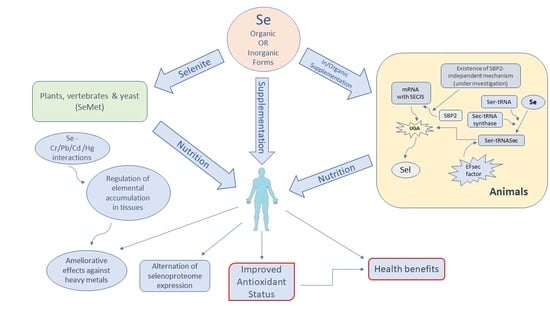
Selenium-Dependent Antioxidant Enzymes: Actions and Properties of Selenoproteins
Abstract
1. Introduction
2. Synthesis of Selenoproteins from Dietary Selenium
3. Selenoproteins and the Antioxidant System
4. Effects of Selenium Supplementation in Chicken
5. Actions and Properties of Selenoproteins
6. Effects of Se Deficiency on mRNA Expression of Selenoproteins and Non-Selenoproteins
7. Ranking in the Hierarchy of Selenoproteins
8. Recent Findings and Main Characteristics of Major Selenoproteins
9. Conclusions
Author Contributions
Conflicts of Interest
References
- Mariotti, M.; Santesmasses, D.; Guigó, R. Evolution of Selenophosphate Synthetase; Springer: New York, NY, USA, 2016. [Google Scholar]
- Hatfield, D.L.; Marla, J.B.; Vadim, N.; Gladyshev, V.N. Selenium: Its Molecular Biology and Role in Human Health; Springer Science & Business Media: Berlin, Germany, 2011. [Google Scholar]
- Mariotti, M. Selenocysteine Extinctions in Insects. In Short Views on Insect Genomics and Proteomics; Springer: New York, NY, USA, 2016. [Google Scholar]
- Mariotti, M.; Lobanov, A.V.; Manta, B.; Santesmasses, D.; Bofill, A.; Guigó, R.; Gabaldón, T.; Gladyshev, V.N. Lokiarchaeota marks the transition between the archaeal and eukaryotic selenocysteine encoding systems. Mol. Biol. Evol. 2016, 33, 2441–2453. [Google Scholar] [CrossRef] [PubMed]
- Sarangi, K.G.; White, L.; Castellano, S. Genetic Adaptation and Selenium Uptake in Vertebrates; Wiley: Chichester, UK, 2017. [Google Scholar]
- Peng, T.; Lin, J.; Xu, Y.-Z.; Zhang, Y. Comparative genomics reveals new evolutionary and ecological patterns of selenium utilization in bacteria. ISME J. 2016, 10, 2048–2059. [Google Scholar] [CrossRef] [PubMed]
- Santesmasses, D.; Mariotti, M.; Guigó, R. Computational identification of the selenocysteine tRNA (tRNASec) in genomes. PLoS Comput. Biol. 2017, 13. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority. Scientific opinion on dietary reference values for selenium. EFSA J. 2014, 12. [Google Scholar] [CrossRef]
- Rayman, M.P. Selenium and human health. Lancet 2012, 379, 1256–1268. [Google Scholar] [CrossRef]
- Steinbrenner, H.; Speckmann, B.; Sies, H. Toward understanding success and failures in the use of selenium for cancer prevention. Antioxid. Redox Signal. 2013, 19, 181–191. [Google Scholar] [CrossRef] [PubMed]
- Méplan, C. Association of single nucleotide polymorphisms in selenoprotein genes with cancer risk. In Selenoproteins; Humana Press: New York, NY, USA, 2018. [Google Scholar]
- Bertz, M.; Kühn, K.; Koeberle, S.C.; Müller, M.F.; Hoelzer, D.; Thies, K.; Deubel, S.; Thierbach, R.; Kipp, A.P. Selenoprotein H controls cell cycle progression and proliferation of human colorectal cancer cells. Free Radic. Biol. Med. 2018, in press. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, L. Selenium, selenoproteins and the thyroid gland: Interactions in health and disease. Nat. Rev. Endocrinol. 2012, 8, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Handy, D.E.; Joseph, L. Selenoproteins in Cardiovascular Redox Pathology; Springer: Cham, Switzerland, 2016. [Google Scholar]
- Pekar, J.; Skolarczyk, J.; Małecka-Massalska, T.; Skórzyńska-Dziduszko, K. Effect of selenium supplementation in thyroid gland diseases. J. Elementol. 2017, 22. [Google Scholar] [CrossRef]
- Misu, H.; Takayama, H.; Saito, Y.; Mita, Y.; Kikuchi, A.; Ishii, K.-A.; Chikamoto, K.; Kanamori, T.; Tajima, N.; Lan, F. Deficiency of the hepatokine selenoprotein P increases responsiveness to exercise in mice through upregulation of reactive oxygen species and AMP-activated protein kinase in muscle. Nat. Med. 2017, 23, 508–516. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, N.; Satoh, K.; Omura, J.; Satoh, T.; Kurosawa, R.; Nogi, M.; Otsuki, T.; Numano, K.; Kozu, K.; Suzuki, K. Selenoprotein P promotes vascular smooth muscle cell proliferation and pulmonary hypertension-a possible novel therapeutic target. Am. Heart Assoc. 2016, 36, A62. [Google Scholar]
- Wright, C.R.; Allsopp, G.L.; Addinsall, A.B.; McRae, N.L.; Andrikopoulos, S.; Stupka, N. A Reduction in selenoprotein S amplifies the inflammatory profile of fast-twitch skeletal muscle in the mdx dystrophic mouse. Med. Inflamm. 2017, 2017. [Google Scholar] [CrossRef] [PubMed]
- Ye, Y.; Fu, F.; Li, X.; Yang, J.; Liu, H. Selenoprotein s is highly expressed in the blood vessels and prevents vascular smooth muscle cells from apoptosis. J. Cell. Biochem. 2016, 117, 106–117. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.; Fan, R.; Zhao, X.; Zhao, W.; Liu, W.; Yang, J.; Sattar, H.; Zhao, J.; Zhang, Z.; Xu, S. Selenoprotein W redox-regulated Ca2+ channels correlate with selenium deficiency-induced muscles Ca2+ leak. Oncotarget 2016, 7, 57618–57632. [Google Scholar] [CrossRef] [PubMed]
- Rocourt, C.R.; Cheng, W.-H. Selenium supranutrition: Are the potential benefits of chemoprevention outweighed by the promotion of diabetes and insulin resistance? Nutrients 2013, 5, 1349–1365. [Google Scholar] [CrossRef] [PubMed]
- Kryukov, G.V.; Castellano, S.; Novoselov, S.V.; Lobanov, A.V.; Zehtab, O.; Guigó, R.; Gladyshev, V.N. Characterization of mammalian selenoproteomes. Science 2003, 300, 1439–1443. [Google Scholar] [CrossRef] [PubMed]
- Gladyshev, V.N.; Arnér, E.S.; Berry, M.J.; Brigelius-Flohé, R.; Bruford, E.A.; Burk, R.F.; Carlson, B.A.; Castellano, S.; Chavatte, L.; Conrad, M. Selenoprotein gene nomenclature. J. Biol. Chem. 2016, 291, 24036–24040. [Google Scholar] [CrossRef] [PubMed]
- Kumar, B.S.; Priyadarsini, K. Selenium nutrition: How important is it? Biomed. Prev. Nutr. 2014, 4, 333–341. [Google Scholar] [CrossRef]
- Roman, M.; Jitaru, P.; Barbante, C. Selenium biochemistry and its role for human health. Metallomics 2014, 6, 25–54. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Wang, T.; Li, Q.; Li, D. Prevention of keshan disease by selenium supplementation: A systematic review and meta-analysis. Biol. Trace Elem. Res. 2018. [Google Scholar] [CrossRef] [PubMed]
- Liu, H.; Yu, F.; Shao, W.; Ding, D.; Yu, Z.; Chen, F.; Geng, D.; Tan, X.; Lammi, M.J.; Guo, X. Associations between selenium content in hair and kashin-beck disease/keshan disease in children in northwestern china: A prospective cohort study. Biol. Trace Elem. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Loscalzo, J. Keshan disease, selenium deficiency, and the selenoproteome. N. Engl. J. Med. 2014, 370, 1756–1760. [Google Scholar] [CrossRef] [PubMed]
- Zhang, R. Polymorphism of antioxidant selenoprotein genes and kashin-beck disease susceptibility, a systematic review and meta-analysis. Osteoarthr. Cartil. 2017, 25, 213–214. [Google Scholar] [CrossRef][Green Version]
- Ventura, M.; Melo, M.; Carrilho, F. Selenium and thyroid disease: From pathophysiology to treatment. Int. J. Endocrinol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Agarwal, A.; Majzoub, A. Role of antioxidants in male infertility. BJUI Knowl. 2016, 1–9. [Google Scholar]
- Pappa, E.C.; Pappas, A.C.; Surai, P.F. Selenium content in selected foods from the greek market and estimation of the daily intake. Sci. Total Environ. 2006, 372, 100–108. [Google Scholar] [CrossRef] [PubMed]
- Wasowicz, W.; Gromadzinska, J.; Rydzynski, K.; Tomczak, J. Selenium status of low-selenium area residents: Polish experience. Toxicol. Lett. 2003, 137, 95–101. [Google Scholar] [CrossRef]
- Barclay, M.N.I.; MacPherson, A.; Dixon, J. Selenium content of a range of UK foods. J. Food Compos. Anal. 1995, 8, 307–318. [Google Scholar] [CrossRef]
- Diaz-Alarcon, J.; Navarro-Alarcón, M.; de la Serrana, H.L.-G.; Lopez-Martinez, M. Determination of selenium in cereals, legumes and dry fruits from southeastern spain for calculation of daily dietary intake. Sci. Total Environ. 1996, 184, 183–189. [Google Scholar] [CrossRef]
- Sigrist, M.; Brusa, L.; Campagnoli, D.; Beldoménico, H. Determination of selenium in selected food samples from argentina and estimation of their contribution to the Se dietary intake. Food Chem. 2012, 134, 1932–1937. [Google Scholar] [CrossRef] [PubMed]
- Kipp, A.P.; Strohm, D.; Brigelius-Flohé, R.; Schomburg, L.; Bechthold, A.; Leschik-Bonnet, E.; Heseker, H. German Nutrition Society (DGE). Revised reference values for selenium intake. J. Trace Elem. Med. Biol. 2015, 32, 195–199. [Google Scholar] [CrossRef] [PubMed]
- Fisinin, V.I.; Papazyan, T.T.; Surai, P.F. Producing selenium-enriched eggs and meat to improve the selenium status of the general population. Crit. Rev. Biotechnol. 2009, 29, 18–28. [Google Scholar] [CrossRef] [PubMed]
- Combs, G.; Combs, S. The nutritional biochemistry of selenium. Annu. Rev. Nutr. 1984, 4, 257–280. [Google Scholar] [CrossRef] [PubMed]
- Heras, I.L.; Palomo, M.; Madrid, Y. Selenoproteins: The key factor in selenium essentiality. State of the art analytical techniques for selenoprotein studies. Anal. Bioanal. Chem. 2011, 400, 1717–1727. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, D.; Schlautman, B.; Steffan, S.; Polashock, J.; Vorsa, N.; Zalapa, J. The american cranberry mitochondrial genome reveals the presence of selenocysteine (tRNA-Sec and SECIS) insertion machinery in land plants. Gene 2014, 536, 336–343. [Google Scholar] [CrossRef] [PubMed]
- Rayman, M.P. The use of high-selenium yeast to raise selenium status: How does it measure up? Br. J. Nutr. 2004, 92, 557–573. [Google Scholar] [CrossRef] [PubMed]
- Combs, G.F. Selenium in global food systems. Br. J. Nutr. 2001, 85, 517–547. [Google Scholar] [CrossRef] [PubMed]
- Pesti, G.; Combs, G., Jr. Studies on the enteric absorption of selenium in the chick using localized coccidial infections. Poult. Sci. 1976, 55, 2265–2274. [Google Scholar] [CrossRef] [PubMed]
- Wolffram, S. Absorption and metabolism of selenium: Difference between organic and inorganic sources. In Biotechnology in the Feed Industry; Nottingham University Press: Nottingham, UK, 1999; Volume 15, pp. 547–566. [Google Scholar]
- Würmli, R.; Wolffram, S.; Stingelin, Y.; Scharrer, E. Stimulation of mucosal uptake of selenium from selenite by l-cysteine in sheep small intestine. Biol. Trace Elem. Res. 1989, 20, 75–85. [Google Scholar] [CrossRef] [PubMed]
- Huang, K.; Lauridsen, E.; Clausen, J. The uptake of Na-selenite in rat brain. Biol. Trace Elem. Res. 1994, 46, 91–102. [Google Scholar] [CrossRef] [PubMed]
- Vendeland, S.; Deagen, J.; Butler, J.; Whanger, P. Uptake of selenite, selenomethionine and selenate by brush border membrane vesicles isolated from rat small intestine. Biometals 1994, 7, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Mariotti, M.; Guigó, R. Selenoprofiles: Profile-based scanning of eukaryotic genome sequences for selenoprotein genes. Bioinformatics 2010, 26, 2656–2663. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Holmgren, A. Selenoproteins. J. Biol. Chem. 2009, 284, 723–727. [Google Scholar] [CrossRef] [PubMed]
- Rayman, M.P. The importance of selenium to human health. Lancet 2000, 356, 233–241. [Google Scholar] [CrossRef]
- Carlson, B.A.; Lee, B.J.; Tsuji, P.A.; Copeland, P.R.; Schweizer, U.; Gladyshev, V.N.; Hatfield, D.L. Selenocysteine tRNA [Ser] Sec, the central component of selenoprotein biosynthesis: Isolation, identification, modification, and sequencing. In Selenoproteins; Humana Press: New York, NY, USA, 2018. [Google Scholar]
- Böck, A. Biosynthesis of selenoproteins—An overview. Biofactors 2000, 11, 77–78. [Google Scholar] [CrossRef] [PubMed]
- Lacourciere, G.M.; Stadtman, T.C. Utilization of selenocysteine as a source of selenium for selenophosphate biosynthesis. Biofactors 2001, 14, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Schoenmakers, E.; Carlson, B.; Agostini, M.; Moran, C.; Rajanayagam, O.; Bochukova, E.; Tobe, R.; Peat, R.; Gevers, E.; Muntoni, F. Mutation in human selenocysteine transfer RNA selectively disrupts selenoprotein synthesis. J. Clin. Investig. 2016, 126, 992–996. [Google Scholar] [CrossRef] [PubMed]
- Gribling-Burrer, A.-S.; Leichter, M.; Wurth, L.; Huttin, A.; Schlotter, F.; Troffer-Charlier, N.; Cura, V.; Barkats, M.; Cavarelli, J.; Massenet, S. SECIS-binding protein 2 interacts with the SMN complex and the methylosome for selenoprotein mRNP assembly and translation. Nucleic Acids Res. 2017, 45, 5399–5413. [Google Scholar] [CrossRef] [PubMed]
- Chambers, I.; Frampton, J.; Goldfarb, P.; Affara, N.; McBain, W.; Harrison, P.R. The structure of the mouse glutathione peroxidase gene: The selenocysteine in the active site is encoded by the ‘termination’ codon, TGA. EMBO J. 1986, 5, 1221–1227. [Google Scholar] [PubMed]
- Copeland, P.R.; Driscoll, D.M. RNA binding proteins and selenocysteine. Biofactors 2001, 14, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.-R.; Burnap, R.L.; Inoue, Y. An independent role of cytochrome c-550 in cyanobacterial photosystem II as revealed by double-deletion mutagenesis of the psbO and psbV genes in Synechocystis sp. PCC 6803. Biochemistry 1995, 34, 12661–12668. [Google Scholar] [CrossRef] [PubMed]
- Low, S.C.; Berry, M.J. Knowing when not to stop: Selenocysteine incorporation in eukaryotes. Trends Biochem. Sci. 1996, 21, 203–208. [Google Scholar] [CrossRef]
- Bellinger, F.P.; Raman, A.V.; Reeves, M.A.; Berry, M.J. Regulation and function of selenoproteins in human disease. Biochem. J. 2009, 422, 11–22. [Google Scholar] [CrossRef] [PubMed]
- Copeland, P.R.; Fletcher, J.E.; Carlson, B.A.; Hatfield, D.L.; Driscoll, D.M. A novel RNA binding protein, SBP2, is required for the translation of mammalian selenoprotein mRNAs. EMBO J. 2000, 19, 306–314. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, L.; Schweizer, U.; Köhrle, J. Selenium and selenoproteins in mammals: Extraordinary, essential, enigmatic. Cell. Mol. Life Sci. 2004, 61, 1988–1995. [Google Scholar] [CrossRef] [PubMed]
- Fujiwara, T.; Busch, K.; Gross, H.J.; Mizutani, T. A SECIS binding protein (SBP) is distinct from selenocysteyl-tRNA protecting factor (SePF). Biochimie 1999, 81, 213–218. [Google Scholar] [CrossRef]
- Fletcher, J.E.; Copeland, P.R.; Driscoll, D.M. Polysome distribution of phospholipid hydroperoxide glutathione peroxidase mRNA: Evidence for a block in elongation at the UGA/selenocysteine codon. RNA 2000, 6, 1573–1584. [Google Scholar] [CrossRef] [PubMed]
- Martin, G.W., III; Berry, M.J. Selenocysteine codons decrease polysome association on endogenous selenoprotein mRNAs. Genes Cells 2001, 6, 121–129. [Google Scholar] [CrossRef] [PubMed]
- Hatfield, D.; Lee, B.J.; Hampton, L.; Diamond, A.M. Selenium induces changes in the selenocysteine tRNA [Ser] Sec population in mammalian cells. Nucleic Acids Res. 1991, 19, 939–943. [Google Scholar] [CrossRef] [PubMed]
- Chittum, H.S.; Baek, H.J.; Diamond, A.M.; Fernandez-Salguero, P.; Gonzalez, F.; Ohama, T.; Hatfield, D.L.; Kuehn, M.; Lee, B.J. Selenocysteine tRNA [Ser] Sec levels and selenium-dependent glutathione peroxidase activity in mouse embryonic stem cells heterozygous for a targeted mutation in the tRNA [Ser] Sec gene. Biochemistry 1997, 36, 8634–8639. [Google Scholar] [CrossRef] [PubMed]
- Jameson, R.R.; Carlson, B.A.; Butz, M.; Esser, K.; Hatfield, D.L.; Diamond, A.M. Selenium influences the turnover of selenocysteine tRNA [Ser] Sec in chinese hamster ovary cells. J. Nutr. 2002, 132, 1830–1835. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Carlson, B.A.; Xu, X.-M.; Gladyshev, V.N.; Hatfield, D.L. Selective rescue of selenoprotein expression in mice lacking a highly specialized methyl group in selenocysteine tRNA. J. Biol. Chem. 2005, 280, 5542–5548. [Google Scholar] [CrossRef] [PubMed]
- Jameson, R.R.; Diamond, A.M. A regulatory role for sec tRNA [Ser] Sec in selenoprotein synthesis. RNA 2004, 10, 1142–1152. [Google Scholar] [CrossRef] [PubMed]
- Seeher, S.; Atassi, T.; Mahdi, Y.; Carlson, B.A.; Braun, D.; Wirth, E.K.; Klein, M.O.; Reix, N.; Miniard, A.C.; Schomburg, L. Secisbp2 is essential for embryonic development and enhances selenoprotein expression. Antioxid. Redox Signal. 2014, 21, 835–849. [Google Scholar] [CrossRef] [PubMed]
- Papp, L.V.; Lu, J.; Holmgren, A.; Khanna, K.K. From selenium to selenoproteins: Synthesis, identity, and their role in human health. Antioxid. Redox Signal. 2007, 9, 775–806. [Google Scholar] [CrossRef] [PubMed]
- Mills, G.C. The purification and properties of glutathione peroxidase of erythrocytes. J. Biol. Chem. 1959, 234, 502–506. [Google Scholar] [PubMed]
- Schacter, L. Generation of superoxide anion and hydrogen peroxide by erythrocytes from individuals with sickle trait or normal haemoglobin. Eur. J. Clin. Investig. 1986, 16, 204–210. [Google Scholar] [CrossRef]
- Persson, T.; Popescu, B.O.; Cedazo-Minguez, A. Oxidative stress in alzheimer’s disease: Why did antioxidant therapy fail? Oxid. Med. Cell. Longev. 2014. [Google Scholar] [CrossRef] [PubMed]
- Ramoutar, R.R.; Brumaghim, J.L. Antioxidant and anticancer properties and mechanisms of inorganic selenium, oxo-sulfur, and oxo-selenium compounds. Cell Biochem. Biophys. 2010, 58, 1–23. [Google Scholar] [CrossRef] [PubMed]
- Surai, P.F. Selenium in Nutrition and Health; Nottingham University Press: Nottingham, UK, 2006. [Google Scholar]
- Ayala, A.; Muñoz, M.F.; Argüelles, S. Lipid peroxidation: Production, metabolism, and signaling mechanisms of malondialdehyde and 4-hydroxy-2-nonenal. Oxid. Med. Cell. Longev. 2014. [Google Scholar] [CrossRef] [PubMed]
- Surai, P.F. Vitamin e in Avian Reproduction; FAO: Rome, Italy, 1999. [Google Scholar]
- Ames, B.N.; Gold, L.S. The causes and prevention of cancer: Gaining perspective. Environ. Health Perspect. 1997, 105, 865–873. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Helbock, H.J.; Beckman, K.B.; Shigenaga, M.K.; Walter, P.B.; Woodall, A.A.; Yeo, H.C.; Ames, B.N. DNA oxidation matters: The HPLC-electrochemical detection assay of 8-oxo-deoxyguanosine and 8-oxo-guanine. Proc. Natl. Acad. Sci. USA 1998, 95, 288–293. [Google Scholar] [CrossRef] [PubMed]
- Ames, B.N. An enthusiasm for metabolism. J. Biol. Chem. 2003, 278, 4369–4380. [Google Scholar] [CrossRef] [PubMed]
- Lobo, V.; Patil, A.; Phatak, A.; Chandra, N. Free radicals, antioxidants and functional foods: Impact on human health. Pharmacogn. Rev. 2010, 4, 118–126. [Google Scholar] [CrossRef] [PubMed]
- Lagouge, M.; Larsson, N.G. The role of mitochondrial DNA mutations and free radicals in disease and ageing. J. Intern. Med. 2013, 273, 529–543. [Google Scholar] [CrossRef] [PubMed]
- Diplock, A. Antioxidants and Disease Prevention Molecular Aspects of Medicine; Pergamon Press: Oxford, UK, 1994. [Google Scholar]
- Halliwell, B. Free radicals and antioxidants: A personal view. Nutr. Rev. 1994, 52, 253–265. [Google Scholar] [CrossRef] [PubMed]
- Davies, M.J. Protein oxidation and peroxidation. Biochem. J. 2016, 473, 805–825. [Google Scholar] [CrossRef] [PubMed]
- Hawkins, C.L.; Morgan, P.E.; Davies, M.J. Quantification of protein modification by oxidants. Free Radic. Biol. Med. 2009, 46, 965–988. [Google Scholar] [CrossRef] [PubMed]
- Berlett, B.S.; Stadtman, E.R. Protein oxidation in aging, disease, and oxidative stress. J. Biol. Chem. 1997, 272, 20313–20316. [Google Scholar] [CrossRef] [PubMed]
- Tramutola, A.; Lanzillotta, C.; Perluigi, M.; Butterfield, D.A. Oxidative stress, protein modification and alzheimer disease. Brain Res. Bull. 2017, 133, 88–96. [Google Scholar] [CrossRef] [PubMed]
- Stadtman, E.R. Protein oxidation and aging. Science 1992, 257, 1220–1224. [Google Scholar] [CrossRef] [PubMed]
- Stadtman, E.R.; Levine, R.L. Protein oxidation. Ann. N. Y. Acad. Sci. 2000, 899, 191–208. [Google Scholar] [CrossRef] [PubMed]
- Sultana, R.; Perluigi, M.; Butterfield, D.A. Protein oxidation and lipid peroxidation in brain of subjects with alzheimer’s disease: Insights into mechanism of neurodegeneration from redox proteomics. Antioxid. Redox Signal. 2006, 8, 2021–2037. [Google Scholar] [CrossRef] [PubMed]
- Steinbrenner, H.; Speckmann, B.; Klotz, L.-O. Selenoproteins: Antioxidant selenoenzymes and beyond. Arch. Biochem. Biophys. 2016, 595, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Storkey, C.; Pattison, D.I.; Ignasiak, M.T.; Schiesser, C.H.; Davies, M.J. Kinetics of reaction of peroxynitrite with selenium-and sulfur-containing compounds: Absolute rate constants and assessment of biological significance. Free Radic. Biol. Med. 2015, 89, 1049–1056. [Google Scholar] [CrossRef] [PubMed]
- Pappas, A.; Zoidis, E.; Surai, P.; Zervas, G. Selenoproteins and maternal nutrition. Comp. Biochem. Physiol. B 2008, 151, 361–372. [Google Scholar] [CrossRef] [PubMed]
- Toppo, S.; Flohé, L.; Ursini, F.; Vanin, S.; Maiorino, M. Catalytic mechanisms and specificities of glutathione peroxidases: Variations of a basic scheme. BBA-Gen. Subj. 2009, 1790, 1486–1500. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.-A.; Kirnarsky, L.; Sherman, S.; Gladyshev, V.N. Selenoprotein oxidoreductase with specificity for thioredoxin and glutathione systems. Proc. Natl. Acad. Sci. USA 2001, 98, 3673–3678. [Google Scholar] [CrossRef] [PubMed]
- Arnér, E.S. Selenoproteins—What unique properties can arise with selenocysteine in place of cysteine? Exp. Cell Res. 2010, 316, 1296–1303. [Google Scholar] [CrossRef] [PubMed]
- Brzezińska-Ślebodzińska, E.; Ślebodziński, A.B.; Pietras, B.; Wieczorek, G. Antioxidant effect of vitamin E and glutathione on lipid peroxidation in boar semen plasma. Biol. Trace Elem. Res. 1995, 47, 69–74. [Google Scholar] [CrossRef]
- Kagan, V.; Nohl, H.; Quinn, P.; Cadenas, E.; Packer, L. Handbook of Antioxidants; Marcel Dekker: New York, NY, USA, 1996. [Google Scholar]
- Surai, P.F. Natural Antioxidants in Avian Nutrition and Reproduction; Nottingham University Press: Nottingham, UK, 2002. [Google Scholar]
- Młochowski, J.; Brząszcz, M.; Giurg, M.; Palus, J.; Wójtowicz, H. Selenium-promoted oxidation of organic compounds: Reactions and mechanisms. Eur. J. Org. Chem. 2003, 2003, 4329–4339. [Google Scholar] [CrossRef]
- Bowie, A.; O’Neill, L.A. Oxidative stress and nuclear factor-κb activation: A reassessment of the evidence in the light of recent discoveries. Biochem. Pharmacol. 2000, 59, 13–23. [Google Scholar] [CrossRef]
- Thundathil, J.; de Lamirande, E.; Gagnon, C. Nitric oxide regulates the phosphorylation of the threonine-glutamine-tyrosine motif in proteins of human spermatozoa during capacitation. Biol. Reprod. 2003, 68, 1291–1298. [Google Scholar] [CrossRef] [PubMed]
- Halliwell, B.; Gutteridge, J.M. Free Radicals in Biology and Medicine; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- McCord, J.M. The evolution of free radicals and oxidative stress. Am. J. Med. 2000, 108, 652–659. [Google Scholar] [CrossRef]
- Dröge, W. Free radicals in the physiological control of cell function. Physiol. Rev. 2002, 82, 47–95. [Google Scholar] [CrossRef] [PubMed]
- Yoon, S.-O.; Yun, C.-H.; Chung, A.-S. Dose effect of oxidative stress on signal transduction in aging. Mech. Ageing Dev. 2002, 123, 1597–1604. [Google Scholar] [CrossRef]
- Seifried, H.E.; Anderson, D.E.; Fisher, E.I.; Milner, J.A. A review of the interaction among dietary antioxidants and reactive oxygen species. J. Nutr. Biochem. 2007, 18, 567–579. [Google Scholar] [CrossRef] [PubMed]
- Steinbrenner, H.; Sies, H. Protection against reactive oxygen species by selenoproteins. Biochim. Biophys. Acta 2009, 1790, 1478–1485. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Limphong, P.; Pieper, J.; Liu, Q.; Rodesch, C.K.; Christians, E.; Benjamin, I.J. Glutathione-dependent reductive stress triggers mitochondrial oxidation and cytotoxicity. FASEB J. 2012, 26, 1442–1451. [Google Scholar] [CrossRef] [PubMed]
- Meister, A. On the antioxidant effects of ascorbic acid and glutathione. Biochem. Pharmacol. 1992, 44, 1905–1915. [Google Scholar] [CrossRef]
- Dannenmann, B.; Lehle, S.; Hildebrand, D.G.; Kübler, A.; Grondona, P.; Schmid, V.; Holzer, K.; Fröschl, M.; Essmann, F.; Rothfuss, O.; et al. High glutathione and glutathione peroxidase-2 levels mediate cell-type-specific DNA damage protection in human induced pluripotent stem cells. Stem Cell Rep. 2015, 4, 886–898. [Google Scholar]
- Kanwal, R.; Pandey, M.; Bhaskaran, N.; MacLennan, G.T.; Fu, P.; Ponsky, L.E.; Gupta, S. Protection against oxidative DNA damage and stress in human prostate by glutathione S-transferase P1. Mol. Carcinog. 2014, 53, 8–18. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Vivancos, P.; de Simone, A.; Kiddle, G.; Foyer, C.H. Glutathione—Linking cell proliferation to oxidative stress. Free Radic. Biol. Med. 2015, 89, 1154–1164. [Google Scholar] [CrossRef] [PubMed]
- Jin, L.; Li, D.; Alesi, G.N.; Fan, J.; Kang, H.-B.; Lu, Z.; Boggon, T.J.; Jin, P.; Yi, H.; Wright, E.R.; et al. Glutamate dehydrogenase 1 signals through antioxidant glutathione peroxidase 1 to regulate redox homeostasis and tumor growth. Cancer Cell 2015, 27, 257–270. [Google Scholar] [CrossRef] [PubMed]
- Geltink, R.I.K.; O’Sullivan, D.; Pearce, E.L. Caught in the crossfire: GSH controls T cell metabolic reprogramming. Immunity 2017, 46, 525–527. [Google Scholar] [CrossRef] [PubMed]
- Mak, T.W.; Grusdat, M.; Duncan, G.S.; Dostert, C.; Nonnenmacher, Y.; Cox, M.; Binsfeld, C.; Hao, Z.; Brüstle, A.; Itsumi, M. Glutathione primes T cell metabolism for inflammation. Immunity 2017, 46, 675–689. [Google Scholar] [CrossRef] [PubMed]
- Sen, C.K.; Packer, L. Thiol homeostasis and supplements in physical exercise. Am. J. Clin. Nutr. 2000, 72, 653s–669s. [Google Scholar] [CrossRef] [PubMed]
- Lenzi, A.; Gandini, L.; Picardo, M.; Tramer, F.; Sandri, G.; Panfili, E. Lipoperoxidation damage of spermatozoa polyunsaturated fatty acids (PUFA): Scavenger mechanisms and possible scavenger therapies. Front. Biosci. 2000, 5, 1–15. [Google Scholar]
- Meister, A.; Anderson, M.E. Glutathione. Annu. Rev. Biochem. 1983, 52, 711–760. [Google Scholar] [CrossRef] [PubMed]
- Bains, J.S.; Shaw, C.A. Neurodegenerative disorders in humans: The role of glutathione in oxidative stress-mediated neuronal death. Brain Res. Rev. 1997, 25, 335–358. [Google Scholar] [CrossRef]
- Thompson, K.H.; Godin, D.V.; Lee, M. Tissue antioxidant status in streptozotocin-induced diabetes in rats. Biol. Trace Elem. Res. 1992, 35, 213–224. [Google Scholar] [CrossRef] [PubMed]
- Palamanda, J.R.; Kehrer, J.P. Involvement of vitamin E and protein thiols in the inhibition of microsomal lipid peroxidation by glutathione. Lipids 1993, 28, 427–431. [Google Scholar] [CrossRef] [PubMed]
- Elliott, S.J.; Koliwad, S.K. Redox control of ion channel activity in vascular endothelial cells by glutathione. Microcirculation 1997, 4, 341–347. [Google Scholar] [CrossRef] [PubMed]
- Brigelius-Flohé, R.; Maiorino, M. Glutathione peroxidases. BBA-Gen. Subj. 2013, 1830, 3289–3303. [Google Scholar] [CrossRef] [PubMed]
- Pedrero, Z.; Madrid, Y. Novel approaches for selenium speciation in foodstuffs and biological specimens: A review. Anal. Chim. Acta 2009, 634, 135–152. [Google Scholar] [CrossRef] [PubMed]
- Winterbourn, C.C.; Hampton, M.B. Thiol chemistry and specificity in redox signaling. Free Radic. Biol. Med. 2008, 45, 549–561. [Google Scholar] [CrossRef] [PubMed]
- Sies, H. Strategies of antioxidant defense. Eur. J. Biochem. 1993, 215, 213–219. [Google Scholar] [CrossRef] [PubMed]
- Buttke, T.M.; Sandstrom, P.A. Oxidative stress as a mediator of apoptosis. Immunol. Today 1994, 15, 7–10. [Google Scholar] [CrossRef]
- Huang, J.-Q.; Ren, F.-Z.; Jiang, Y.-Y.; Xiao, C.; Lei, X.G. Selenoproteins protect against avian nutritional muscular dystrophy by metabolizing peroxides and regulating redox/apoptotic signaling. Free Radic. Biol. Med. 2015, 83, 129–138. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Liu, C.; Liu, C.; Teng, X.; Li, S. Selenium deficiency mainly influences antioxidant selenoproteins expression in broiler immune organs. Biol. Trace Elem. Res. 2016, 172, 209–221. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.-X.; Zhang, C.; Cao, C.-Y.; Zhu, S.-Y.; Li, H.; Sun, Y.-C.; Li, J.-L. Dietary selenium status regulates the transcriptions of selenoproteome and activities of selenoenzymes in chicken kidney at low or super-nutritional levels. Biol. Trace Elem. Res. 2016, 170, 438–448. [Google Scholar] [CrossRef] [PubMed]
- Tang, J.; Huang, X.; Wang, L.; Li, Q.; Xu, J.; Jia, G.; Liu, G.; Chen, X.; Shang, H.; Zhao, H. Supranutritional dietary selenium depressed expression of selenoprotein genes in three immune organs of broilers. Anim. Sci. J. 2017, 88, 331–338. [Google Scholar] [CrossRef] [PubMed]
- Xiang, L.-R.; Li, W.; Wang, L.-L.; Cao, C.-Y.; Li, N.; Li, X.-N.; Jiang, X.-Q.; Li, J.-L. The supranutritional selenium status alters blood glucose and pancreatic redox homeostasis via a modulated selenotranscriptome in chickens (Gallus gallus). RSC Adv. 2017, 7, 24438–24445. [Google Scholar] [CrossRef]
- Khan, A.Z.; Kumbhar, S.; Hamid, M.; Afzal, S.; Parveen, F.; Liu, Y.; Shu, H.; Mengistu, B.M.; Huang, K. Effects of selenium-enriched probiotics on heart lesions by influencing the mRNA expressions of selenoproteins and heat shock proteins in heat stressed broiler chickens. Pak. Vet. J. 2016, 36, 460–464. [Google Scholar]
- Khan, A.Z.; Kumbhar, S.; Liu, Y.; Hamid, M.; Pan, C.; Nido, S.A.; Parveen, F.; Huang, K. Dietary supplementation of selenium-enriched probiotics enhances meat quality of broiler chickens (Gallus gallus domesticus) raised under high ambient temperature. Biol. Trace Elem. Res. 2018, 182, 328–338. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Zhang, Z.; Dong, H.; Jiang, X. Molecular cloning and sequence analysis of selenoprotein W gene and its mRNA expression patterns in response to metabolic status and cadmium exposure in goldfish, carassius auratus. Comp. Biochem. Physiol. B 2015, 184, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Gan, L.; Liu, Q.; Xu, H.-B.; Zhu, Y.-S.; Yang, X.-L. Effects of selenium overexposure on glutathione peroxidase and thioredoxin reductase gene expressions and activities. Biol. Trace Elem. Res. 2002, 89, 165–175. [Google Scholar] [CrossRef]
- El-Sharaky, A.; Newairy, A.; Badreldeen, M.; Eweda, S.; Sheweita, S. Protective role of selenium against renal toxicity induced by cadmium in rats. Toxicology 2007, 235, 185–193. [Google Scholar] [CrossRef] [PubMed]
- Jamba, L.; Nehru, B.; Bansal, M. Effect of selenium supplementation on the influence of cadmium on glutathione and glutathione peroxidase system in mouse liver. J. Trace Elem. Exp. Med. 2000, 13, 299–304. [Google Scholar] [CrossRef]
- Al-Waeli, A.; Zoidis, E.; Pappas, A.; Demiris, N.; Zervas, G.; Fegeros, K. The role of organic selenium in cadmium toxicity: Effects on broiler performance and health status. Animal 2013, 7, 386–393. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Chen, P.; Wan, H.; Wang, Y.; Hao, P.; Liu, Y.; Liu, J. Selenium–chromium (VI) interaction regulates the contents and correlations of trace elements in chicken brain and serum. Biol. Trace Elem. Res. 2018, 181, 154–163. [Google Scholar] [CrossRef] [PubMed]
- Gao, H.; Liu, C.; Song, S.; Fu, J. Effects of dietary selenium against lead toxicity on mRNA levels of 25 selenoprotein genes in the cartilage tissue of broiler chicken. Biol. Trace Elem. Res. 2016, 172, 234–241. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Wang, Y.; An, Y.; Tian, Y.; Li, S.; Teng, X. Selenium for the mitigation of toxicity induced by lead in chicken testes through regulating mRNA expressions of HSPs and selenoproteins. Environ. Sci. Pollut. Res. 2017, 24, 14312–14321. [Google Scholar] [CrossRef] [PubMed]
- Li, X.; Xing, M.; Chen, M.; Zhao, J.; Fan, R.; Zhao, X.; Cao, C.; Yang, J.; Zhang, Z.; Xu, S. Effects of selenium-lead interaction on the gene expression of inflammatory factors and selenoproteins in chicken neutrophils. Ecotoxicol. Environ. Saf. 2017, 139, 447–453. [Google Scholar] [CrossRef] [PubMed]
- Cai, J.; Zhang, Y.; Yang, J.; Liu, Q.; Zhao, R.; Hamid, S.; Wang, H.; Xu, S.; Zhang, Z. Antagonistic effects of selenium against necroptosis injury via adiponectin-necrotic pathway induced by cadmium in heart of chicken. RSC Adv. 2017, 7, 44438–44446. [Google Scholar] [CrossRef]
- Bao, R.; Wang, X.; Zheng, S.; Zhang, Q.; Lin, H.; Li, S. Selenium supplementation changes the ion profile in the pancreas of chickens treated with cadmium. Biol. Trace Elem. Res. 2018, 181, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Berggren, M.M.; Mangin, J.F.; Gasdaska, J.R.; Powis, G. Effect of selenium on rat thioredoxin reductase activity: Increase by supranutritional selenium and decrease by selenium deficiency. Biochem. Pharmacol. 1999, 57, 187–193. [Google Scholar] [CrossRef]
- Rayman, M.P. Selenium in cancer prevention: A review of the evidence and mechanism of action. Proc. Nutr. Soc. 2005, 64, 527–542. [Google Scholar] [CrossRef] [PubMed]
- Pappas, A.; Zoidis, E.; Papadomichelakis, G.; Fegeros, K. Supranutritional selenium level affects fatty acid composition and oxidative stability of chicken breast muscle tissue. J. Anim. Physiol. Anim. Nutr. 2012, 96, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Seo, Y.R.; Kelley, M.R.; Smith, M.L. Selenomethionine regulation of p53 by a ref1-dependent redox mechanism. Proc. Natl. Acad. Sci. USA 2002, 99, 14548–14553. [Google Scholar] [CrossRef] [PubMed]
- Kibriya, M.G.; Jasmine, F.; Argos, M.; Verret, W.J.; Rakibuz-Zaman, M.; Ahmed, A.; Parvez, F.; Ahsan, H. Changes in gene expression profiles in response to selenium supplementation among individuals with arsenic-induced pre-malignant skin lesions. Toxicol. Lett. 2007, 169, 162–176. [Google Scholar] [CrossRef] [PubMed]
- Whanger, P.D. Selenium and the brain: A review. Nutr. Neurosci. 2001, 4, 81–97. [Google Scholar] [CrossRef] [PubMed]
- Birt, D.F.; Julius, A.D.; Runice, C.E.; White, L.T.; Lawson, T.; Pour, P.M. Enhancement of bop-induced pancreatic carcinogenesis in selenium-fed syrian golden hamsters under specific dietary conditions. Nutr. Cancer 1988, 11, 21–33. [Google Scholar] [CrossRef] [PubMed]
- Seo, Y.R.; Sweeney, C.; Smith, M.L. Selenomethionine induction of DNA repair response in human fibroblasts. Oncogene 2002, 21, 3663–3669. [Google Scholar] [CrossRef] [PubMed]
- Christmann, M.; Tomicic, M.T.; Roos, W.P.; Kaina, B. Mechanisms of human DNA repair: An update. Toxicology 2003, 193, 3–34. [Google Scholar] [CrossRef]
- Kennedy, A.R.; Ware, J.H.; Guan, J.; Donahue, J.J.; Biaglow, J.E.; Zhou, Z.; Stewart, J.; Vazquez, M.; Wan, X.S. Selenomethionine protects against adverse biological effects induced by space radiation. Free Radic. Biol. Med. 2004, 36, 259–266. [Google Scholar] [CrossRef] [PubMed]
- Turanov, A.A.; Xu, X.-M.; Carlson, B.A.; Yoo, M.-H.; Gladyshev, V.N.; Hatfield, D.L. Biosynthesis of selenocysteine, the 21st amino acid in the genetic code, and a novel pathway for cysteine biosynthesis. Adv. Nutr. 2011, 2, 122–128. [Google Scholar] [CrossRef] [PubMed]
- Araiso, Y.; Palioura, S.; Ishitani, R.; Sherrer, R.L.; O’Donoghue, P.; Yuan, J.; Oshikane, H.; Domae, N.; DeFranco, J.; Söll, D. Structural insights into RNA-dependent eukaryal and archaeal selenocysteine formation. Nucl. Acids Res. 2007, 36, 1187–1199. [Google Scholar] [CrossRef] [PubMed]
- Fairweather-Tait, S.J.; Collings, R.; Hurst, R. Selenium bioavailability: Current knowledge and future research requirements. Am. J. Clin. Nutr. 2010, 91, 1484S–1491S. [Google Scholar] [CrossRef] [PubMed]
- Short, S.P.; Williams, C.S. Selenoproteins in tumorigenesis and cancer progression. In Advances in Cancer Research; Academic Press Inc.: Cambridge, MA, USA, 2017. [Google Scholar]
- Schomburg, L. Dietary selenium and human health. Nutrients 2017, 9. [Google Scholar] [CrossRef] [PubMed]
- Shchedrina, V.A.; Zhang, Y.; Labunskyy, V.M.; Hatfield, D.L.; Gladyshev, V.N. Structure–function relations, physiological roles, and evolution of mammalian er-resident selenoproteins. Antioxid. Redox Signal. 2010, 12, 839–849. [Google Scholar] [CrossRef] [PubMed]
- Labunskyy, V.M.; Yoo, M.-H.; Hatfield, D.L.; Gladyshev, V.N. Sep15, a thioredoxin-like selenoprotein, is involved in the unfolded protein response and differentially regulated by adaptive and acute ER stresses. Biochemistry 2009, 48, 8458–8465. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, A.D.; Labunskyy, V.M.; Fomenko, D.E.; Araç, D.; Chelliah, Y.; Amezcua, C.A.; Rizo, J.; Gladyshev, V.N.; Deisenhofer, J. NMR structures of the selenoproteins Sep15 and SelM reveal redox activity of a new thioredoxin-like family. J. Biol. Chem. 2006, 281, 3536–3543. [Google Scholar] [CrossRef] [PubMed]
- Metanis, N.; Hilvert, D. Natural and synthetic selenoproteins. Curr. Opin. Chem. Biol. 2014, 22, 27–34. [Google Scholar] [CrossRef] [PubMed]
- Gromer, S.; Johansson, L.; Bauer, H.; Arscott, L.D.; Rauch, S.; Ballou, D.P.; Williams, C.H.; Schirmer, R.H.; Arnér, E.S. Active sites of thioredoxin reductases: Why selenoproteins? Proc. Natl. Acad. Sci. USA 2003, 100, 12618–12623. [Google Scholar] [CrossRef] [PubMed]
- Ishizaki, A. Onitai-itai (ouch-ouch) disease. Asian Med. J. 1971, 14, 421–436. [Google Scholar]
- Bianco, A.C.; Larsen, P.R. Cellular and structural biology of the deiodinases. Thyroid 2005, 15, 777–786. [Google Scholar] [CrossRef] [PubMed]
- Jurynec, M.J.; Xia, R.; Mackrill, J.J.; Gunther, D.; Crawford, T.; Flanigan, K.M.; Abramson, J.J.; Howard, M.T.; Grunwald, D.J. Selenoprotein N is required for ryanodine receptor calcium release channel activity in human and zebrafish muscle. Proc. Natl. Acad. Sci. USA 2008, 105, 12485–12490. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, B.R.; Roberts, B.R.; Bush, A.I.; Hare, D.J. Selenium, selenoproteins and neurodegenerative diseases. Metallomics 2015, 7, 1213–1228. [Google Scholar] [CrossRef] [PubMed]
- Habibian, M.; Sadeghi, G.; Ghazi, S.; Moeini, M.M. Selenium as a feed supplement for heat-stressed poultry: A review. Biol. Trace Elem. Res. 2015, 165, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Dalgaard, T.S.; Briens, M.; Engberg, R.M.; Lauridsen, C. The influence of selenium and selenoproteins on immune responses of poultry and pigs. Anim. Feed Sci. Technol. 2018, 238, 73–83. [Google Scholar] [CrossRef]
- Zhu, S.-Y.; Li, X.-N.; Sun, X.-C.; Lin, J.; Li, W.; Zhang, C.; Li, J.-L. Biochemical characterization of the selenoproteome in gallus gallus via bioinformatics analysis: Structure–function relationships and interactions of binding molecules. Metallomics 2017, 9, 124–131. [Google Scholar] [CrossRef] [PubMed]
- Reeg, S.; Grune, T. Protein oxidation in aging: Does it play a role in aging progression? Antioxid. Redox Signal. 2015, 23, 239–255. [Google Scholar] [CrossRef] [PubMed]
- Khoso, P.A.; Yang, Z.; Liu, C.; Li, S. Selenoproteins and heat shock proteins play important roles in immunosuppression in the bursa of fabricius of chickens with selenium deficiency. Cell Stress Chaperones 2015, 20, 967–978. [Google Scholar] [CrossRef] [PubMed]
- Du, Q.; Yao, H.; Yao, L.; Zhang, Z.; Lei, X.; Xu, S. Selenium deficiency influences the expression of selenoproteins and inflammatory cytokines in chicken aorta vessels. Biol. Trace Elem. Res. 2016, 173, 501–513. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Huang, J.; Zhang, H.; Lei, X.; Du, Z.; Xiao, C.; Chen, S.; Ren, F. Selenium deficiency-induced apoptosis of chick embryonic vascular smooth muscle cells and correlations with 25 selenoproteins. Biol. Trace Elem. Res. 2017, 176, 407–415. [Google Scholar] [CrossRef] [PubMed]
- Gao, Y.; Zhang, J.; Huang, X.; Zhang, G. Glutathione peroxidase 1, selenoprotein K, and selenoprotein H may play important roles in chicken testes in response to selenium deficiency. Biol. Trace Elem. Res. 2017, 179, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.; Sun, B.; Zhang, J.; Gao, Y.; Li, G.; Chang, Y. Selenium deficiency induced injury in chicken muscular stomach by downregulating selenoproteins. Biol. Trace Elem. Res. 2017, 179, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Sun, Z.; Liu, C.; Pan, T.; Yao, H.; Li, S. Selenium accelerates chicken dendritic cells differentiation and affects selenoproteins expression. Dev. Comp. Immunol. 2017, 77, 30–37. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; An, Y.; Jiao, W.; Zhang, Z.; Han, H.; Gu, X.; Teng, X. Selenium protects against lead-induced apoptosis via endoplasmic reticulum stress in chicken kidneys. Biol. Trace Elem. Res. 2017, 182, 354–363. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.-Q.; Ren, F.-Z.; Jiang, Y.-Y.; Lei, X. Characterization of selenoprotein M and its response to selenium deficiency in chicken brain. Biol. Trace Elem. Res. 2016, 170, 449–458. [Google Scholar] [CrossRef] [PubMed]
- Zoidis, E.; Pappas, A.; Georgiou, C.; Komaitis, E.; Feggeros, K. Selenium affects the expression of GPx4 and catalase in the liver of chicken. Comp. Biochem. Physiol. B 2010, 155, 294–300. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Z.-H.; Lin, H.-J.; Yao, H.-D.; Zhang, Z.-W.; Fu, J.; Xu, S.-W. Selw protects against H2O2-induced liver injury in chickens via inhibiting inflammation and apoptosis. RSC Adv. 2017, 7, 15158–15167. [Google Scholar] [CrossRef]
- Fan, R.; Yao, H.; Zhao, X.; Cao, C.; Yang, T.; Luan, Y.; Zhang, Z.; Xu, S. Gene expression of selenoproteins can be regulated by selenoprotein k silencing in chicken myoblasts. BioMetals 2016, 29, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Jiang, Y.; Guo, H.; Zhang, H.; Lei, X.; Ren, F. Association Analysis between Selenoprotein Genes (Selw and Seln) and Meat Quality Traits in Chicken. Available online: https://www.researchgate.net/publication/312119322_Association_analysis_between_selenoprotein_genes_Selw_and_Seln_and_meat_quality_traits_in_chicken (accessed on 23 March 2018).
- Luan, Y.; Zhao, J.; Yao, H.; Zhao, X.; Fan, R.; Zhao, W.; Zhang, Z.; Xu, S. Selenium deficiency influences the mrna expression of selenoproteins and cytokines in chicken erythrocytes. Biol. Trace Elem. Res. 2016, 171, 427–436. [Google Scholar] [CrossRef] [PubMed]
- Khoso, P.A.; Yang, Z.; Liu, C.; Li, S. Selenium deficiency downregulates selenoproteins and suppresses immune function in chicken thymus. Biol. Trace Elem. Res. 2015, 167, 48–55. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.; Zhang, H.; Wang, S.; Ruan, D.; Xie, X.; Yu, D.; Lin, Y. Estimation of dietary selenium requirement for chinese egg-laying ducks. Anim. Prod. Sci. 2015, 55, 1056–1063. [Google Scholar] [CrossRef]
- Li, J.-L.; Sunde, R.A. Selenoprotein transcript level and enzyme activity as biomarkers for selenium status and selenium requirements of chickens (Gallus gallus). PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.M.; Sunde, R.A. Selenoprotein transcript level and enzyme activity as biomarkers for selenium status and selenium requirements in the turkey (Meleagris gallopavo). PLoS ONE 2016, 11. [Google Scholar] [CrossRef] [PubMed]
- Dalia, A.; Loh, T.; Sazili, A.; Jahromi, M.; Samsudin, A. The effect of dietary bacterial organic selenium on growth performance, antioxidant capacity, and selenoproteins gene expression in broiler chickens. BMC Vet. Res. 2017, 13. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Sun, L.-H.; Huang, J.-Q.; Briens, M.; Qi, D.-S.; Xu, S.-W.; Lei, X.G. A novel organic selenium compound exerts unique regulation of selenium speciation, selenogenome, and selenoproteins in broiler chicks. J. Nutr. 2017, 147, 789–797. [Google Scholar] [CrossRef] [PubMed]
- Sunde, R.A.; Raines, A.M. Selenium regulation of the selenoprotein and nonselenoprotein transcriptomes in rodents. Adv. Nutr. 2011, 2, 138–150. [Google Scholar] [CrossRef] [PubMed]
- Behne, D.; Kyriakopoulos, A. Mammalian selenium-containing proteins. Annu. Rev. Nutr. 2001, 21, 453–473. [Google Scholar] [CrossRef] [PubMed]
- Hatfield, D.L.; Gladyshev, V.N. How selenium has altered our understanding of the genetic code. Mol. Cell. Biol. 2002, 22, 3565–3576. [Google Scholar] [CrossRef] [PubMed]
- O’Driscoll, M.; Ruiz-Perez, V.L.; Woods, C.G.; Jeggo, P.A.; Goodship, J.A. A splicing mutation affecting expression of ataxia–telangiectasia and Rad3–related protein (ATR) results in Seckel syndrome. Nat. Genet. 2003, 33, 497–501. [Google Scholar] [CrossRef] [PubMed]
- Schomburg, I.; Chang, A.; Ebeling, C.; Gremse, M.; Heldt, C.; Huhn, G.; Schomburg, D. Brenda, the enzyme database: Updates and major new developments. Nucl. Acids Res. 2004, 32, D431–D433. [Google Scholar] [CrossRef] [PubMed]
- Brigelius-Flohe, R.; Traber, M.G. Vitamin e: Function and metabolism. FASEB J. 1999, 13, 1145–1155. [Google Scholar] [CrossRef] [PubMed]
- Moriarty, P.M.; Reddy, C.C.; Maquat, L.E. Selenium deficiency reduces the abundance of mRNA for se-dependent glutathione peroxidase 1 by a UGA-dependent mechanism likely to be nonsense codon-mediated decay of cytoplasmic mRNA. Mol. Cell. Biol. 1998, 18, 2932–2939. [Google Scholar] [CrossRef] [PubMed]
- Maquat, L.E. Evidence that selenium deficiency results in the cytoplasmic decay of GPx1 mRNA dependent on pre-mRNA splicing proteins bound to the mRNA exon-exon junction. Biofactors 2001, 14, 37–42. [Google Scholar] [CrossRef] [PubMed]
- Sachdev, S.W.; Sunde, R.A. Selenium regulation of transcript abundance and translational efficiency of glutathione peroxidase-1 and -4 in rat liver. Biochem. J. 2001, 357, 851–858. [Google Scholar]
- Jiang, X.-Q.; Cao, C.-Y.; Li, Z.-Y.; Li, W.; Zhang, C.; Lin, J.; Li, X.-N.; Li, J.-L. Delineating hierarchy of selenotranscriptome expression and their response to selenium status in chicken central nervous system. J. Inorg. Biochem. 2017, 169, 13–22. [Google Scholar] [CrossRef] [PubMed]
- Jiang, Y.-Y.; Huang, J.-Q.; Lin, G.-C.; Guo, H.-Y.; Ren, F.-Z.; Zhang, H. Characterization and expression of chicken selenoprotein U. Biol. Trace Elem. Res. 2015, 166, 216–224. [Google Scholar] [CrossRef] [PubMed]
- Sunde, R.A.; Sunde, G.R.; Sunde, C.M.; Sunde, M.L.; Evenson, J.K. Cloning, sequencing, and expression of selenoprotein transcripts in the turkey (Meleagris gallopavo). PLoS ONE 2015, 10. [Google Scholar] [CrossRef] [PubMed]
- Kumaraswamy, E.; Malykh, A.; Korotkov, K.V.; Kozyavkin, S.; Hu, Y.; Kwon, S.Y.; Moustafa, M.E.; Carlson, B.A.; Berry, M.J.; Lee, B.J. Structure-expression relationships of the 15-kDa selenoprotein gene possible role of the protein in cancer etiology. J. Biol. Chem. 2000, 275, 35540–35547. [Google Scholar] [CrossRef] [PubMed]
- Yoboue, E.D.; Rimessi, A.; Anelli, T.; Pinton, P.; Sitia, R. Regulation of calcium fluxes by GPX8, a type-II transmembrane peroxidase enriched at the mitochondria-associated endoplasmic reticulum membrane. Antioxid. Redox Signal. 2017, 27, 583–595. [Google Scholar] [CrossRef] [PubMed]
- Marino, M.; Stoilova, T.; Giorgi, C.; Bachi, A.; Cattaneo, A.; Auricchio, A.; Pinton, P.; Zito, E. SEPN1, an endoplasmic reticulum-localized selenoprotein linked to skeletal muscle pathology, counteracts hyperoxidation by means of redox-regulating SERCA2 pump activity. Hum. Mol. Genet. 2015, 24, 1843–1855. [Google Scholar] [CrossRef] [PubMed]
- Arbogast, S.; Beuvin, M.; Fraysse, B.; Zhou, H.; Muntoni, F.; Ferreiro, A. Oxidative stress in SEPN1-related myopathy: From pathophysiology to treatment. Ann. Neurol. 2009, 65, 677–686. [Google Scholar] [CrossRef] [PubMed]
- Zalk, R.; Marks, A.R. Ca2+ Release Channels Join the ‘Resolution Revolution’. Trends Biochem. Sci. 2017, 42, 543–555. [Google Scholar] [CrossRef] [PubMed]
- Program, N.T.; Program, N.T. Report on carcinogens. In Public Health Service, National Toxicology Program; U.S. Department of Health and Human Services: Research Triangle Park, NC, USA, 2011. [Google Scholar]
- Castets, P.; Bertrand, A.T.; Beuvin, M.; Ferry, A.; Le Grand, F.; Castets, M.; Chazot, G.; Rederstorff, M.; Krol, A.; Lescure, A.; et al. Satellite cell loss and impaired muscle regeneration in selenoprotein n deficiency. Hum. Mol. Genet. 2011, 20, 694–704. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Li, C.; Gu, G.; Wang, Q.; Guo, M. Selenoprotein n was required for the regulation of selenium on the uterine smooth muscle contraction in mice. Biol. Trace Elem. Res. 2017, 183, 138–146. [Google Scholar] [CrossRef] [PubMed]
- Grumolato, L.; Ghzili, H.; Montero-Hadjadje, M.; Gasman, S.; Lesage, J.; Tanguy, Y.; Galas, L.; Ait-Ali, D.; Leprince, J.; Guérineau, N.C. Selenoprotein T is a PACAP-regulated gene involved in intracellular Ca2+ mobilization and neuroendocrine secretion. FASEB J. 2008, 22, 1756–1768. [Google Scholar] [CrossRef] [PubMed]
- Boukhzar, L.; Hamieh, A.; Cartier, D.; Tanguy, Y.; Alsharif, I.; Castex, M.; Arabo, A.; El Hajji, S.; Bonnet, J.J.; Errami, M.; et al. Selenoprotein t exerts an essential oxidoreductase activity that protects dopaminergic neurons in mouse models of parkinson’s disease. Antioxid. Redox Signal. 2016, 24, 557–574. [Google Scholar] [CrossRef] [PubMed]
- Gladyshev, V.N.; Factor, V.M.; Housseau, F.; Hatfield, D.L. Contrasting patterns of regulation of the antioxidant selenoproteins, thioredoxin reductase, and glutathione peroxidase, in cancer cells. Biochem. Biophys. Res. Commun. 1998, 251, 488–493. [Google Scholar] [CrossRef] [PubMed]
- Korotkov, K.V.; Kumaraswamy, E.; Zhou, Y.; Hatfield, D.L.; Gladyshev, V.N. Association between the 15-kDa selenoprotein and UDP-glucose: Glycoprotein glucosyltransferase in the endoplasmic reticulum of mammalian cells. J. Biol. Chem. 2001, 276, 15330–15336. [Google Scholar] [CrossRef] [PubMed]
- Guariniello, S.; Colonna, G.; Raucci, R.; Costantini, M.; Di Bernardo, G.; Bergantino, F.; Castello, G.; Costantini, S. Structure–function relationship and evolutionary history of the human selenoprotein M (SelM) found over-expressed in hepatocellular carcinoma. Biochim. Biophys. Acta Proteins Proteom. 2014, 1844, 447–456. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Lee, S.-G.; Choo, M.-K.; Kim, J.H.; Lee, H.M.; Kim, S.; Fomenko, D.E.; Kim, H.-Y.; Park, J.M.; Gladyshev, V.N. Selenoprotein MsrB1 promotes anti-inflammatory cytokine gene expression in macrophages and controls immune response in vivo. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-Y.; Gladyshev, V.N. Different catalytic mechanisms in mammalian selenocysteine-and cysteine-containing methionine-R-sulfoxide reductases. PLoS Biol. 2005, 3. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Kaya, A.; Weerapana, E.; Marino, S.M.; Gladyshev, V.N. Methionine sulfoxide reductases preferentially reduce unfolded oxidized proteins and protect cells from oxidative protein unfolding. J. Biol. Chem. 2012, 287, 24448–24459. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Péterfi, Z.; Hoffmann, F.W.; Moore, R.E.; Kaya, A.; Avanesov, A.; Tarrago, L.; Zhou, Y.; Weerapana, E.; Fomenko, D.E. MsrB1 and MICALs regulate actin assembly and macrophage function via reversible stereoselective methionine oxidation. Mol. Cell 2013, 51, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Mehmeti, I.; Lortz, S.; Avezov, E.; Jörns, A.; Lenzen, S. ER-resident antioxidative GPx7 and GPx8 enzyme isoforms protect insulin-secreting INS-1E β-cells against lipotoxicity by improving the ER antioxidative capacity. Free Radic. Biol. Med. 2017, 112, 121–130. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Cheng, W.; Nie, T.; Lai, H.; Hu, X.; Luo, J.; Li, F.; Li, H. Selenoprotein K mediates the proliferation, migration, and invasion of human choriocarcinoma cells by negatively regulating human chorionic gonadotropin expression via ERK, p38 MAPK, and Akt signaling pathway. Biol. Trace Elem. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Rozovsky, S.; Liu, J.; Zhang, Z. The intrinsically disordered membrane enzymes selenoprotein S and selenoprotein K. FASEB J. 2017, 31. [Google Scholar] [CrossRef]
- Gao, Y.; Feng, H.C.; Walder, K.; Bolton, K.; Sunderland, T.; Bishara, N.; Quick, M.; Kantham, L.; Collier, G.R. Regulation of the selenoprotein SelS by glucose deprivation and endoplasmic reticulum stress—SelS is a novel glucose-regulated protein. FEBS Lett. 2004, 563, 185–190. [Google Scholar] [CrossRef]
- Curran, J.E.; Jowett, J.B.; Elliott, K.S.; Gao, Y.; Gluschenko, K.; Wang, J.; Azim, D.M.A.; Cai, G.; Mahaney, M.C.; Comuzzie, A.G. Genetic variation in selenoprotein S influences inflammatory response. Nat. Genet. 2005, 37, 1234–1241. [Google Scholar] [CrossRef] [PubMed]
- Chabory, E.; Damon, C.; Lenoir, A.; Henry-Berger, J.; Vernet, P.; Cadet, R.; Saez, F.; Drevet, J. Mammalian glutathione peroxidases control acquisition and maintenance of spermatozoa integrity. J. Anim. Sci. 2010, 88, 1321–1331. [Google Scholar] [CrossRef] [PubMed]
- Li, M.Z.; Elledge, S.J. Slic: A method for sequence-and ligation-independent cloning. Methods Mol. Biol. 2012, 852, 51–59. [Google Scholar] [PubMed]
- Shema, R.; Kulicke, R.; Cowley, G.S.; Stein, R.; Root, D.E.; Heiman, M. Synthetic lethal screening in the mammalian central nervous system identifies Gpx6 as a modulator of huntington’s disease. Proc. Natl. Acad. Sci. USA 2015, 112, 268–272. [Google Scholar] [CrossRef] [PubMed]
- Ursini, F. The world of glutathione peroxidases. J. Trace Elem. Med. Biol. 2000, 14, 116. [Google Scholar]
- Ingold, I.; Berndt, C.; Schmitt, S.; Doll, S.; Poschmann, G.; Buday, K.; Roveri, A.; Peng, X.; Freitas, F.P.; Seibt, T. Selenium utilization by GPX4 is required to prevent hydroperoxide-induced ferroptosis. Cell 2017, 21, 409–422. [Google Scholar] [CrossRef] [PubMed]
- Jiao, Y.; Wang, Y.; Guo, S.; Wang, G. Glutathione peroxidases as oncotargets. Oncotarget 2017, 8, 80093–80102. [Google Scholar] [CrossRef] [PubMed]
- Delaunay-Moisan, A.; Ponsero, A.; Toledano, M.B. Reexamining the function of glutathione in oxidative protein folding and secretion. Antioxid. Redox Signal. 2017, 27, 1178–1199. [Google Scholar] [CrossRef] [PubMed]
- Flohé, L.; Brigelius-Flohé, R. Basics and News on Glutathione Peroxidases. In Selenium; Springer Nature: New York, NY, USA, 2016; pp. 211–222. [Google Scholar]
- Utomo, A.; Jiang, X.; Furuta, S.; Yun, J.; Levin, D.S.; Wang, Y.-C.J.; Desai, K.V.; Green, J.E.; Chen, P.-L.; Lee, W.-H. Identification of a novel putative non-selenocysteine containing phospholipid hydroperoxide glutathione peroxidase (NPGPx) essential for alleviating oxidative stress generated from polyunsaturated fatty acids in breast cancer cells. J. Biol. Chem. 2004, 279, 43522–43529. [Google Scholar] [CrossRef] [PubMed]
- Toppo, S.; Vanin, S.; Bosello, V.; Tosatto, S.C. Evolutionary and structural insights into the multifaceted glutathione peroxidase (Gpx) superfamily. Antioxid. Redox Signal. 2008, 10, 1501–1514. [Google Scholar] [CrossRef] [PubMed]
- Bianco, A.C.; Larsen, P.R. Selenium, Deiodinases and Endocrine Function. Available online: https://link.springer.com/chapter/10.1007/0-387-33827-6_19 (accessed on 23 March 2018).
- Köhrle, J.; Gärtner, R. Selenium and thyroid. Best Pract. Res. Clin. Endocrinol. Metab. 2009, 23, 815–827. [Google Scholar] [CrossRef] [PubMed]
- Beckett, G.J.; Arthur, J.R. Selenium and endocrine systems. J. Endocrinol. 2005, 184, 455–465. [Google Scholar] [CrossRef] [PubMed]
- Panee, J.; Stoytcheva, Z.R.; Liu, W.; Berry, M.J. Selenoprotein h is a redox-sensing high mobility group family DNA-binding protein that up-regulates genes involved in glutathione synthesis and phase II detoxification. J. Biol. Chem. 2007, 282, 23759–23765. [Google Scholar] [CrossRef] [PubMed]
- Mendelev, N.; Mehta, S.L.; Witherspoon, S.; He, Q.; Sexton, J.Z.; Li, P.A. Upregulation of human selenoprotein H in murine hippocampal neuronal cells promotes mitochondrial biogenesis and functional performance. Mitochondrion 2011, 11, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.Y.; Al-Khayat, A.; Al-Murshedi, F.; Al-Futaisi, A.; Chioza, B.A.; Pedro Fernandez-Murray, J.; Self, J.E.; Salter, C.G.; Harlalka, G.V.; Rawlins, L.E. A mutation of EPT1 (SELENOI) underlies a new disorder of kennedy pathway phospholipid biosynthesis. Brain 2017, 140, 547–554. [Google Scholar] [PubMed]
- Nogly, P.; Gushchin, I.; Remeeva, A.; Esteves, A.M.; Borges, N.; Ma, P.; Ishchenko, A.; Grudinin, S.; Round, E.; Moraes, I. X-ray structure of a CDP-alcohol phosphatidyltransferase membrane enzyme and insights into its catalytic mechanism. Nat. Commun. 2014, 5. [Google Scholar] [CrossRef] [PubMed]
- Han, S.-J.; Lee, B.C.; Yim, S.H.; Gladyshev, V.N.; Lee, S.-R. Characterization of mammalian selenoprotein o: A redox-active mitochondrial protein. PLoS ONE 2014, 9. [Google Scholar] [CrossRef] [PubMed]
- Gladyshev, M.I.; Sushchik, N.N.; Gubanenko, G.A.; Demirchieva, S.M.; Kalachova, G.S. Effect of way of cooking on content of essential polyunsaturated fatty acids in muscle tissue of humpback salmon (Oncorhynchus gorbuscha). Food Chem. 2006, 96, 446–451. [Google Scholar] [CrossRef]
- Burk, R.F.; Hill, K.E. Selenoprotein P: An extracellular protein with unique physical characteristics and a role in selenium homeostasis. Annu. Rev. Nutr. 2005, 25, 215–235. [Google Scholar] [CrossRef] [PubMed]
- Rayman, M.P. Selenoproteins and human health: Insights from epidemiological data. BBA-Gen. Subj. 2009, 1790, 1533–1540. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Hill, K.E.; Byrne, D.W.; Xu, J.; Burk, R.F. Effectiveness of selenium supplements in a low-selenium area of china. Am. J. Clin. Nutr. 2005, 81, 829–834. [Google Scholar] [CrossRef] [PubMed]
- Yao, H.-D.; Wu, Q.; Zhang, Z.-W.; Li, S.; Wang, X.-L.; Lei, X.-G.; Xu, S.-W. Selenoprotein W serves as an antioxidant in chicken myoblasts. BBA-Gen. Subj. 2013, 1830, 3112–3120. [Google Scholar] [CrossRef] [PubMed]
- Whanger, P. Selenoprotein W: A review. Cell. Mol. Life Sci. 2000, 57, 1846–1852. [Google Scholar] [CrossRef] [PubMed]
- McAllister, K.N.; Bouillaut, L.; Kahn, J.N.; Self, W.T.; Sorg, J.A. Using CRISPR-Cas9-mediated genome editing to generate C. difficile mutants defective in selenoproteins synthesis. Sci. Rep. 2017, 7. [Google Scholar] [CrossRef] [PubMed]
| Radicals | Non-Radicals |
|---|---|
| RO•, Alkoxyl | H2O2, Hydrogen peroxide |
| HOO•, Hydroperoxyl | HOCl, Hypochlorous acid |
| HO•, Hydroxyl | O3, Ozone |
| ROO•, Peroxyl | 1O2, Singlet oxygen |
| O2•¯, Superoxide | ONOO¯, Peroxynitrite |
| NO•, Nitric oxide | NO¯, Nitroxyl anion |
| NO2•, Nitrogen dioxide | HNO2, Nitrous acid |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zoidis, E.; Seremelis, I.; Kontopoulos, N.; Danezis, G.P. Selenium-Dependent Antioxidant Enzymes: Actions and Properties of Selenoproteins. Antioxidants 2018, 7, 66. https://doi.org/10.3390/antiox7050066
Zoidis E, Seremelis I, Kontopoulos N, Danezis GP. Selenium-Dependent Antioxidant Enzymes: Actions and Properties of Selenoproteins. Antioxidants. 2018; 7(5):66. https://doi.org/10.3390/antiox7050066
Chicago/Turabian StyleZoidis, Evangelos, Isidoros Seremelis, Nikolaos Kontopoulos, and Georgios P. Danezis. 2018. "Selenium-Dependent Antioxidant Enzymes: Actions and Properties of Selenoproteins" Antioxidants 7, no. 5: 66. https://doi.org/10.3390/antiox7050066
APA StyleZoidis, E., Seremelis, I., Kontopoulos, N., & Danezis, G. P. (2018). Selenium-Dependent Antioxidant Enzymes: Actions and Properties of Selenoproteins. Antioxidants, 7(5), 66. https://doi.org/10.3390/antiox7050066