Specificity of CD8+ T-Cell Responses Following Vaccination with Conserved Regions of HIV-1 in Nairobi, Kenya
Abstract
1. Introduction
2. Materials and Methods
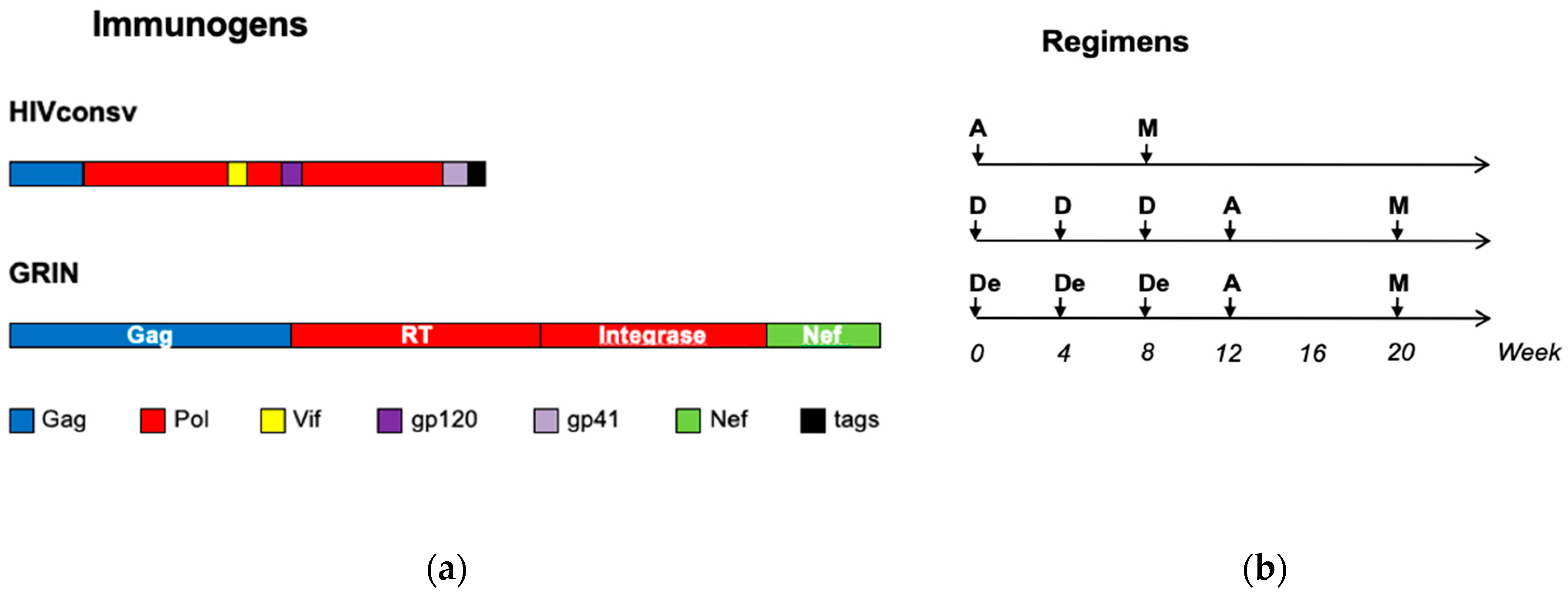
2.1. Trial HIV-CORE 004
2.2. Cryopreserved PBMC Samples
2.3. Peptides
2.4. HLA-Transfected Cell Lines
2.5. Cultured IFN-γ ELISPOT Assay
2.6. Intracellular Cytokine Staining (ICS) Assay and HLA Restriction
2.7. In Vitro Assay for Peptide-HLA-Class I Complex Formation
2.8. Cytotoxicity Assay
3. Results
3.1. The Study Subjects and Vaccination
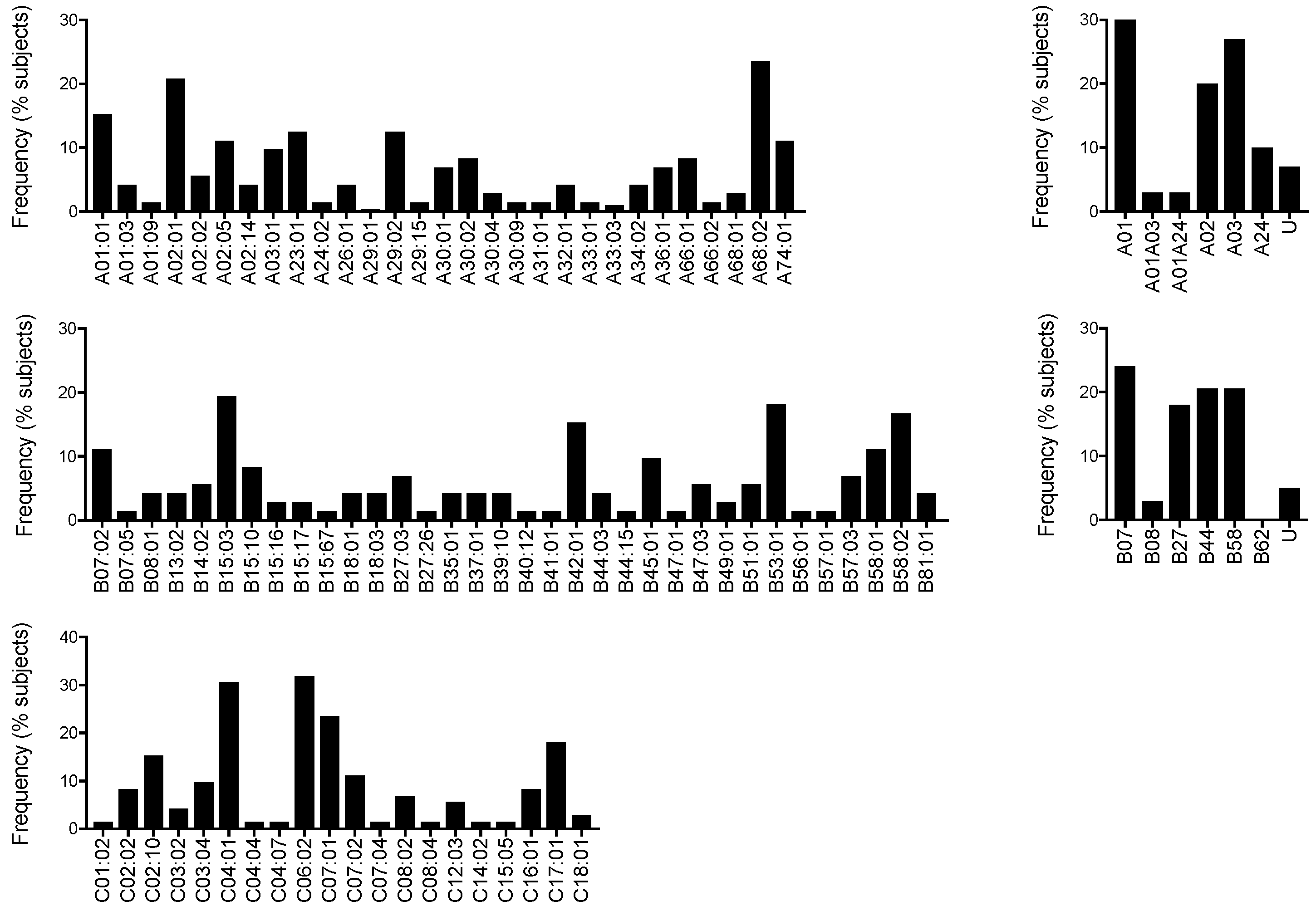
3.2. HLA Allele Frequency in Nairobi Volunteers
3.3. Definition of Optimal CD8+ T-Cell Epitopes Restricted by African HLAs
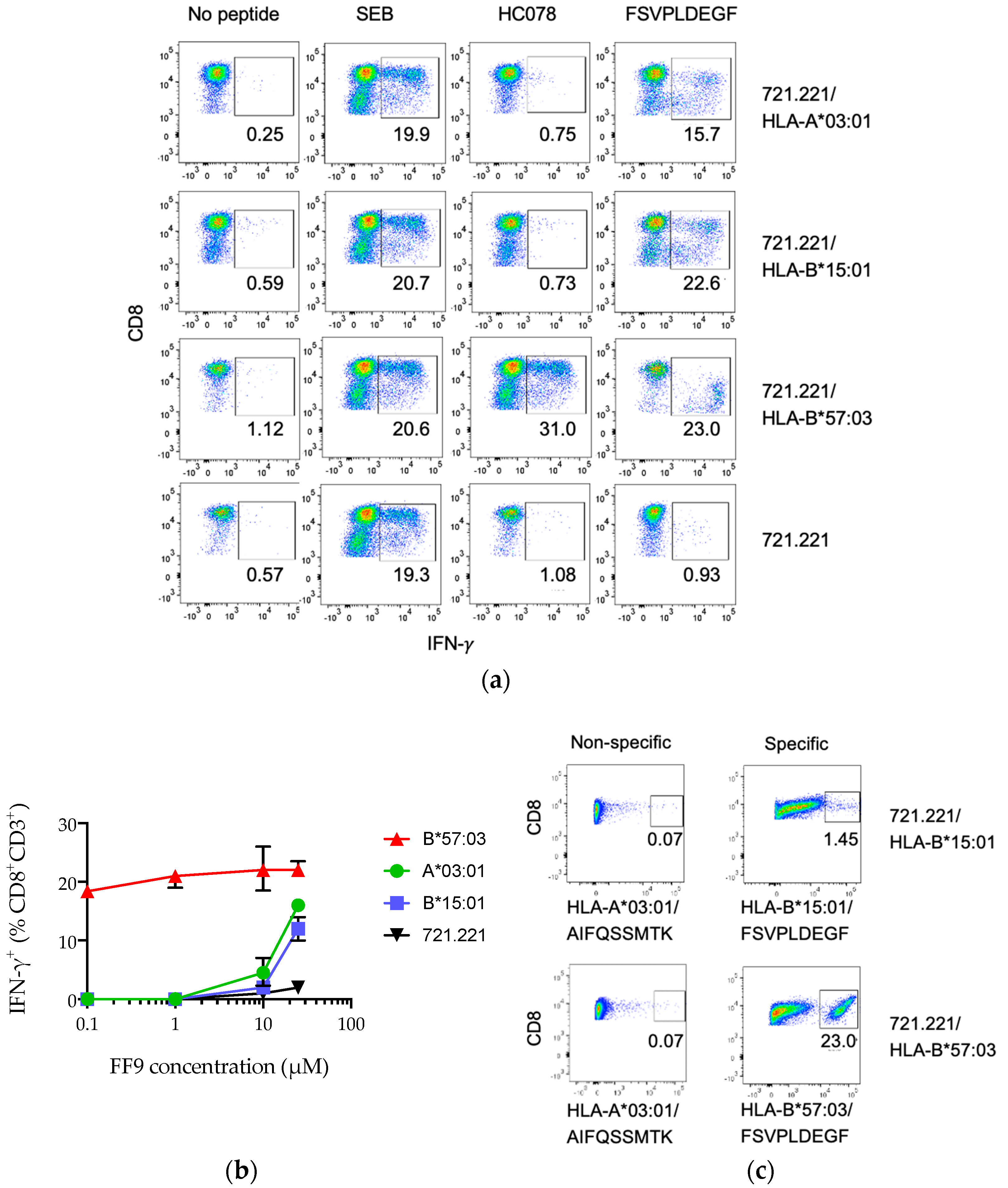
3.4. Peptide FF9 Stimulates Functional T-Cell Response via Three HLA Supertypes
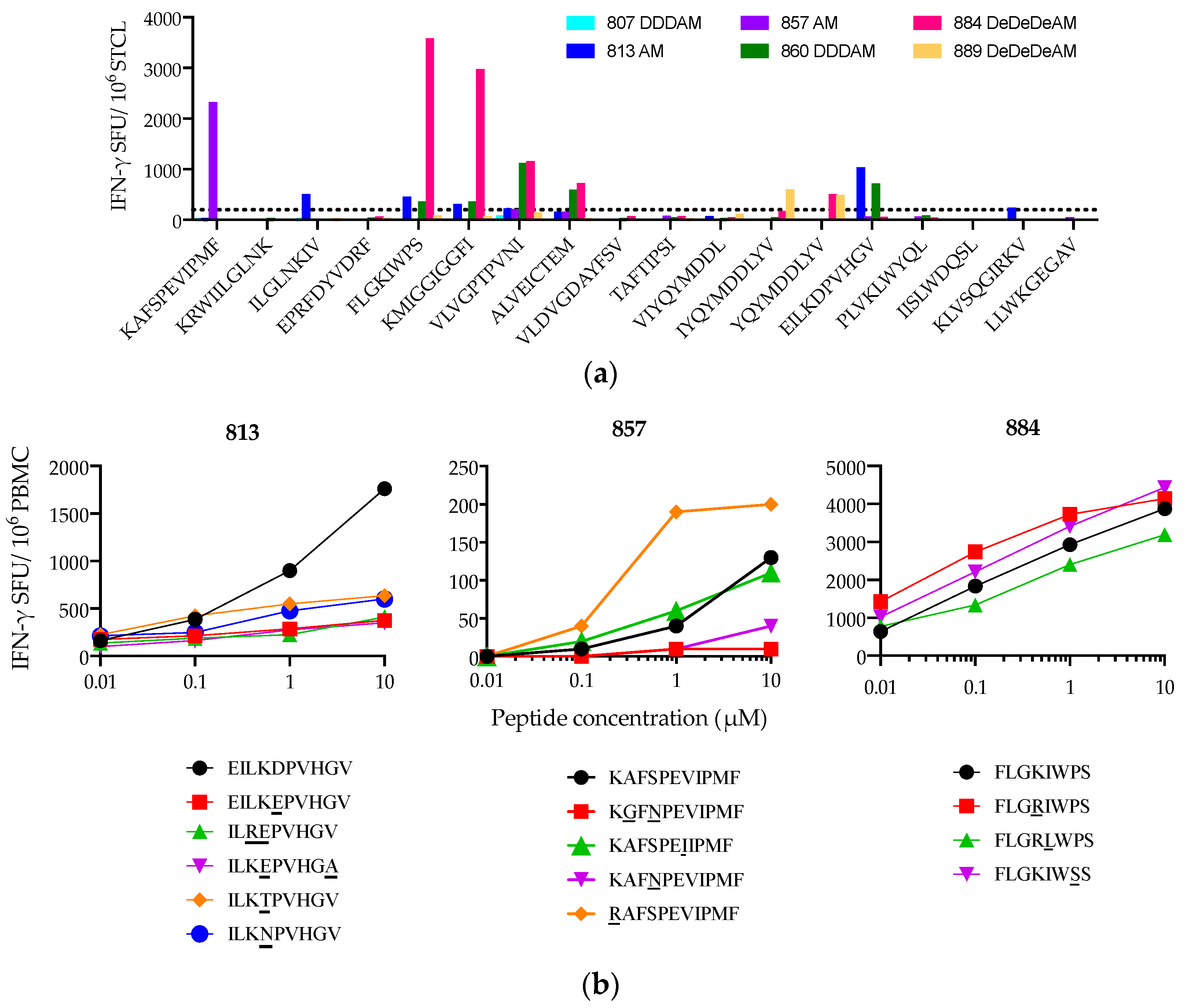
3.5. Heterogeneity of Vaccine-Elicited T-Cell Responses Restricted through HLA-A*02:01
3.6. Suboptimal In Vivo Response Rescued by Index Epitope Boost
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Townsend, A.; Bodmer, H. Antigen recognition by class I-restricted T lymphocytes. Annu. Rev. Immunol. 1989, 7, 601–624. [Google Scholar] [CrossRef] [PubMed]
- Trowsdale, J.; Campbell, R.D. Complexity in the major histocompatibility complex. Eur. J. Immunogenet. 1992, 19, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Bjorkman, P.; Parham, J.P. Structure, function, and diversity of class I major histocompatibility complex molecules. Annu. Rev. Biochem. 1990, 59, 253–288. [Google Scholar] [CrossRef] [PubMed]
- Hewitt, E.W. The MHC class I antigen presentation pathway: Strategies for viral immune evasion. Immunology 2003, 110, 163–169. [Google Scholar] [CrossRef] [PubMed]
- Fellay, J.; Frahm, N.; Shianna, K.V.; Cirulli, E.T.; Casimiro, D.R.; Robertson, M.N.; Haynes, B.F.; Geraghty, D.E.; McElrath, M.J.; Goldstein, D.B. Host genetic determinants of T cell responses to the MRKAd5 HIV-1 gag/pol/nef vaccine in the step trial. J. Infect. Dis. 2012, 203, 773–779. [Google Scholar] [CrossRef] [PubMed]
- Fellay, J.; Shianna, K.V.; Ge, D.; Colombo, S.; Ledergerber, B.; Weale, M.; Zhang, K.; Gumbs, C.; Castagna, A.; Cossarizza, A.; et al. A whole-genome association study of major determinants for host control of HIV-1. Science 2007, 317, 944–947. [Google Scholar] [CrossRef]
- Carrington, M.; O’Brien, S.J. The influence of HLA genotype on AIDS. Annu. Rev. Med. 2003, 54, 535–551. [Google Scholar] [CrossRef]
- Dalmasso, C.; Carpentier, W.; Meyer, L.; Rouzioux, C.; Goujard, C.; Chaix, M.L.; Lambotte, O.; Avettand-Fenoel, V.; Le Clerc, S.; de Senneville, L.D.; et al. Distinct genetic loci control plasma HIV-RNA and cellular HIV-DNA levels in HIV-1 infection: The ANRS Genome Wide Association 01 study. PLoS ONE 2008, 3, e3907. [Google Scholar] [CrossRef]
- Goonetilleke, N.; Liu, M.K.; Salazar-Gonzalez, J.F.; Ferrari, G.; Giorgi, E.; Ganusov, V.V.; Keele, B.F.; Learn, G.H.; Turnbull, E.L.; Salazar, M.G.; et al. The first T cell response to transmitted/founder virus contributes to the control of acute viremia in HIV-1 infection. J. Exp. Med. 2009, 206, 1253–1272. [Google Scholar] [CrossRef]
- Salazar-Gonzalez, J.F.; Salazar, M.G.; Keele, B.F.; Learn, G.H.; Giorgi, E.E.; Li, H.; Decker, J.M.; Wang, S.; Baalwa, J.; Kraus, M.H.; et al. Genetic identity, biological phenotype, and evolutionary pathways of transmitted/founder viruses in acute and early HIV-1 infection. J. Exp. Med. 2009, 206, 1273–1289. [Google Scholar] [CrossRef]
- Juarez-Molina, C.I.; Payne, R.; Soto-Nava, M.; Avila-Rios, S.; Valenzuela-Ponce, H.; Adland, E.; Leitman, E.; Brener, J.; Muenchhoff, M.; Branch, S.; et al. Impact of HLA selection pressure on HIV fitness at a population level in Mexico and Barbados. J. Virol. 2014, 88, 10392–10398. [Google Scholar] [CrossRef] [PubMed]
- Kawashima, Y.; Pfafferott, K.; Frater, J.; Matthews, P.; Payne, R.; Addo, M.; Gatanaga, H.; Fujiwara, M.; Hachiya, A.; Koizumi, H.; et al. Adaptation of HIV-1 to human leukocyte antigen class I. Nature 2009, 458, 641–645. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Picado, J.; Prado, J.G.; Fry, E.E.; Pfafferott, K.; Leslie, A.; Chetty, S.; Thobakgale, C.; Honeyborne, I.; Crawford, H.; Matthews, P.; et al. Fitness cost of escape mutations in p24 Gag in association with control of human immunodeficiency virus type 1. J. Virol. 2006, 80, 3617–3623. [Google Scholar] [CrossRef] [PubMed]
- Payne, R.; Muenchhoff, M.; Mann, J.; Roberts, H.E.; Matthews, P.; Adland, E.; Hempenstall, A.; Huang, K.H.; Brockman, M.; Brumme, Z.; et al. Impact of HLA-driven HIV adaptation on virulence in populations of high HIV seroprevalence. Proc. Nat. Acad. Sci. USA 2014, 111, E5393–E5400. [Google Scholar] [CrossRef] [PubMed]
- Yue, L.; Pfafferott, K.J.; Baalwa, J.; Conrod, K.; Dong, C.C.; Chui, C.; Rong, R.; Claiborne, D.T.; Prince, J.L.; Tang, J.; et al. Transmitted virus fitness and host T cell responses collectively define divergent infection outcomes in two HIV-1 recipients. PLoS Pathog. 2015, 11, e1004565. [Google Scholar] [CrossRef]
- Haynes, B.F.; Pantaleo, G.; Fauci, A.S. Toward an understanding of the correlates of protective immunity to HIV infection. Science 1996, 271, 324–328. [Google Scholar] [CrossRef]
- Kaur, G.; Mehra, N. Genetic determinants of HIV-1 infection and progression to AIDS: Immune response genes. Tissue Antigens 2009, 74, 373–385. [Google Scholar] [CrossRef]
- Leslie, A.; Price, D.A.; Mkhize, P.; Bishop, K.; Rathod, A.; Day, C.; Crawford, H.; Honeyborne, I.; Asher, T.E.; Luzzi, G.; et al. Differential selection pressure exerted on HIV by CTL targeting identical epitopes but restricted by distinct HLA alleles from the same HLA supertype. J. Immunol. 2006, 177, 4699–4708. [Google Scholar] [CrossRef]
- Leslie, A.J.; Pfafferott, K.J.; Chett, P.; Draenert, R.; Addo, M.M.; Feeney, M.; Tang, Y.; Holmes, E.C.; Allen, T.; Prado, J.G.; et al. HIV evolution: CTL escape mutation and reversion after transmission. Nat. Med. 2004, 10, 282–289. [Google Scholar] [CrossRef]
- Sette, A.; Sidney, J. Nine major HLA class I supertypes account for the vast preponderance of HLA-A and -B polymorphism. Immunogenetics 1999, 50, 201–212. [Google Scholar] [CrossRef]
- Sidney, J.; Peters, B.; Frahm, N.; Brander, C.; Sette, A. HLA class I supertypes: A revised and updated classification. BMC Immunol. 2008, 9, 1. [Google Scholar] [CrossRef]
- Trachtenberg, E.; Korber, B.; Sollars, C.; Kepler, T.B.; Hraber, P.T.; Hayes, E.; Funkhouser, R.; Fugate, M.; Theiler, J.; Hsu, Y.S.; et al. Advantage of rare HLA supertype in HIV disease progression. Nat. Med. 2003, 9, 928–935. [Google Scholar] [CrossRef]
- Bodmer, W.F. Evolutionary significance of the HL-A system. Nature 1972, 237, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Hill, A.V.; Allsopp, C.E.; Kwiatkowski, D.; Anstey, N.M.; Twumasi, P.; Rowe, P.A.; Bennett, S.; Brewster, D.; McMichael, A.J.; Greenwood, B.M. Common west African HLA antigens are associated with protection from severe malaria. Nature 1991, 352, 595–600. [Google Scholar] [PubMed]
- Hughes, A.L.; Ota, T.; Nei, M. Positive Darwinian selection promotes charge profile diversity in the antigen-binding cleft of class I major-histocompatibility-complex molecules. Mol. Biol. Evol. 1990, 7, 515–524. [Google Scholar] [CrossRef] [PubMed]
- Little, A.M.; Parham, P. Polymorphism and evolution of HLA class I and II genes and molecules. Rev. Immunogenet. 1999, 1, 105–123. [Google Scholar]
- Demers, K.R.; Reuter, M.A.; Betts, M.R. CD8(+) T-cell effector function and transcriptional regulation during HIV pathogenesis. Immunol. Rev. 2013, 254, 190–206. [Google Scholar] [CrossRef]
- Eller, M.A.; Goonetilleke, N.; Tassaneetrithep, B.; Eller, L.A.; Costanzo, M.C.; Johnson, S.; Betts, M.R.; Krebs, S.J.; Slike, B.M.; Nitayaphan, S.; et al. Expansion of Inefficient HIV-Specific CD8 T Cells during Acute Infection. J. Virol. 2016, 90, 4005–4016. [Google Scholar] [CrossRef]
- Makedonas, G.; Betts, M.R. Living in a house of cards: Re-evaluating CD8+ T-cell immune correlates against HIV. Immunol. Rev. 2011, 239, 109–124. [Google Scholar] [CrossRef]
- Walker, B.; McMichael, A. The T-Cell Response to HIV. Cold Spring Harb. Perspect. Med. 2012, 2, 1–19. [Google Scholar] [CrossRef]
- Walker, B.D.; Korber, B.T. Immune control of HIV: The obstacles of HLA and viral diversity. Nat. Immunol. 2001, 2, 473–475. [Google Scholar] [CrossRef] [PubMed]
- Betts, M.R.; Ambrozak, D.R.; Douek, D.C.; Bonhoeffer, S.; Brenchley, J.M.; Casazza, J.P.; Koup, R.A.; Picker, L.J. Analysis of total human immunodeficiency virus (HIV)-specific CD4(+) and CD8(+) T-cell responses: Relationship to viral load in untreated HIV infection. J. Virol. 2001, 75, 11983–11991. [Google Scholar] [CrossRef] [PubMed]
- Betts, M.R.; Nason, M.C.; West, S.M.; De Rosa, S.C.; Migueles, S.A.; Abraham, J.; Lederman, M.M.; Benito, J.M.; Goepfert, P.A.; Connors, M.; et al. HIV nonprogressors preferentially maintain highly functional HIV-specific CD8+ T cells. Blood 2006, 107, 4781–4789. [Google Scholar]
- Ferrari, G.; Korber, B.; Goonetilleke, N.; Liu, M.K.; Turnbull, E.L.; Salazar-Gonzalez, J.F.; Hawkins, N.; Self, S.; Watson, S.; Betts, M.R.; et al. Relationship between functional profile of HIV-1 specific CD8 T cells and epitope variability with the selection of escape mutants in acute HIV-1 infection. PLoS Pathog 2011, 7, e1001273. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, S.; Deleage, C.; Darko, S.; Ransier, A.; Truong, D.P.; Agarwal, D.; Japp, A.S.; Wu, V.H.; Kuri-Cervantes, L.; Abdel-Mohsen, M.; et al. Elite control of HIV is associated with distinct functional and transcriptional signatures in lymphoid tissue CD8(+) T cells. Sci. Transl. Med. 2019, 11. [Google Scholar] [CrossRef] [PubMed]
- Price, D.A.; Brenchley, J.M.; Ruff, L.E.; Betts, M.R.; Hill, B.J.; Roederer, M.; Koup, R.A.; Migueles, R.A.; Gostick, E.; Wooldridge, L.; et al. Avidity for antigen shapes clonal dominance in CD8+ T cell populations specific for persistent DNA viruses. J. Exp. Med. 2005, 202, 1349–1361. [Google Scholar] [CrossRef] [PubMed]
- Frahm, N.; Kiepiela, P.; Adams, S.; Linde, C.H.; Hewitt, H.S.; Sango, K.; Feeney, M.E.; Addo, M.M.; Lichterfeld, M.; Lahaie, M.P.; et al. Control of human immunodeficiency virus replication by cytotoxic T lymphocytes targeting subdominant epitopes. Nat. Immunol. 2006, 7, 173–178. [Google Scholar] [CrossRef]
- Kunwar, P.; Hawkins, N.; Dinges, W.L.; Liu, Y.; Gabriel, E.E.; Swan, D.A.; Stevens, C.E.; Maenza, J.; Collier, A.C.; Mullins, J.I.; et al. Superior Control of HIV-1 Replication by CD8+ T Cells Targeting Conserved Epitopes: Implications for HIV Vaccine Design. PLoS ONE 2013, 8, e64405. [Google Scholar] [CrossRef]
- Mothe, B.; Llano, A.; Ibarrondo, J.; Daniels, M.; Miranda, C.; Zamarreno, J.; Bach, V.; Zuniga, R.; Perez-Alvarez, S.; Berger, C.T.; et al. Definition of the viral targets of protective HIV-1-specific T cell responses. J. Transl. Med. 2011, 9, 208. [Google Scholar] [CrossRef]
- Murakoshi, H.; Akahoshi, T.; Koyanagi, M.; Chikata, T.; Naruto, T.; Maruyama, R.; Tamura, Y.; Ishizuka, N.; Gatanaga, H.; Oka, S.; et al. Clinical Control of HIV-1 by Cytotoxic T Cells Specific for Multiple Conserved Epitopes. J. Virol. 2015, 89, 5330–5339. [Google Scholar] [CrossRef]
- Murakoshi, H.; Zou, C.; Kuse, N.; Akahoshi, T.; Chikata, T.; Gatanaga, H.; Oka, S.; Hanke, T.; Takiguchi, M. CD8+ T cells specific for conserved, cross-reactive Gag epitopes with strong ability to suppress HIV-1 replication. Retrovirology 2018, 15, 46. [Google Scholar] [CrossRef] [PubMed]
- Ondondo, B.; Murakoshi, H.; Clutton, G.; Abdul-Jawad, S.; Wee, E.G.; Gatanaga, H.; Oka, S.; McMichael, A.J.; Takiguchi, M.; Korber, B.; et al. Novel Conserved-region T-cell Mosaic Vaccine With High Global HIV-1 Coverage Is Recognized by Protective Responses in Untreated Infection. Mol. Ther. 2016, 24, 832–842. [Google Scholar] [CrossRef] [PubMed]
- Zou, C.; Murakoshi, H.; Kuse, N.; Akahoshi, T.; Chikata, T.; Gatanaga, H.; Oka, S.; Hanke, T.; Takiguchi, M. Effective Suppression of HIV-1 Replication by Cytotoxic T Lymphocytes Specific for Pol Epitopes in Conserved Mosaic Vaccine Immunogens. J. Virol. 2019, 93, e02142–e02218. [Google Scholar] [CrossRef] [PubMed]
- Altfeld, M.; Addo, M.M.; Rosenberg, E.S.; Hecht, F.M.; Lee, P.K.; Vogel, M.; Yu, X.G.; Draenert, R.; Johnston, M.N.; Strick, D.; et al. Influence of HLA-B57 on clinical presentation and viral control during acute HIV-1 infection. AIDS 2003, 17, 2581–2591. [Google Scholar]
- Altfeld, M.; Kalife, E.T.; Qi, Y.; Streeck, H.; Lichterfeld, M.; Johnston, M.N.; Burgett, N.; Swartz, M.E.; Yang, A.; Alter, G.; et al. HLA Alleles Associated with Delayed Progression to AIDS Contribute Strongly to the Initial CD8(+) T Cell Response against HIV-1. PLoS. Med. 2006, 3, e403. [Google Scholar] [CrossRef]
- Bihl, F.; Frahm, N.; Di Giammarino, L.; Sidney, J.; John, M.; Yusim, K.; Woodberry, T.; Sango, K.; Hewitt, H.S.; Henry, L.; et al. Impact of HLA-B alleles, epitope binding affinity, functional avidity, and viral coinfection on the immunodominance of virus-specific CTL responses. J. Immunol. 2006, 176, 4094–4101. [Google Scholar]
- Crawford, H.; Lumm, W.; Leslie, A.; Schaefer, M.; Boeras, D.; Prado, J.G.; Tang, J.; Farmer, P.; Ndung’u, T.; Lakhi, S.; et al. Evolution of HLA-B*5703 HIV-1 escape mutations in HLA-B*5703-positive individuals and their transmission recipients. J. Exp. Med. 2009, 206, 909–921. [Google Scholar] [CrossRef]
- Flores-Villanueva, P.O.; Yunis, E.J.; Delgado, J.C.; Vittinghoff, E.; Buchbinder, S.; Leung, J.Y.; Uglialoro, A.M.; Clavijo, O.P.; Rosenberg, E.S.; Kalams, S.A.; et al. Control of HIV-1 viremia and protection from AIDS are associated with HLA-Bw4 homozygosity. Proc. Nat. Acad. Sci. USA 2001, 98, 5140–5145. [Google Scholar] [CrossRef]
- Goulder, P.J.; Walker, B.D. HIV and HLA class I: An evolving relationship. Immunity 2012, 37, 426–440. [Google Scholar] [CrossRef]
- Borthwick, N.; Ahmed, T.; Ondondo, B.; Hayes, P.; Rose, A.; Ebrahimsa, U.; Hayton, E.J.; Black, A.; Bridgeman, A.; Rosario, M.; et al. Vaccine-elicited human T cells recognizing conserved protein regions inhibit HIV-1. Mol. Ther. 2014, 22, 464–475. [Google Scholar] [CrossRef]
- Hanke, T. Conserved immunogens in prime-boost strategies for the next-generation HIV-1 vaccines. Expert. Opin. Biol. Ther. 2014, 14, 601–616. [Google Scholar] [CrossRef] [PubMed]
- Hanke, T. Aiming for protective T-cell responses: A focus on the first generation conserved-region HIVconsv vaccines in preventive and therapeutic clinical trials. Expert. Rev. Vaccines 2019, 18, 1029–1041. [Google Scholar] [CrossRef] [PubMed]
- Letourneau, S.; Im, E.-J.; Mashishi, T.; Brereton, C.; Bridgeman, A.; Yang, H.; Dorrell, L.; Dong, T.; Korber, B.; McMichael, A.J.; et al. Design and pre-clinical evaluation of a universal HIV-1 vaccine. PLoS ONE 2007, 2, e984. [Google Scholar] [CrossRef] [PubMed]
- Murakoshi, H.; Kuse, N.; Akahoshi, T.; Zhang, Y.; Chikata, T.; Borghan, M.A.; Gatanaga, H.; Oka, S.; Sakai, K.; Takiguchi, M. Broad Recognition of Circulating HIV-1 by HIV-1-Specific Cytotoxic T-Lymphocytes with Strong Ability to Suppress HIV-1 Replication. J. Virol. 2019, 93, e01480–e01518. [Google Scholar] [CrossRef]
- Rolland, M.; Nickle, D.C.; Mullins, J.I. HIV-1 group M conserved elements vaccine. PLoS Pathog. 2007, 3, e157. [Google Scholar] [CrossRef]
- Borthwick, N.J.; Lane, T.; Moyo, N.; Crook, A.; Shim, J.M.; Baines, I.; Wee, E.G.; Hawkins, P.H.; Gillmore, J.D.; Hanke, T.; et al. Randomized phase I trial HIV-CORE 003: Depletion of serum amyloid P component and immunogenicity of DNA vaccination against HIV-1. PLoS ONE 2018, 13, e0197299. [Google Scholar] [CrossRef]
- Fidler, S.; Stohr, W.; Pace, M.; Dorrell, L.; Lever, A.; Pett, S.; Kinloch-de Loes, S.; Fox, J.; Clarke, A.; Nelson, M.; et al. Antiretroviral therapy alone versus antiretroviral therapy with a kick and kill approach, on measures of the HIV reservoir in participants with recent HIV infection (the RIVER trial): A phase 2, randomised trial. Lancet 2020, 395, 888–898. [Google Scholar] [CrossRef]
- Mothe, B.; Manzardo, C.; Snachez-Bernabeau, A.; Coll, P.; Moron-Lopez, S.; Puertas, M.C.; Rosas, M.; Cobarsi, P.; Escrig, N.; Perez-Alvarez, N.; et al. Therapeutic vaccination refocused T-cell responses to conserved regions of HIV-1 in early reated individuals (BCN 01 study). Lancet eClinMed 2019, 1, 65–80. [Google Scholar] [CrossRef]
- Mothe, B.; Rosas-Umbert, M.; Coll, P.; Manzardo, C.; Puertas, M.C.; Morón-López, S.; Llano, A.; Miranda, C.; Cedeño, S.; López, M.; et al. HIVconsv vaccines and romidepsin in early-treated HIV-1-infected individuals: Safety, immunogenicity and effect on the viral reservoir (study BCN 02). Front. Immunol. 2020. [Google Scholar] [CrossRef]
- Mutua, G.; Farah, B.; Langat, R.; Indangasi, J.; Ogola, S.; Onsembe, B.; Kopycinski, J.T.; Hayes, P.; Ashraf, A.; Borthwick, N.J.; et al. Broad HIV-1 inhibition in vitro by vaccine-elicited CD8+ T cells in African adults. Mol. Ther. Methods Clin. Dev. 2016, 3, 16061. [Google Scholar] [CrossRef]
- Ahmed, T.; Borthwick, N.J.; Gilmour, J.; Hayes, P.; Dorrell, L.; Hanke, T. Control of HIV-1 replication in vitro by vaccine-induced human CD8 T cells through conserved subdominant Pol epitopes. Vaccine 2016, 34, 1215–1224. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Borthwick, N.; Lin, Z.; Akahoshi, T.; Llano, A.; Silva-Arrieta, S.; Ahmed, T.; Dorrell, L.; Brander, C.; Murakoshi, H.; Takiguchi, M.; et al. Novel, in-natural-infection subdominant HIV-1 CD8+ T-cell epitopes revealed in human recipients of conserved-region T-cell vaccines. PLoS ONE 2017, 12, e0176418. [Google Scholar] [CrossRef] [PubMed]
- Borthwick, N.; Silva-Arrieta, S.; Llano, A.; Takiguchi, M.; Brander, C.; Hanke, T. Novel Nested Peptide Epitopes Recognized by CD4(+) T Cells Induced by HIV-1 Conserved-Region Vaccines. Vaccines (Basel) 2020, 8, 28. [Google Scholar] [CrossRef] [PubMed]
- Falk, K.; Rotzschke, O.; Takiguchi, M.; Gnau, V.; Stevanovic, S.; Jung, G.; Rammensee, H.G. Peptide motifs of HLA-B58, B60, B61, and B62 molecules. Immunogenetics 1995, 41, 165–168. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, J.; Kariyone, A.; Akiyama, N.; Kano, K.; Takiguchi, M. Presentation of human minor histocompatibility antigens by HLA-B35 and HLA-B38 molecules. Proc. Natl. Acad. Sci. USA 1990, 87, 2583–2587. [Google Scholar] [CrossRef] [PubMed]
- Mpendo, J.; Mutua, G.; Nyombayire, J.; Ingabire, R.; Nanvubya, A.; Anzala, O.; Karita, E.; Hayes, P.; Kopycinski, J.; Dally, L.; et al. A Phase I Double Blind, Placebo-Controlled, Randomized Study of the Safety and Immunogenicity of Electroporated HIV DNA with or without Interleukin 12 in Prime-Boost Combinations with an Ad35 HIV Vaccine in Healthy HIV-Seronegative African Adults. PLoS ONE 2015, 10, e0134287. [Google Scholar] [CrossRef]
- Omosa-Manyonyi, G.; Mpendo, J.; Ruzagira, E.; Kilembe, W.; Chomba, E.; Roman, F.; Bourguignon, P.; Koutsoukos, M.; Collard, A.; Voss, G.; et al. A Phase I Double Blind, Placebo-Controlled, Randomized Study of the Safety and Immunogenicity of an Adjuvanted HIV-1 Gag-Pol-Nef Fusion Protein and Adenovirus 35 Gag-RT-Int-Nef Vaccine in Healthy HIV-Uninfected African Adults. PLoS ONE 2015, 10, e0125954. [Google Scholar] [CrossRef]
- LANL HMID CTL/CD8+ Epitope Summary. 2020. Available online: https://www.hiv.lanl.gov/content/immunology/tables/ctl_summary.html (accessed on 3 May 2020).
- Abdul-Jawad, S.; Ondondo, B.; van Hateren, A.; Gardner, A.; Elliott, T.; Korber, B.; Hanke, T. Increased Valency of Conserved-mosaic Vaccines Enhances the Breadth and Depth of Epitope Recognition. Mol. Ther. 2016, 24, 375–384. [Google Scholar] [CrossRef]
- Sliz, P.; Michielin, O.; Cerottini, J.C.; Luescher, I.; Romero, P.; Karplus, M.; Wiley, D.C. Crystal structures of two closely related but antigenically distinct HLA-A2/melanocyte-melanoma tumor-antigen peptide complexes. J. Immunol. 2001, 167, 3276–3284. [Google Scholar] [CrossRef]
- Roider, J.; Meissner, T.; Kraut, F.; Vollbrecht, T.; Stirner, R.; Bogner, J.R.; Draenert, R. Comparison of experimental fine-mapping to in silico prediction results of HIV-1 epitopes reveals ongoing need for mapping experiments. Immunology 2014, 143, 193–201. [Google Scholar] [CrossRef]
- Samri, A.; Haas, G.; Duntze, J.; Bouley, J.M.; Calvez, V.; Katlama, C.; Autran, B. Immunogenicity of mutations induced by nucleoside reverse transcriptase inhibitors for human immunodeficiency virus type 1-specific cytotoxic T cells. J. Virol. 2000, 74, 9306–9312. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Vollbrecht, T.; Eberle, J.; Roider, J.; Bühler, S.; Stirner, R.; Henrich, N.; Seybold, U.; Bogner, J.R.; Draenert, R. Control of M184V HIV-1 mutants by CD8 T-cell responses. Med. Microbiol. Immunol. 2012, 201, 201–211. [Google Scholar] [CrossRef] [PubMed]
- Johnson, R.P.; Trocha, A.; Buchanan, T.M.; Walker, B.D. Identification of overlapping HLA class I-restricted cytotoxic T cell epitopes in a conserved region of the human immunodeficiency virus type 1 envelope glycoprotein: Definition of minimum epitopes and analysis of the effects of sequence variation. J. Exp. Med. 1992, 175, 961–971. [Google Scholar]
- Johnson, R.P.; Trocha, A.; Buchanan, T.M.; Walker, B.D. Recognition of a highly conserved region of human immunodeficiency virus type 1 gp120 by an HLA-Cw4-restricted cytotoxic T-lymphocyte clone. J. Virol. 1993, 67, 438–445. [Google Scholar] [PubMed]
- Kloverpris, H.N.; Stryhn, A.; Harndahl, M.; Payne, R.; Towers, G.J.; Chen, F.; Riddell, L.; Walker, B.D.; Ndung’u, T.; Leslie, A.; et al. HLA-specific intracellular epitope processing shapes an immunodominance pattern for HLA-B*57 that is distinct from HLA-B*58:01. J. Virol. 2013, 87, 10889–10894. [Google Scholar] [CrossRef]
- Kloverpris, H.N.; Stryhn, A.; Harndahl, M.; van der Stok, M.; Payne, R.P.; Matthews, P.C.; Chen, F.; Riddell, L.; Walker, B.D.; Ndung’u, T.; et al. HLA-B*57 Micropolymorphism shapes HLA allele-specific epitope immunogenicity, selection pressure, and HIV immune control. J. Virol. 2012, 86, 919–929. [Google Scholar] [CrossRef]
- Llano, A.; Cedeño, S.; Arrieta, S.S.; Brander, C. Optimal HIV CTL epitopes update: Growing diversity in epitope length and HLA restriction; HIV Immunology and HIV/SIV Vaccine Databases; Theoretical Biology and Biophysics Group, Los Alamos National Laboratory: Los Alamos, NM, USA, 2019.
- Lano, A.; Williams, A.; Olvera, A.; Silva-Arrieta, S.; Brander, D. Best-Characterized HIV-1 CTL Epitopes: The 2013 Update; HIV Immunology and HIV/SIV Vaccine Databases; Theoretical Biology and Biophysics Group, Los Alamos National Laboratory: Los Alamos, NM, USA, 2013.
- Abidi, S.H.; Shahid, A.; Lakhani, L.S.; Shah, R.; Okinda, N.; Ojwang, P.; Abbas, F.; Rowland-Jones, S.; Ali, S. HIV-1 progression links with viral genetic variability and subtype, and patient’s HLA type: Analysis of a Nairobi-Kenyan cohort. Med. Microbiol. Immunol. 2014, 203, 57–63. [Google Scholar] [CrossRef]
- Bird, T.G.; Kaul, R.; Rostron, T.; Kimani, J.; Embree, J.; Dunn, P.P.; Bwayo, J.J.; Plummer, F.A.; Rowland-Jones, S.L.; Dong, T.; et al. HLA typing in a Kenyan cohort identifies novel class I alleles that restrict cytotoxic T-cell responses to local HIV-1 clades. AIDS 2002, 16, 1899–1904. [Google Scholar] [CrossRef][Green Version]
- Cao, K.; Moormann, A.M.; Lyke, K.E.; Masaberg, C.; Sumba, O.P.; Doumbo, O.K.; Koech, D.; Lancaster, A.; Nelson, M.; Meyer, D.; et al. Differentiation between African populations is evidenced by the diversity of alleles and haplotypes of HLA class I loci. Tissue Antigens 2004, 63, 293–325. [Google Scholar] [CrossRef]
- Khoja, S.; Ojwang, P.; Khan, S.; Okinda, N.; Harania, R.; Ali, S. Genetic analysis of HIV-1 subtypes in Nairobi, Kenya. PLoS ONE 2008, 3, e3191. [Google Scholar] [CrossRef]
- Tang, J.; Tang, S.; Lobashevsky, E.; Myracle, A.D.; Fideli, U.; Aldrovandi, G.; Allen, S.; Musonda, R.; Kaslow, R.A.; Zambia-UAB HIV Research Project. Favorable and unfavorable HLA class I alleles and haplotypes in Zambians predominantly infected with clade C human immunodeficiency virus type 1. J. Virol. 2002, 76, 8276–8284. [Google Scholar] [CrossRef] [PubMed]
- Los Alamos National Laboratory HIV Molecular Immunoloy Database. 2019. Available online: www.hiv.lanl.gov (accessed on 5 May 2019).
- Stewart-Jones, G.B.; Gillespie, G.; Overton, I.M.; Kaul, R.; Roche, P.; McMichael, A.J.; Rowland-Jones, S.; Jones, E.Y. Structures of three HIV-1 HLA-B*5703-peptide complexes and identification of related HLAs potentially associated with long-term nonprogression. J. Immunol. 2005, 175, 2459–2468. [Google Scholar] [CrossRef] [PubMed]
- Hansen, S.G.; Ford, J.C.; Lewis, M.S.; Ventura, A.B.; Hughes, C.M.; Coyne-Johnson, L.; Whizin, N.; Oswald, K.; Shoemaker, R.; Swanson, T.; et al. Profound early control of highly pathogenic SIV by an effector memory T-cell vaccine. Nature 2011, 473, 523–527. [Google Scholar] [CrossRef] [PubMed]
- Hansen, S.G.; Piatak, M.; Ventura, A.B.; Hughes, C.M.; Gilbride, R.M.; Ford, J.C.; Oswald, K.; Shoemaker, R.; Li, Y.; Lewis, M.S.; et al. Immune clearance of highly pathogenic SIV infection. Nature 2013, 502, 100–104. [Google Scholar] [CrossRef] [PubMed]
- Hansen, S.G.; Sacha, J.B.; Hughes, C.M.; Ford, J.C.; Burwitz, B.J.; Scholz, I.; Gilbride, R.M.; Lewis, M.S.; Gilliam, A.N.; Ventura, A.B.; et al. Cytomegalovirus vectors violate CD8+ T cell epitope recognition paradigms. Science 2013, 340, 1237874. [Google Scholar] [CrossRef] [PubMed]
- Hansen, S.G.; Vieville, C.; Whizin, N.; Coyne-Johnson, L.; Siess, D.C.; Drummond, D.D.; Legasse, A.W.; Axthelm, M.K.; Oswald, K.; Trubey, C.M.; et al. Effector memory T cell responses are associated with protection of rhesus monkeys from mucosal simian immunodeficiency virus challenge. Nat. Med. 2009, 15, 293–299. [Google Scholar] [CrossRef]
- Hansen, S.G.; Wu, H.L.; Burwitz, B.J.; Hughes, C.M.; Hammond, K.B.; Ventura, A.B.; Reed, J.S.; Gilbride, R.M.; Ainslie, E.; Morrow, D.W.; et al. Broadly targeted CD8(+) T cell responses restricted by major histocompatibility complex E. Science 2016, 351, 714–720. [Google Scholar] [CrossRef]
- Jönsson, P.; Southcombe, J.H.; Santos, A.M.; Huo, J.; Fernandes, R.A.; McColl, J.; Lever, M.; Evans, E.J.; Hudson, A.; Chang, V.T.; et al. Remarkably low affinity of CD4/peptide-major histocompatibility complex class II protein interactions. Proc. Natl. Acad. Sci. USA 2016, 113, 5682–5687. [Google Scholar] [CrossRef]
- Phetsouphanh, C.; Aldridge, D.; Marchi, E.; Munier, C.M.L.; Meyerowitz, J.; Murray, L.; Van Vuuren, C.; Goedhals, D.; Fidler, S.; Kelleher, A.; et al. Maintenance of Functional CD57+ Cytolytic CD4+ T Cells in HIV+ Elite Controllers. Front. Immunol. 2019, 10, 1844. [Google Scholar] [CrossRef]
- Assarsson, E.; Sidney, J.; Oseroff, C.; Pasquetto, V.; Bui, H.H.; Frahm, N.; Brander, C.; Peters, B.; Grey, H.; Sette, A. A quantitative analysis of the variables affecting the repertoire of T cell specificities recognized after vaccinia virus infection. J. Immunol. 2007, 178, 7890–7901. [Google Scholar] [CrossRef]
- Yewdell, J.W. Confronting complexity: Real-world immunodominance in antiviral CD8+ T cell responses. Immunity 2006, 25, 533–543. [Google Scholar] [CrossRef]
- Yewdell, J.W. Designing CD8+ T cell vaccines: It’s not rocket science (yet). Curr. Opin. Immunol. 2010, 2, 402–410. [Google Scholar] [CrossRef] [PubMed]
- Yewdell, J.W.; Bennink, J.R. Immunodominance in major histocompatibility complex class I-restricted T lymphocyte responses. Annu. Rev. Immunol. 1999, 17, 51–88. [Google Scholar] [CrossRef] [PubMed]
- Brehm, M.A.; Pinto, A.K.; Daniels, K.A.; Schneck, J.P.; Welsh, R.M.; Selin, L.K. T cell immunodominance and maintenance of memory regulated by unexpectedly cross-reactive pathogens. Nat. Immunol. 2002, 3, 627–634. [Google Scholar] [CrossRef] [PubMed]
- Probst, H.C.; Tschannen, K.; Gallimore, A.; Martinic, M.; Basler, M.; Dumrese, T.; Jones, E.; van den Broek, M.F. Immunodominance of an antiviral cytotoxic T cell response is shaped by the kinetics of viral protein expression. J. Immunol. 2003, 171, 5415–5422. [Google Scholar] [CrossRef]
- Van der Most, R.G.; Concepcion, R.J.; Oseroff, C.; Alexander, J.; Southwood, S.; Sidney, J.; Chesnut, R.W.; Ahmed, R.; Sette, A. Uncovering subdominant cytotoxic T-lymphocyte responses in lymphocytic choriomeningitis virus-infected BALB/c mice. J. Virol. 1997, 71, 5110–5114. [Google Scholar] [CrossRef]
- Van der Most, R.G.; Murali-Krishna, K.; Lanier, J.G.; Wherry, E.J.; Puglielli, M.T.; Blattman, J.N.; Sette, A.; Ahmed, R. Changing immunodominance patterns in antiviral CD8 T-cell responses after loss of epitope presentation or chronic antigenic stimulation. Virology 2003, 315, 93–102. [Google Scholar] [CrossRef]
- Frahm, N.; Yusim, K.; Suscovich, T.J.; Adams, S.; Sidney, J.; Hraber, P.; Hewitt, H.S.; Linde, C.H.; Kavanagh, D.G.; Woodberry, T.; et al. Extensive HLA class I allele promiscuity among viral CTL epitopes. Eur. J. Immunol. 2007, 37, 2419–2433. [Google Scholar] [CrossRef]
- Douek, D.C.; Betts, M.R.; Brenchley, J.M.; Hill, B.J.; Ambrozak, D.R.; Ngai, K.L.; Karandikar, N.J.; Casazza, J.P.; Koup, R.A. A novel approach to the analysis of specificity, clonality, and frequency of HIV-specific T cell responses reveals a potential mechanism for control of viral escape. J. Immunol. 2002, 168, 3099–3104. [Google Scholar] [CrossRef]
- Barouch, D.H.; O’Brien, K.L.; Simmons, N.L.; King, S.L.; Abbink, P.; Maxfield, L.F.; Sun, Y.H.; La Porte, A.; Riggs, A.M.; Lynch, D.M.; et al. Mosaic HIV-1 vaccines expand the breadth and depth of cellular immune responses in rhesus monkeys. Nat. Med. 2010, 16, 319–323. [Google Scholar] [CrossRef]
- Fischer, W.; Perkins, S.; Theiler, J.; Bhattacharya, T.; Yusim, K.; Funkhouser, R.; Kuiken, C.; Haynes, B.; Letvin, N.L.; Walker, B.D.; et al. Polyvalent vaccines for optimal coverage of potential T-cell epitopes in global HIV-1 variants. Nat. Med. 2007, 13, 100–106. [Google Scholar] [CrossRef] [PubMed]
- Theiler, J.; Yoon, H.; Yusim, K.; Picker, L.J.; Fruh, K.; Korber, B. Epigraph: A Vaccine Design Tool Applied to an HIV Therapeutic Vaccine and a Pan-Filovirus Vaccine. Sci. Rep. 2016, 6, 33987. [Google Scholar] [CrossRef] [PubMed]

| No. | Parental | VID 2 | Name | Shorter | Reported | Predicted 1 | Confirmed |
|---|---|---|---|---|---|---|---|
| Sequence | Sequence | HLA | HLA | HLA | |||
| HC014 | EWDRIYKRWIILGLN | 829 | YN10 | YKRWIILGLN | Not rep’d | B*27:03 | |
| HC049 | (K)NFPISPIETVPVKLK | 831 | SL10 | SPIETVPVKL | B*81:01 | B*81:01 | |
| IL9 | IETVPVKL | B*40:01 | |||||
| B*53:01 | |||||||
| HC078 | YFSVPLDEGFRKYTA | 873 | FF9 | FSVPLDEGF | B*57:03 | B*57:03 | B*57:03 |
| B*15:01 | B*15:01 | ||||||
| A*03:01 | A*03:01 | ||||||
| A*26:01 | |||||||
| HC088 | GSPAIFQSSMTKILE | 830 | AK9 | AIFQSSMTK | A*03:01 | A*03:01 | A*03:01 |
| A*11:01 | |||||||
| II9 | IFQSSMTKI | Not rep’d | B*51:01 | ||||
| 866 | SK11 | SPAIFQSSMTK | A*11:01 | B*07:05 ♣ | |||
| A*11:01 | |||||||
| B*53:01 ♣ | |||||||
| GM9 | GSPAIFQSSM | Not rep’d | B*07:05 | ||||
| SM9 | SPAIFQSSM | B7 | B*07:05 | B*07:05 ♣ | |||
| B*53:01 | B*53:01 ♣ | ||||||
| C*04:02 | |||||||
| HC164 | VQMAVFIHNFKRKGGI | 830 | AR9 | AVFIHNFKR | A*03:01 | A*03:01 | |
| A*74:01 | |||||||
| MR10 | MAVFIHNFKR | Not rep’d | A*03:01 | ||||
| A*66:01 | |||||||
| A*68:01 | |||||||
| MK9 | MAVFIHNFK | A*03:01 | A*03:01 | ||||
| A*74:01 | |||||||
| B*51:01 | |||||||
| VR8 | VFIHNFKR | Not rep’d | A*66:01 | ||||
| A*68:01 | |||||||
| 889 | MR10 | MAVFIHNFKR | Not rep’d | ||||
| MK9 | MAVFIHNFK | Not rep’d | C*02:02 | ||||
| VR8 | VFIHNFKR | Not rep’d | |||||
| HC176 | VVPRRKAKIIRDYGK | 831 | KK10 | KAKIIRDYGK | Not rep’d | ||
| 889 | KK8 | KIIRDYGK | Not rep’d | ||||
| KY8 | KAKIIRDY | Not rep’d | |||||
| HC177 | RKAKIIRDYGKQMAG | 810 | RK11 | RKAKIIRDYGK | Not rep’d | ||
| RY9 | RKAKIIRDY | B*15:03 | B*15:03 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mohamed, Y.S.; Borthwick, N.J.; Moyo, N.; Murakoshi, H.; Akahoshi, T.; Siliquini, F.; Hannoun, Z.; Crook, A.; Hayes, P.; Fast, P.E.; et al. Specificity of CD8+ T-Cell Responses Following Vaccination with Conserved Regions of HIV-1 in Nairobi, Kenya. Vaccines 2020, 8, 260. https://doi.org/10.3390/vaccines8020260
Mohamed YS, Borthwick NJ, Moyo N, Murakoshi H, Akahoshi T, Siliquini F, Hannoun Z, Crook A, Hayes P, Fast PE, et al. Specificity of CD8+ T-Cell Responses Following Vaccination with Conserved Regions of HIV-1 in Nairobi, Kenya. Vaccines. 2020; 8(2):260. https://doi.org/10.3390/vaccines8020260
Chicago/Turabian StyleMohamed, Yehia S., Nicola J. Borthwick, Nathifa Moyo, Hayato Murakoshi, Tomohiro Akahoshi, Francesca Siliquini, Zara Hannoun, Alison Crook, Peter Hayes, Patricia E. Fast, and et al. 2020. "Specificity of CD8+ T-Cell Responses Following Vaccination with Conserved Regions of HIV-1 in Nairobi, Kenya" Vaccines 8, no. 2: 260. https://doi.org/10.3390/vaccines8020260
APA StyleMohamed, Y. S., Borthwick, N. J., Moyo, N., Murakoshi, H., Akahoshi, T., Siliquini, F., Hannoun, Z., Crook, A., Hayes, P., Fast, P. E., Mutua, G., Jaoko, W., Silva-Arrieta, S., Llano, A., Brander, C., Takiguchi, M., & Hanke, T. (2020). Specificity of CD8+ T-Cell Responses Following Vaccination with Conserved Regions of HIV-1 in Nairobi, Kenya. Vaccines, 8(2), 260. https://doi.org/10.3390/vaccines8020260






