Differential Genome Replication of a Unique Single-Amino-Acid Mutation in the Adenovirus-4 Component of the Live Oral Adenovirus Type 4 and Type 7 Vaccine
Abstract
1. Introduction
2. Methods
2.1. Ethics Approval
2.2. Cell Lines and Adenovirus
2.3. Virus Quantification and Plaque Purification
2.4. In Vitro Infection of Human Lung Fibroblast WI-38 Cells
2.5. Human Fecal Specimens for HAdV-4 SNP Analysis
2.6. Next-Generation Sequencing (NGS) and Single-Nucleotide Polymorphism (SNP) Quantitative PCR Assay
2.7. Statistical Analysis
3. Results
3.1. Discovery of Mutant Virus in Oral Live HAdV-4 Vaccine
3.2. Mutant HAdV-4 Generates More Viral DNA than Wild-Type in WI-38 Cells
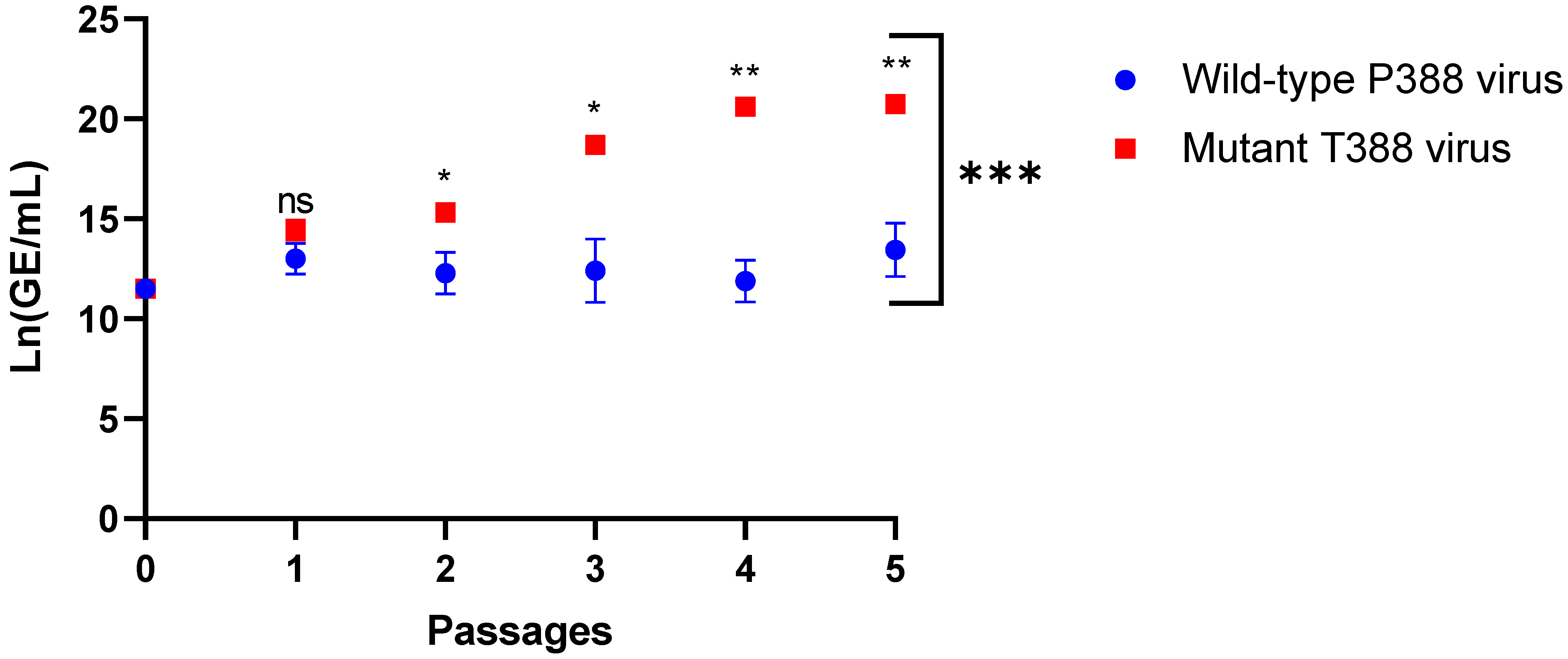
3.3. HAdV-4 Mutant T388 Outcompeted Wild-Type P388 in WI-38 Cells
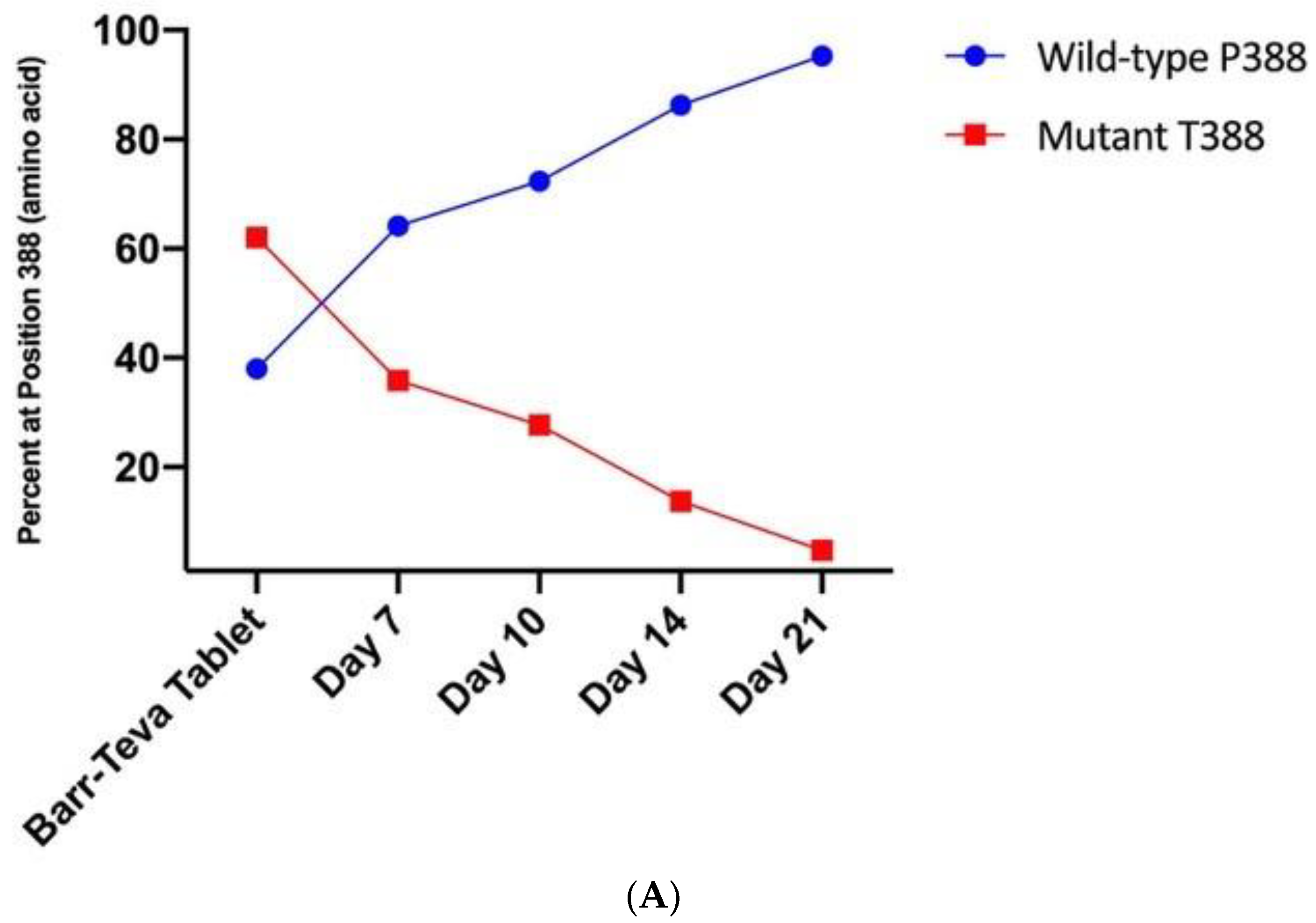
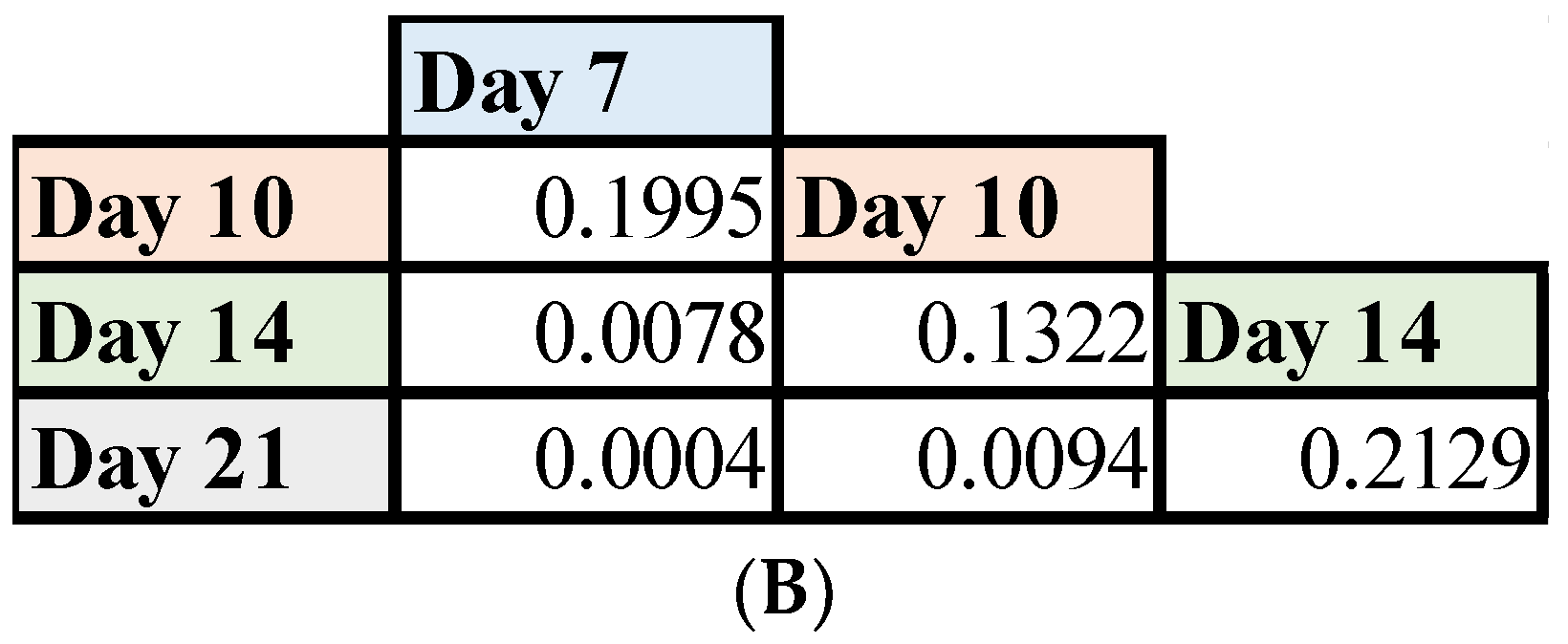
3.4. Mutant HAdV-4 Was Replication Deficient in Human Gastrointestinal Tract
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Disclaimer
References
- Probst, V.; Datyner, E.K.; Haddadin, Z.; Rankin, D.A.; Hamdan, L.; Rahman, H.K.; Spieker, A.; Stewart, L.S.; Guevara, C.; Yepsen, E.; et al. Human adenovirus species in children with acute respiratory illnesses. J. Clin. Virol. 2021, 134, 104716. [Google Scholar] [CrossRef] [PubMed]
- Mennechet, F.J.D.; Paris, O.; Ouoba, A.R.; Arenas, S.S.; Sirima, S.B.; Dzomo, G.R.T.; Diarra, A.; Traore, I.T.; Kania, D.; Eichholz, K.; et al. A review of 65 years of human adenovirus seroprevalence. Expert Rev. Vaccines 2019, 18, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Sivan, A.V.; Lee, T.; Binn, L.N.; Gaydos, J.C. Adenovirus-associated acute respiratory disease in healthy adolescents and adults: A literature review. Mil. Med. 2007, 172, 1198–1203. [Google Scholar] [CrossRef]
- Sanchez, J.L.; Cooper, M.J.; Myers, C.A.; Cummings, J.F.; Vest, K.G.; Russell, K.L.; Hiser, M.J.; Gaydos, C.A. Respiratory Infections in the U.S. Military: Recent Experience and Control. Clin. Microbiol. Rev. 2015, 28, 743–800. [Google Scholar] [CrossRef] [PubMed]
- Kolavic-Gray, S.A.; Binn, L.N.; Sanchez, J.L.; Cersovsky, S.B.; Polyak, C.S.; Mitchell-Raymundo, F.; Asher, L.V.; Vaughn, D.W.; Feighner, B.H.; Innis, B.L. Large Epidemic of Adenovirus Type 4 Infection among Military Trainees: Epidemiological, Clinical, and Laboratory Studies. Clin. Infect. Dis. 2002, 35, 808–818. [Google Scholar] [CrossRef] [PubMed]
- Top, F.H.; Buescher, E.L.; Bancroft, W.H.; Russell, P.K. Immunization with live types 7 and 4 adenovirus vaccines. II. Antibody response and protective effect against acute respiratory disease due to adenovirus type 7. J. Infect. Dis. 1971, 124, 155–160. [Google Scholar] [CrossRef]
- Top, F.H.; Dudding, B.A.; Russell, P.K.; Buescher, E.L. Control of respiratory disease in recruits with types 4 and 7 adenovirus vaccines. Am. J. Epidemiol. 1971, 94, 142–146. [Google Scholar] [CrossRef]
- Gaydos, C.A.; Gaydos, J.C. Adenovirus vaccines in the U.S. military. Mil. Med. 1995, 160, 300–304. [Google Scholar] [CrossRef]
- Russell, K.L.; Hawksworth, A.W.; Ryan, M.A.; Strickler, J.; Irvine, M.; Hansen, C.J.; Gray, G.C.; Gaydos, J.C. Vaccine-preventable adenoviral respiratory illness in US military recruits, 1999–2004. Vaccine 2006, 24, 2835–2842. [Google Scholar] [CrossRef]
- Hoke, C.H.; Snyder, C.E. History of the restoration of adenovirus type 4 and type 7 vaccine, live oral (Adenovirus Vaccine) in the context of the Department of Defense acquisition system. Vaccine 2013, 31, 1623–1632. [Google Scholar] [CrossRef]
- Hoke, C.H.; Hawksworth, A.; Snyder, C.E. Initial assessment of impact of adenovirus type 4 and type 7 vaccine on febrile respiratory illness and virus transmission in military basic trainees, March 2012. MSMR 2012, 19, 2–4. [Google Scholar] [PubMed]
- Kuschner, R.A.; Russell, K.L.; Abuja, M.; Bauer, K.M.; Faix, D.J.; Hait, H.; Henrick, J.; Jacobs, M.; Liss, A.; Lynch, J.A.; et al. A phase 3, randomized, double-blind, placebo-controlled study of the safety and efficacy of the live, oral adenovirus type 4 and type 7 vaccine, in U.S. military recruits. Vaccine 2013, 31, 2963–2971. [Google Scholar] [CrossRef] [PubMed]
- Choudhry, A.; Mathena, J.; Albano, J.D.; Yacovone, M.; Collins, L. Safety evaluation of adenovirus type 4 and type 7 vaccine live, oral in military recruits. Vaccine 2016, 34, 4558–4564. [Google Scholar] [CrossRef]
- Bandyopadhyay, A.S.; Garon, J.; Seib, K.; Orenstein, W.A. Polio vaccination: Past, present and future. Futur. Microbiol. 2015, 10, 791–808. [Google Scholar] [CrossRef]
- McNall, R.J.; Wharton, A.K.; Anderson, R.; Clemmons, N.; Lopareva, E.N.; Gonzalez, C.; Espinosa, A.; Probert, W.S.; Hacker, J.K.; Liu, G.; et al. Genetic characterization of mumps viruses associated with the resurgence of mumps in the United States: 2015–2017. Virus Res. 2020, 281, 197935. [Google Scholar] [CrossRef] [PubMed]
- Wright, K.E.; Dimock, K.; Brown, E.G. Biological characteristics of genetic variants of Urabe AM9 mumps vaccine virus. Virus Res. 2000, 67, 49–57. [Google Scholar] [CrossRef] [PubMed]
- Amexis, G.; Fineschi, N.; Chumakov, K. Correlation of Genetic Variability with Safety of Mumps Vaccine Urabe AM9 Strain. Virology 2001, 287, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Ryman, K.D.; Ledger, T.N.; Campbell, G.A.; Watowich, S.J.; Barrett, A.D.T. Mutation in a 17D-204 Vaccine Substrain-Specific Envelope Protein Epitope Alters the Pathogenesis of Yellow Fever Virus in Mice. Virology 1998, 244, 59–65. [Google Scholar] [CrossRef]
- Collins, N.D.; Beck, A.; Widen, S.G.; Wood, T.G.; Higgs, S.; Barrett, A.D.T. Structural and Nonstructural Genes Contribute to the Genetic Diversity of RNA Viruses. mBio 2018, 9, e01871-18. [Google Scholar] [CrossRef]
- Webster, A.; Leith, I.R.; Nicholson, J.; Hounsell, J.; Hay, R.T. Role of preterminal protein processing in adenovirus replication. J. Virol. 1997, 71, 6381–6389. [Google Scholar] [CrossRef]
- Botting, C.H.; Hay, R.T. Role of conserved residues in the activity of adenovirus preterminal protein. J. Gen. Virol. 2001, 82, 1917–1927. [Google Scholar] [CrossRef] [PubMed]
- Kato, S.E.; Chahal, J.S.; Flint, S.J. Reduced Infectivity of Adenovirus Type 5 Particles and Degradation of Entering Viral Genomes Associated with Incomplete Processing of the Preterminal Protein. J. Virol. 2012, 86, 13554–13565. [Google Scholar] [CrossRef] [PubMed]
- Houng, H.-S.H.; Clavio, S.; Graham, K.; Kuschner, R.; Sun, W.; Russell, K.L.; Binn, L.N. Emergence of a new human adenovirus type 4 (Ad4) genotype: Identification of a novel inverted terminal repeated (ITR) sequence from majority of Ad4 isolates from US military recruits. J. Clin. Virol. 2006, 35, 381–387. [Google Scholar] [CrossRef] [PubMed]
- Hang, J.; Vento, T.J.; Norby, E.A.; Jarman, R.G.; Keiser, P.B.; Kuschner, R.A.; Binn, L.N. Adenovirus type 4 respiratory infections with a concurrent outbreak of coxsackievirus A21 among United States Army Basic Trainees, a retrospective viral etiology study using next-generation sequencing. J. Med. Virol. 2017, 89, 1387–1394. [Google Scholar] [CrossRef] [PubMed]
- Purkayastha, A.; Su, J.; McGraw, J.; Ditty, S.E.; Hadfield, T.L.; Seto, J.; Russell, K.L.; Tibbetts, C.; Seto, D. Genomic and Bioinformatics Analyses of HAdV-4vac and HAdV-7vac, Two Human Adenovirus (HAdV) Strains That Constituted Original Prophylaxis against HAdV-Related Acute Respiratory Disease, a Reemerging Epidemic Disease. J. Clin. Microbiol. 2005, 43, 3083–3094. [Google Scholar] [CrossRef]
- Risso-Ballester, J.; Cuevas, J.M.; Sanjuán, R. Genome-Wide Estimation of the Spontaneous Mutation Rate of Human Adenovirus 5 by High-Fidelity Deep Sequencing. PLoS Pathog. 2016, 12, e1006013. [Google Scholar] [CrossRef]
- Lyons, A.; Longfield, J.; Kuschner, R.; Straight, T.; Binn, L.; Seriwatana, J.; Reitstetter, R.; Froh, I.B.; Craft, D.; McNabb, K.; et al. A double-blind, placebo-controlled study of the safety and immunogenicity of live, oral type 4 and type 7 adenovirus vaccines in adults. Vaccine 2008, 26, 2890–2898. [Google Scholar] [CrossRef]
- Chanock, R.M.; Ludwig, W.; Heubner, R.J.; Cate, T.R.; Chu, L.-W. Immunization by Selective Infection with Type 4 Adenovirus Grown in Human Diploid Tissue Culture: I. Safety and Lack of Oncogenicity and Tests for Potency in Volunteers. JAMA 1966, 195, 445–452. [Google Scholar] [CrossRef]
- Paris, R.; Kuschner, R.A.; Binn, L.; Thomas, S.J.; Colloca, S.; Nicosia, A.; Cortese, R.; Bailer, R.T.; Sullivan, N.; Koup, R.A. Adenovirus Type 4 and 7 Vaccination or Adenovirus Type 4 Respiratory Infection Elicits Minimal Cross-Reactive Antibody Responses to Nonhuman Adenovirus Vaccine Vectors. Clin. Vaccine Immunol. 2014, 21, 783–786. [Google Scholar] [CrossRef]
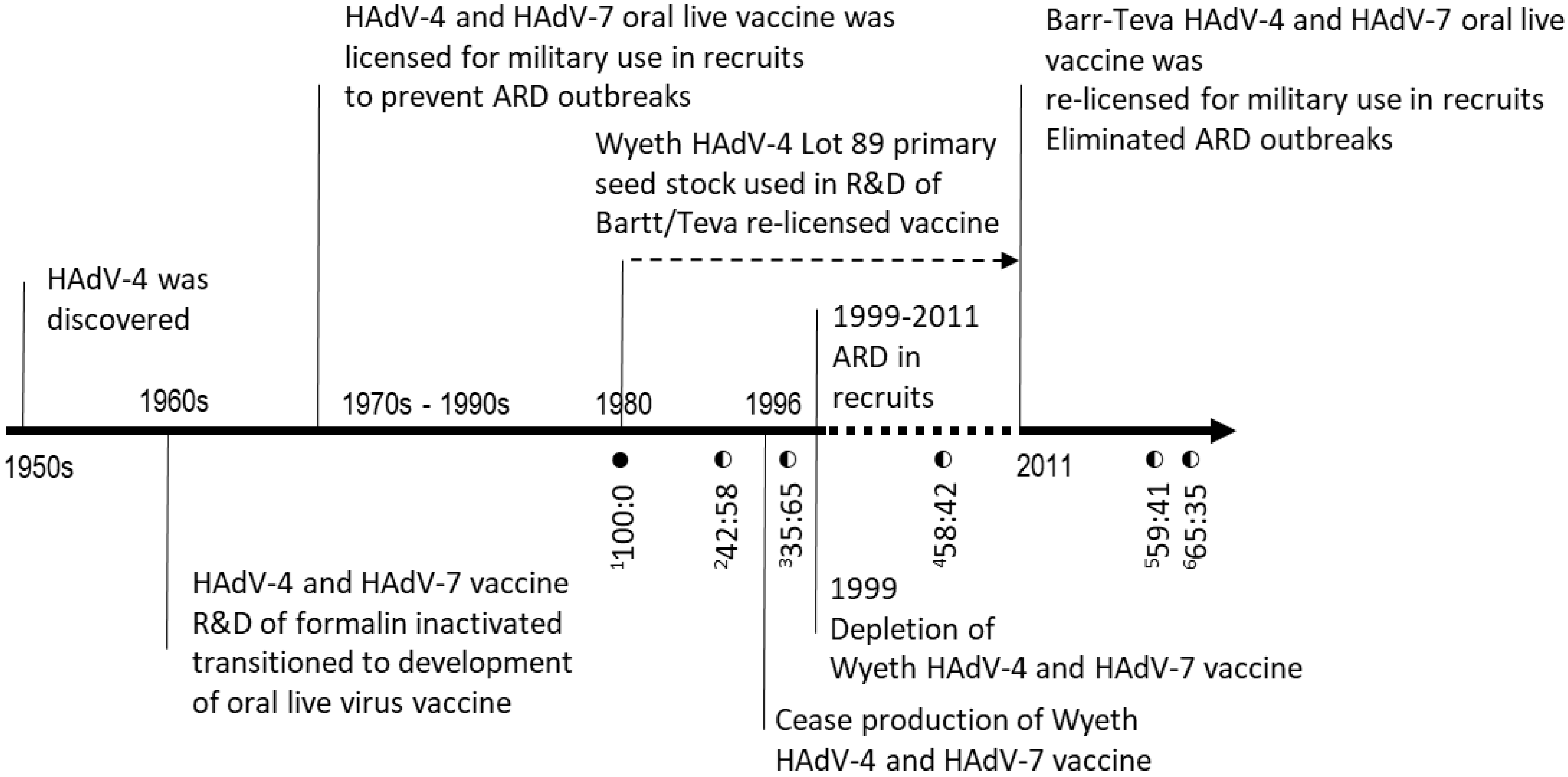


| Material ID | Material Type | Manufacturer | Nucleotide Percentages (%) at Position 9171 | |
|---|---|---|---|---|
| G9171 | T9171 | |||
| HAdV-4 CL68578 | Vaccine reference strain | Wyeth | 100 | 0 |
| Lot 089 | Seed stock | Wyeth | 100 | 0 |
| Lot 4948232 | Vaccine tablet | Wyeth | 42 | 58 |
| Lot 4958221 | Vaccine tablet | Wyeth | 35 | 65 |
| Lot F4A023015A | Vaccine tablet | Teva | 58 | 42 |
| Lot 34600363 | Vaccine tablet | Teva | 65 | 35 |
| Lot 34600390A | Vaccine tablet | Teva | 59 | 41 |
| Days | Mean Ln(GE/mL) P388 | Mean Ln(GE/mL) T388 | Mean Ln(GE/mL) Difference | Mean Difference StdErr * | p-Value * |
|---|---|---|---|---|---|
| 1 | 8.96 | 9.9 | 0.63 | 0.18 | 0.06 |
| 2 | 8.37 | 10.38 | 2.02 | 0.63 | 0.09 |
| 3 | 9.57 | 12.48 | 2.91 | 1.13 | 0.11 |
| 4 | 9.87 | 13.63 | 3.76 | 1.77 | 0.27 |
| 5 | 12.02 | 15.01 | 2.99 | 0.15 | <0.0001 |
| 6 | 13.09 | 16.18 | 3.09 | 0.26 | 0.001 |
| 7 | 13.34 | 16.42 | 3.08 | 0.38 | 0.01 |
| 8 | 14.22 | 17.17 | 2.95 | 0.21 | 0.0005 |
| 9 | 14.29 | 17.98 | 3.70 | 0.46 | 0.004 |
| 10 | 14.75 | 18.63 | 3.88 | 0.49 | 0.01 |
| Passage | Mean Ln(GE/mL) P388 | Mean Ln(GE/mL) T388 | Mean Ln(GE/mL) Difference | Mean Difference Standard Error | p * Value |
|---|---|---|---|---|---|
| 0 | 11.51 | 11.51 | 0.00 | 0.00 | 1.0 |
| 1 | 13.02 | 14.44 | 1.42 | 0.53 | 0.07 |
| 2 | 12.27 | 15.31 | 3.04 | 0.61 | 0.04 |
| 3 | 12.40 | 18.70 | 6.31 | 0.92 | 0.02 |
| 4 | 11.88 | 20.61 | 8.73 | 0.63 | 0.003 |
| 5 | 13.44 | 20.73 | 7.29 | 0.77 | 0.01 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Collins, N.D.; Beaty, S.; Wallace, E.; Li, Y.; Sanborn, M.; Yang, Y.; Adhikari, A.; Shabram, P.; Warfield, K.; Karasavvas, N.; et al. Differential Genome Replication of a Unique Single-Amino-Acid Mutation in the Adenovirus-4 Component of the Live Oral Adenovirus Type 4 and Type 7 Vaccine. Vaccines 2023, 11, 1144. https://doi.org/10.3390/vaccines11071144
Collins ND, Beaty S, Wallace E, Li Y, Sanborn M, Yang Y, Adhikari A, Shabram P, Warfield K, Karasavvas N, et al. Differential Genome Replication of a Unique Single-Amino-Acid Mutation in the Adenovirus-4 Component of the Live Oral Adenovirus Type 4 and Type 7 Vaccine. Vaccines. 2023; 11(7):1144. https://doi.org/10.3390/vaccines11071144
Chicago/Turabian StyleCollins, Natalie D., Shannon Beaty, Elana Wallace, Yuanzhang Li, Mark Sanborn, Yu Yang, Anima Adhikari, Paul Shabram, Kelly Warfield, Nicos Karasavvas, and et al. 2023. "Differential Genome Replication of a Unique Single-Amino-Acid Mutation in the Adenovirus-4 Component of the Live Oral Adenovirus Type 4 and Type 7 Vaccine" Vaccines 11, no. 7: 1144. https://doi.org/10.3390/vaccines11071144
APA StyleCollins, N. D., Beaty, S., Wallace, E., Li, Y., Sanborn, M., Yang, Y., Adhikari, A., Shabram, P., Warfield, K., Karasavvas, N., Kuschner, R. A., & Hang, J. (2023). Differential Genome Replication of a Unique Single-Amino-Acid Mutation in the Adenovirus-4 Component of the Live Oral Adenovirus Type 4 and Type 7 Vaccine. Vaccines, 11(7), 1144. https://doi.org/10.3390/vaccines11071144






