Emergency FMD Serotype O Vaccines Protect Cattle against Heterologous Challenge with a Variant Foot-and-Mouth Disease Virus from the O/ME-SA/Ind2001 Lineage
Abstract
:1. Introduction
2. Materials and Methods
2.1. Experimental Animals
2.2. Viruses and Vaccines
2.3. Immunisation, Challenge, Clinical Score and Sample Collection
2.4. Serological Assays to Measure Antibodies
2.5. Viraemia and Virus Excretion
2.6. Statistical Analysis
3. Results
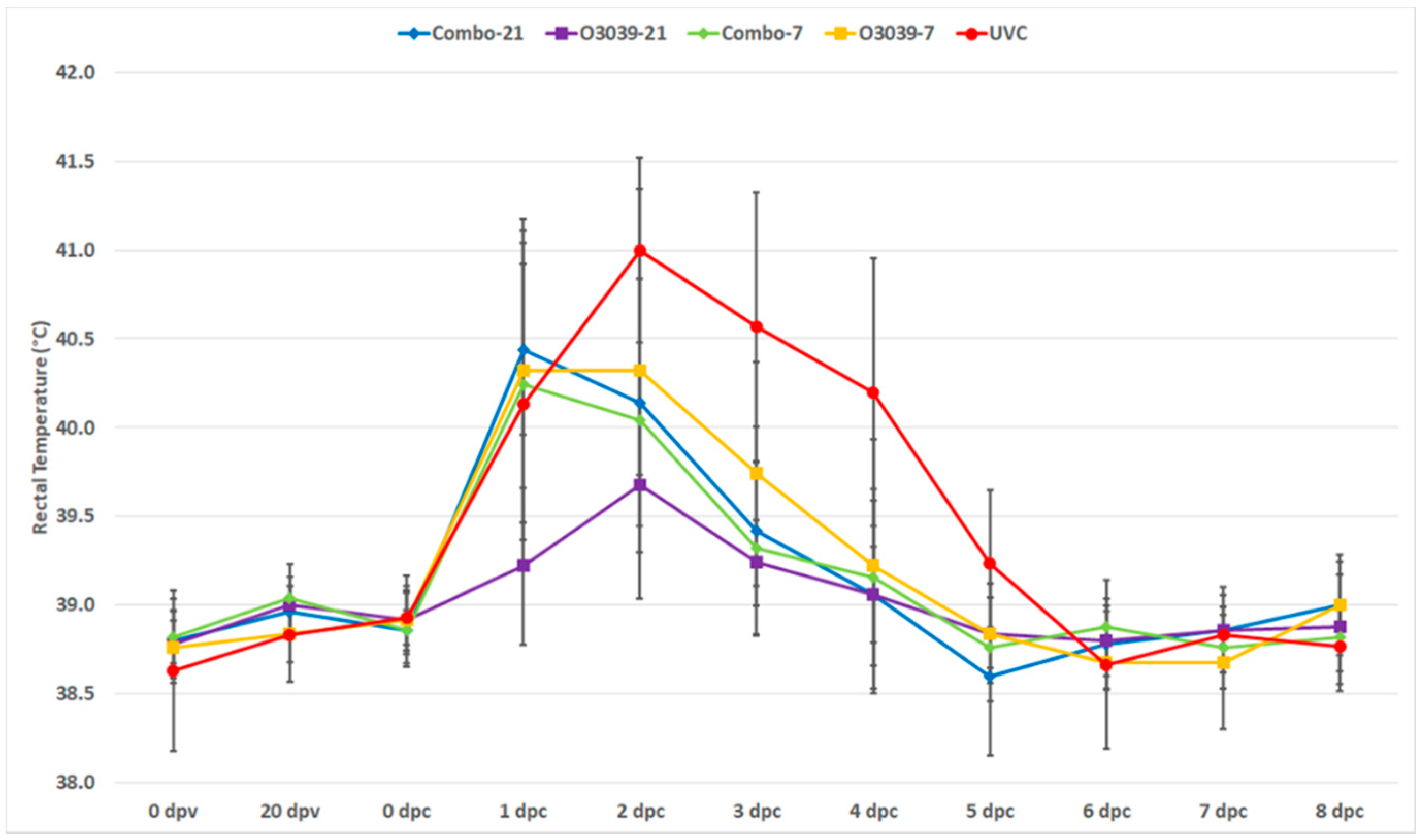
3.1. Post Challenge Outcomes (Temperature and Clinical Disease)
3.2. Serological Response to Vaccination at the Time of Challenge
3.3. NSP Antibody Responses
3.4. Viraemia and Virus Excretion in Clinical Samples
3.5. Virus Persistence and Carrier Status
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Rodrigo, K.J.; Dopazo, J. Evolutionary Analysis of the Picornavirus Family. J. Mol. Evol. 1995, 40, 362–371. [Google Scholar] [CrossRef] [PubMed]
- Domingo, E.; Escarmis, C.; Baranowski, E.; Ruiz-Jarabo, C.M.; Carrillo, E.; Nunez, J.I.; Sobrino, F. Evolution of Foot-and-Mouth Disease Virus. Virus Res. 2003, 91, 47–63. [Google Scholar] [CrossRef]
- Knight-Jones, T.J.; Rushton, J. The Economic Impacts of Foot and Mouth Disease—What Are They, How Big Are They and Where Do They Occur? Prev. Vet. Med. 2013, 112, 161–173. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Buetre, B.; Wicks, S.; Kruger, H.; Millist, N.; Yainshet, A.; Garner, G.; Duncan, A.; Abdalla, A.; Trestrail, C.; Hatt, M.; et al. Potential Socio-Economic Impacts of an Outbreak of Foot-and-Mouth Disease in Australia; Commonwealth of Australia: Canberra, Australia, 2013. [Google Scholar]
- Brehm, K.E.; Kumar, N.; Thulke, H.H.; Haas, B. High Potency Vaccines Induce Protection against Heterologous Challenge with Foot-and-Mouth Disease Virus. Vaccine 2008, 26, 1681–1687. [Google Scholar] [CrossRef]
- Clavijo, A.; Sanchez-Vazquez, M.J.; Buzanovsky, L.P.; Martini, M.; Pompei, J.C.; Cosivi, O. Current Status and Future Prospects to Achieve Foot-and-Mouth Disease Eradication in South America. Transbound. Emerg. Dis. 2017, 64, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Geale, D.W.; Barnett, P.V.; Clarke, G.W.; Davis, J.; Kasari, T.R. A Review of Oie Country Status Recovery Using Vaccinate-to-Live Versus Vaccinate-to-Die Foot-and-Mouth Disease Response Policies II: Waiting Periods after Emergency Vaccination in FMD Free Countries. Transbound. Emerg. Dis. 2015, 62, 388–406. [Google Scholar] [CrossRef]
- Garland, A.J.; de Clercq, K. Cattle, Sheep and Pigs Vaccinated against Foot and Mouth Disease: Does Trade in These Animals and Their Products Present a Risk of Transmitting the Disease? Rev. Sci. Tech. 2011, 30, 189–206. [Google Scholar] [CrossRef] [Green Version]
- OIE/-FAO Foot-and-Mouth Disease Reference Laboratory Network. WRL_FMD_Reports. Annual Report 2015; World Reference Laboratory for FMD: Surrey, UK, 2015; Available online: OIE-FAO FMD Ref Lab Network Report 2015.pdf (wrlfmd.org) (accessed on 21 August 2021).
- Knowles, N.J.; Bachanek-Bankowska, K.; Wadsworth, J.; Mioulet, V.; Valdazo-Gonzalez, B.; Eldaghayes, I.M.; Dayhum, A.S.; Kammon, A.M.; Sharif, M.A.; Waight, S.; et al. Outbreaks of Foot-and-Mouth Disease in Libya and Saudi Arabia During 2013 Due to an Exotic O/ME-SA/Ind-2001 Lineage Virus. Transbound. Emerg. Dis. 2016, 63, e431–e435. [Google Scholar] [CrossRef] [Green Version]
- Fishbourne, E.; Ludi, A.B.; Wilsden, G.; Hamblin, P.; Statham, B.; Bin-Tarif, A.; Brocchi, E.; Grazioli, S.; Dekker, A.; Eble, P.; et al. Efficacy of a High Potency O1 Manisa Foot-and-Mouth Disease Vaccine in Cattle against Heterologous Challenge with a Field Virus from the O/ME-SA/Ind-2001 Lineage Collected in North Africa. Vaccine 2017, 35, 2761–2765. [Google Scholar] [CrossRef]
- Qui, Y.; Abila, R.; Rodtian, P.; King, D.P.; Knowles, N.J.; Ngo, N.T.; Le, V.T.; Khounsy, S.; Bounma, P.; Lwin, S.; et al. Emergence of Exotic Serotype O Foot-and-Mouth Disease Virus O/ME-SA/Ind-2001d in South-East Asia in 2015. Transbound. Emerg. Dis. 2018, 65, e104–e112. [Google Scholar]
- Hicks, H.M.; Wadsworth, J.; Azhar, M.; Afzal, M.; Manzoor, S.; Abubakar, M.; Khan, E.H.; king, D.P.; Knowles, N.J. Genome Sequences of Foot-and-Mouth Disease Virus O/ME-SA/Ind-2001e Strains Isolated in Pakistan. Microbiol. Resour. Announc. 2020, 9, e00165-20. [Google Scholar] [CrossRef] [PubMed]
- Jamal, S.M.; Khan, S.; Knowles, N.J.; Wadsworth, J.; Hicks, H.M.; Mioulet, V.; Bin-Tarif, A.; Ludi, A.B.; Shah, S.A.A.; Abubakar, M.; et al. Foot-and-Mouth Disease Viruses of the O/ME-SA/Ind-2001e Sublineage in Pakistan. Transbound. Emerg. Dis. 2021. online ahead of print. [Google Scholar] [CrossRef] [PubMed]
- OIE. FMD_Manual. Foot and Mouth Disease (Infection with Foot and Mouth Disease Virus). In OIE Terestrial Manual: Chapter 3.1.8; OIE: Paris, France, 2017. [Google Scholar]
- Singanallur, N.B.; Dekker, A.; Eble, P.; Kluitenberg, F.v.; Weerdmeester, K.; Horsington, J.; Vosloo, W. Comparing the Efficacy of Two FMDV Serotype A Emergency Vaccines against Heterologous Challenge with an A/ASIA/G IX/SEA-97 Variant Virus. Vaccines 2020, 8, 80. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dekker, A.; Sanz-Bernado, B.; Phaedra, E.; King, D.; Singanallur, N.B.; Vosloo, W. Good Quality A Malaysia 97 Protects against A/ASIA/G-VII (A/IRN/22/2015). Vaccines 2020, 8, 24. [Google Scholar] [CrossRef] [Green Version]
- Horsington, J.; Perez, C.B.; Maradei, E.; Novo, S.G.; Gonzales, J.L.; Singanallur, N.B.; Bonastre, P.; Vosloo, W. Protective Effects of High-Potency FMDV O1 Manisa Monovalent Vaccine in Cattle Challenged with FMDV O/SKR/2010 at 7 or 4 Days Post Vaccination. Vaccine 2017, 35, 5179–5185. [Google Scholar] [CrossRef] [PubMed]
- Vosloo, W.; Nguyen, H.T.T.; Fosgate, G.T.; Morris, J.M.; Wang, J.; Kim, P.V.; Quach, N.V.; Le, P.T.T.; Hung, D.; Tran, H.X.; et al. Efficacy of a High Potency O1 Manisa Monovalent Vaccine against Heterologous Challenge with a FMDV O Mya98 Lineage Virus in Pigs 4 and 7 Days Post Vaccination. Vaccine 2015, 33, 2778–2785. [Google Scholar]
- Singanallur, N.B.; Nguyen, H.T.T.; Fosgate, G.T.; Morris, J.M.; Davis, A.; Giles, M.; Kim, P.V.; Quach, N.V.; Le, P.T.T.; Nguyen, P.N.H.; et al. A Malaysia 97 Monovalent Foot-and-Mouth Disease Vaccine (>6 pd50/Dose) Protects Pigs against Challenge with a Variant FMDV A SEA-97 Lineage Virus, 4- and 7- Days Post Vaccination. Vaccine 2015, 33, 4513–4519. [Google Scholar]
- Moonen, P.; Jacobs, L.; Crienen, A.; Dekker, A. Detection of Carriers of Foot-and-Mouth Disease Virus among Vaccinated Cattle. Vet. Microbiol. 2004, 103, 151–160. [Google Scholar] [CrossRef]
- Golding, S.M.; Hedger, R.S.; Talbot, P. Radial Immuno-Diffusion and Serum Neutralisation Techniques for the Assay of Antibodies to Swine Vesicular Disease. Res. Vet. Sci. 1976, 20, 6. [Google Scholar] [CrossRef]
- Bouma, A.; Dekker, A.; de Jong, M.C. No Foot-and-Mouth Disease Virus Transmission between Individually Housed Calves. Vet. Microbiol. 2004, 98, 29–36. [Google Scholar] [CrossRef]
- Moonen, P.; Boonstra, J.; van der Honing, R.H.; Leendertse, C.B.; Jacobs, L.; Dekker, A. Validation of a Lightcycler-Based Reverse Transcription Polymerase Chain Reaction for the Detection of Foot-and-Mouth Disease Virus. J. Virol. Methods 2003, 113, 35–41. [Google Scholar] [CrossRef]
- Bates, D.; Maechler, M.; Bolker, B.; Walker, S. Fitting Linear Mixed-Effects Models Using Lme4. J. Stat. Softw. 2015, 67, 48. [Google Scholar] [CrossRef]
- R-Core-Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2019; Available online: https://www.R-Project.Org/ (accessed on 21 August 2021).
- OIE/FAO Foot-and-Mouth Disease Reference Laboratory Network. WRL_FMD_Reports. Annual Report 2016; World Reference Laboratory for FMD: Surrey, UK, 2016; Available online: OIE-FAO FMD Ref Lab Network Report 2016.pdf (wrlfmd.org) (accessed on 21 August 2021).
- Cox, S.J.; Barnett, P.V.; Dani, P.; Salt, J.S. Emergency Vaccination of Sheep against Foot-and-Mouth Disease: Protection against Disease and Reduction in Contact Transmission. Vaccine 1999, 17, 1858–1868. [Google Scholar] [CrossRef]
- Doel, T.R.; Williams, L.; Barnett, P.V. Emergency Vaccination against Foot-and-Mouth Disease: Rate of Development of Immunity and Its Implications for the Carrier State. Vaccine 1994, 12, 592–600. [Google Scholar] [CrossRef]
- Horsington, J.; Zhang, Z.; Bittner, H.; Hole, K.; Singanallur, N.B.; Alexandersen, S.; Vosloo, W. Early Protection in Sheep against Intratypic Heterologous Challenge with Serotype O Foot-and-Mouth Disease Virus Using High-Potency, Emergency Vaccine. Vaccine 2015, 33, 422–429. [Google Scholar] [CrossRef]
- Arzt, J.; Juleff, N.; Zhang, Z.; Rodriguez, L.L. The Pathogenesis of Foot-and-Mouth Disease I: Viral Pathways in Cattle. Transbound. Emerg. Dis. 2011, 58, 291–304. [Google Scholar] [CrossRef]
- Orsel, K.; de Jong, M.C.; Bouma, A.; Stegeman, J.A.; Dekker, A. The Effect of Vaccination on Foot and Mouth Disease Virus Transmission among Dairy Cows. Vaccine 2007, 25, 327–335. [Google Scholar] [CrossRef]
- Sutmoller, P.; McVicar, J.W.; Cottral, G.E. The Epizootiological Importance of Foot-and-Mouth Disease Carriers. I. Experimentally Produced Foot-and-Mouth Disease Carriers in Susceptible and Immune Cattle. Arch Gesamte Virusforsch 1968, 23, 227–235. [Google Scholar] [CrossRef]
- Backer, J.A.; Engel, B.; Dekker, A.; van Roermund, H.J.W. Vaccination against Foot-and-Mouth Disease II: Regaining FMD-Free Status. Prev. Vet. Med. 2012, 107, 41–50. [Google Scholar] [CrossRef]
- Backer, J.A.; Hagenaars, T.J.; Nodelijk, G.; van Roermund, H.J.W. Vaccination against Foot-and-Mouth Disease I: Epidemiological Consequences. Prev. Vet. Med. 2012, 107, 27–40. [Google Scholar] [CrossRef]
- Tenzin, A.D.; Vernooij, H.; Bouma, A.; Stegeman, A. Rate of Foot-and-Mouth Disease Virus Transmission by Carriers Quantified from Experimental Data. Risk Anal. 2008, 28, 303–309. [Google Scholar] [PubMed]
- Bertram, R.M.; Vu, L.T.; Pauszek, S.J.; Brito, B.P.; Hartwig, E.J.; Smoliga, G.R.; Hoang, B.H.; Phuong, N.T.; Stenfeldt, C.; Fish, I.H.; et al. Lack of Transmission of Foot-and-Mouth Disease Virus from Persistently Infected Cattle to Naïve Cattle under Field Conditions in Vietnam. Fontiers Vet. Sci. 2018, 5, 174. [Google Scholar] [CrossRef] [Green Version]
- Orsel, K.; Roest, H.I.; Elzinga-Bril, E.M.; van Hemert-Kluitenberg, F.; Dekker, A. Detection of Foot-and-Mouth Disease Virus in Infected Pigs by Rt-Pcr Four Weeks after Challenge. Vet. Rec. 2008, 162, 753–754. [Google Scholar] [CrossRef] [PubMed] [Green Version]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Singanallur, N.B.; Dekker, A.; Eblé, P.L.; van Hemert-Kluitenberg, F.; Weerdmeester, K.; Horsington, J.J.; Vosloo, W. Emergency FMD Serotype O Vaccines Protect Cattle against Heterologous Challenge with a Variant Foot-and-Mouth Disease Virus from the O/ME-SA/Ind2001 Lineage. Vaccines 2021, 9, 1110. https://doi.org/10.3390/vaccines9101110
Singanallur NB, Dekker A, Eblé PL, van Hemert-Kluitenberg F, Weerdmeester K, Horsington JJ, Vosloo W. Emergency FMD Serotype O Vaccines Protect Cattle against Heterologous Challenge with a Variant Foot-and-Mouth Disease Virus from the O/ME-SA/Ind2001 Lineage. Vaccines. 2021; 9(10):1110. https://doi.org/10.3390/vaccines9101110
Chicago/Turabian StyleSinganallur, Nagendrakumar Balasubramanian, Aldo Dekker, Phaedra Lydia Eblé, Froukje van Hemert-Kluitenberg, Klaas Weerdmeester, Jacquelyn J Horsington, and Wilna Vosloo. 2021. "Emergency FMD Serotype O Vaccines Protect Cattle against Heterologous Challenge with a Variant Foot-and-Mouth Disease Virus from the O/ME-SA/Ind2001 Lineage" Vaccines 9, no. 10: 1110. https://doi.org/10.3390/vaccines9101110
APA StyleSinganallur, N. B., Dekker, A., Eblé, P. L., van Hemert-Kluitenberg, F., Weerdmeester, K., Horsington, J. J., & Vosloo, W. (2021). Emergency FMD Serotype O Vaccines Protect Cattle against Heterologous Challenge with a Variant Foot-and-Mouth Disease Virus from the O/ME-SA/Ind2001 Lineage. Vaccines, 9(10), 1110. https://doi.org/10.3390/vaccines9101110





