Chimeric Antigen by the Fusion of SARS-CoV-2 Receptor Binding Domain with the Extracellular Domain of Human CD154: A Promising Improved Vaccine Candidate
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plasmids
2.2. Biological Reagents
2.3. Mammalian Cell Lines and Cell Culture Conditions
2.4. Plasmid Transient Expression
2.5. Production and Quantification of LVs
2.6. Transduction of HEK-293 Cells Using LVs and Generation of RBD-CD-Expressing Cell Pools
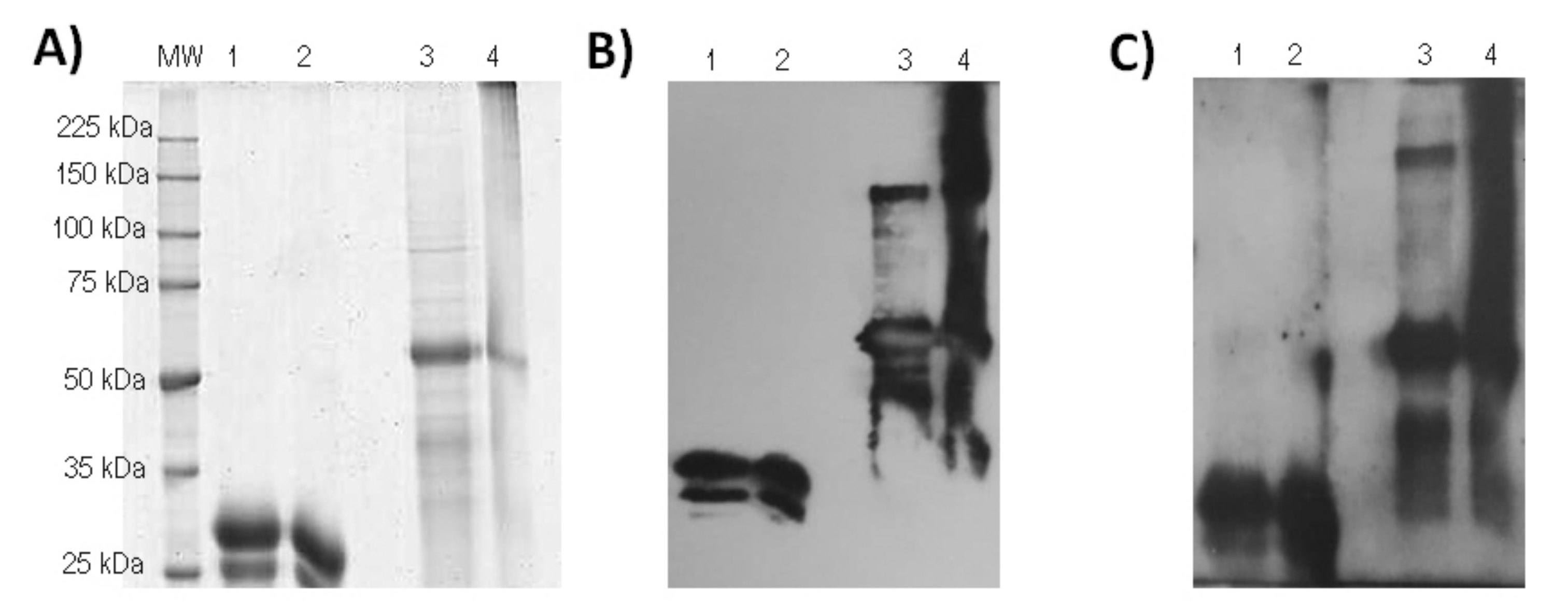
2.7. Cloning by Limiting Dilution and Evaluation of RBD-CD Expression Levels in Cell Pools and Clones
2.8. SDS-PAGE
2.9. Immunoblotting Analysis
2.10. Evaluation of RBD-CD Protein in HEK-293 Cell Culture Supernatant by ELISA
2.11. Production of RBD-CD Protein in Batch Culture
2.12. Protein Purification
2.13. ESI-MS/MS Analysis
2.13.1. Buffer-Free Digestion
2.13.2. Mass Spectrometry Analysis
2.13.3. Protein Identification
2.14. Surface Plasmon Resonance Experimental Procedure
2.15. Evaluation of RBD-CD and RBD-H6-HEK Binding to Human CD40
2.16. In Vitro Proliferation Assay in NCI-H460 Cells
2.17. RBD-CD and RBD-H6-HEK Immunoreaction by Samples from Convalescent Patients
2.18. Animals and Immunization Schedules
2.18.1. Immunogenicity in Mice
2.18.2. Comparison of Immunogenicy of RBD-CD and RBD-H6-HEK in Mice
2.18.3. Immunogenicity in Non-Human Primates
2.19. Serum Antibodies Evaluation by ELISA
2.20. RBD to ACE2 Plate-Based Binding Assay
2.21. Microneutralization of Live SARS-CoV-2 Virus in Vero E6 Cells
2.22. Statistical Analysis
3. Results
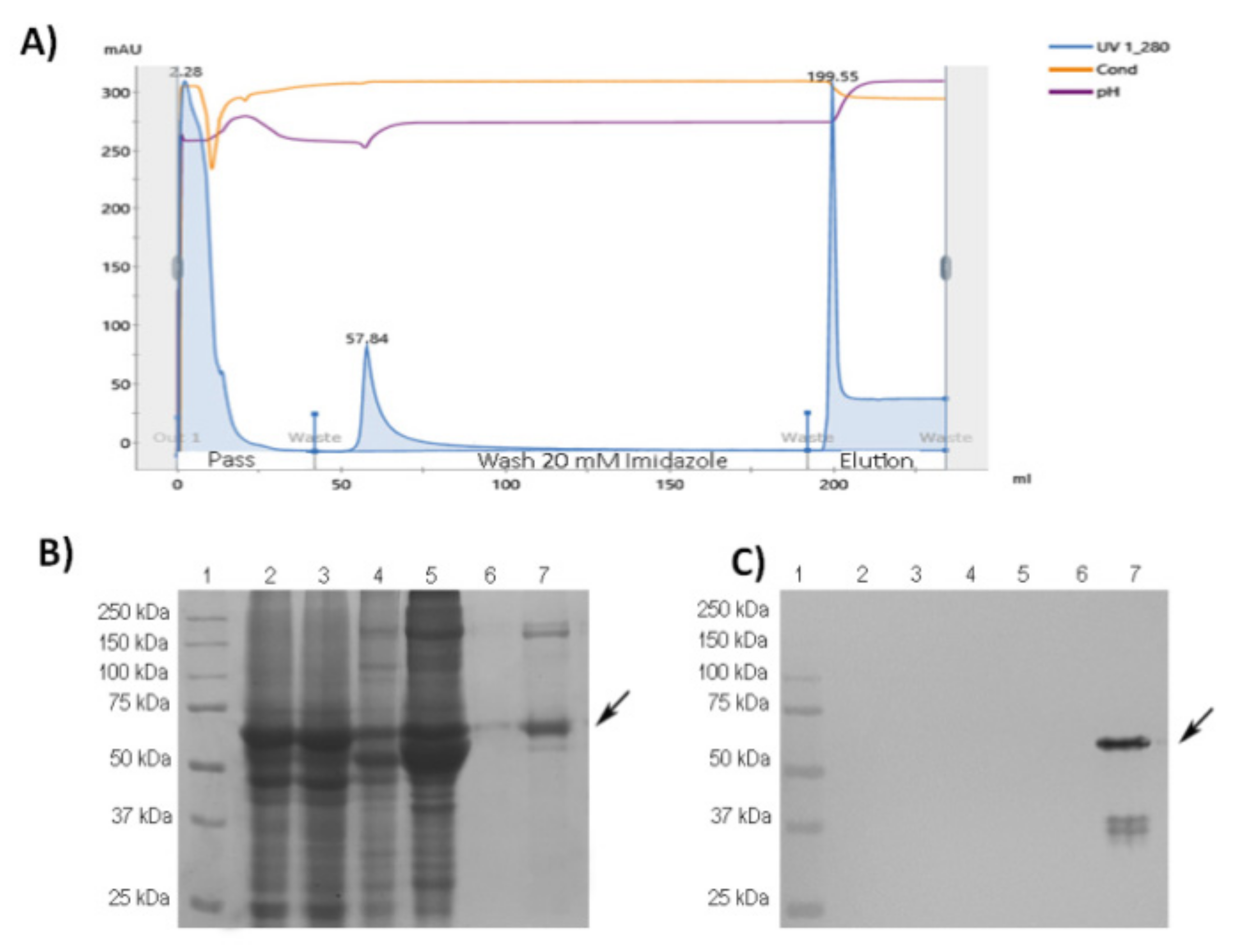
3.1. Purification and Characterization of RBD-CD from Recombinant HEK-293 Culture Supernatant
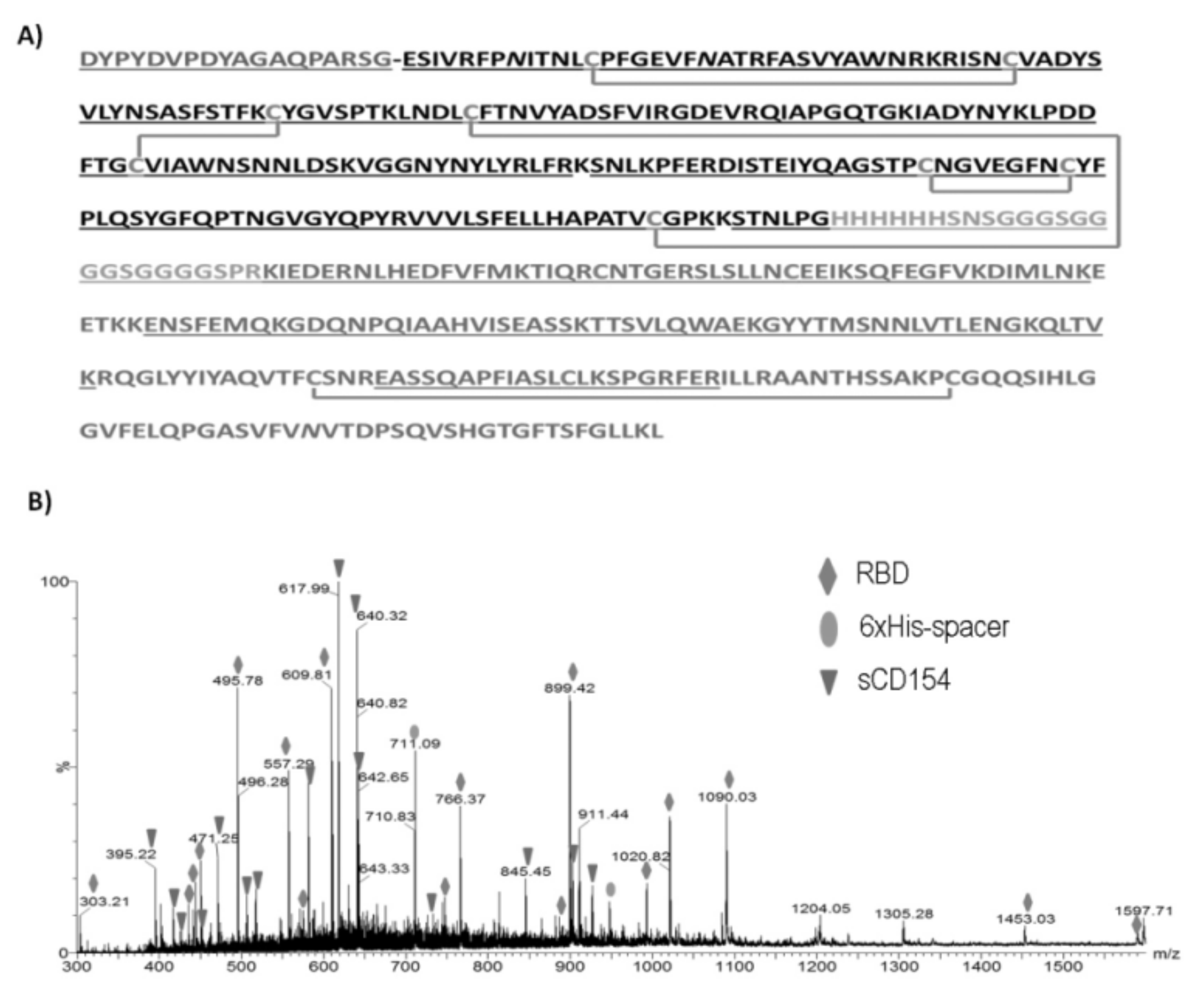
3.2. ESI-MS Analysis of the Trypsin Digestion
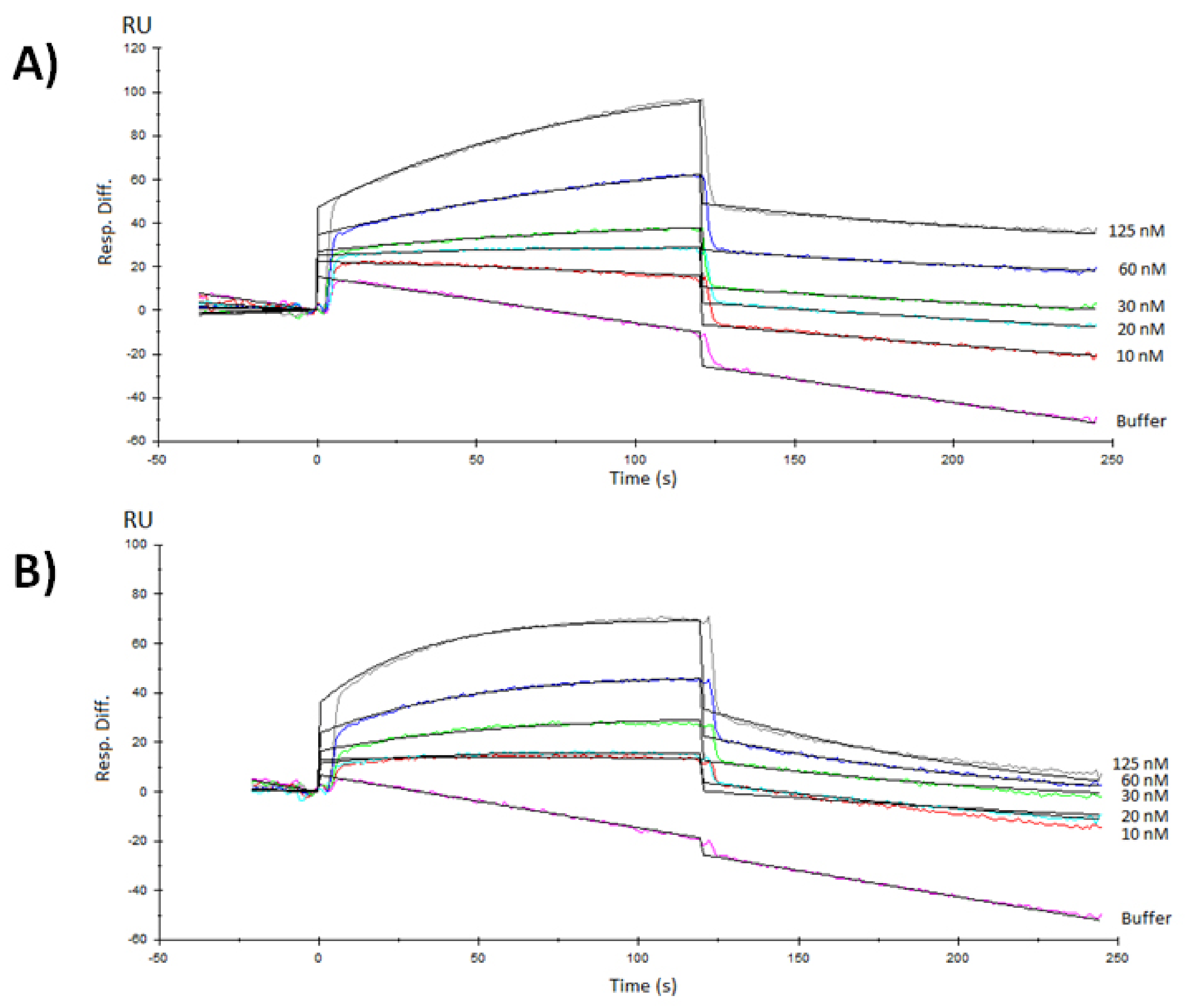
3.3. Surface Plasmon Resonance Characterization of the RBD-ACE2 Binding
3.4. CD40 Binding and Signaling in NCI-H460 Cells
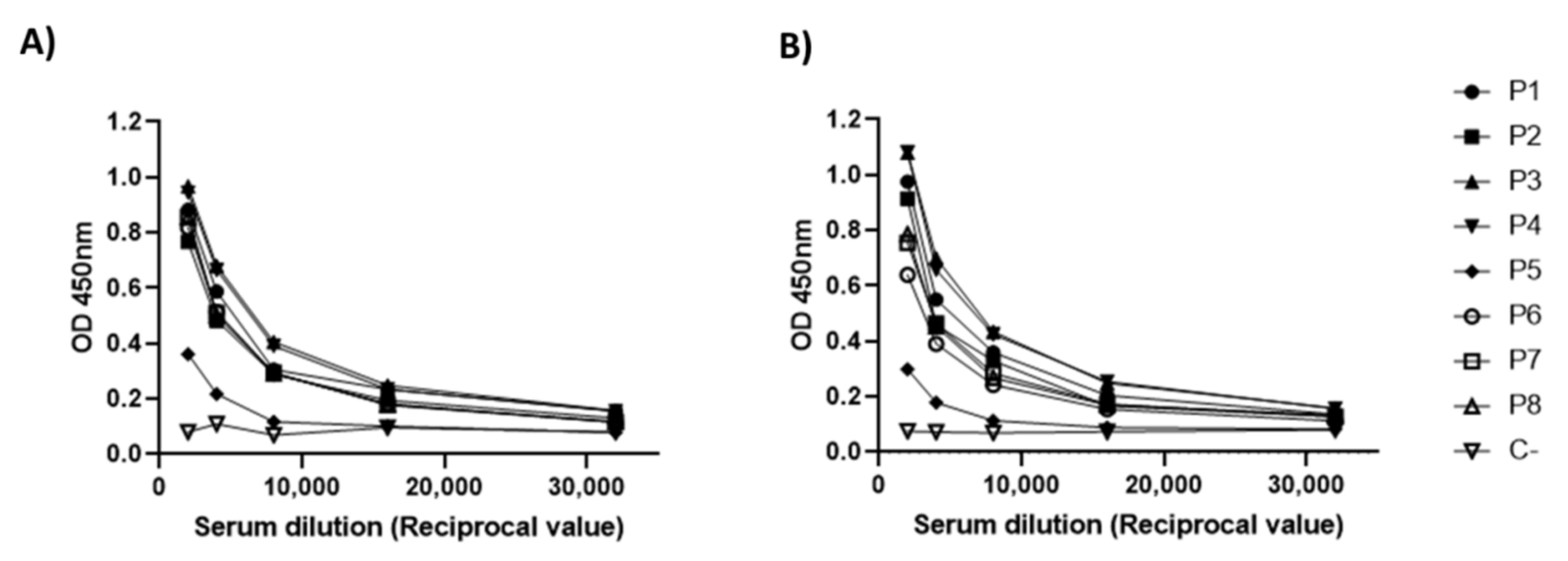
3.5. Antigenicity Analysis of RBD-CD
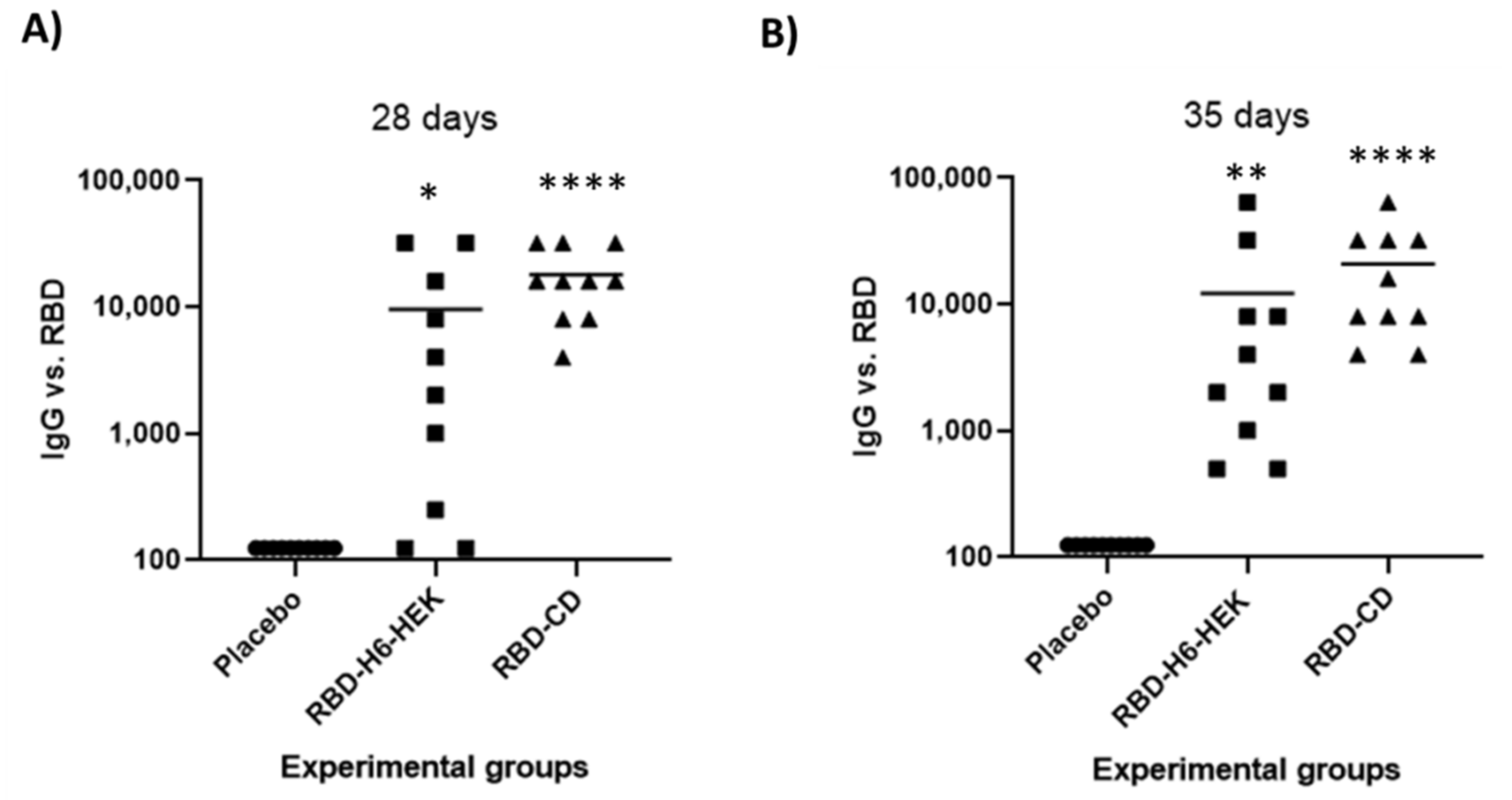
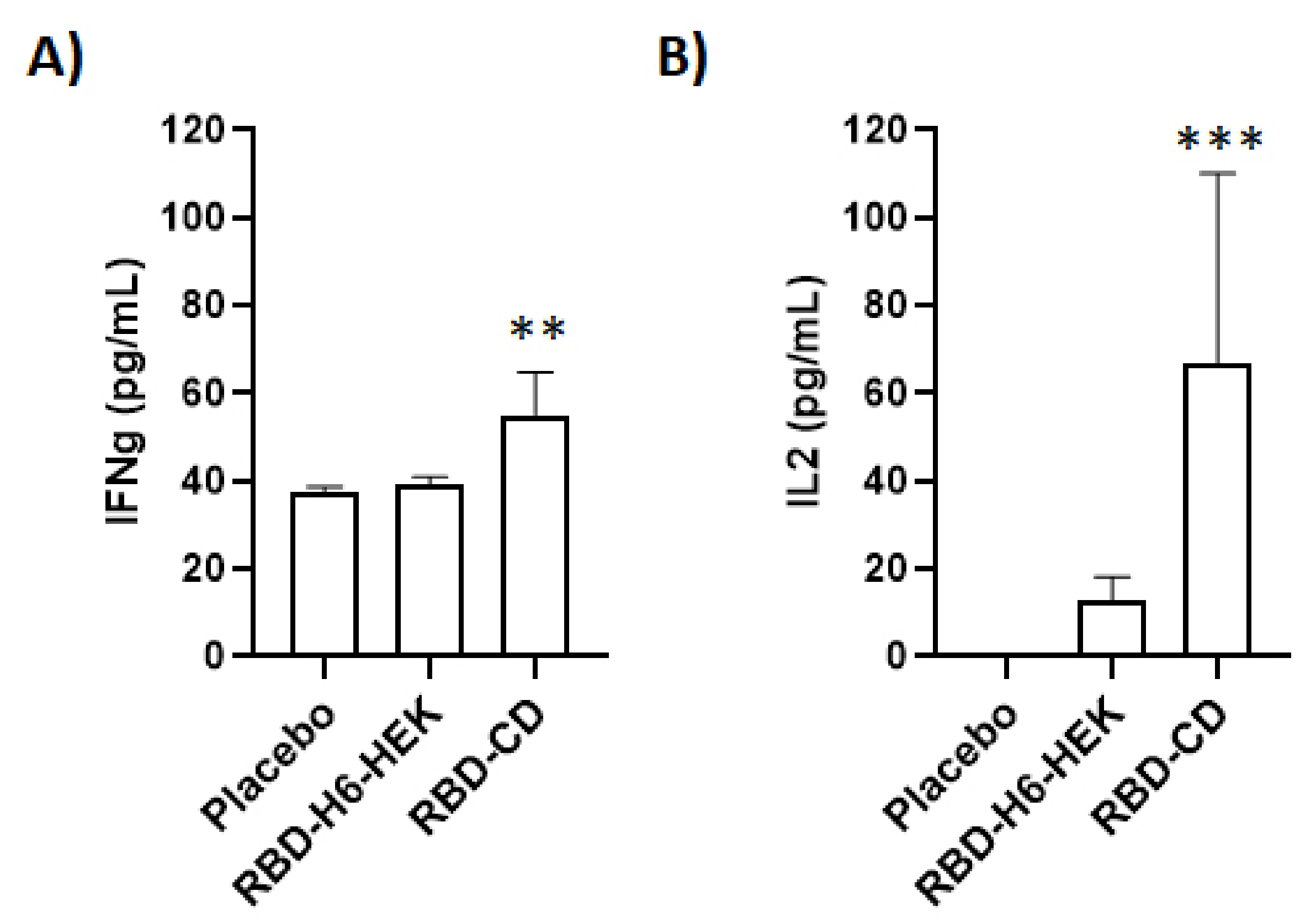
3.6. Comparison between RBD-CD and RBD-H6-HEK in Mice
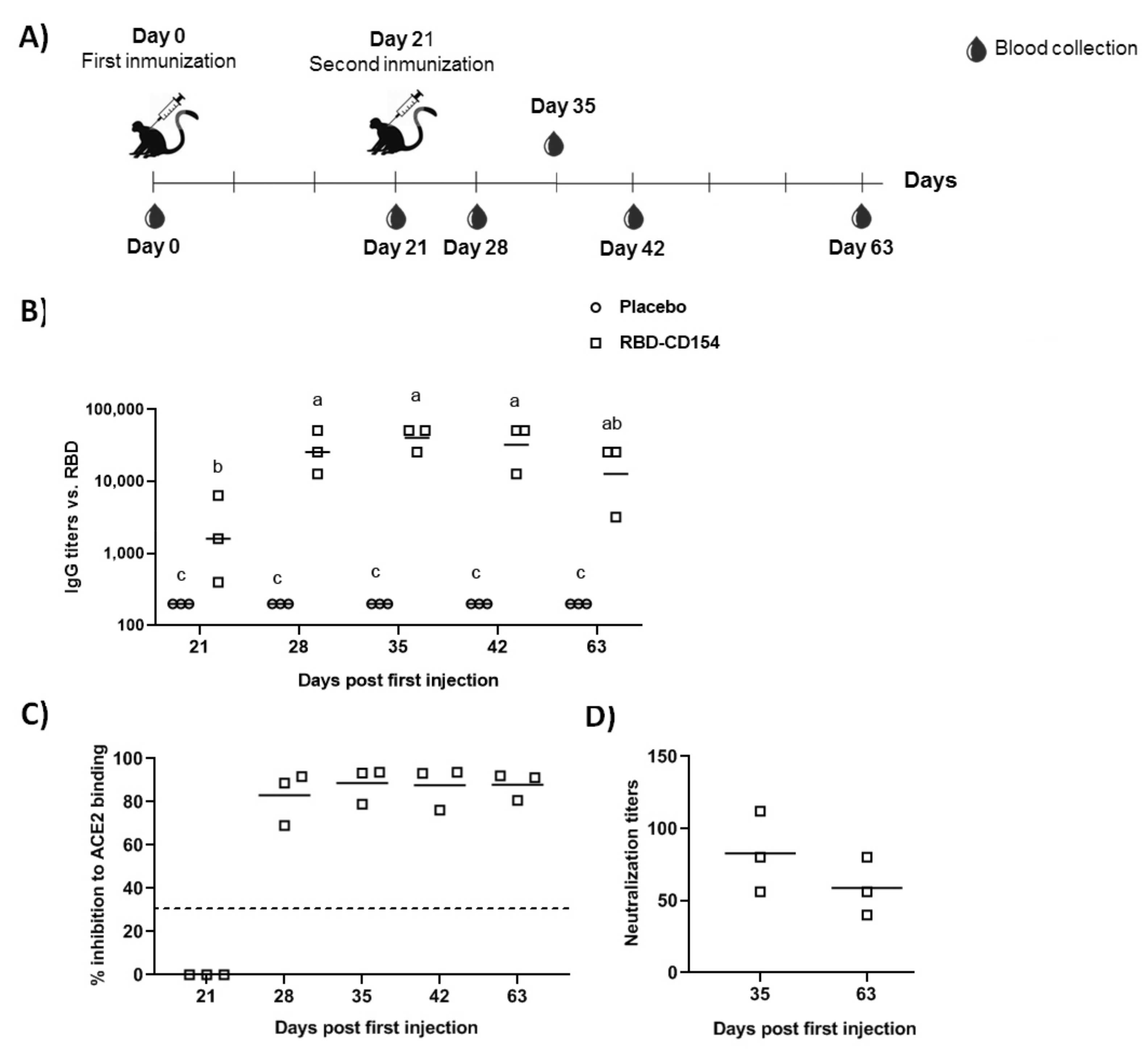
3.7. Study in Non-Human Primates
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Astuti, I. Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2): An overview of viral structure and host response. Diabetes Metab. Syndr. Clin. Res. Rev. 2020, 14, 407–412. [Google Scholar] [CrossRef] [PubMed]
- Brant, A.C.; Tian, W.; Majerciak, V.; Yang, W.; Zheng, Z.-M. SARS-CoV-2: From its discovery to genome structure, transcription, and replication. Cell Biosci. 2021, 11, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Valdes-Balbin, Y.; Santana-Mederos, D.; Paquet, F.; Fernandez, S.; Climent, Y.; Chiodo, F.; Rodríguez, L.; Sanchez Ramirez, B.; Leon, K.; Hernandez, T. Molecular aspects concerning the use of the SARS-CoV-2 receptor binding domain as a target for preventive vaccines. ACS Cent. Sci. 2021, 7, 757–767. [Google Scholar] [CrossRef]
- Liu, Z.; Xu, W.; Xia, S.; Gu, C.; Wang, X.; Wang, Q.; Zhou, J.; Wu, Y.; Cai, X.; Qu, D. RBD-Fc-based COVID-19 vaccine candidate induces highly potent SARS-CoV-2 neutralizing antibody response. Signal Transduct. Target. Ther. 2020, 5, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Lan, J.; Ge, J.; Yu, J.; Shan, S.; Zhou, H.; Fan, S.; Zhang, Q.; Shi, X.; Wang, Q.; Zhang, L. Structure of the SARS-CoV-2 spike receptor-binding domain bound to the ACE2 receptor. Nature 2020, 581, 215–220. [Google Scholar] [CrossRef] [Green Version]
- Bisht, H.; Roberts, A.; Vogel, L.; Bukreyev, A.; Collins, P.L.; Murphy, B.R.; Subbarao, K.; Moss, B. Severe acute respiratory syndrome coronavirus spike protein expressed by attenuated vaccinia virus protectively immunizes mice. Proc. Natl. Acad. Sci. USA 2004, 101, 6641–6646. [Google Scholar] [CrossRef] [Green Version]
- Yang, Z.-Y.; Kong, W.-P.; Huang, Y.; Roberts, A.; Murphy, B.R.; Subbarao, K.; Nabel, G.J. A DNA vaccine induces SARS coronavirus neutralization and protective immunity in mice. Nature 2004, 428, 561–564. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Yang, J.; Wang, W.; Chen, Z.; Lu, S.; Yang, F.; Bi, Z.; Bao, L.; Mo, F.; Li, X.; Huang, Y. A vaccine targeting the RBD of the S protein of SARS-CoV-2 induces protective immunity. Nature 2020, 586, 572–577. [Google Scholar] [CrossRef]
- van Doremalen, N.; Lambe, T.; Spencer, A.; Belij-Rammerstorfer, S.; Purushotham, J.N.; Port, J.R.; Avanzato, V.A.; Bushmaker, T.; Flaxman, A.; Ulaszewska, M. ChAdOx1 nCoV-19 vaccine prevents SARS-CoV-2 pneumonia in rhesus macaques. Nature 2020, 586, 578–582. [Google Scholar] [CrossRef]
- Laczkó, D.; Hogan, M.J.; Toulmin, S.A.; Hicks, P.; Lederer, K.; Gaudette, B.T.; Castaño, D.; Amanat, F.; Muramatsu, H.; Oguin III, T.H. A single immunization with nucleoside-modified mRNA vaccines elicits strong cellular and humoral immune responses against SARS-CoV-2 in mice. Immunity 2020, 53, 724–732.e7. [Google Scholar] [CrossRef]
- Zhu, F.-C.; Li, Y.-H.; Guan, X.-H.; Hou, L.-H.; Wang, W.-J.; Li, J.-X.; Wu, S.-P.; Wang, B.-S.; Wang, Z.; Wang, L. Safety, tolerability, and immunogenicity of a recombinant adenovirus type-5 vectored COVID-19 vaccine: A dose-escalation, open-label, non-randomised, first-in-human trial. Lancet 2020, 395, 1845–1854. [Google Scholar] [CrossRef]
- Corbett, K.S.; Edwards, D.K.; Leist, S.R.; Abiona, O.M.; Boyoglu-Barnum, S.; Gillespie, R.A.; Himansu, S.; Schäfer, A.; Ziwawo, C.T.; DiPiazza, A.T. SARS-CoV-2 mRNA vaccine design enabled by prototype pathogen preparedness. Nature 2020, 586, 567–571. [Google Scholar] [CrossRef]
- Gao, Q.; Bao, L.; Mao, H.; Wang, L.; Xu, K.; Yang, M.; Li, Y.; Zhu, L.; Wang, N.; Lv, Z. Development of an inactivated vaccine candidate for SARS-CoV-2. Science 2020, 369, 77–81. [Google Scholar] [CrossRef]
- Mulligan, M.J.; Lyke, K.E.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Raabe, V.; Bailey, R.; Swanson, K.A. Phase I/II study of COVID-19 RNA vaccine BNT162b1 in adults. Nature 2020, 586, 589–593. [Google Scholar] [CrossRef]
- Shen, X.; Tang, H.; McDanal, C.; Wagh, K.; Fischer, W.; Theiler, J.; Yoon, H.; Li, D.; Haynes, B.F.; Sanders, K.O. SARS-CoV-2 variant B. 1.1. 7 is susceptible to neutralizing antibodies elicited by ancestral spike vaccines. Cell Host Microbe 2021, 29, 529–539.e3. [Google Scholar] [CrossRef]
- Supasa, P.; Zhou, D.; Dejnirattisai, W.; Liu, C.; Mentzer, A.J.; Ginn, H.M.; Zhao, Y.; Duyvesteyn, H.M.; Nutalai, R.; Tuekprakhon, A. Reduced neutralization of SARS-CoV-2 B. 1.1. 7 variant by convalescent and vaccine sera. Cell 2021, 184, 2201–2211.e7. [Google Scholar] [CrossRef]
- Marlin, R.; Godot, V.; Cardinaud, S.; Galhaut, M.; Coleon, S.; Zurawski, S.; Dereuddre-Bosquet, N.; Cavarelli, M.; Gallouët, A.-S.; Maisonnasse, P. Targeting SARS-CoV-2 receptor-binding domain to cells expressing CD40 improves protection to infection in convalescent macaques. Nat. Commun. 2021, 12, 5215. [Google Scholar] [CrossRef]
- Zou, J.; Xie, X.; Fontes-Garfias, C.R.; Swanson, K.A.; Kanevsky, I.; Tompkins, K.; Cutler, M.; Cooper, D.; Dormitzer, P.R.; Shi, P.-Y. The effect of SARS-CoV-2 D614G mutation on BNT162b2 vaccine-elicited neutralization. npj Vaccines 2021, 6, 44. [Google Scholar] [CrossRef]
- Plante, J.A.; Liu, Y.; Liu, J.; Xia, H.; Johnson, B.A.; Lokugamage, K.G.; Zhang, X.; Muruato, A.E.; Zou, J.; Fontes-Garfias, C.R. Spike mutation D614G alters SARS-CoV-2 fitness. Nature 2021, 592, 116–121.e2379. [Google Scholar] [CrossRef]
- Garcia-Beltran, W.F.; Lam, E.C.; Denis, K.S.; Nitido, A.D.; Garcia, Z.H.; Hauser, B.M.; Feldman, J.; Pavlovic, M.N.; Gregory, D.J.; Poznansky, M.C. Multiple SARS-CoV-2 variants escape neutralization by vaccine-induced humoral immunity. Cell 2021, 184, 2372–2383.e9. [Google Scholar] [CrossRef]
- Schiller, J.T.; Lowy, D.R. Raising expectations for subunit vaccine. J. Infect. Dis. 2015, 211, 1373–1375. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, X.; Zou, F.; Yu, F.; Li, R.; Yuan, Y.; Zhang, Y.; Zhang, X.; Deng, J.; Chen, T.; Song, Z. Nanoparticle vaccines based on the receptor binding domain (RBD) and heptad repeat (HR) of SARS-CoV-2 elicit robust protective immune responses. Immunity 2020, 53, 1315–1330.e9. [Google Scholar] [CrossRef] [PubMed]
- Brouwer, P.J.; Brinkkemper, M.; Maisonnasse, P.; Dereuddre-Bosquet, N.; Grobben, M.; Claireaux, M.; de Gast, M.; Marlin, R.; Chesnais, V.; Diry, S. Two-component spike nanoparticle vaccine protects macaques from SARS-CoV-2 infection. Cell 2021, 184, 1188–1200.e19. [Google Scholar] [CrossRef] [PubMed]
- McMahan, K.; Yu, J.; Mercado, N.B.; Loos, C.; Tostanoski, L.H.; Chandrashekar, A.; Liu, J.; Peter, L.; Atyeo, C.; Zhu, A. Correlates of protection against SARS-CoV-2 in rhesus macaques. Nature 2021, 590, 630–634. [Google Scholar] [CrossRef]
- Ramos, O.S.; Pose, A.G.; Gómez-Puerta, S.; Gomez, J.N.; Redondo, A.V.; Benites, J.C.Á.; Amarán, L.S.; Parra, N.C.; Alonso, J.R.T. Avian CD154 enhances humoral and cellular immune responses induced by an adenovirus vector-based vaccine in chickens. Comp. Immunol. Microbiol. Infect. Dis. 2011, 34, 259–265. [Google Scholar] [CrossRef]
- Kornuta, C.A.; Langellotti, C.A.; Bidart, J.E.; Soria, I.; Quattrocchi, V.; Gammella, M.; Valenzuela, F.C.; Mignaqui, A.C.; Ferraris, S.; Charleston, B. A plasmid encoding the extracellular domain of CD40 ligand and Montanide™ GEL01 as adjuvants enhance the immunogenicity and the protection induced by a DNA vaccine against BoHV-1. Vaccine 2021, 39, 1007–1017. [Google Scholar] [CrossRef]
- Hashem, A.M.; Algaissi, A.; Agrawal, A.S.; Al-Amri, S.S.; Alhabbab, R.Y.; Sohrab, S.S.; Almasoud, A.S.; Alharbi, N.K.; Peng, B.-H.; Russell, M. A highly immunogenic, protective, and safe adenovirus-based vaccine expressing Middle East respiratory syndrome coronavirus S1-CD40L fusion protein in a transgenic human dipeptidyl peptidase 4 mouse model. J. Infect. Dis. 2019, 220, 1558–1567. [Google Scholar] [CrossRef]
- Cao, J.; Wang, X.; Du, Y.; Li, Y.; Wang, X.; Jiang, P. CD40 ligand expressed in adenovirus can improve the immunogenicity of the GP3 and GP5 of porcine reproductive and respiratory syndrome virus in swine. Vaccine 2010, 28, 7514–7522. [Google Scholar] [CrossRef]
- Henn, V.; Steinbach, S.; Büchner, K.; Presek, P.; Kroczek, R.A. The inflammatory action of CD40 ligand (CD154) expressed on activated human platelets is temporally limited by coexpressed CD40. Blood J. Am. Soc. Hematol. 2001, 98, 1047–1054. [Google Scholar] [CrossRef]
- Suárez, M.; Sordo, Y.; Prieto, Y.; Rodríguez, M.P.; Méndez, L.; Rodríguez, E.M.; Rodríguez-Mallon, A.; Lorenzo, E.; Santana, E.; González, N. A single dose of the novel chimeric subunit vaccine E2-CD154 confers early full protection against classical swine fever virus. Vaccine 2017, 35, 4437–4443. [Google Scholar] [CrossRef]
- Limonta-Fernández, M.; Chinea-Santiago, G.; Martín-Dunn, A.M.; Gonzalez-Roche, D.; Bequet-Romero, M.; Marquez-Perera, G.; González-Moya, I.; Canaan-Haden-Ayala, C.; Cabrales-Rico, A.; Espinosa-Rodríguez, L.A. The SARS-CoV-2 receptor-binding domain expressed in Pichia pastoris as a candidate vaccine antigen. medRxiv 2021. [Google Scholar] [CrossRef]
- Lao-Gonzalez, T.; Bueno-Soler, A.; Duran-Hernandez, A.; Sosa-Aguiar, K.; Hinojosa-Puerta, L.E.; Hernandez-Garcia, T.; de la Luz-Hernandez, K.R.; Palacios-Oliva, J.; Boggiano-Ayo, T. Screening and selection strategy for the establishment of biosimilar to trastuzumab-expressing CHO-K1 cell lines. AMB Express 2021, 11, 1–17. [Google Scholar] [CrossRef]
- Bae, D.H.; Marino, M.; Iaffaldano, B.; Fenstermaker, S.; Afione, S.; Argaw, T.; McCright, J.; Kwilas, A.; Chiorini, J.A.; Timmons, A.E. Design and testing of vector-producing HEK293T cells bearing a genomic deletion of the SV40 T antigen coding region. Mol. Ther.-Methods Clin. Dev. 2020, 18, 631–638. [Google Scholar] [CrossRef]
- Laemmli, U.K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 1970, 227, 680–685. [Google Scholar] [CrossRef]
- Giron, C.C.; Laaksonen, A.; da Silva, F.L.B. On the interactions of the receptor-binding domain of SARS-CoV-1 and SARS-CoV-2 spike proteins with monoclonal antibodies and the receptor ACE2. Virus Res. 2020, 285, 198021. [Google Scholar] [CrossRef]
- Yang, Y.; Zhang, Y.; Qu, Y.; Zhang, C.; Liu, X.-W.; Zhao, M.; Mu, Y.; Li, W. Key residues of the receptor binding domain in the spike protein of SARS-CoV-2 mediating the interactions with ACE2: A molecular dynamics study. Nanoscale 2021, 13, 9364–9370. [Google Scholar] [CrossRef]
- Du, L.; Zhao, G.; Chan, C.C.; Li, L.; He, Y.; Zhou, Y.; Zheng, B.-J.; Jiang, S. A 219-mer CHO-expressing receptor-binding domain of SARS-CoV S protein induces potent immune responses and protective immunity. Viral Immunol. 2010, 23, 211–219. [Google Scholar] [CrossRef] [Green Version]
- Baruah, V.; Bose, S. Immunoinformatics-aided identification of T cell and B cell epitopes in the surface glycoprotein of 2019-nCoV. J. Med. Virol. 2020, 92, 495–500. [Google Scholar] [CrossRef] [Green Version]
- Wang, D.; Mai, J.; Zhou, W.; Yu, W.; Zhan, Y.; Wang, N.; Epstein, N.D.; Yang, Y. Immunoinformatic analysis of T-and B-cell epitopes for SARS-CoV-2 vaccine design. Vaccines 2020, 8, 355. [Google Scholar] [CrossRef]
- Rojas, J.M.; Pascual, E.; Wattegedera, S.R.; Avia, M.; Santiago, C.; Martín, V.; Entrican, G.; Sevilla, N. Hemagglutinin protein of Peste des Petits Ruminants virus (PPRV) activates the innate immune response via Toll-like receptor 2 signaling. Virulence 2021, 12, 690–703. [Google Scholar] [CrossRef]
- Kurup, D.; Wirblich, C.; Ramage, H.; Schnell, M.J. Rabies virus-based COVID-19 vaccine CORAVAX™ induces high levels of neutralizing antibodies against SARS-CoV-2. npj Vaccines 2020, 5, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.; Pilkington, E.H.; Kelly, H.G.; Tan, H.-X.; Juno, J.A.; Wheatley, A.K.; Kent, S.J. Aggregation by peptide conjugation rescues poor immunogenicity of the HA stem. PLoS ONE 2020, 15, e0241649. [Google Scholar] [CrossRef] [PubMed]
- Pham, N.B.; Meng, W.S. Protein aggregation and immunogenicity of biotherapeutics. Int. J. Pharm. 2020, 585, 119523. [Google Scholar] [CrossRef] [PubMed]
- Espinosa, L.A.; Ramos, Y.; Andújar, I.; Torres, E.O.; Cabrera, G.; Martín, A.; González, D.; Chinea, G.; Becquet, M.; González, I. In-solution buffer-free digestion for the analysis of SARS-CoV-2 RBD proteins allows a full sequence coverage and detection of post-translational modifications in a single ESI-MS spectrum. BioRxiv 2021. [Google Scholar] [CrossRef]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.-Y. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef]
- Sui, J.; Deming, M.; Rockx, B.; Liddington, R.C.; Zhu, Q.K.; Baric, R.S.; Marasco, W.A. Effects of human anti-spike protein receptor binding domain antibodies on severe acute respiratory syndrome coronavirus neutralization escape and fitness. J. Virol. 2014, 88, 13769–13780. [Google Scholar] [CrossRef] [Green Version]
- Shang, J.; Ye, G.; Shi, K.; Wan, Y.; Luo, C.; Aihara, H.; Geng, Q.; Auerbach, A.; Li, F. Structural basis of receptor recognition by SARS-CoV-2. Nature 2020, 581, 221–224. [Google Scholar] [CrossRef] [Green Version]
- Lu, X.-D.; Zhu, X.-L.; Chen, C.; Zhang, G.-B.; Zhang, X.-G.; Huang, J.-A. Inhibition of lung cancer cell proliferation by CD40 signaling through tumor necrosis factor I. Ai Zheng = Aizheng = Chin. J. Cancer 2009, 28, 20–23. [Google Scholar]
- Perera, Y.; Costales, H.C.; Diaz, Y.; Reyes, O.; Farina, H.G.; Mendez, L.; Gómez, R.E.; Acevedo, B.E.; Gomez, D.E.; Alonso, D.F. Sensitivity of tumor cells towards CIGB-300 anticancer peptide relies on its nucleolar localization. J. Pept. Sci. 2012, 18, 215–223. [Google Scholar] [CrossRef]
- Perea, S.E.; Reyes, O.; Puchades, Y.; Mendoza, O.; Vispo, N.S.; Torrens, I.; Santos, A.; Silva, R.; Acevedo, B.; López, E. Antitumor effect of a novel proapoptotic peptide that impairs the phosphorylation by the protein kinase 2 (casein kinase 2). Cancer Res. 2004, 64, 7127–7129. [Google Scholar] [CrossRef] [Green Version]
- Walsh, E.E.; Frenck, R.W., Jr.; Falsey, A.R.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Neuzil, K.; Mulligan, M.J.; Bailey, R. Safety and immunogenicity of two RNA-based Covid-19 vaccine candidates. N. Engl. J. Med. 2020, 383, 2439–2450. [Google Scholar] [CrossRef]
- Mandolesi, M.; Sheward, D.J.; Hanke, L.; Ma, J.; Pushparaj, P.; Vidakovics, L.P.; Kim, C.; Loré, K.; Dopico, X.C.; Coquet, J.M. SARS-CoV-2 protein subunit vaccination elicits potent neutralizing antibody responses. BioRxiv 2020. [Google Scholar]
- Tan, H.-X.; Juno, J.A.; Lee, W.S.; Barber-Axthelm, I.; Kelly, H.G.; Wragg, K.M.; Esterbauer, R.; Amarasena, T.; Mordant, F.L.; Subbarao, K. Immunogenicity of prime-boost protein subunit vaccine strategies against SARS-CoV-2 in mice and macaques. Nat. Commun. 2021, 12, 1403. [Google Scholar] [CrossRef]
- Yin, W.; Gorvel, L.; Zurawski, S.; Li, D.; Ni, L.; Duluc, D.; Upchurch, K.; Kim, J.; Gu, C.; Ouedraogo, R. Functional specialty of CD40 and dendritic cell surface lectins for exogenous antigen presentation to CD8+ and CD4+ T cells. EBioMedicine 2016, 5, 46–58. [Google Scholar] [CrossRef] [Green Version]
- Godot, V.; Tcherakian, C.; Gil, L.; Cervera-Marzal, I.; Li, G.; Cheng, L.; Ortonne, N.; Lelièvre, J.-D.; Pantaleo, G.; Fenwick, C. TLR-9 agonist and CD40-targeting vaccination induces HIV-1 envelope-specific B cells with a diversified immunoglobulin repertoire in humanized mice. PLoS Pathog. 2020, 16, e1009025. [Google Scholar] [CrossRef]
- Cheng, L.; Wang, Q.; Li, G.; Banga, R.; Ma, J.; Yu, H.; Yasui, F.; Zhang, Z.; Pantaleo, G.; Perreau, M. TLR3 agonist and CD40-targeting vaccination induces immune responses and reduces HIV-1 reservoirs. J. Clin. Investig. 2018, 128, 4387–4396. [Google Scholar] [CrossRef] [Green Version]
- Zurawski, G.; Shen, X.; Zurawski, S.; Tomaras, G.D.; Montefiori, D.C.; Roederer, M.; Ferrari, G.; Lacabaratz, C.; Klucar, P.; Wang, Z. Superiority in rhesus macaques of targeting HIV-1 Env gp140 to CD40 versus LOX-1 in combination with replication-competent NYVAC-KC for induction of Env-specific antibody and T cell responses. J. Virol. 2017, 91, e01596-16. [Google Scholar] [CrossRef] [Green Version]
- Li, W. Synergistic antibody induction by antigen–CD40 ligand fusion protein as improved immunogen. Immunology 2005, 115, 215–222. [Google Scholar] [CrossRef]
- Pose, A.G.; Gómez, J.N.; Sánchez, A.V.; Redondo, A.V.; Rodríguez, E.R.; Seguí, R.M.; Ramos, E.M.G.; Moltó, M.P.R.; Rodríguez, E.S.; Cordero, L.R. Subunit influenza vaccine candidate based on CD154 fused to HAH5 increases the antibody titers and cellular immune response in chickens. Vet. Microbiol. 2011, 152, 328–337. [Google Scholar] [CrossRef]
- Wienhold, D.; Armengol, E.; Marquardt, A.; Marquardt, C.; Voigt, H.; Büttner, M.; Saalmüller, A.; Pfaff, E. Immunomodulatory effect of plasmids co-expressing cytokines in classical swine fever virus subunit gp55/E2-DNA vaccination. Vet. Res. 2005, 36, 571–587. [Google Scholar] [CrossRef] [Green Version]
- Kelly, H.G.; Tan, H.-X.; Juno, J.A.; Esterbauer, R.; Ju, Y.; Jiang, W.; Wimmer, V.C.; Duckworth, B.C.; Groom, J.R.; Caruso, F. Self-assembling influenza nanoparticle vaccines drive extended germinal center activity and memory B cell maturation. Jci Insight 2020, 5, e136653. [Google Scholar] [CrossRef]
- He, Q.; Mao, Q.; Peng, X.; He, Z.; Lu, S.; Zhang, J.; Gao, F.; Bian, L.; An, C.; Yu, W. Immunogenicity and protective efficacy of a recombinant protein subunit vaccine and an inactivated vaccine against SARS-CoV-2 variants in non-human primates. Signal Transduct. Target. Ther. 2022, 7, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Sun, S.; He, L.; Zhao, Z.; Gu, H.; Fang, X.; Wang, T.; Yang, X.; Chen, S.; Deng, Y.; Li, J. Recombinant vaccine containing an RBD-Fc fusion induced protection against SARS-CoV-2 in nonhuman primates and mice. Cell. Mol. Immunol. 2021, 18, 1070–1073. [Google Scholar] [CrossRef] [PubMed]
- Suárez-Pedroso, M.; Sordo-Puga, Y.; Sosa-Teste, I.; Rodriguez-Molto, M.P.; Naranjo-Valdés, P.; Sardina-González, T.; Santana-Rodríguez, E.; Montero-Espinosa, C.; Frías-Laporeaux, M.T.; Fuentes-Rodríguez, Y. Novel chimeric E2CD154 subunit vaccine is safe and confers long lasting protection against classical swine fever virus. Vet. Immunol. Immunopathol. 2021, 234, 110222. [Google Scholar] [CrossRef] [PubMed]
- Sordo, Y.; Suárez, M.; Caraballo, R.; Sardina, T.; Brown, E.; Duarte, C.; Lugo, J.; Gil, L.; Perez, D.; Oliva, A. Humoral and cellular immune response in mice induced by the classical swine fever virus E2 protein fused to the porcine CD154 antigen. Biologicals 2018, 52, 67–71. [Google Scholar] [CrossRef] [PubMed]
- Laffins, M.M.; Mellal, N.; Almlie, C.L.; Regalia, D.E. Evaluation of infrared thermometry in cynomolgus macaques (Macaca fascicularis). J. Am. Assoc. Lab. Anim. Sci. 2017, 56, 84–89. [Google Scholar] [PubMed]
- Koo, B.-S.; Lee, D.-H.; Kang, P.; Jeong, K.-J.; Lee, S.; Kim, K.; Lee, Y.; Huh, J.-W.; Kim, Y.-H.; Park, S.-J. Reference values of hematological and biochemical parameters in young-adult cynomolgus monkey (Macaca fascicularis) and rhesus monkey (Macaca mulatta) anesthetized with ketamine hydrochloride. Lab. Anim. Res. 2019, 35, 1–6. [Google Scholar] [CrossRef] [Green Version]
- Wierda, W.; Castro, J.; Aguillon, R.; Sampath, D.; Jalayer, A.; McMannis, J.; Prussak, C.; Keating, M.; Kipps, T. A phase I study of immune gene therapy for patients with CLL using a membrane-stable, humanized CD154. Leukemia 2010, 24, 1893–1900. [Google Scholar] [CrossRef] [Green Version]
- Wierda, W.G.; Cantwell, M.J.; Woods, S.J.; Rassenti, L.Z.; Prussak, C.E.; Kipps, T.J. CD40-ligand (CD154) gene therapy for chronic lymphocytic leukemia. Blood J. Am. Soc. Hematol. 2000, 96, 2917–2924. [Google Scholar]
- Han, M.; Lee, J.Y.; Wu, V.; Sakurada, K.; Nguyen, B.; Canton, D.A.; Twitty, C.G. Intra tumoral electroporation of IL-12 and SARS-Cov-2 spike plasmids drives a coordinated vaccine response and elicits robust anti-tumor immunity. Cancer Immunol. Res. 2022, 10, 1. [Google Scholar]
- Jensen, S.; Twitty, C.; Paustian, C.; Laws, M.; McDonnell, G.; Wegmann, K.; Moudgil, T.; Afentoulis, M.; Han, M.; Foerter, K.M. 480 Preliminary evaluation of a novel coronavirus vaccine (CORVax) using electroporation of plasmid DNA encoding a stabilized prefusion SARS-CoV-2 spike protein alone or with transfection of plasmid IL-12. J. Immuno. Therapy Cancer 2020. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ávalos, I.; Lao, T.; Rodríguez, E.M.; Zamora, Y.; Rodríguez, A.; Ramón, A.; Lemos, G.; Cabrales, A.; Bequet-Romero, M.; Casillas, D.; et al. Chimeric Antigen by the Fusion of SARS-CoV-2 Receptor Binding Domain with the Extracellular Domain of Human CD154: A Promising Improved Vaccine Candidate. Vaccines 2022, 10, 897. https://doi.org/10.3390/vaccines10060897
Ávalos I, Lao T, Rodríguez EM, Zamora Y, Rodríguez A, Ramón A, Lemos G, Cabrales A, Bequet-Romero M, Casillas D, et al. Chimeric Antigen by the Fusion of SARS-CoV-2 Receptor Binding Domain with the Extracellular Domain of Human CD154: A Promising Improved Vaccine Candidate. Vaccines. 2022; 10(6):897. https://doi.org/10.3390/vaccines10060897
Chicago/Turabian StyleÁvalos, Ileanet, Thailin Lao, Elsa María Rodríguez, Yasser Zamora, Alianet Rodríguez, Ailyn Ramón, Gilda Lemos, Ania Cabrales, Monica Bequet-Romero, Dionne Casillas, and et al. 2022. "Chimeric Antigen by the Fusion of SARS-CoV-2 Receptor Binding Domain with the Extracellular Domain of Human CD154: A Promising Improved Vaccine Candidate" Vaccines 10, no. 6: 897. https://doi.org/10.3390/vaccines10060897
APA StyleÁvalos, I., Lao, T., Rodríguez, E. M., Zamora, Y., Rodríguez, A., Ramón, A., Lemos, G., Cabrales, A., Bequet-Romero, M., Casillas, D., Andújar, I., Espinosa, L. A., González, L. J., Alvarez, Y., Carpio, Y., & Estrada, M. P. (2022). Chimeric Antigen by the Fusion of SARS-CoV-2 Receptor Binding Domain with the Extracellular Domain of Human CD154: A Promising Improved Vaccine Candidate. Vaccines, 10(6), 897. https://doi.org/10.3390/vaccines10060897






