The Role of Dietary Phenolic Compounds in Epigenetic Modulation Involved in Inflammatory Processes
Abstract
1. Introduction
2. Dietary Phenolic Compounds
2.1. Phenolic Acids
2.2. Flavonoids
2.2.1. Flavonols
2.2.2. Flavones
2.2.3. Flavanones
2.2.4. Isoflavones
2.2.5. Flavanols
2.2.6. Anthocyanins
2.3. Lignans
2.4. Stilbenes
2.5. Phenolic Alcohols
2.6. Curcuminoids
3. Mechanisms of Epigenetic Regulation
3.1. DNA Methylation
3.2. Histone Modification
3.2.1. Histone Acetylation
3.2.2. Other Histone Modifications
3.3. Non-Coding RNAs
4. Epigenetic Modifications in Inflammation
4.1. NF-κB Signaling
4.2. Chronic Inflammation
5. Epigenetics, Inflammation, and Phenolic Compounds
5.1. Phenolic Acids
5.2. Stilbenes
5.3. Flavonols
5.4. Flavones
5.5. Flavanols
5.6. Isoflavones
5.7. Curcuminoids
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
List of Abbreviations
| CBP | CREB-binding protein |
| CpG | 5’-cytosine-phosphate-guanine-3’ |
| COX | cyclooxygenase |
| DHCA | dihydrocaffeic acid |
| DNMT | DNA methyltransferase |
| EGCG | epigallocatechin gallate |
| ERK | extracellular signal-regulated kinase |
| HAT | histone acetyltransferase |
| HDAC | histone deacetylase |
| HMT | histone methyltransferases |
| HUVEC | umbilical vein endothelial cells |
| IKK | IκB kinase |
| IL | interleukin |
| iNOS | inducible nitric oxide synthase |
| IP-10 | interferon-γ-inducible protein |
| LPS | lipopolysaccharide |
| MIP | macrophage inflammatory protein |
| miRNA | microRNA |
| NF-κB | nuclear factor kappa B |
| Nrf-2 | nuclear factor 2 (erythroid-derived 2)-like 2 |
| piRNA | PIWI-interacting RNA |
| RNS | reactive nitrogen species |
| ROS | reactive oxygen species |
| siRNA | short interfering RNA |
| SIRT | sirtuin |
| snRNA | small nuclear RNA |
| SOCS | suppressor of the cytokine signaling |
| TNF-α | tumor necrosis factor alpha |
| Tregs | regulatory T cells |
| TREM-1 | triggering receptor expressed on myeloid cells 1 |
References
- Owona, B.A.; Ebrahimi, A.; Schluesenner, H. Epigenetic effects of natural polyphenols: A focus on SIRT1-mediated mechanisms. Mol. Nutr. Food Res. 2013, 58, 22–32. [Google Scholar] [CrossRef]
- Manach, C.; Scalbert, A.; Morand, C.; Rémésy, C.; Jiménez, L. Polyphenols: Food sources and bioavailability. Am. J. Clin. Nutr. 2004, 79, 727–747. [Google Scholar] [CrossRef] [PubMed]
- Bravo, L. Polyphenols: Chemistry, dietary sources, metabolism, and nutritional significance. Nutr. Rev. 1998, 56, 317–333. [Google Scholar] [CrossRef] [PubMed]
- Scalbert, A.; Manach, C.; Morand, C.; Rémésy, C.; Jimenez, L. Dietary Polyphenols and the Prevention of Diseases. Crit. Rev. Food Sci. Nutr. 2005, 45, 287–306. [Google Scholar] [CrossRef] [PubMed]
- Číž, M.; Čížová, H.; Denev, P.; Kratchanova, M.; Slavov, A.; Lojek, A. Different methods for control and comparison of the antioxidant properties of vegetables. Food Control. 2010, 21, 518–523. [Google Scholar] [CrossRef]
- Denev, P.; Kratchanov, C.G.; Ciz, M.; Lojek, A.; Kratchanova, M.G. Bioavailability and Antioxidant Activity of Black Chokeberry (Aronia melanocarpa) Polyphenols: in vitro and in vivo Evidences and Possible Mechanisms of Action: A Review. Compr. Rev. Food Sci. Food Saf. 2012, 11, 471–489. [Google Scholar] [CrossRef]
- Denev, P.; Kratchanova, M.; Číž, M.; Lojek, A.; Vasicek, O.; Nedelcheva, P.; Blazheva, D.; Toshkova, R.; Gardeva, E.; Yossifova, L.; et al. Biological activities of selected polyphenol-rich fruits related to immunity and gastrointestinal health. Food Chem. 2014, 157, 37–44. [Google Scholar] [CrossRef]
- Cory, H.; Passarelli, S.; Szeto, J.; Tamez, M.; Mattei, J. The Role of Polyphenols in Human Health and Food Systems: A Mini-Review. Front. Nutr. 2018, 5, 87. [Google Scholar] [CrossRef]
- Zhang, H.; Tsao, R. Dietary polyphenols, oxidative stress and antioxidant and anti-inflammatory effects. Curr. Opin. Food Sci. 2016, 8, 33–42. [Google Scholar] [CrossRef]
- Joven, J.; Micol, V.; Segura-Carretero, A.; Alonso-Villaverde, C.; Menendez, J.A.; Platform, B.F.C. Polyphenols and the Modulation of Gene Expression Pathways: Can We Eat Our Way Out of the Danger of Chronic Disease? Crit. Rev. Food Sci. Nutr. 2014, 54, 985–1001. [Google Scholar] [CrossRef]
- Hardy, T.M.; Tollefsbol, T.O. Epigenetic diet: Impact on the epigenome and cancer. Epigenomics 2011, 3, 503–518. [Google Scholar] [CrossRef] [PubMed]
- Kiss, A.K.; Granica, S.; Stolarczyk, M.; Melzig, M.F. Epigenetic modulation of mechanisms involved in inflammation: Influence of selected polyphenolic substances on histone acetylation state. Food Chem. 2012, 131, 1015–1020. [Google Scholar] [CrossRef]
- Blaze, J.; Wang, J.; Ho, L.; Mendelev, N.; Haghighi, F.; Pasinetti, G.M. Polyphenolic Compounds Alter Stress-Induced Patterns of Global DNA Methylation in Brain and Blood. Mol. Nutr. Food Res. 2018, 62. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.J.; Kim, S.H.; Yun, J.-M. Fisetin Inhibits Hyperglycemia-Induced Proinflammatory Cytokine Production by Epigenetic Mechanisms. Evidence-Based Complement. Altern. Med. 2012, 2012, 1–10. [Google Scholar] [CrossRef]
- Xiao, X.; Shi, D.; Liu, L.; Wang, J.; Xie, X.; Kang, T.; Deng, W. Quercetin Suppresses Cyclooxygenase-2 Expression and Angiogenesis through Inactivation of P300 Signaling. PLoS ONE 2011, 6, e22934. [Google Scholar] [CrossRef]
- Kim, H.J.; Lee, W.; Yun, J.-M. Luteolin Inhibits Hyperglycemia-Induced Proinflammatory Cytokine Production and Its Epigenetic Mechanism in Human Monocytes. Phytotherapy Res. 2014, 28, 1383–1391. [Google Scholar] [CrossRef]
- Zhang, H.P.; Zhao, Z.X.; Pang, X.F.; Yang, J.; Yu, H.X.; Zhang, Y.H.; Zhou, H.; Zhao, J.H. Genistein Protects Against Ox-LDL-Induced Inflammation Through MicroRNA-155/SOCS1-Mediated Repression of NF-AB Signaling Pathway in HUVECs. Inflammation 2017, 40, 1450–1459. [Google Scholar] [CrossRef]
- Yousefi, H.; Alihemmati, A.; Karimi, P.; Alipour, M.R.; Habibi, P.; Ahmadiasl, N. Effect of genistein on expression of pancreatic SIRT1, inflammatory cytokines and histological changes in ovariectomized diabetic rat. Iran. J. Basic Med. Sci. 2017, 20, 423–429. [Google Scholar]
- Majid, S.; Dar, A.A.; Shahryari, V.; Hirata, H.; Ahmad, A.; Saini, S.; Tanaka, Y.; Dahiya, A.V.; Dahiya, R. Genistein reverses hypermethylation and induces active histone modifications in tumor suppressor gene B-Cell translocation gene 3 in prostate cancer. Cancer 2009, 116, 66–76. [Google Scholar] [CrossRef]
- Fang, M.Z.; Wang, Y.; Ai, N.; Hou, Z.; Sun, Y.; Lu, H.; Welsh, W.; Yang, C.S. Tea polyphenol (-)-epigallocatechin-3-gallate inhibits DNA methyltransferase and reactivates methylation-silenced genes in cancer cell lines. Cancer Res. 2003, 63, 7563–7570. [Google Scholar]
- Choi, K.C.; Jung, M.G.; Lee, Y.H.; Yoon, J.C.; Kwon, S.H.; Kang, H.B.; Kim, M.J.; Cha, J.H.; Kim, Y.J.; Jun, W.J.; et al. Epigallocatechin-3-Gallate, a Histone Acetyltransferase Inhibitor, Inhibits EBV-Induced B Lymphocyte Transformation via Suppression of ReIA Acetylation. Cancer Res. 2009, 69, 583–592. [Google Scholar] [CrossRef] [PubMed]
- Cordero-Herrera, I.; Chen, X.P.; Ramos, S.; Devaraj, S. (-)-Epicatechin attenuates high-glucose-induced inflammation by epigenetic modulation in human monocytes. Eur. J. Nutr. 2017, 56, 1369–1373. [Google Scholar] [CrossRef] [PubMed]
- Borra, M.T.; Smith, B.C.; Denu, J.M. Mechanism of Human SIRT1 Activation by Resveratrol. J. Boil. Chem. 2005, 280, 17187–17195. [Google Scholar] [CrossRef] [PubMed]
- Pan, W.; Yu, H.; Huang, S.; Zhu, P.-L. Resveratrol Protects against TNF-α-Induced Injury in Human Umbilical Endothelial Cells through Promoting Sirtuin-1-Induced Repression of NF-KB and p38 MAPK. PLoS ONE 2016, 11, e0147034. [Google Scholar] [CrossRef] [PubMed]
- Lou, X.-D.; Wang, H.; Xia, S.-J.; Skog, S.; Sun, J. Effects of Resveratrol on the Expression and DNA Methylation of Cytokine Genes in Diabetic Rat Aortas. Arch. Immunol. Ther. Exp. 2014, 62, 329–340. [Google Scholar] [CrossRef]
- Kumar, A.; Rimando, A.M.; Levenson, A.S. Resveratrol and pterostilbene as a microRNA-mediated chemopreventive and therapeutic strategy in prostate cancer. Ann. NY Acad. Sci. 2017, 1403, 15–26. [Google Scholar] [CrossRef]
- Neckers, L.; Trepel, J.; Lee, S.; Chung, E.-J.; Lee, M.-J.; Jung, Y.-J.; Marcu, M.G. Curcumin is an Inhibitor of p300 Histone Acetylatransferase. Med. Chem. 2006, 2, 169–174. [Google Scholar] [CrossRef]
- Sun, M.; Estrov, Z.; Ji, Y.; Coombes, K.R.; Harris, D.H.; Kurzrock, R. Curcumin (diferuloylmethane) alters the expression profiles of microRNAs in human pancreatic cancer cells. Mol. Cancer Ther. 2008, 7, 464–473. [Google Scholar] [CrossRef]
- Guo, Y.; Shu, L.; Zhang, C.; Su, Z.-Y.; Kong, A.-N. Curcumin inhibits anchorage-independent growth of HT29 human colon cancer cells by targeting epigenetic restoration of the tumor suppressor gene DLEC1. Biochem. Pharmacol. 2015, 94, 69–78. [Google Scholar] [CrossRef]
- Maugeri, A.; Mazzone, M.G.; Giuliano, F.; Vinciguerra, M.; Basile, G.; Barchitta, M.; Agodi, A. Curcumin Modulates DNA Methyltransferase Functions in a Cellular Model of Diabetic Retinopathy. Oxidative Med. Cell. Longev. 2018, 2018, 1–12. [Google Scholar] [CrossRef]
- Vattem, D.; Shetty, K. biological functionality of ellagic acid: A review. J. Food Biochem. 2005, 29, 234–266. [Google Scholar] [CrossRef]
- Crupi, P.; Bleve, G.; Tufariello, M.; Corbo, F.; Clodoveo, M.L.; Tarricone, L. Comprehensive identification and quantification of chlorogenic acids in sweet cherry by tandem mass spectrometry techniques. J. Food Compos. Anal. 2018, 73, 103–111. [Google Scholar] [CrossRef]
- Fonayet, J.V.; Millán, S.; Martí, M.P.; Borràs, E.; Arola, L. Advanced separation methods of food anthocyanins, isoflavones and flavanols. J. Chromatogr. A 2009, 1216, 7143–7172. [Google Scholar] [CrossRef]
- Harborne, J.B.; A Williams, C. Advances in flavonoid research since 1992. Phytochemistry 2000, 55, 481–504. [Google Scholar] [CrossRef]
- Coward, L.; Smith, M.; Kirk, M.; Barnes, S. Chemical modification of isoflavones in soyfoods during cooking and processing. Am. J. Clin. Nutr. 1998, 68, 1486S–1491S. [Google Scholar] [CrossRef]
- Setchell, K.D.R.; Brown, N.M.; Lydeking-Olsen, E. The Clinical Importance of the Metabolite Equol—A Clue to the Effectiveness of Soy and Its Isoflavones. J. Nutr. 2002, 132, 3577–3584. [Google Scholar] [CrossRef]
- Lakenbrink, C.; Lapczynski, S.; Maiwald, B.; Engelhardt, U.H. Flavonoids and other polyphenols in consumer brews of tea and other caffeinated beverages. J. Agric. Food Chem. 2000, 48, 2848–2852. [Google Scholar] [CrossRef]
- Graham, H.N. Green tea composition, consumption, and polyphenol chemistry. Prev. Med. 1992, 21, 334–350. [Google Scholar] [CrossRef]
- Guyot, S.; Marnet, N.; Drilleau, J.-F. Thiolysis-HPLC characterization of apple procyanidins covering a large range of polymerization states. J. Agric. Food Chem. 2001, 49, 14–20. [Google Scholar] [CrossRef]
- Thompson, L.U.; Robb, P.; Serraino, M.; Cheung, F. Mammalian lignan production from various foods. Nutr. Cancer 1991, 16, 43–52. [Google Scholar] [CrossRef]
- Adlercreutz, H.; Mazur, W. Phyto-oestrogens and Western diseases. Ann. Med. 1997, 29, 95–120. [Google Scholar] [CrossRef] [PubMed]
- Heinonen, S.; Salonen, J.T.; Liukkonen, K.; Poutanen, K.; Wähälä, K.; Deyama, T.; Nishibe, S.; Adlercreutz, H. In vitro metabolism of plant lignans: New precursors of mammalian lignans enterolactone and enterodiol. J. Agric. Food Chem. 2001, 49, 3178–3186. [Google Scholar] [CrossRef] [PubMed]
- Bavaresco, L. Role of viticultural factors on stilbene concentrations of grapes and wine. Drugs Under Exp. Clin. Res. 2003, 29, 181–187. [Google Scholar]
- Delmas, D.; Lançon, A.; Colin, D.; Jannin, B.; Latruffe, N. Resveratrol as a Chemopreventive Agent: A Promising Molecule for Fighting Cancer. Curr. Drug Targets 2006, 7, 423–442. [Google Scholar] [CrossRef]
- Das, D.K.; Mukherjee, S.; Ray, D. Resveratrol and red wine, healthy heart and longevity. Hear. Fail. Rev. 2010, 15, 467–477. [Google Scholar] [CrossRef]
- Lee, P.-S.; Chiou, Y.-S.; Ho, C.-T.; Pan, M.-H. Chemoprevention by resveratrol and pterostilbene: Targeting on epigenetic regulation. BioFactors 2017, 44, 26–35. [Google Scholar] [CrossRef]
- Cabrini, L.; Barzanti, V.; Cipollone, M.; Fiorentini, D.; Grossi, G.; Tolomelli, B.; Zambonin, L.; Landi, L. Antioxidants and total peroxyl radical-trapping ability of olive and seed oils. J. Agric. Food Chem. 2001, 49, 6026–6032. [Google Scholar] [CrossRef]
- Covas, M.I.; Miró-Casas, E.; Fitó, M.; Farré-Albadalejo, M.; Gimeno, E.; Marrugat, J.; De La Torre, R. Bioavailability of tyrosol, an antioxidant phenolic compound present in wine and olive oil, in humans. Drugs Under Exp. Clin. Res. 2003, 29, 203–206. [Google Scholar]
- De La Torre, R.; Covas, M.I.; Pujadas, M.A.; Fitó, M.; Farré, M. Is dopamine behind the health benefits of red wine? Eur. J. Nutr. 2006, 45, 307–310. [Google Scholar] [CrossRef]
- Amalraj, A.; Pius, A.; Gopi, S.; Gopi, S. Biological activities of curcuminoids, other biomolecules from turmeric and their derivatives—A review. J. Tradit. Complement. Med. 2016, 7, 205–233. [Google Scholar] [CrossRef]
- Pan, M.-H.; Lai, C.-S.; Wu, J.-C.; Ho, C.-T. Epigenetic and disease targets by polyphenols. Curr. Pharm. Des. 2013, 19, 6156–6185. [Google Scholar] [CrossRef] [PubMed]
- Suárez-Álvarez, B.; Raneros, A.B.; Ortega, F.; López-Soto, A. Epigenetic modulation of the immune function: A potential target for tolerance. Epigenetics 2013, 8, 694–702. [Google Scholar] [CrossRef]
- Ospelt, C.; Gay, S.; Klein, K. Epigenetics in the pathogenesis of RA. Semin. Immunopathol. 2017, 39, 409–419. [Google Scholar] [CrossRef] [PubMed]
- Fuhrmann, J.; Thompson, P.R. Protein Arginine Methylation and Citrullination in Epigenetic Regulation. ACS Chem. Boil. 2015, 11, 654–668. [Google Scholar] [CrossRef] [PubMed]
- Morandini, A.C.; Santos, C.F.; Yilmaz, O. Role of epigenetics in modulation of immune response at the junction of host–pathogen interaction and danger molecule signaling. Pathog. Dis. 2016, 74. [Google Scholar] [CrossRef] [PubMed]
- Gibney, E.R.; Nolan, C.M. Epigenetics and gene expression. Heredity 2010, 105, 4–13. [Google Scholar] [CrossRef]
- Castillo-Aguilera, O.; Depreux, P.; Halby, L.; Arimondo, P.B.; Goossens, L. DNA Methylation Targeting: The DNMT/HMT Crosstalk Challenge. Biomolecules 2017, 7, 3. [Google Scholar] [CrossRef]
- Fernandes, G.F.S.; Silva, G.D.B.; Pavan, A.R.; Chiba, D.E.; Chin, C.M.; Dos Santos, J.L. Epigenetic Regulatory Mechanisms Induced by Resveratrol. Nutrients 2017, 9, 1201. [Google Scholar] [CrossRef]
- Leshner, M.; Wang, S.; Lewis, C.; Zheng, H.; Chen, X.A.; Santy, L.; Wang, Y. PAD4 mediated histone hypercitrullination induces heterochromatin decondensation and chromatin unfolding to form neutrophil extracellular trap-like structures. Front. Immunol. 2012, 3. [Google Scholar] [CrossRef]
- Chuang, J.C.; Jones, P.A. Epigenetics and microRNAs. Pediatr. Res. 2007, 61, 24r–29r. [Google Scholar] [CrossRef]
- Xu, Y.-M.; Du, J.-Y.; Lau, A.T.Y. Posttranslational modifications of human histone H3: An update. Proteomics 2014, 14, 2047–2060. [Google Scholar] [CrossRef] [PubMed]
- Frías-Lasserre, D.; Villagra, C.A. The Importance of ncRNAs as Epigenetic Mechanisms in Phenotypic Variation and Organic Evolution. Front. Microbiol. 2017, 8. [Google Scholar] [CrossRef] [PubMed]
- Yekta, S.; Shih, I.-H.; Bartel, B. MicroRNA-Directed Cleavage of HOXB8 mRNA. Science 2004, 304, 594–596. [Google Scholar] [CrossRef] [PubMed]
- Agrawal, N.; Dasaradhi, P.V.N.; Mohmmed, A.; Malhotra, P.; Bhatnagar, R.K.; Mukherjee, S.K. RNA Interference: Biology, Mechanism, and Applications. Microbiol. Mol. Boil. Rev. 2003, 67, 657–685. [Google Scholar] [CrossRef] [PubMed]
- Muniz, L.; Egloff, S.; Kiss, T. RNA elements directing in vivo assembly of the 7SK/MePCE/Larp7 transcriptional regulatory snRNP. Nucleic Acids Res. 2013, 41, 4686–4698. [Google Scholar] [CrossRef]
- Siomi, M.C.; Sato, K.; Pezic, D.; Aravin, A.A. PIWI-interacting small RNAs: The vanguard of genome defence. Nat. Rev. Mol. Cell Boil. 2011, 12, 246–258. [Google Scholar] [CrossRef]
- Milagro, F.I.; Mansego, M.L.; De Miguel, C.; Martínez, J.A. Dietary factors, epigenetic modifications and obesity outcomes: Progresses and perspectives. Mol. Asp. Med. 2013, 34, 782–812. [Google Scholar] [CrossRef]
- Adler, V.; Yin, Z.; Tew, K.D.; Ronai, Z.A. Role of redox potential and reactive oxygen species in stress signaling. Oncogene 1999, 18, 6104–6111. [Google Scholar] [CrossRef]
- Olefsky, J.M. IKK epsilon: A Bridge between Obesity and Inflammation. Cell 2009, 138, 834–836. [Google Scholar] [CrossRef]
- Yun, J.-M.; Jialal, I.; Devaraj, S. Epigenetic regulation of high glucose-induced proinflammatory cytokine production in monocytes by curcumin. J. Nutr. Biochem. 2011, 22, 450–458. [Google Scholar] [CrossRef]
- Shakibaei, M.; Buhrmann, C.; Mobasheri, A. Resveratrol-mediated SIRT-1 Interactions with p300 Modulate Receptor Activator of NF-kappa B Ligand (RANKL) Activation of NF-kappa B Signaling and Inhibit Osteoclastogenesis in Bone-derived Cells. J. Biol. Chem. 2011, 286, 11492–11505. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, A.; Saavedra, N.; Salazar, L.A.; Abdalla, D.S. Modulation of Immune Function by Polyphenols: Possible Contribution of Epigenetic Factors. Nutrients 2013, 5, 2314–2332. [Google Scholar] [CrossRef] [PubMed]
- Chen, S.; Jiang, H.; Wu, X.; Fang, J. Therapeutic Effects of Quercetin on Inflammation, Obesity, and Type 2 Diabetes. Mediat. Inflamm. 2016, 2016, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Z.; Syed, M.A.; Panchal, D.; Rogers, D.; Joo, M.; Sadikot, R.T. Curcumin mediated epigenetic modulation inhibits TREM-1 expression in response to lipopolysaccharide. Int. J. Biochem. Cell Boil. 2012, 44, 2032–2043. [Google Scholar] [CrossRef]
- Boyanapalli, S.S.; Huang, Y.; Su, Z.-Y.; Cheng, D.; Zhang, C.; Guo, Y.; Rao, R.; Androulakis, I.P.; Kong, A.-N. Pharmacokinetics and Pharmacodynamics of Curcumin in regulating anti-inflammatory and epigenetic gene expression. Biopharm. Drug Dispos. 2018, 39, 289–297. [Google Scholar] [CrossRef]
- Bo, S.; Togliatto, G.; Gambino, R.; Ponzo, V.; Lombardo, G.; Rosato, R.; Cassader, M.; Brizzi, M.F. Impact of sirtuin-1 expression on H3K56 acetylation and oxidative stress: A double-blind randomized controlled trial with resveratrol supplementation. Acta Diabetol. 2018, 55, 331–340. [Google Scholar] [CrossRef]
- Crescenti, A.; Solà, R.; Valls, R.M.; Caimari, A.; Del Bas, J.M.; Anguera, A.; Anglès, N.; Arola, L. Cocoa Consumption Alters the Global DNA Methylation of Peripheral Leukocytes in Humans with Cardiovascular Disease Risk Factors: A Randomized Controlled Trial. PLoS ONE 2013, 8, e65744. [Google Scholar] [CrossRef]
- Uddin, M.; Aiello, A.E.; Wildman, D.E.; Koenen, K.C.; Pawelec, G.; Santos, R.D.L.; Goldmann, E.; Galea, S. Epigenetic and immune function profiles associated with posttraumatic stress disorder. Proc. Natl. Acad. Sci. USA 2010, 107, 9470–9475. [Google Scholar] [CrossRef] [PubMed]
- Klengel, T.; Mehta, D.; Anacker, C.; Rex-Haffner, M.; Pruessner, J.C.; Pariante, C.M.; Pace, T.W.W.; Mercer, K.B.; Mayberg, H.S.; Bradley, B.; et al. Faculty of 1000 evaluation for Allele-specific FKBP5 DNA demethylation mediates gene-childhood trauma interactions. Nat. Neurosci. 2013, 16, 33–41. [Google Scholar] [CrossRef]
- Gao, Y.; Tollefsbol, T.O. Impact of Epigenetic Dietary Components on Cancer through Histone Modifications. Curr. Med. Chem. 2015, 22, 2051–2064. [Google Scholar] [CrossRef]
- Ding, S.; Jang, H.; Fang, J. Regulation of Immune Function by Polyphenols. J. Immunol. Res. 2018, 2018, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Maugeri, A.; Barchitta, M.; Mazzone, M.G.; Giuliano, F.; Basile, G.; Agodi, A. Resveratrol Modulates SIRT1 and DNMT Functions and Restores LINE-1 Methylation Levels in ARPE-19 Cells under Oxidative Stress and Inflammation. Int. J. Mol. Sci. 2018, 19, 2118. [Google Scholar] [CrossRef] [PubMed]
- Kong, S.; Yeung, P.; Fang, D. The class III histone deacetylase sirtuin 1 in immune suppression and its therapeutic potential in rheumatoid arthritis. J. Genet. Genom. 2013, 40, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Kala, R.; Shah, H.N.; Martin, S.L.; Tollefsbol, T.O. Epigenetic-based combinatorial resveratrol and pterostilbene alters DNA damage response by affecting SIRT1 and DNMT enzyme expression, including SIRT1-dependent γ-H2AX and telomerase regulation in triple-negative breast cancer. BMC Cancer 2015, 15, 672. [Google Scholar] [CrossRef]
- Yang, H.; Zhang, W.; Pan, H.; Feldser, H.G.; Lainez, E.; Miller, C.; Leung, S.; Zhong, Z.; Zhao, H.; Sweitzer, S.; et al. SIRT1 Activators Suppress Inflammatory Responses through Promotion of p65 Deacetylation and Inhibition of NF-κB Activity. PLoS ONE 2012, 7, e46364. [Google Scholar] [CrossRef]
- Schug, T.T.; Xu, Q.; Gao, H.; Peres-Da-Silva, A.; Draper, D.W.; Fessler, M.B.; Purushotham, A.; Li, X. Myeloid Deletion of SIRT1 Induces Inflammatory Signaling in Response to Environmental Stress. Mol. Cell. Boil. 2010, 30, 4712–4721. [Google Scholar] [CrossRef]
- Tili, E.; Michaille, J.-J. Promiscuous Effects of Some Phenolic Natural Products on Inflammation at Least in Part Arise from Their Ability to Modulate the Expression of Global Regulators, Namely microRNAs. Molecules 2016, 21, 1263. [Google Scholar] [CrossRef]
- Tseng, B.; Lee, W.-J.; Chen, Y.-R.; Tseng, T.-H. Quercetin induces FasL-related apoptosis, in part, through promotion of histone H3 acetylation in human leukemia HL-60 cells. Oncol. Rep. 2011, 25, 583–591. [Google Scholar] [CrossRef]
- Abdulla, A.; Zhao, X.; Yang, F. Natural Polyphenols Inhibit Lysine-Specific Demethylase-1 in vitro. J. Biochem. Pharmacol. Res. 2013, 1, 56–63. [Google Scholar]
- Ruiz, P.A.; Braune, A.; Hölzlwimmer, G.; Quintanilla-Fend, L.; Haller, D. Quercetin inhibits TNF-induced NF-kappaB transcription factor recruitment to proinflammatory gene promoters in murine intestinal epithelial cells. J. Nutr. 2007, 137, 1208–1215. [Google Scholar] [CrossRef]
- Reichard, J.F.; Motz, G.T.; Puga, A. Heme oxygenase-1 induction by NRF2 requires inactivation of the transcriptional repressor BACH1. Nucleic Acids Res. 2007, 35, 7074–7086. [Google Scholar] [CrossRef] [PubMed]
- Rangarajan, P.; Karthikeyan, A.; Dheen, S.T. Role of dietary phenols in mitigating microglia-mediated neuroinflammation. Neuromolecular Med. 2016, 18, 453–464. [Google Scholar] [CrossRef] [PubMed]
- Kim, A.; Yun, J.-M. Combination Treatments with Luteolin and Fisetin Enhance Anti-Inflammatory Effects in High Glucose-Treated THP-1 Cells Through Histone Acetyltransferase/Histone Deacetylase Regulation. J. Med. Food 2017, 20, 782–789. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Perkins, J.T.; Hennig, B. EGCG prevents PCB-126-induced endothelial cell inflammation via epigenetic modifications of NF-κB target genes in human endothelial cells. J. Nutr. Biochem. 2015, 28, 164–170. [Google Scholar] [CrossRef]
- Yun, J.-M.; Jialal, I.; Devaraj, S. Effects of epigallocatechin gallate on regulatory T cell number and function in obese v. lean volunteers. Br. J. Nutr. 2010, 103, 1771–1777. [Google Scholar] [CrossRef]
- Kim, M.; Long, T.I.; Arakawa, K.; Wang, R.; Yu, M.C.; Laird, P.W. DNA Methylation as a Biomarker for Cardiovascular Disease Risk. PLoS ONE 2010, 5, e9692. [Google Scholar] [CrossRef]
- Zhao, J.Y.; Goldberg, J.; Bremner, J.D.; Vaccarino, V. Global DNA Methylation Is Associated with Insulin Resistance A Monozygotic Twin Study. Diabetes 2012, 61, 542–546. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, H. Genistein, an epigenome modifier during cancer prevention. Epigenetics 2011, 6, 888–891. [Google Scholar] [CrossRef]
- Boyanapalli, S.S.S.; Kong, A.-N.T. “Curcumin, the King of Spices”: Epigenetic Regulatory Mechanisms in the Prevention of Cancer, Neurological, and Inflammatory Diseases. Curr. Pharmacol. Rep. 2015, 1, 129–139. [Google Scholar] [CrossRef]
- Wada, T.T.; Araki, Y.; Sato, K.; Aizaki, Y.; Yokota, K.; Kim, Y.T.; Oda, H.; Kurokawa, R.; Mimura, T. Aberrant histone acetylation contributes to elevated interleukin-6 production in rheumatoid arthritis synovial fibroblasts. Biochem. Biophys. Res. Commun. 2014, 444, 682–686. [Google Scholar] [CrossRef]
- Karunaweera, N.; Raju, R.; Gyengesi, E.; Munch, G. Plant polyphenols as inhibitors of NF-kappa B induced cytokine production a potential anti-inflammatory treatment for Alzheimer’s disease? Front. Mol. Neurosci. 2015, 8. [Google Scholar] [CrossRef] [PubMed]
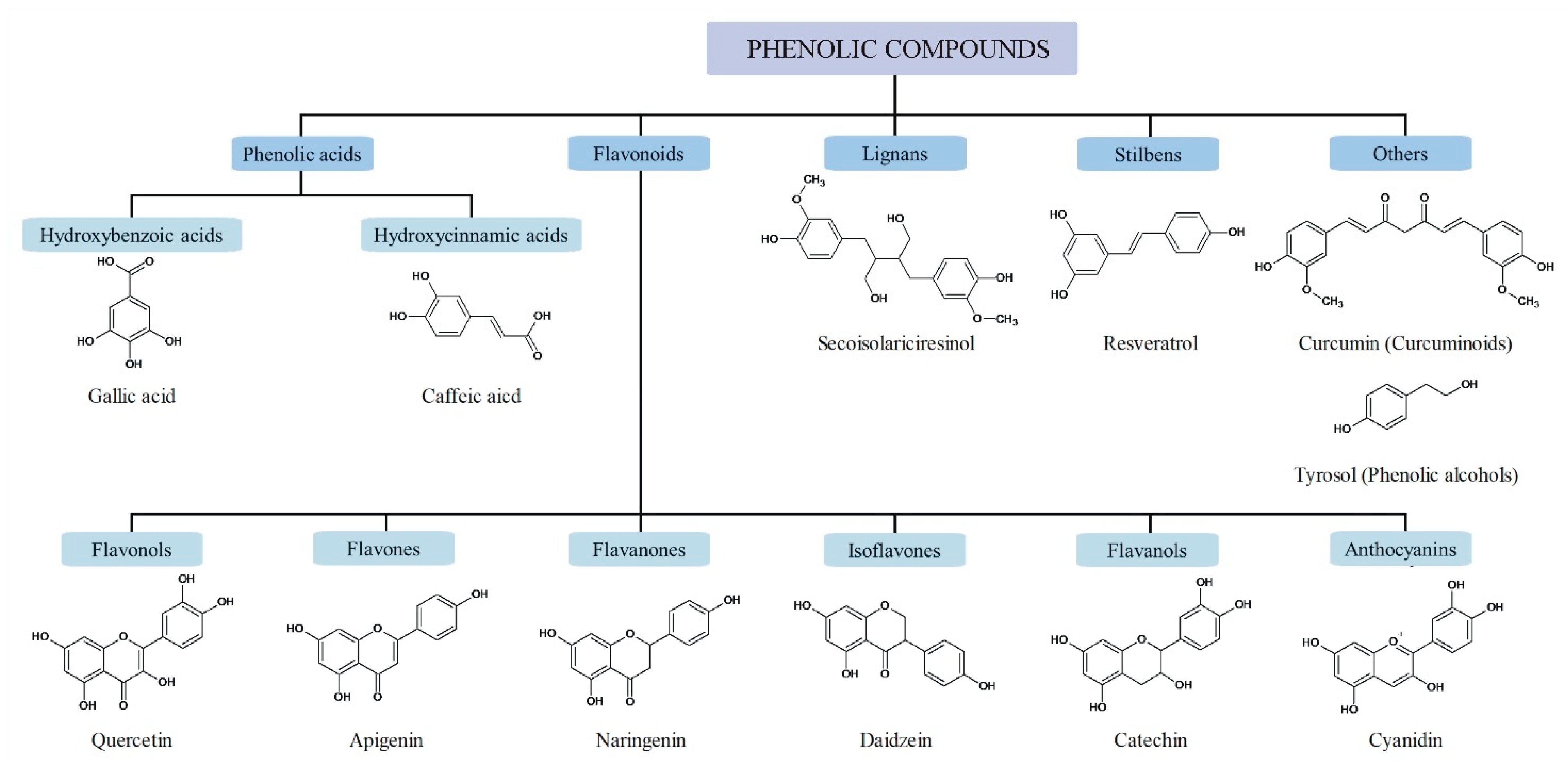
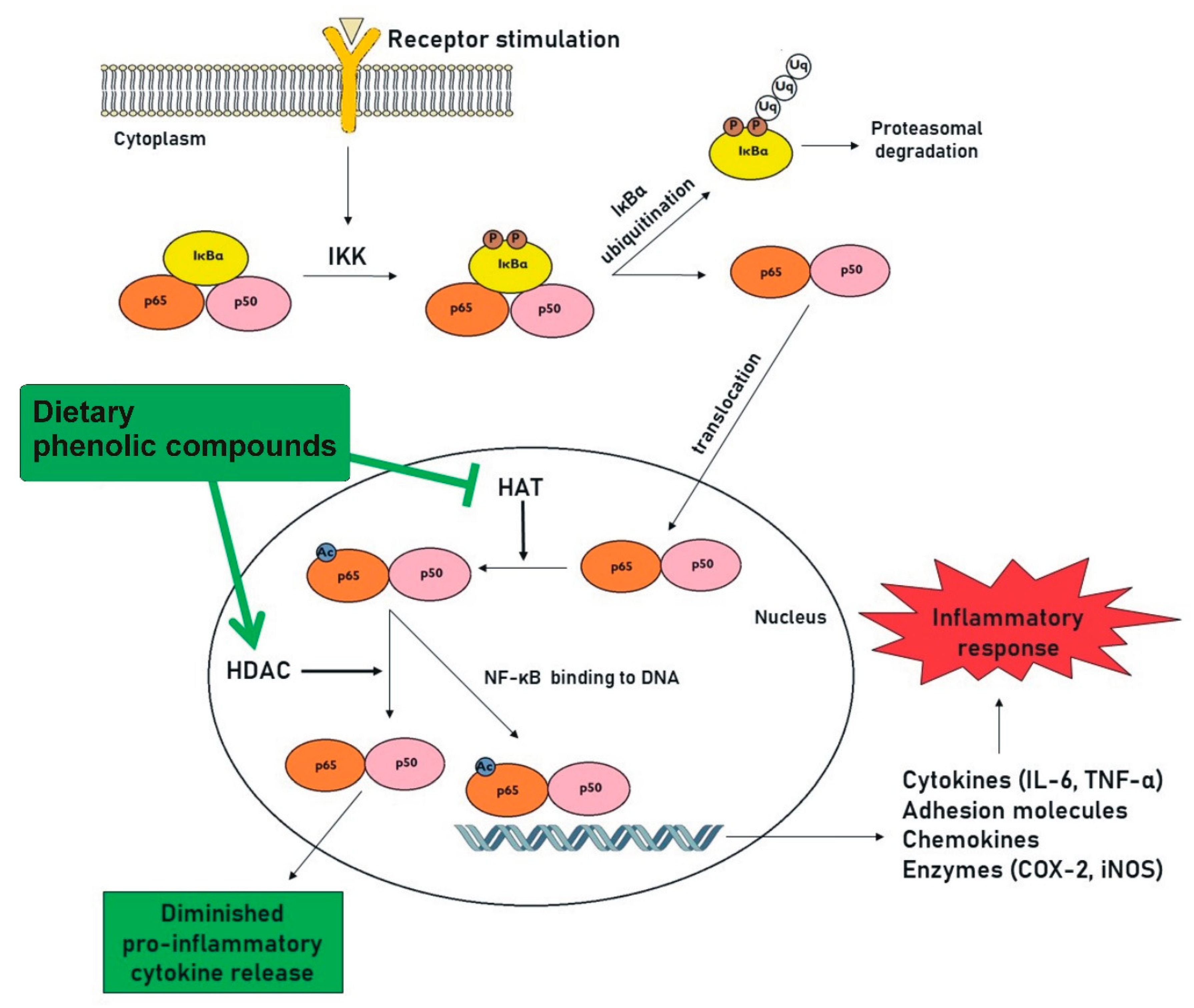
| Classification | Examples of Phenolic Compounds | Main Sources | Possible Targets | References | |
|---|---|---|---|---|---|
| Phenolic Acids | Hydroxybenzoic acids | gallic, protocatechuic, ellagic acids | berries, olive oil, tea | HATs, HDACs | [12] |
| Hydroxycinnamic acids | coumaric, ferulic, sinapic, caffeic acids | fruits (blueberries, cherries, apples), grains | DNMTs | [13] | |
| Flavonoids | Flavonols | quercetin, kaempferol, fisetin | vegetables (leek, broccoli), blueberries | HATs, HDACs | [14,15] |
| Flavones | luteolin, apigenin | parsley, celery | HATs, HDACs | [16] | |
| Flavanones | naringenin, hesperetin, eriodictyol | citrus fruits | not determined | ||
| Isoflavones | genistein, daidzein, glycitein | legumes (soya) | DNMTs, HATs, SIRT1, miRNA | [17,18,19] | |
| Flavanols | catechin, epicatechin, gallocatechin, epigallocatechin, epigallocatechin-3-gallate | green tea, cocoa | DNMTs, HATs. HDACs, HMTs | [20,21,22] | |
| Anthocyanins | cyanidin | fruits (blackcurrants, blackberries) | not determined | ||
| Lignans | secoisolariciresinol, matairesinol | linseed | not determined | ||
| Stilbenes | resveratrol, pterostilbene | grapes, red wine, blueberries | SIRT1, DNMTs, miRNA | [23,24,25,26] | |
| Phenolic alcohols | tyrosol, hydroxytyrosol | virgin olive oil | not determined | ||
| Curcuminoids | curcumin | turmeric | DNMTs, miRNA, HATs, HDACs | [27,28,29,30] | |
| Species | Strain Characteristics | Sample Size (per Group) | Pathological Condition | Type of Study | Phenolic Compound Intervention | Reference |
|---|---|---|---|---|---|---|
| Mus musculus | CD45.2 + C57BL/6 | n = 9–10 | repeated social defeat stress | observational, cross-sectional | dihydrocaffeic acid 5 mg/kg/d for 2 weeks, delivered through drinking water, treatment repeated 4 months later | [13] |
| Mus musculus | BALB/c 6 weeks old 20–25 g body weight | n = 3 | intervention | epigallocatechin-3-gallate 100 mmol/L, 50 mmol/L | [21] | |
| Mus musculus | C57BL/6J males 6–8 weeks old 20–30 g body weight | n = 12 | inflammation | intervention | curcumin 200 mg/kg (i.p.) | [74] |
| Rattus norvegicus | Sprague–Dawley females 250–300 g body weight | n = 6 | inflammation | intervention | curcumin 40 mg/kg (final injection volume of 0.2 mL) | [75] |
| Rattus norvegicus | Wistar females 10 weeks old 180–220 g body weight | n = 10 | ovariectomized diabetic rat | intervention | genistein 1 mg/kg/day, subcutaneously, for 8 weeks | [18] |
| Rattus norvegicus | Sprague–Dawley males two weeks old | n = 19 (normal group), 16 (normal intervention group), 29 (diabetic group), 25 (diabetic intervention group) | diabetes | intervention | resveratrol intraperitoneally 10 mg/kg/day, for 10 weeks | [25] |
| Type of Study | Sample Size | Participant Characteristics | Dietary Intervention | Disease | Reference |
|---|---|---|---|---|---|
| intervention | n = 128 | age ≥40 years | one capsule/day of resveratrol 500 mg/day (n = 43), 40 mg/day (n = 43) or placebo (n = 42), for 6 months | type 2 diabetes mellitus | [76] |
| intervention | n = 214 | men and women 20 years of age | Treated group (45 men and 65 women) received the cocoa cream product (13 g/unit; 1 g cocoa/unit, 6 units/d; 6 g cocoa/d) for 2 weeks. Control group consisted of 46 men and 58 women. | cardiovascular disease risk | [77] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Číž, M.; Dvořáková, A.; Skočková, V.; Kubala, L. The Role of Dietary Phenolic Compounds in Epigenetic Modulation Involved in Inflammatory Processes. Antioxidants 2020, 9, 691. https://doi.org/10.3390/antiox9080691
Číž M, Dvořáková A, Skočková V, Kubala L. The Role of Dietary Phenolic Compounds in Epigenetic Modulation Involved in Inflammatory Processes. Antioxidants. 2020; 9(8):691. https://doi.org/10.3390/antiox9080691
Chicago/Turabian StyleČíž, Milan, Adéla Dvořáková, Veronika Skočková, and Lukáš Kubala. 2020. "The Role of Dietary Phenolic Compounds in Epigenetic Modulation Involved in Inflammatory Processes" Antioxidants 9, no. 8: 691. https://doi.org/10.3390/antiox9080691
APA StyleČíž, M., Dvořáková, A., Skočková, V., & Kubala, L. (2020). The Role of Dietary Phenolic Compounds in Epigenetic Modulation Involved in Inflammatory Processes. Antioxidants, 9(8), 691. https://doi.org/10.3390/antiox9080691







