; Giovanetti, M.; Iranzadeh, A.; Fonseca, V.; Giandhari, J.; Doolabh, D.; Pillay, S.; San, E.J.; Msomi, N.; et al. Detection of a SARS-CoV-2 variant of concern in South Africa. Nature 2021, 592, 438–443. [Google Scholar] [CrossRef]
- Hoffmann, M.; Arora, P.; Groß, R.; Seidel, A.; Hörnich, B.F.; Hahn, A.S.; Krüger, N.; Graichen, L.; Hofmann-Winkler, H.; Kempf, A.; et al. SARS-CoV-2 variants B.1.351 and P.1 escape from neutralizing antibodies. Cell 2021, 184, 2384–2393.e12. [Google Scholar] [CrossRef]
- Tada, T.; Dcosta, B.M.; Samanovic-Golden, M.; Herati, R.S.; Cornelius, A.; Mulligan, M.J.; Landau, N.R. Neutralization of viruses with European, South African, and United States SARS-CoV-2 variant spike proteins by convalescent sera and BNT162b2 mRNA vaccine-elicited antibodies. bioRxiv 2021. [Google Scholar] [CrossRef]
- Wang, P.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; Graham, B.S.; et al. Increased Resistance of SARS-CoV-2 Variants B.1.351 and B.1.1.7 to Antibody Neutralization. bioRxiv 2021. [Google Scholar] [CrossRef]
- Wang, Z.; Schmidt, F.; Weisblum, Y.; Muecksch, F.; Barnes, C.O.; Finkin, S.; Schaefer-Babajew, D.; Cipolla, M.; Gaebler, C.; Lieberman, J.A.; et al. mRNA vaccine-elicited antibodies to SARS-CoV-2 and circulating variants. Nature 2021, 592, 616–622. [Google Scholar] [CrossRef] [PubMed]
- Wibmer, C.K.; Ayres, F.; Hermanus, T.; Madzivhandila, M.; Kgagudi, P.; Oosthuysen, B.; Lambson, B.E.; de Oliveira, T.; Vermeulen, M.; van der Berg, K.; et al. SARS-CoV-2 501Y.V2 escapes neutralization by South African COVID-19 donor plasma. Nat. Med. 2021, 27, 622–625. [Google Scholar] [CrossRef] [PubMed]
- Cele, S.; Gazy, I.; Jackson, L.; Hwa, S.-H.; Tegally, H.; Lustig, G.; Giandhari, J.; Pillay, S.; Wilkinson, E.; Naidoo, Y.; et al. Escape of SARS-CoV-2 501Y.V2 variants from neutralization by convalescent plasma. medRxiv 2021. [Google Scholar] [CrossRef]
- European Centre for Disease Prevention and Control. SARS-CoV-2—Increased Circulation of Variants of Concern and Vaccine Rollout in the EU/EEA, 14th Update – 15 February 2021. ECDC: Stockholm. 2021. Available online: https://www.ecdc.europa.eu/sites/default/files/documents/RRA-covid-19-14th-update-15-feb-2021.pdf (accessed on 13 October 2021).
- CDC. Science Brief: Emerging SARS-CoV-2 Variants. Available online: https://www.cdc.gov/coronavirus/2019-ncov/science/science-briefs/scientific-brief-emerging-variants.html (accessed on 13 October 2021).
- Faria, N.R.; Claro, I.M.; Candido, D.; Moyses Franco, L.A.; Andrade, P.S.; Coletti, T.M.; Silva, C.A.M.; Sales, F.C.; Manuli, E.R.; Aguiar, R.S.; et al. Genomic characterisation of an emergent SARS-CoV-2 lineage in Manaus: Preliminary findings. Available online: https://virological.org/t/genomic-characterisation-of-an-emergent-sars-cov-2-lineage-in-manaus-preliminary-findings/586 (accessed on 13 October 2021).
- Sabino, E.C.; Buss, L.F.; Carvalho, M.P.S.; Prete, C.A., Jr.; Crispim, M.A.E.; Fraiji, N.A.; Pereira, R.H.M.; Parag, K.V.; da Silva Peixoto, P.; Kraemer, M.U.G.; et al. Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence. Lancet 2021, 397, 452–455. [Google Scholar] [CrossRef]
- Buss, L.F.; Prete, C.A., Jr.; Abrahim, C.M.M.; Mendrone, A., Jr.; Salomon, T.; de Almeida-Neto, C.; França, R.F.O.; Belotti, M.C.; Carvalho, M.; Costa, A.G.; et al. Three-quarters attack rate of SARS-CoV-2 in the Brazilian Amazon during a largely unmitigated epidemic. Science 2021, 371, 288–292. [Google Scholar] [CrossRef]
- Wang, P.; Casner, R.G.; Nair, M.S.; Wang, M.; Yu, J.; Cerutti, G.; Liu, L.; Kwong, P.D.; Huang, Y.; Shapiro, L.; et al. Increased Resistance of SARS-CoV-2 Variant P.1 to Antibody Neutralization. bioRxiv 2021. [Google Scholar] [CrossRef]
- Jangra, S.; Ye, C.; Rathnasinghe, R.; Stadlbauer, D.; Krammer, F.; Simon, V.; Martinez-Sobrido, L.; García-Sastre, A.; Schotsaert, M. The E484K mutation in the SARS-CoV-2 spike protein reduces but does not abolish neutralizing activity of human convalescent and post-vaccination sera. medRxiv 2021. [Google Scholar] [CrossRef]
- Tchesnokova, V.; Kulakesara, H.; Larson, L.; Bowers, V.; Rechkina, E.; Kisiela, D.; Sledneva, Y.; Choudhury, D.; Maslova, I.; Deng, K.; et al. Acquisition of the L452R mutation in the ACE2-binding interface of Spike protein triggers recent massive expansion of SARS-Cov-2 variants. bioRxiv 2021. [Google Scholar] [CrossRef]
- Cantón, R.; De Lucas Ramos, P.; García-Botella, A.; García-Lledó, A.; Gómez-Pavón, J.; González Del Castillo, J.; Hernández-Sampelayo, T.; Martín-Delgado, M.C.; Martín Sánchez, F.J.; Martínez-Sellés, M.; et al. New variants of SARS-CoV-2. Rev. Esp Quim. 2021. [Google Scholar] [CrossRef]
- Cherian, S.; Potdar, V.; Jadhav, S.; Yadav, P.; Gupta, N.; Das, M.; Das, S.; Agarwal, A.; Singh, S.; Abraham, P.; et al. Convergent evolution of SARS-CoV-2 spike mutations, L452R, E484Q and P681R, in the second wave of COVID-19 in Maharashtra, India. bioRxiv 2021. [Google Scholar] [CrossRef]
- Yadav, P.D.; Sapkal, G.N.; Abraham, P.; Ella, R.; Deshpande, G.; Patil, D.Y.; Nyayanit, D.A.; Gupta, N.; Sahay, R.R.; Shete, A.M.; et al. Neutralization of variant under investigation B.1.617 with sera of BBV152 vaccinees. Clin. Infect. Dis. 2021. [Google Scholar] [CrossRef] [PubMed]
- Yadav, P.D.; Sapkal, G.N.; Abraham, P.; Deshpande, G.; Nyayanit, D.A.; Patil, D.Y.; Gupta, N.; Sahay, R.R.; Shete, A.M.; Kumar, S.; et al. Neutralization potential of Covishield vaccinated individuals sera against B.1.617.1. bioRxiv 2021. [Google Scholar] [CrossRef]
- Bolze, A.; Cirulli, E.T.; Luo, S.; White, S.; Cassens, T.; Jacobs, S.; Nguyen, J.; Ramirez, J.M.; Sandoval, E.; Wang, X.; et al. Rapid displacement of SARS-CoV-2 variant B.1.1.7 by B.1.617.2 and P.1 in the United States. medRxiv 2021. [Google Scholar] [CrossRef]
- Ferreira, I.; Datir, R.; Papa, G.; Kemp, S.; Meng, B.; Rakshit, P.; Singh, S.; Pandey, R.; Ponnusamy, K.; Radhakrishnan, V.S.; et al. SARS-CoV-2 B.1.617 emergence and sensitivity to vaccine-elicited antibodies. bioRxiv 2021. [Google Scholar] [CrossRef]
- Hoffmann, M.; Hofmann-Winkler, H.; Krüger, N.; Kempf, A.; Nehlmeier, I.; Graichen, L.; Sidarovich, A.; Moldenhauer, A.-S.; Winkler, M.S.; Schulz, S.; et al. SARS-CoV-2 variant B.1.617 is resistant to Bamlanivimab and evades antibodies induced by infection and vaccination. bioRxiv 2021. [Google Scholar] [CrossRef]
- Frampton, D.; Rampling, T.; Cross, A.; Bailey, H.; Heaney, J.; Byott, M.; Scott, R.; Sconza, R.; Price, J.; Margaritis, M.; et al. Genomic characteristics and clinical effect of the emergent SARS-CoV-2 B.1.1.7 lineage in London, UK: A whole-genome sequencing and hospital-based cohort study. Lancet Infect Dis. 2021, 21, 1246–1256. [Google Scholar] [CrossRef]
- Gallais, F.; Gantner, P.; Bruel, T.; Velay, A.; Planas, D.; Wendling, M.-J.; Bayer, S.; Solis, M.; Laugel, E.; Reix, N.; et al. Anti-SARS-CoV-2 Antibodies Persist for up to 13 Months and Reduce Risk of Reinfection. medRxiv 2021. [Google Scholar] [CrossRef]
- Shen, X.; Tang, H.; McDanal, C.; Wagh, K.; Fischer, W.; Theiler, J.; Yoon, H.; Li, D.; Haynes, B.F.; Sanders, K.O.; et al. SARS-CoV-2 variant B.1.1.7 is susceptible to neutralizing antibodies elicited by ancestral Spike vaccines. bioRxiv 2021. [Google Scholar] [CrossRef]
- Edara, V.V.; Floyd, K.; Lai, L.; Gardner, M.; Hudson, W.; Piantadosi, A.; Waggoner, J.J.; Babiker, A.; Ahmed, R.; Xie, X.; et al. Infection and mRNA-1273 vaccine antibodies neutralize SARS-CoV-2 UK variant. medRxiv 2021. [Google Scholar] [CrossRef]
- Collier, D.A.; De Marco, A.; Ferreira, I.; Meng, B.; Datir, R.P.; Walls, A.C.; Kemp, S.A.; Bassi, J.; Pinto, D.; Silacci-Fregni, C.; et al. Sensitivity of SARS-CoV-2 B.1.1.7 to mRNA vaccine-elicited antibodies. Nature 2021, 593, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Emary, K.R.W.; Golubchik, T.; Aley, P.K.; Ariani, C.V.; Angus, B.; Bibi, S.; Blane, B.; Bonsall, D.; Cicconi, P.; Charlton, S.; et al. Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 variant of concern 202012/01 (B.1.1.7): An exploratory analysis of a randomised controlled trial. Lancet 2021, 397, 1351–1362. [Google Scholar] [CrossRef]
- Pearson, C.A.B.; Russell, T.W.; Davies, N.; Kucharski, A.J.; Edmunds, W.J.; Eggo, R.M.; CMMID COVID-19 working group. Estimates of Severity and Transmissibility of Novel SARS-CoV-2 Variant 501Y.V2 in South Africa. Available online: https://cmmid.github.io/topics/covid19/sa-novel-variant.html (accessed on 24 September 2021).
- Jassat, W.; Mudara, C.; Ozougwu, L.; Tempia, S.; Blumberg, L.; Davies, M.-A.; Pillay, Y.; Carter, T.; Morewane, R.; Wolmarans, M.; et al. Increased mortality among individuals hospitalised with COVID-19 during the second wave in South Africa. medRxiv 2021. [Google Scholar] [CrossRef]
- Coutinho, R.M.; Marquitti, F.M.D.; Ferreira, L.S.; Borges, M.E.; da Silva, R.L.P.; Canton, O.; Portella, T.P.; Lyra, S.P.; Franco, C.; da Silva, A.A.M.; et al. Model-based estimation of transmissibility and reinfection of SARS-CoV-2 P.1 variant. medRxiv 2021. [Google Scholar] [CrossRef]
- Funk, T.; Pharris, A.; Spiteri, G.; Bundle, N.; Melidou, A.; Carr, M.; Gonzalez, G.; Garcia-Leon, A.; Crispie, F.; O’Connor, L.; et al. Characteristics of SARS-CoV-2 variants of concern B.1.1.7, B.1.351 or P.1: Data from seven EU/EEA countries, weeks 38/2020 to 10/2021. Eurosurveillance 2021, 26, 2100348. [Google Scholar] [CrossRef]
- Yao, W.; Wang, Y.; Ma, D.; Tang, X.; Wang, H.; Li, C.; Lin, H.; Li, Y.; Zhong, G. Circulating SARS-CoV-2 variants B.1.1.7, 501Y.V2, and P.1 have gained ability to utilize rat and mouse Ace2 and altered in vitro sensitivity to neutralizing antibodies and ACE2-Ig. bioRxiv 2021. [Google Scholar] [CrossRef]
- Hester Allen, A.; Vusirikala, A.; Flannagan, J.; Katherine, A.; Twohig, K.A.; Zaidi, A.; Harris, R.; Charlett, A.; Dabrera, G.; Kall, M. Increased Household Transmission of COVID-19 Cases Associated with SARS-CoV-2 Variant of Concern B.1.617.2: A National Casecontrol Study; Public Health England: London, UK, 2021. [Google Scholar]
- England, P.H. SARS-CoV-2 Variants of Concern and Variants under Investigation in England. Available online: https://assets.publishing.service.gov.uk/government/uploads/system/uploads/attachment_data/file/993879/Variants_of_Concern_VOC_Technical_Briefing_15.pdf (accessed on 24 September 2021).
- Planas, D.; Veyer, D.; Baidaliuk, A.; Staropoli, I.; Guivel-Benhassine, F.; Rajah, M.M.; Planchais, C.; Porrot, F.; Robillard, N.; Puech, J.; et al. Reduced sensitivity of infectious SARS-CoV-2 variant B.1.617.2 to monoclonal antibodies and sera from convalescent and vaccinated individuals. bioRxiv 2021. [Google Scholar] [CrossRef]
- Zhou, H.; Dcosta, B.M.; Samanovic, M.I.; Mulligan, M.J.; Landau, N.R.; Tada, T.B. 1.526 SARS-CoV-2 variants identified in New York City are neutralized by vaccine-elicited and therapeutic monoclonal antibodies. bioRxiv 2021. [Google Scholar] [CrossRef]
- Gupta, R.K. Will SARS-CoV-2 variants of concern affect the promise of vaccines? Nat. Rev. Immunol. 2021, 21, 340–341. [Google Scholar] [CrossRef] [PubMed]
- Mercado, N.B.; Zahn, R.; Wegmann, F.; Loos, C.; Chandrashekar, A.; Yu, J.; Liu, J.; Peter, L.; McMahan, K.; Tostanoski, L.H.; et al. Single-shot Ad26 vaccine protects against SARS-CoV-2 in rhesus macaques. Nature 2020, 586, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Sahin, U.; Muik, A.; Vogler, I.; Derhovanessian, E.; Kranz, L.M.; Vormehr, M.; Quandt, J.; Bidmon, N.; Ulges, A.; Baum, A.; et al. BNT162b2 induces SARS-CoV-2-neutralising antibodies and T cells in humans. medRxiv 2020. [Google Scholar] [CrossRef]
- Jackson, L.A.; Anderson, E.J.; Rouphael, N.G.; Roberts, P.C.; Makhene, M.; Coler, R.N.; McCullough, M.P.; Chappell, J.D.; Denison, M.R.; Stevens, L.J.; et al. An mRNA Vaccine against SARS-CoV-2—Preliminary Report. N. Engl. J. Med. 2020, 383, 1920–1931. [Google Scholar] [CrossRef]
- Folegatti, P.M.; Ewer, K.J.; Aley, P.K.; Angus, B.; Becker, S.; Belij-Rammerstorfer, S.; Bellamy, D.; Bibi, S.; Bittaye, M.; Clutterbuck, E.A.; et al. Safety and immunogenicity of the ChAdOx1 nCoV-19 vaccine against SARS-CoV-2: A preliminary report of a phase 1/2, single-blind, randomised controlled trial. Lancet 2020, 396, 467–478. [Google Scholar] [CrossRef]
- Ewer, K.J.; Barrett, J.R.; Belij-Rammerstorfer, S.; Sharpe, H.; Makinson, R.; Morter, R.; Flaxman, A.; Wright, D.; Bellamy, D.; Bittaye, M.; et al. T cell and antibody responses induced by a single dose of ChAdOx1 nCoV-19 (AZD1222) vaccine in a phase 1/2 clinical trial. Nat. Med. 2021, 27, 270–278. [Google Scholar] [CrossRef]
- Sadoff, J.; Le Gars, M.; Shukarev, G.; Heerwegh, D.; Truyers, C.; de Groot, A.M.; Stoop, J.; Tete, S.; Van Damme, W.; Leroux-Roels, I.; et al. Interim Results of a Phase 1-2a Trial of Ad26.COV2.S Covid-19 Vaccine. N. Engl. J. Med. 2021, 384, 1824–1835. [Google Scholar] [CrossRef]
- Custers, J.; Kim, D.; Leyssen, M.; Gurwith, M.; Tomaka, F.; Robertson, J.; Heijnen, E.; Condit, R.; Shukarev, G.; Heerwegh, D.; et al. Vaccines based on replication incompetent Ad26 viral vectors: Standardized template with key considerations for a risk/benefit assessment. Vaccine 2021, 39, 3081–3101. [Google Scholar] [CrossRef]
- Stephenson, K.E.; Le Gars, M.; Sadoff, J.; de Groot, A.M.; Heerwegh, D.; Truyers, C.; Atyeo, C.; Loos, C.; Chandrashekar, A.; McMahan, K.; et al. Immunogenicity of the Ad26.COV2.S Vaccine for COVID-19. JAMA 2021, 325, 1535–1544. [Google Scholar] [CrossRef] [PubMed]
- Logunov, D.Y.; Dolzhikova, I.V.; Zubkova, O.V.; Tukhvatullin, A.I.; Shcheblyakov, D.V.; Dzharullaeva, A.S.; Grousova, D.M.; Erokhova, A.S.; Kovyrshina, A.V.; Botikov, A.G. Safety and immunogenicity of an rAd26 and rAd5 vector-based heterologous prime-boost COVID-19 vaccine in two formulations: Two open, non-randomised phase 1/2 studies from Russia. Lancet 2020, 396, 887–897. [Google Scholar] [CrossRef]
- Geers, D.; Shamier, M.C.; Bodger, S.; Hartog, G.D.; Gommers, L. SARS-CoV-2 variants of conern partially escape humoral but not T cell responses in COVID-19 convalescent donors and vaccine recipients. Sci. Immunol. 2021, 6, eabj1750. [Google Scholar] [CrossRef] [PubMed]
- Muik, A.; Wallisch, A.K.; Sänger, B.; Swanson, K.A.; Mühl, J.; Chen, W.; Cai, H.; Maurus, D.; Sarkar, R.; Türeci, Ö.; et al. Neutralization of SARS-CoV-2 lineage B.1.1.7 pseudovirus by BNT162b2 vaccine-elicited human sera. Science 2021, 371, 1152–1153. [Google Scholar] [CrossRef]
- Wang, P.; Nair, M.S.; Liu, L.; Iketani, S.; Luo, Y.; Guo, Y.; Wang, M.; Yu, J.; Zhang, B.; Kwong, P.D.; et al. Antibody resistance of SARS-CoV-2 variants B.1.351 and B.1.1.7. Nature 2021, 593, 130–135. [Google Scholar] [CrossRef] [PubMed]
- Madhi, S.A.; Baillie, V.; Cutland, C.L.; Voysey, M.; Koen, A.L.; Fairlie, L.; Padayachee, S.D.; Dheda, K.; Barnabas, S.L.; Bhorat, Q.E.; et al. Efficacy of the ChAdOx1 nCoV-19 Covid-19 Vaccine against the B.1.351 Variant. N. Engl. J. Med. 2021, 384, 1885–1898. [Google Scholar] [CrossRef]
- Planas, D.; Bruel, T.; Grzelak, L.; Guivel-Benhassine, F.; Staropoli, I.; Porrot, F.; Planchais, C.; Buchrieser, J.; Rajah, M.M.; Bishop, E.; et al. Sensitivity of infectious SARS-CoV-2 B.1.1.7 and B.1.351 variants to neutralizing antibodies. Nat. Med. 2021, 27, 917–924. [Google Scholar] [CrossRef]
- Zhou, D.; Dejnirattisai, W.; Supasa, P.; Liu, C.; Mentzer, A.J.; Ginn, H.M.; Zhao, Y.; Duyvesteyn, H.M.E.; Tuekprakhon, A.; Nutalai, R.; et al. Evidence of escape of SARS-CoV-2 variant B.1.351 from natural and vaccine-induced sera. Cell 2021, 184, 2348–2361.e6. [Google Scholar] [CrossRef]
- Sadoff, J.; Gray, G.; Vandebosch, A.; Cárdenas, V.; Shukarev, G.; Grinsztejn, B.; Goepfert, P.A.; Truyers, C.; Fennema, H.; Spiessens, B.; et al. Safety and Efficacy of Single-Dose Ad26.COV2.S Vaccine against COVID-19. N. Engl. J. Med. 2021, 384, 2187–2201. [Google Scholar] [CrossRef]
- Alter, G.; Yu, J.; Liu, J.; Chandrashekar, A.; Borducchi, E.N.; Tostanoski, L.H.; McMahan, K.; Jacob-Dolan, C.; Martinez, D.R.; Chang, A.; et al. Immunogenicity of Ad26.COV2.S vaccine against SARS-CoV-2 variants in humans. Nature 2021, 596, 268–272. [Google Scholar] [CrossRef] [PubMed]
- Kuzmina, A.; Khalaila, Y.; Voloshin, O.; Keren-Naus, A.; Boehm-Cohen, L.; Raviv, Y.; Shemer-Avni, Y.; Rosenberg, E.; Taube, R. SARS-CoV-2 spike variants exhibit differential infectivity and neutralization resistance to convalescent or post-vaccination sera. Cell Host Microbe 2021, 29, 522–528.e2. [Google Scholar] [CrossRef]
- Liu, Y.; Liu, J.; Xia, H.; Zhang, X.; Fontes-Garfias, C.R.; Swanson, K.A.; Cai, H.; Sarkar, R.; Chen, W.; Cutler, M.; et al. Neutralizing Activity of BNT162b2-Elicited Serum. N. Engl. J. Med. 2021, 384, 1466–1468. [Google Scholar] [CrossRef]
- Kustin, T.; Harel, N.; Finkel, U.; Perchik, S.; Harari, S.; Tahor, M.; Caspi, I.; Levy, R.; Leshchinsky, M.; Ken Dror, S.; et al. Evidence for increased breakthrough rates of SARS-CoV-2 variants of concern in BNT162b2-mRNA-vaccinated individuals. Nat. Med. 2021, 27, 1379–1384. [Google Scholar] [CrossRef]
- Planas, D.; Veyer, D.; Baidaliuk, A.; Staropoli, I.; Guivel-Benhassine, F.; Rajah, M.M.; Planchais, C.; Porrot, F.; Robillard, N.; Puech, J.; et al. Reduced sensitivity of SARS-CoV-2 variant Delta to antibody neutralization. Nature 2021, 596, 276–280. [Google Scholar] [CrossRef]
- Lopez Bernal, J.; Andrews, N.; Gower, C.; Gallagher, E.; Simmons, R.; Thelwall, S.; Stowe, J.; Tessier, E.; Groves, N.; Dabrera, G.; et al. Effectiveness of Covid-19 Vaccines against the B.1.617.2 (Delta) Variant. N. Engl. J. Med. 2021, 385, 585–594. [Google Scholar] [CrossRef] [PubMed]
- Sheikh, A.; McMenamin, J.; Taylor, B.; Robertson, C. SARS-CoV-2 Delta VOC in Scotland: Demographics, risk of hospital admission, and vaccine effectiveness. Lancet 2021, 397, 2461–2462. [Google Scholar] [CrossRef]
- Wall, E.C.; Wu, M.; Harvey, R.; Kelly, G.; Warchal, S.; Sawyer, C.; Daniels, R.; Hobson, P.; Hatipoglu, E.; Ngai, Y.; et al. Neutralising antibody activity against SARS-CoV-2 VOCs B.1.617.2 and B.1.351 by BNT162b2 vaccination. Lancet 2021, 397, 2331–2333. [Google Scholar] [CrossRef]
- Liu, C.; Ginn, H.M.; Dejnirattisai, W.; Supasa, P.; Wang, B.; Tuekprakhon, A.; Nutalai, R.; Zhou, D.; Mentzer, A.J.; Zhao, Y.; et al. Reduced neutralization of SARS-CoV-2 B.1.617 by vaccine and convalescent serum. Cell 2021, 184, 4220–4236.e13. [Google Scholar] [CrossRef] [PubMed]
- Mlcochova, P.; Kemp, S.; Dhar, M.S.; Papa, G.; Meng, B.; Ferreira, I.A.T.M.; Datir, R.; Collier, D.A.; Albecka, A.; Singh, S.; et al. SARS-CoV-2 B.1.617.2 Delta variant replication and immune evasion. Nature 2021. [Google Scholar] [CrossRef]
- Hammerschmidt, S.I.; Bosnjak, B.; Bernhardt, G.; Friedrichsen, M.; Ravens, I.; Dopfer-Jablonka, A.; Hoffmann, M.; Pöhlmann, S.; Behrens, G.M.N.; Förster, R. Neutralization of the SARS-CoV-2 Delta variant after heterologous and homologous BNT162b2 or ChAdOx1 nCoV-19 vaccination. Cell. Mol. Immunol. 2021, 18, 2455–2456. [Google Scholar] [CrossRef]
- Liu, W.J.; Zhao, M.; Liu, K.; Xu, K.; Wong, G.; Tan, W.; Gao, G.F. T-cell immunity of SARS-CoV: Implications for vaccine development against MERS-CoV. Antivir. Res. 2017, 137, 82–92. [Google Scholar] [CrossRef]
- Cox, R.J.; Brokstad, K.A. Not just antibodies: B cells and T cells mediate immunity to COVID-19. Nat. Rev. Immunol. 2020, 20, 581–582. [Google Scholar] [CrossRef]
- Sekine, T.; Perez-Potti, A.; Rivera-Ballesteros, O.; Strålin, K.; Gorin, J.B.; Olsson, A.; Llewellyn-Lacey, S.; Kamal, H.; Bogdanovic, G.; Muschiol, S.; et al. Robust T Cell Immunity in Convalescent Individuals with Asymptomatic or Mild COVID-19. Cell 2020, 183, 158–168.e14. [Google Scholar] [CrossRef]
- Crotty, S. Follicular helper CD4 T cells (TFH). Annu. Rev. Immunol. 2011, 29, 621–663. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J. T helper 2 (Th2) cell differentiation, type 2 innate lymphoid cell (ILC2) development and regulation of interleukin-4 (IL-4) and IL-13 production. Cytokine 2015, 75, 14–24. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bolles, M.; Deming, D.; Long, K.; Agnihothram, S.; Whitmore, A.; Ferris, M.; Funkhouser, W.; Gralinski, L.; Totura, A.; Heise, M.; et al. A double-inactivated severe acute respiratory syndrome coronavirus vaccine provides incomplete protection in mice and induces increased eosinophilic proinflammatory pulmonary response upon challenge. J. Virol. 2011, 85, 12201–12215. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lee, W.S.; Wheatley, A.K.; Kent, S.J.; DeKosky, B.J. Antibody-dependent enhancement and SARS-CoV-2 vaccines and therapies. Nat. Microbiol. 2020, 5, 1185–1191. [Google Scholar] [CrossRef]
- Noh, J.Y.; Jeong, H.W.; Shin, E.-C. SARS-CoV-2 mutations, vaccines, and immunity: Implication of variants of concern. Signal Transduct. Target. Ther. 2021, 6, 203. [Google Scholar] [CrossRef]
- Falk, K.; Rötzschke, O.; Stevanovié, S.; Jung, G.; Rammensee, H.-G. Allele-specific motifs revealed by sequencing of self-peptides eluted from MHC molecules. Nature 1991, 351, 290–296. [Google Scholar] [CrossRef]
- Tarke, A.; Sidney, J.; Methot, N.; Zhang, Y.; Dan, J.M.; Goodwin, B.; Rubiro, P.; Sutherland, A.; da Silva Antunes, R.; Frazier, A.; et al. Negligible impact of SARS-CoV-2 variants on CD4+ and CD8+ T cell reactivity in COVID-19 exposed donors and vaccinees. bioRxiv 2021. [Google Scholar] [CrossRef]
- Redd, A.D.; Nardin, A.; Kared, H.; Bloch, E.M.; Pekosz, A.; Laeyendecker, O.; Abel, B.; Fehlings, M.; Quinn, T.C.; Tobian, A.A. CD8+ T cell responses in COVID-19 convalescent individuals target conserved epitopes from multiple prominent SARS-CoV-2 circulating variants. medRxiv 2021, 8. [Google Scholar] [CrossRef]
- Jordan, S.C.; Shin, B.H.; Gadsden, T.M.; Chu, M.; Petrosyan, A.; Le, C.N.; Zabner, R.; Oft, J.; Pedraza, I.; Cheng, S.; et al. T cell immune responses to SARS-CoV-2 and variants of concern (Alpha and Delta) in infected and vaccinated individuals. Cell. Mol. Immunol. 2021, 1–3. [Google Scholar] [CrossRef]
- Barouch, D.H.; Stephenson, K.E.; Sadoff, J.; Yu, J.; Chang, A.; Gebre, M.; McMahan, K.; Liu, J.; Chandrashekar, A.; Patel, S.; et al. Durable Humoral and Cellular Immune Responses 8 Months after Ad26.COV2.S Vaccination. N. Engl. J. Med. 2021, 385, 951–953. [Google Scholar] [CrossRef]
- Keech, C.; Albert, G.; Cho, I.; Robertson, A.; Reed, P.; Neal, S.; Plested, J.S.; Zhu, M.; Cloney-Clark, S.; Zhou, H.; et al. Phase 1–2 Trial of a SARS-CoV-2 Recombinant Spike Protein Nanoparticle Vaccine. N. Engl. J. Med. 2020, 383, 2320–2332. [Google Scholar] [CrossRef]
- Nasreen, S.; He, S.; Chung, H.; Brown, K.A.; Gubbay, J.B.; Buchan, S.A.; Wilson, S.E.; Sundaram, M.E.; Fell, D.B.; Chen, B.; et al. Effectiveness of COVID-19 vaccines against variants of concern, Canada. medRxiv 2021. [Google Scholar] [CrossRef]
- Wu, K.; Werner, A.P.; Koch, M.; Choi, A.; Narayanan, E.; Stewart-Jones, G.B.E.; Colpitts, T.; Bennett, H.; Boyoglu-Barnum, S.; Shi, W.; et al. Serum Neutralizing Activity Elicited by mRNA-1273 Vaccine. N. Engl. J. Med. 2021, 384, 1468–1470. [Google Scholar] [CrossRef] [PubMed]
- Shinde, V.; Bhikha, S.; Hoosain, Z.; Archary, M.; Bhorat, Q.; Fairlie, L.; Lalloo, U.; Masilela, M.S.L.; Moodley, D.; Hanley, S.; et al. Preliminary Efficacy of the NVX-CoV2373 Covid-19 Vaccine against the B.1.351 Variant. medRxiv 2021. [Google Scholar] [CrossRef]
- Moderna COVID-19 Vaccine Retains Neutralizing Activity against Emerging Variants First Identified in the U.K. and the Republic of South Africa. Available online: https://investors.modernatx.com/news-releases/news-release-details/moderna-covid-19-vaccine-retains-neutralizing-activity-against (accessed on 24 September 2021).
- Abdool Karim, S.S.; de Oliveira, T. New SARS-CoV-2 Variants—Clinical, Public Health, and Vaccine Implications. N. Engl. J. Med. 2021, 384, 1866–1868. [Google Scholar] [CrossRef]
- Stowe, J.; Andrews, N.; Gower, C.; Gallagher, E.; Utsi, L.; Simmons, R.; Thelwall, S.; Tessier, E.; Groves, N.; Dabrera, G.; et al. Effectiveness of COVID-19 Vaccines against Hospital Admission with the Delta (B.1.617.2) Variant. Available online: https://media.tghn.org/articles/Effectiveness_of_COVID19_vaccines_against_hospital_admission_with_the_Delta_B._G6gnnqJ.pdf (accessed on 13 October 2021).
- Abu-Raddad, L.J.; Chemaitelly, H.; Butt, A.A. Effectiveness of the BNT162b2 COVID-19 Vaccine against the B.1.1.7 and B.1.351 Variants. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef]
- Polack, F.P.; Thomas, S.J.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Moreira, E.D.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 mRNA COVID-19 Vaccine. N. Engl. J. Med. 2020, 383, 2603–2615. [Google Scholar] [CrossRef]
- Lopez Bernal, J.; Andrews, N.; Gower, C.; Robertson, C.; Stowe, J.; Tessier, E.; Simmons, R.; Cottrell, S.; Roberts, R.; O’Doherty, M.; et al. Effectiveness of the Pfizer-BioNTech and Oxford-AstraZeneca vaccines on covid-19 related symptoms, hospital admissions, and mortality in older adults in England: Test negative case-control study. BMJ 2021, 373, n1088. [Google Scholar] [CrossRef]
- Dagan, N.; Barda, N.; Kepten, E.; Miron, O.; Perchik, S.; Katz, M.A.; Hernán, M.A.; Lipsitch, M.; Reis, B.; Balicer, R.D. BNT162b2 mRNA Covid-19 Vaccine in a Nationwide Mass Vaccination Setting. N. Engl. J. Med. 2021, 384, 1412–1423. [Google Scholar] [CrossRef]
- Ismail, S.A.; Garcia Vilaplana, T.; Elgohari, S.; Stowe, J.; Tessier, E.; Andrews, N.; Vusirikala, A.; Ramsay, M.; Mandal, S.; Lopez Bernal, J. Effectiveness of BNT162b2 mRNA and ChAdOx1 adenovirus vector COVID-19 vaccines on risk of hospitalisation among older adults in England: An observational study using surveillance data. Public Health Engl. 2021. [Google Scholar]
- Hall, V.J.; Foulkes, S.; Saei, A.; Andrews, N.; Oguti, B.; Charlett, A.; Wellington, E.; Stowe, J.; Gillson, N.; Atti, A.; et al. Effectiveness of BNT162b2 mRNA Vaccine against Infection and COVID-19 Vaccine Coverage in Healthcare Workers in England, Multicentre Prospective Cohort Study (the SIREN Study). SSRN Electron. J. 2021. [Google Scholar] [CrossRef]
- Bernal, J.L.; Andrews, N.; Gower, C.; Stowe, J.; Robertson, C.; Tessier, E.; Simmons, R.; Cottrell, S.; Roberts, R.; O’Doherty, M.; et al. Early effectiveness of COVID-19 vaccination with BNT162b2 mRNA vaccine and ChAdOx1 adenovirus vector vaccine on symptomatic disease, hospitalisations and mortality in older adults in England. medRxiv 2021. [Google Scholar] [CrossRef]
- Haas, E.J.; Angulo, F.J.; McLaughlin, J.M.; Anis, E.; Singer, S.R.; Khan, F.; Brooks, N.; Smaja, M.; Mircus, G.; Pan, K.; et al. Impact and effectiveness of mRNA BNT162b2 vaccine against SARS-CoV-2 infections and COVID-19 cases, hospitalisations, and deaths following a nationwide vaccination campaign in Israel: An observational study using national surveillance data. Lancet 2021, 397, 1819–1829. [Google Scholar] [CrossRef]
- Baden, L.R.; El Sahly, H.M.; Essink, B.; Kotloff, K.; Frey, S.; Novak, R.; Diemert, D.; Spector, S.A.; Rouphael, N.; Creech, C.B.; et al. Efficacy and Safety of the mRNA-1273 SARS-CoV-2 Vaccine. N. Engl. J. Med. 2021, 384, 403–416. [Google Scholar] [CrossRef] [PubMed]
- Moderna. Moderna Announces Publication of Results from the Pivotal Phase 3 Trial of the Moderna COVID-19 Vaccine in The N. Engl. J. Med. 2021. Available online: https://investors.modernatx.com/news-releases/news-release-details/moderna-announces-publication-results-pivotal-phase-3-trial (accessed on 3 July 2021).
- A Study to Evaluate the Safety, Reactogenicity, and Effectiveness of mRNA-1273 Vaccine in Adolescents 12 to <18 Years Old to Prevent COVID-19 (TeenCove). Available online: https://clinicaltrials.gov/ct2/show/NCT04649151 (accessed on 5 July 2021).
- Thompson, M.G.; Burgess, J.L.; Naleway, A.L.; Tyner, H.L.; Yoon, S.K.; Meece, J.; Olsho, L.E.W.; Caban-Martinez, A.J.; Fowlkes, A.; Lutrick, K.; et al. Interim Estimates of Vaccine Effectiveness of BNT162b2 and mRNA-1273 COVID-19 Vaccines in Preventing SARS-CoV-2 Infection Among Health Care Personnel, First Responders, and Other Essential and Frontline Workers—Eight U.S. Locations, December 2020–March 2021. MMWR Morb. Mortal. Wkly Rep. 2021, 70, 495–500. [Google Scholar] [CrossRef] [PubMed]
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E. Single-dose administration and the influence of the timing of the booster dose on immunogenicity and efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine: A pooled analysis of four randomised trials. Lancet 2021, 397, 881–891. [Google Scholar] [CrossRef]
- Dejnirattisai, W.; Zhou, D.; Supasa, P.; Liu, C.; Mentzer, A.J.; Ginn, H.M.; Zhao, Y.; Duyvesteyn, H.M.E.; Tuekprakhon, A.; Nutalai, R.; et al. Antibody evasion by the Brazilian P.1 strain of SARS-CoV-2. bioRxiv 2021, 9. [Google Scholar] [CrossRef]
- AstraZeneca. AZD1222 US Phase III Primary Analysis Confirms Safety and Efficacy. Available online: https://www.astrazeneca.com/content/astraz/media-centre/press-releases/2021/azd1222-us-phase-iii-primary-analysis-confirms-safety-and-efficacy.html (accessed on 24 September 2021).
- Voysey, M.; Clemens, S.A.C.; Madhi, S.A.; Weckx, L.Y.; Folegatti, P.M.; Aley, P.K.; Angus, B.; Baillie, V.L.; Barnabas, S.L.; Bhorat, Q.E.; et al. Safety and efficacy of the ChAdOx1 nCoV-19 vaccine (AZD1222) against SARS-CoV-2: An interim analysis of four randomised controlled trials in Brazil, South Africa, and the UK. Lancet 2021, 397, 99–111. [Google Scholar] [CrossRef]
- Liu, J.; Liu, Y.; Xia, H.; Zou, J.; Weaver, S.C.; Swanson, K.A.; Cai, H.; Cutler, M.; Cooper, D.; Muik, A.; et al. BNT162b2-elicited neutralization of B.1.617 and other SARS-CoV-2 variants. Nature 2021, 596, 273–275. [Google Scholar] [CrossRef]
- Pritchard, E.; Matthews, P.C.; Stoesser, N.; Eyre, D.W.; Gethings, O.; Vihta, K.-D.; Jones, J.; House, T.; VanSteenHouse, H.; Bell, I.; et al. Impact of vaccination on new SARS-CoV-2 infections in the UK. medRxiv 2021. [Google Scholar] [CrossRef]
- Ledford, H. J&J’s one-shot COVID vaccine offers hope for faster protection. Nature 2021. [Google Scholar] [CrossRef]
- Corchado-Garcia, J.; Puyraimond-Zemmour, D.; Hughes, T.; Cristea-Platon, T.; Lenehan, P.; Pawlowski, C.; Bade, S.; O’Horo, J.C.; Gores, G.J.; Williams, A.W.; et al. Real-world effectiveness of Ad26.COV2.S adenoviral vector vaccine for COVID-19. medRxiv 2021. [Google Scholar] [CrossRef]
- Gottlieb Says Delta Virus Variant likely to Become Dominant U.S. Strain. Available online: https://www.cbsnews.com/news/covid-19-delta-varient-dominant-strain-likely/ (accessed on 25 September 2021).
- NEWSWIRE, C.P. Novavax Confirms High Levels of Efficacy against Original and Variant COVID-19 Strains in United Kingdom and South Africa Trials. CISION PR NEWSWIRE 2021. [Google Scholar]
- Mahase, E. COVID-19: Novavax vaccine efficacy is 86% against UK variant and 60% against South African variant. BMJ 2021, 372, n296. [Google Scholar] [CrossRef]
- Heath, P.T.; Galiza, E.P.; Baxter, D.N.; Boffito, M.; Browne, D.; Burns, F.; Chadwick, D.R.; Clark, R.; Cosgrove, C.; Galloway, J.; et al. Safety and Efficacy of NVX-CoV2373 Covid-19 Vaccine. N. Engl. J. Med. 2021, 385, 1172–1183. [Google Scholar] [CrossRef] [PubMed]
- Callaway, E.; Mallapaty, S. Novavax offers first evidence that COVID vaccines protect people against variants. Nature 2021, 590, 17. [Google Scholar] [CrossRef]
- Logunov, D.Y.; Dolzhikova, I.V.; Shcheblyakov, D.V.; Tukhvatulin, A.I.; Zubkova, O.V.; Dzharullaeva, A.S.; Kovyrshina, A.V.; Lubenets, N.L.; Grousova, D.M.; Erokhova, A.S.; et al. Safety and efficacy of an rAd26 and rAd5 vector-based heterologous prime-boost COVID-19 vaccine: An interim analysis of a randomised controlled phase 3 trial in Russia. Lancet 2021, 397, 671–681. [Google Scholar] [CrossRef]
- Jones, I.; Roy, P. Sputnik V COVID-19 vaccine candidate appears safe and effective. Lancet 2021, 397, 642–643. [Google Scholar] [CrossRef]
- CDC. COVID-19 Vaccine Breakthrough Case Investigation and Reporting. Available online: https://www.cdc.gov/vaccines/covid-19/health-departments/breakthrough-cases.html (accessed on 6 July 2021).
- Comparing COVID-19 Booster Vaccinations (COV-BOOST). Available online: https://www.covboost.org.uk/about (accessed on 6 July 2021).
- Oxford Leads First Trial Investigating Dosing with Alternating Vaccines. Available online: https://www.ox.ac.uk/news/2021-02-04-oxford-leads-first-trial-investigating-dosing-alternating-vaccines (accessed on 6 July 2021).
- Russia Ready to Trial Combined AstraZeneca, Sputnik V Vaccine in Ukraine. Available online: https://www.reuters.com/article/us-health-coronavirus-russia-ukraine/russia-ready-to-trial-combined-astrazeneca-sputnik-v-vaccine-in-ukraine-idUSKBN2970JP (accessed on 6 July 2021).
- FDA Statement on Following the Authorized Dosing Schedules for COVID-19 Vaccines. Available online: https://www.fda.gov/news-events/press-announcements/fda-statement-following-authorized-dosing-schedules-covid-19-vaccines (accessed on 6 July 2021).
- GSK and CureVac to Develop Next Generation mRNA COVID-19 Vaccines. Available online: https://www.gsk.com/en-gb/media/press-releases/gsk-and-curevac-to-develop-next-generation-mrna-covid-19-vaccines/ (accessed on 6 July 2021).
- Collier, D.A.; Ferreira, I.A.T.M.; Datir, R.; Kotagiri, P.; Lim, E.; Meng, B.; Bergamaschi, L.; Collaboration, C.N.; Elmer, A.; Kingston, N.; et al. Age-related heterogeneity in immune responses to SARS-CoV-2 vaccine BNT162b2. medRxiv 2021. [Google Scholar] [CrossRef]
- Le Bert, N.; Tan, A.T.; Kunasegaran, K.; Tham, C.Y.L.; Hafezi, M.; Chia, A.; Chng, M.H.Y.; Lin, M.; Tan, N.; Linster, M.; et al. SARS-CoV-2-specific T cell immunity in cases of COVID-19 and SARS, and uninfected controls. Nature 2020, 584, 457–462. [Google Scholar] [CrossRef]
- Hansen, J.; Baum, A.; Pascal, K.E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B.O.; Yan, Y.; Koon, K.; Patel, K.; et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 2020, 369, 1010–1014. [Google Scholar] [CrossRef] [PubMed]
- Shannon, A.; Selisko, B.; Le, N.-T.-T.; Huchting, J.; Touret, F.; Piorkowski, G.; Fattorini, V.; Ferron, F.; Decroly, E.; Meier, C.; et al. Rapid incorporation of Favipiravir by the fast and permissive viral RNA polymerase complex results in SARS-CoV-2 lethal mutagenesis. Nat. Commun. 2020, 11, 4682. [Google Scholar] [CrossRef] [PubMed]
| Pango Lineage and Corresponding WHO Naming | Attributes * | |
|---|---|---|
| Variants of Interest |
|
|
| Variants of concern |
|
|
| Variants of high consequences |
|
|
| Name of the Mutation | Location in Spike Protein | Important Effects |
|---|---|---|
| N501Y | Receptor-Binding Domain |
|
| K417N and K417T | Receptor-Binding Domain |
|
| E484K, E484Q, and E484P | Receptor-Binding Domain | |
| L452R | Receptor-Binding Domain | |
| S477G, S44N, and S477R | Receptor-Binding Domain |
|
| R246I/M | N-terminal domain (adjacent to the glycosylation sites) |
|
| 69del and 70del | N-terminal domain | |
| S13I and W152C | N-terminal domain |
|
| L18F | N-terminal domain |
|
| 141–143del | N-terminal domain |
|
| 144del | N-terminal domain |
|
| D614G | Carboxy(C)-terminal region of the S1 domain |
|
| H655Y | Adjacent to the S1/S2 cleavage site |
|
| P681H/R | Adjacent to the S1/S2 cleavage site |
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ramesh, S.; Govindarajulu, M.; Parise, R.S.; Neel, L.; Shankar, T.; Patel, S.; Lowery, P.; Smith, F.; Dhanasekaran, M.; Moore, T. Emerging SARS-CoV-2 Variants: A Review of Its Mutations, Its Implications and Vaccine Efficacy. Vaccines 2021, 9, 1195. https://doi.org/10.3390/vaccines9101195
Ramesh S, Govindarajulu M, Parise RS, Neel L, Shankar T, Patel S, Lowery P, Smith F, Dhanasekaran M, Moore T. Emerging SARS-CoV-2 Variants: A Review of Its Mutations, Its Implications and Vaccine Efficacy. Vaccines. 2021; 9(10):1195. https://doi.org/10.3390/vaccines9101195
Chicago/Turabian StyleRamesh, Sindhu, Manoj Govindarajulu, Rachel S. Parise, Logan Neel, Tharanath Shankar, Shriya Patel, Payton Lowery, Forrest Smith, Muralikrishnan Dhanasekaran, and Timothy Moore. 2021. "Emerging SARS-CoV-2 Variants: A Review of Its Mutations, Its Implications and Vaccine Efficacy" Vaccines 9, no. 10: 1195. https://doi.org/10.3390/vaccines9101195
APA StyleRamesh, S., Govindarajulu, M., Parise, R. S., Neel, L., Shankar, T., Patel, S., Lowery, P., Smith, F., Dhanasekaran, M., & Moore, T. (2021). Emerging SARS-CoV-2 Variants: A Review of Its Mutations, Its Implications and Vaccine Efficacy. Vaccines, 9(10), 1195. https://doi.org/10.3390/vaccines9101195






