Abstract
Cardiovascular diseases hinge on a vicious, self-amplifying cycle in which mitochondrial deoxyribonucleic acid (mtDNA) dysfunction undermines cardiac bioenergetics and unleashes sterile inflammation. The heart’s reliance on oxidative phosphorylation (OXPHOS) makes it exquisitely sensitive to mtDNA insults—mutations, oxidative lesions, copy-number shifts, or aberrant methylation—that impair ATP production, elevate reactive oxygen species (ROS), and further damage the mitochondrial genome. Damaged mtDNA fragments then escape into the cytosol, where they aberrantly engage cGAS–STING, TLR9, and NLRP3 pathways, driving cytokine storms, pyroptosis, and tissue injury. We propose that this cycle represents an almost unifying pathogenic mechanism in a spectrum of mtDNA-driven cardiovascular disorders. In this review, we aim to synthesize the pathophysiological roles of mtDNA in this cycle and its implications for cardiovascular diseases. Furthermore, we seek to evaluate preclinical and clinical strategies aimed at interrupting this cycle—bolstering mtDNA repair and copy-number maintenance, reversing pathogenic methylation, and blocking mtDNA-triggered innate immune activation—and discuss critical gaps that must be bridged to translate these approaches into precision mitochondrial genome medicine for cardiovascular disease.
1. Introduction
Mitochondria are highly specialized, double-membrane organelles with cell- and tissue-specific morphology, dynamics, and functions. They play a central role in metabolism and signaling pathways, including energy generation, calcium homeostasis, ROS production, heme synthesis, steroidogenesis, programmed cell death, and metabolite catalysis [1,2,3]. Each mitochondrion contains multiple copies of circular mitochondrial DNA (mtDNA), organized within nucleoids in the inner mitochondrial matrix, and characterized by its own transcriptional machinery [2,4,5]. Animal tissue cells contain thousands of mtDNA copies spread out over hundreds of mitochondria, encoding 13 polypeptides for the respiratory chain complexes, 22 transfer ribonucleic acids (tRNAs), and 2 ribosomal RNAs (rRNAs), as well as enzymes needed for their transcription and translation [2,3,5,6,7]. Notably, mtDNA copies often exceed what is strictly necessary for oxidative phosphorylation, suggesting additional roles in mitochondrial signaling or immune regulation [2]. Like nuclear DNA (nDNA), mtDNA is vulnerable to mutations and exogenous or endogenous damage, which can drive mitochondrial dysfunction and human pathologies [4,8,9,10,11].
Given the heart’s absolute dependence on OXPHOS, mtDNA damage naturally manifests in cardiac syndromes—either as primary mitochondrial disease or as part of multisystem disorders. Clinically, the vicious cycle of bioenergetic failure and sterile inflammation manifests across cardiovascular diseases. In hypertension [12,13,14,15] and atherosclerosis [16,17], mtDNA mutations have been widely reported, and oxidized mtDNA accelerates vascular senescence and exacerbates plaque inflammation. In myocardial ischemia, damaged mtDNA is released and triggers severe inflammatory responses, exacerbating tissue damage [18,19,20,21,22,23,24,25,26]. In heart failure, mtDNA lesions and depletion are involved in critical ATP deficits and worsen contractile dysfunction [27,28]. In diabetic cardiomyopathy, fatty acid-driven mtDNA release provokes cardiomyocyte pyroptosis via the cGAS–STING pathway [29,30]. Arrhythmias, too, are associated with oxidative lesions and mutations in mtDNA [31,32]. Thus, mtDNA has emerged as a critical target for mechanistic understanding and therapeutic innovation in cardiovascular medicine. Elucidating the molecular pathways linking mtDNA integrity to disease progression may open new avenues for biomarker development and precise interventions.
In this review, we synthesize the physiological and pathological features of mtDNA, propose a unified “vicious cycle” theory linking mtDNA dysfunction to cardiac bioenergetic collapse and sterile inflammation, and highlight emerging therapeutic strategies and remaining challenges. Our goal is to inform the development of systematic, mtDNA-targeted interventions that mitigate myocardial injury, prevent heart failure, and reduce mortality.
2. Overview of mtDNA
The origin of mitochondria as an ancient archaeal endosymbiont endows the organelle with its own double-stranded, circular genome [33]. This autonomous genome retains the machinery needed to replicate, transcribe, and translate its genes, thereby allowing mitochondria to self-regulate critical functions such as energy production, signaling, and apoptosis (Figure 1).
Figure 1.
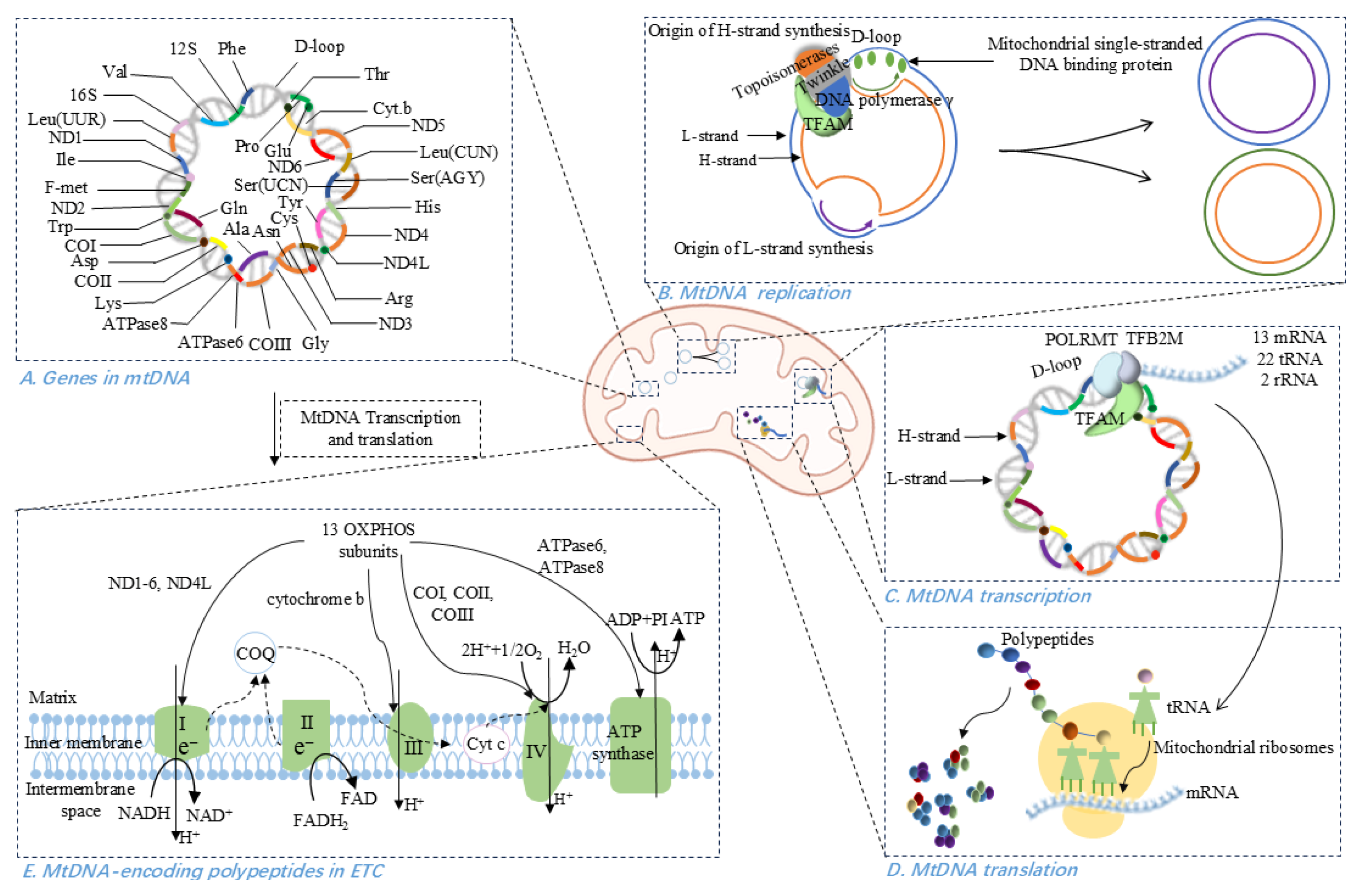
Physiological properties of mitochondrial DNA (mtDNA). (A) Gene distribution in mtDNA, with 28 genes on the heavy (H) strand and 9 on the light (L) strand. (B) MtDNA replication occurs in the ‘replisome’ comprising DNA polymerase γ, mitochondrial single-stranded DNA binding protein, Twinkle helicase, topoisomerases, and TFAM. (C) Transcription of mtDNA initiated at the D-loop region is regulated by POLRMT, TFAM, and TFB2M, producing 13 mRNAs, 22 tRNAs, and 2 rRNAs. (D) MtDNA-directed translation involves initiation complex assembly, mRNA binding to the ribosomal subunit, tRNA-mediated peptide elongation, translation termination, and ribosome recycling. (E) MtDNA-encoded polypeptides contribute to the function of ETC, specifically within complexes I, III, IV, and V.
2.1. Structure of mtDNA
MtDNA exists as a closed, circular duplex with a purine-rich heavy (H) strand and a pyrimidine-rich light (L) strand [3,4]. It contains 37 genes-28 on the H-strand and 9 on the L-strand-that are directly or indirectly involved in ATP synthesis via oxidative phosphorylation [3,5,34]. Unlike the nDNA counterpart, mtDNA is extraordinarily gene-dense (~93% coding), with overlapping reading frames and only one major non-coding region, the displacement loop (D-loop), which harbors the origin of replication and promoters for transcription [3].
Rather than being wrapped around histones, mtDNA is packaged into 100-nm nucleoids through binding to the nuclear-encoded mitochondrial transcription factor A (TFAM) [4,35]. TFAM is imported into mitochondria via an N-terminal mitochondrial targeting peptide that is cleaved upon entry to form mature TFAM. In the mitochondrial matrix, TFAM coats mtDNA to stabilize its structure and binds the D-loop to coordinate the bidirectional transcription from the light-strand promoter (LSP) and two heavy-strand promoters (HSP1, HSP2) [4,36,37].
2.2. MtDNA Replication, Transcription, and Translation
MtDNA replication occurs continuously, independent of the cell-division cycle, and is entirely governed by nuclear-encoded factors [3]. In eukaryotes, mtDNA replication takes place within a “replisome” complex whose catalytic core is DNA polymerase γ (POLγ), accompanied by two accessory subunits [3,5]. This machinery also includes the mitochondrial single-stranded DNA-binding protein (mtSSB), which stabilizes unwound DNA and enhances POLγ activity; the Twinkle helicase/primase; various topoisomerases; and TFAM, which may protect mtDNA from oxidative damage [3]. Mammalian mtDNA replicates mainly by a strand-displacement mechanism: synthesis of the heavy (H) strand initiates at the D-loop origin and proceeds unidirectionally until exposing the light (L)-strand origin, at which point L-strand synthesis begins in the opposite direction. Both strands elongate asynchronously until two daughter molecules are complete [38,39].
Transcription of mtDNA is driven by three promoters: the light-strand promoter (LSP) and two heavy-strand promoters (HSP1, HSP2) [37]. Nuclear-encoded mitochondrial RNA polymerase (POLRMT) forms a pre-initiation complex with TFAM and TFB2M at the D-loop, melting DNA to expose each promoter [40,41]. From HSP2 and LSP, genome-length polycistronic transcripts are produced; the H-strand is additionally transcribed from promoter HSP1 to generate a shorter transcript consisting of two rRNAs and two tRNAs [42]. Transcript elongation is carried out by POLRMT, with assistance from the mitochondrial transcription elongation factor, while termination is regulated by mitochondrial transcription termination factor 1 (MTERF1) [3].
The mitochondrial ribosomes utilize a unique set of translation initiation, elongation, and termination factors, culminating in a translation cycle completely different from the bacterial and cytosolic ones [43,44]. Initiation factor mtIF-3 dissociates ribosomal subunits to allow assembly of the initiation complex, aligning the start codon of mRNA on the small subunit’s P-site [3,45]. Peptide elongation is mediated by nuclear-encoded mitochondrial elongation factor Tu (tRNA delivery) and mitochondrial elongation factor G (mRNA–tRNA translocation and ribosome recycling) [3,46,47,48,49,50]. Termination relies on mitochondrial release factor 1 (mtRF1), which recognizes stop codons and catalyzes peptide release [3,45].
2.3. MtDNA in OXPHOS System
The mitochondrial OXPHOS system serves the fundamental purpose of ATP transduction by oxidizing fuel molecules via the respiratory electron transport chain (ETC) [7]. The mammalian ETC comprises five multimeric complexes (Complexes I–V) embedded in the inner mitochondrial membrane [51]. These complexes harness energy released from electron transfer from NADH or FADH2, largely generated by the tricarboxylic acid cycle-to pump protons across the inner mitochondrial membrane, creating a proton-motive force that Complex V uses to synthesize ATP [51,52,53].
Although the majority of OXPHOS subunits are encoded in the nucleus, translated in the cytosol, and imported into mitochondria [7,37,51], mtDNA encodes 13 indispensable polypeptides that belong to four of the five complexes (I, III, IV, and V) [3,7,37,54]. In total, 92 structural OXPHOS subunits-13 mtDNA-encoded and 79 nDNA-encoded-assemble into a highly coordinated machinery [3,34].
Complex I contains 44 subunits (7 mtDNA-encoded ND1-ND6, ND4L, and 37 nDNA-encoded), Complex II is entirely nDNA-encoded (4 subunits), Complex III includes one mtDNA-encoded cytochrome b plus 10 nDNA-encoded subunits, Complex IV comprises three mtDNA-encoded (COI–COIII) and 11 nDNA-encoded proteins, and Complex V (ATP synthase) contains two mtDNA-encoded (ATPase6, ATPase8) and 17 nDNA-encoded subunits [3,34,55,56]. Together, nDNA- and mtDNA-encoded components orchestrate efficient electron flow and prevent accumulation of harmful assembly intermediates [51].
3. MtDNA Dysfunction in Cardiovascular Diseases
In the heart, mitochondrial oxidative metabolism supplies approximately 90% of cellular ATP, underlining its centrality to cardiovascular function [57]. Cardiac mtDNA governs both the mitochondrial lifecycle and the integrity of the oxidative phosphorylation system [58]. Excess ROS in cardiomyocytes and endothelial cells inflicts irreversible mtDNA damage, leading to mutations that further impair OXPHOS and disrupt mitophagy. Such dysfunction precipitates the escape of mtDNA and mitochondrial proteins into the cytosol, activating innate immune pathways and driving tissue injury [57,58]. These intra- and extra-mitochondrial mtDNA–mediated processes (Figure 2 and Figure 3) highlight mtDNA as a compelling therapeutic target. Table 1 provides representative examples of distinct cardiovascular pathologies driven by mtDNA abnormalities, summarizing their underlying mechanisms and the supporting evidence.
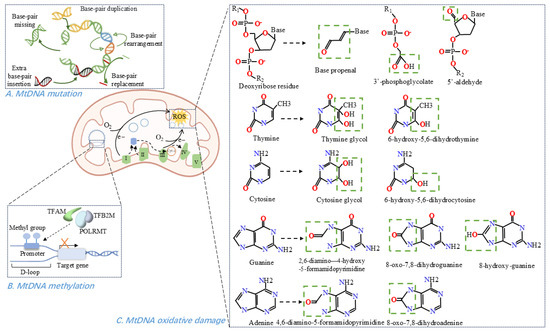
Figure 2.
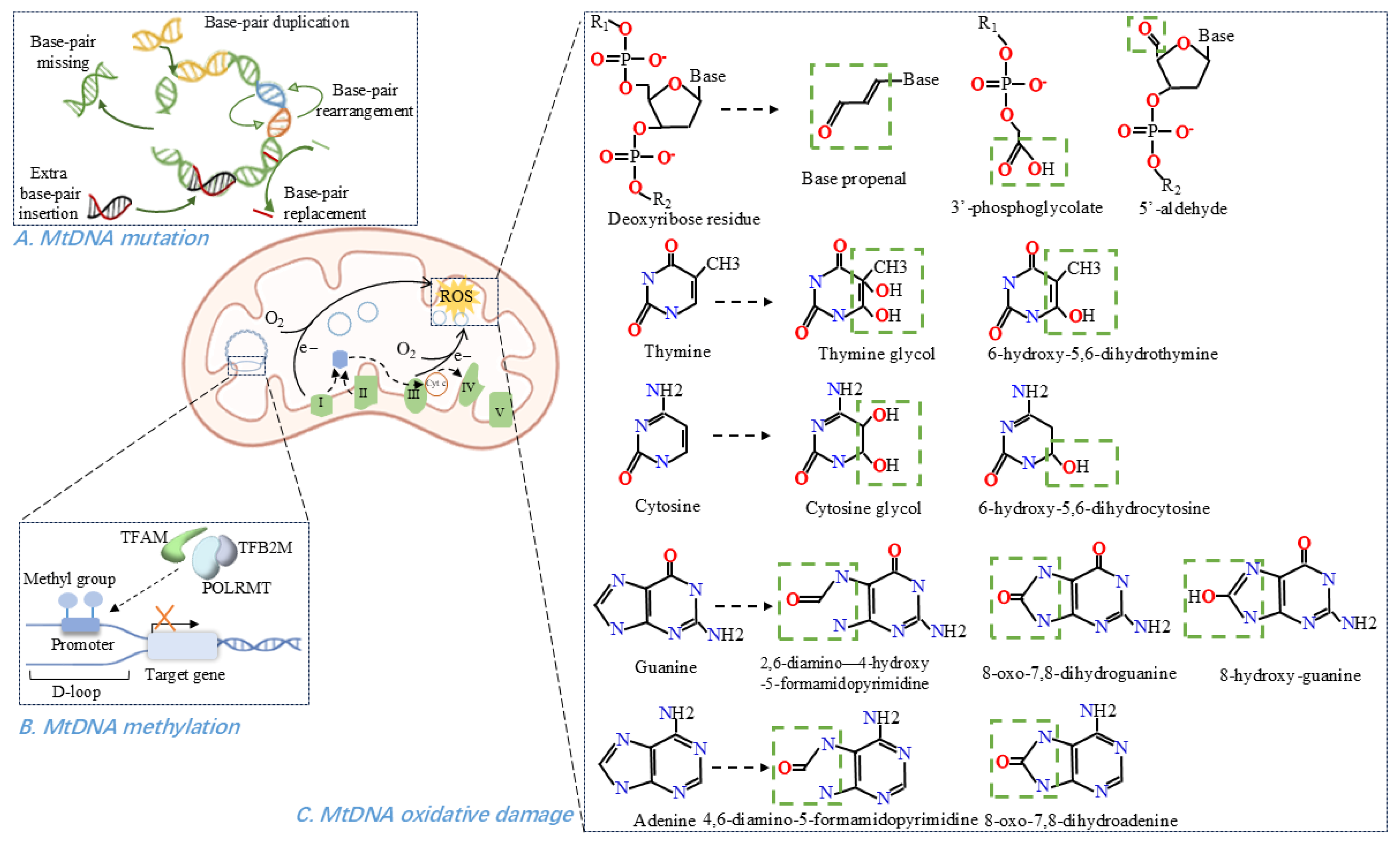
MtDNA-related pathological processes within mitochondria. (A) Forms of mtDNA mutations, include base-pair substitutions, rearrangements, duplications, deletions, and insertions. (B) Methylation of mtDNA in the D-loop region, involves methyl group addition and leads to transcriptional repression. (C) Representative oxidative modifications of mtDNA affect both the sugar-phosphate backbone and the four nucleotide bases.
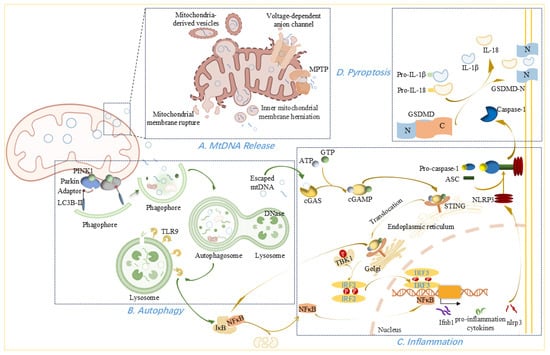
Figure 3.
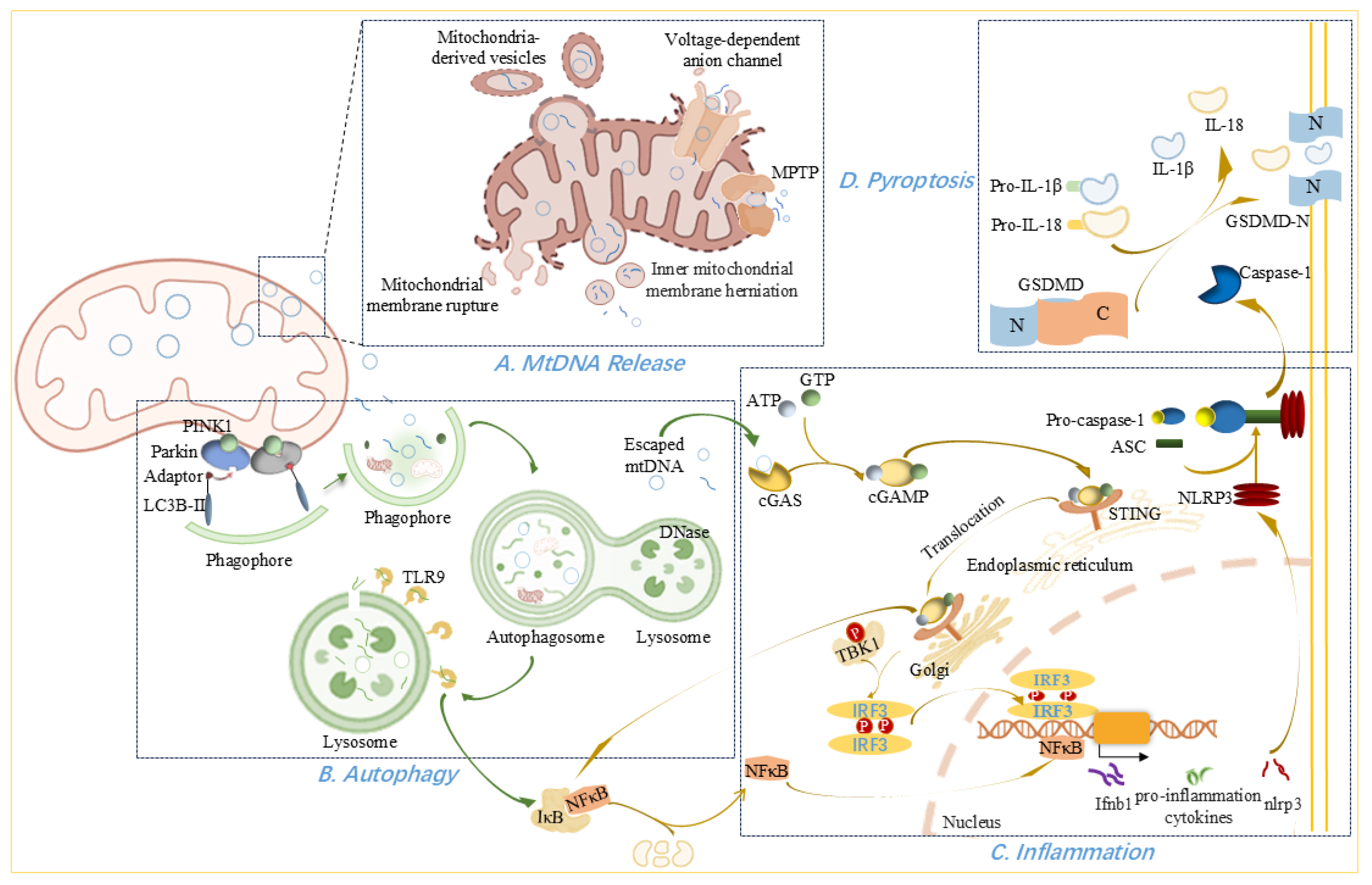
MtDNA release from mitochondria and its physiological and pathological effects outside the organelle. (A) Mechanisms of mtDNA release. MtDNA release occurs via multiple pathways: inner mitochondrial membrane herniation into the cytoplasm; oligomerization of VDACs forming outer membrane pores; leakage through the MPTP; mitochondria-derived vesicles; and excessive mitochondrial fission or rupture. (B) Autophagic Clearance of mtDNA. Autophagy begins with phagophore formation and expansion mediated by PINK1, Parkin, adaptor, and lipidated LC3B-II. The resulting autophagosomes subsequently fuse with lysosomes for mtDNA degradation. (C) mtDNA-Induced Inflammation. Cytosolic mtDNA activates innate immunity via TLR9–NFκB–NLRP3 signaling, leading to elevated inflammatory cytokines, and through the cGAS–cGAMP–STING axis, leading to TBK1/IKK-mediated IRF3 and NFκB activation. This promotes type I interferon and proinflammatory cytokine production (D) Pyroptosis pathway. NLRP3 inflammasome activation cleaves pro-caspase-1 into caspase-1, which processes pro-IL-1β and pro-IL-18 into active forms. Caspase-1 also cleaves GSDMD, generating the N-terminal fragment (GSDMD-N) that forms membrane pores, facilitating the secretion of IL-1β and IL-18 and leading to pyroptotic cell death.
3.1. MtDNA Oxidative Damage
Any damage, especially oxidative damage to mtDNA can alter mitochondrial transcripts and impair oxidative phosphorylation [59]. Mitochondria are significant sources of ROS in cells [60,61] and mtDNA is a major target for ROS-mediated damage due to its proximity to the principal ROS source, the inner mitochondrial membrane [9,62]. Other factors contributing to mtDNA’s susceptibility include the absence of protective histone proteins, limited mtDNA repair activity, retention of superoxide anions within mitochondria, and the negatively charged environment of the mitochondrial matrix [9,59,63,64]. ROS can oxidize bases and the sugar-phosphate backbone of mtDNA or incorporate oxidized bases during DNA polymerization [26,65]. If incompletely repaired, DNA nicks and abasic sites accumulate, blocking mtDNA transcription and replication [26].
Susceptibility to oxidative damage follows the base reactivity order G > C > T >> A, with guanine being the most prone to oxidation, leading to 8-oxo-guanine formation and subsequent G-A mismatches [66,67]. Oxidative-generated DNA modifications have been systematically reviewed elsewhere [64,68]. Hydroxyl radicals readily attack the 2′-deoxyribose, creating carbon-centered sugar radicals that lead to strand incision and unique products (base propenal, 3′-phosphoglycolate, 5′-aldehyde) [64]. Although purine 8,5′-cyclo-2′-deoxyribonucleosides can form via sugar-base radical addition, they have not yet been detected in mtDNA [64]. On pyrimidines, hydroxyl radicals add at C5 or C6 to yield C5–OH–C6 or C6–OH–C5 pyrimidine radicals and downstream redox products [64,68]. The principal purine modification is C8 addition, producing 2,6-diamino-4-hydroxy-5-formamidopyrimidine/4,6-diamino-5-formamidopyrimidine or 8-hydroxy-guanine/8-oxo-7,8-dihydroguanine/8-oxo-7,8-dihydroadenine through different redox pathways [64,69]. Moreover, oxidative stress compromises mitochondrial integrity, promoting the release of modified mtDNA into the cytosol [67].
In most cardiovascular diseases, elevated ROS levels increase mtDNA damage and reduce oxidative phosphorylation [21,31,70,71]. Both clinical and animal studies demonstrate a tight link between mtDNA lesions and mitochondrial dysfunction in myocardial remodeling and heart failure [8,9,63]. Reports indicate oxidative mtDNA damage rises by over 50% in failing hearts [10]. The common mtDNA4977 deletion, a hallmark oxidative lesion, accumulates with age in human hearts and during vascular aging and atherosclerosis [72,73]. In atherosclerosis, conditions such as hyperglycaemia and smoking may further promote mtDNA damage via ROS [74,75], and such damage compromises metabolism, fostering endothelial dysfunction and pro-atherogenic phenotypic changes in vascular smooth muscle cells (VSMC) [11]. Notably, leukocyte levels of 8-hydroxy-2′-deoxyguanosine, an oxidative purine product, predict coronary artery disease risk in type 2 diabetes [69]. As the most common macrovascular complication of diabetes, atherosclerosis also involves ROS-mediated DNA strand breaks [76], with damage accumulating from early lesions and escalating with disease progression [77].
Beyond oxidative insults, non-oxidative mtDNA damage also contributes to cardiovascular diseases: heart failure arises from heart-specific mutant uracil-DNA glycosylase 1 expression [78], and atherosclerosis can result from ROS-independent mtDNA damage in smooth muscle cells and monocytes [75]. Additionally, cardiac hypertrophy, fibrosis, and failure occur following primary mtDNA damage induced by homozygous POLG mutations or prolonged zidovudine treatment [79,80].
3.2. MtDNA Mutations
Mutations in the mitochondrial genome give rise to a wide spectrum of diseases, the majority of which are maternally inherited and involve defects in oxidative energy metabolism [81,82]. MtDNA mutation rate significantly exceeds that of nDNA owing to increased oxidative damage, frequent replication errors, inefficient repair mechanisms, and absence of histone protection [83,84]. In adults, mtDNA mutations account for approximately 70% of mitochondrial diseases [85]. Depending on their location, these mutations may impair specific respiratory-chain proteins or disrupt overall mitochondrial protein synthesis if affecting tRNA or rRNA genes [86].
MtDNA mutations are broadly classified as point mutations or large-scale rearrangements [86]. Most point mutations occur in tRNA genes, with tRNALeuUUR and tRNAIle representing “hotspots” for cardiomyopathy-associated mtDNA variants [7]. Rearrangements are typically heteroplasmic, coexisting with varying proportions of wild-type genomes [7]. Under normal conditions, pathogenic mtDNA variants constitute less than 1% of the total mtDNA pool, rendering tissues effectively homoplasmic for the wild-type sequence [85,86]. MtDNA mutation only manifests clinically once its heteroplasmic load exceeds a tissue-specific threshold-often above 60%—which varies according to the reliance of each tissue on oxidative metabolism [7,85,87]. Consequently, the severity of cellular or tissue dysfunction and the overall phenotypic expression of mtDNA disease are largely determined by the mutant-to-wild-type genome ratio [7,85,88].
MtDNA mutations are strongly associated with cardiovascular disease [7], with specific examples including mtDNA abnormalities in dilated cardiomyopathy [89], increased mtDNA deletions in atrial fibrillation [31], the MTTL1 m.3243A>G mutation and sudden cardiac death [90], and small deletions or heteroplasmic length variants in chronic atrial fibrillation [32]. Notably, approximately 20% of known pathogenic mtDNA point mutations are linked to cardiomyopathy, with tRNA mutations, particularly in tRNALeuUUR and tRNAIle-constituting recognized hotspots [7]. Pathogenic variants in tRNAIle (m.4277T>C, m.4295A>G, m.4300A>G, m.4320C>T) are associated with mitochondrial hypertrophic cardiomyopathy, while m.4269A>G causes encephalocardiomyopathy and m.4317A>G leads to fatal infantile cardiomyopathy [91]. The m.3243A>G mutation in tRNALeuUUR commonly induces MELAS syndrome, frequently involving cardiac manifestations such as hypertrophic cardiomyopathy or Wolff-Parkinson-White syndrome [7,92]. Additionally, transgenic models support a causal role: accelerated accumulation of mtDNA mutations in mice recapitulates human disease phenotypes, including premature aging, dilated cardiac hypertrophy, and fatal congestive heart failure [93]. Kearns–Sayre syndrome—caused by mtDNA rearrangements—demonstrates how impaired energy supply disrupts high-demand tissues [7,94].
Together, these findings highlight the central role of mtDNA mutations-especially those affecting tRNA genes-in the pathogenesis of mitochondrial cardiomyopathies.
3.3. MtDNA Copy-Number Variations
MtDNA copy-number varies among individuals and even among different tissues within the same individual [95], serving as a candidate biomarker of mitochondrial function that reflects both mitochondrial abundance per cell and genome copies per mitochondrion [63,96]. MtDNA depletion, defined as a reduction to less than 30% of normal levels [91], arises from impaired biogenesis or excessive loss due to enzyme deficiencies [97], defective nucleotide metabolism [98], loss of mitochondrial fusion [99], drug toxicity [100,101,102], and other causes. Depletion disrupts the mitochondrial network, reduces cristae membrane content, and leads to a circular morphology of the remaining cristae [103].
Conversely, in response to mitochondrial dysfunction, nuclear-encoded factors may drive disproportionate mitochondrial biogenesis, leading to over-replication of the mtDNA and mitochondrial proliferation [95]. Accordingly, increased mtDNA copy-number in mitochondrial diseases may represent a compensatory mechanism to alleviate bioenergetic deficits, delay disease onset, and modulate phenotypic severity [95,104]. This compensatory increase helps explain why patients harboring high loads of deletion mutants sometimes present with only mild mitochondrial myopathy [95].
In humans, mtDNA content correlates positively with mitochondrial gene transcription and serves as an indicator of mitochondrial dysfunction [80]. Reduced mtDNA copy-number underlies severe cardiac disorders such as cardiomyopathy [91], pathological cardiac remodeling [105,106], atrial fibrillation [107], and heart failure [10,108]. Certain chemotherapeutic agents-including doxorubicin [100,101,102] and nucleoside reverse transcriptase inhibitors [109,110]—exert cardiotoxicity by inhibiting DNA replication and depleting mtDNA. In Chagasic cardiomyopathy, poly (ADP-ribose) polymerase 1 translocates to mitochondria and binds mitochondrial POLγ, impairing mtDNA maintenance and exacerbating mitochondrial dysfunction, oxidative stress, and cardiac remodeling [105]. A case of severe dilated mitochondrial cardiomyopathy was linked to a novel homoplasmic tRNAIle mutation accompanied by profound mtDNA depletion [91]. Additionally, patients with heart failure of diverse etiologies exhibit reduced mtDNA content alongside decreased levels of mtDNA-encoded proteins [111].
3.4. MtDNA Methylation
Epigenetic mechanisms produce heritable phenotypic changes via chromosome modifications rather than alterations in DNA sequence, enabling adaptive responses to environmental stimuli [40,112]. Among these, DNA methylation is the most extensively studied modification and plays a pivotal role in establishing cellular identity. Dysregulated mtDNA methylation has been implicated in aging and various diseases [41,113,114]. MtDNA methylation involves transfer of a methyl group from S-adenosyl methionine to DNA bases, with primarily adenine in the D-loop of the light (L) strand [4,114,115]. Increased methylation in this region generally suppresses transcription [4], thereby affecting mitochondrial quality control and often triggering the integrated stress response [116]. DNA methyltransferases (DNMTs) mediate mtDNA methylation, with DNMT1 being the most abundant and active form throughout adulthood [116]. DNMT1 harbors a mitochondrial targeting sequence, which directs its import into mitochondria, where it binds mtDNA and catalyzes D-loop methylation [117,118]. Upregulation of mitochondrial DNMT1 exerts gene-specific effects: it represses ND6 (the sole protein-coding gene on the L strand) while enhancing ND1 expression on the H strand [41]. Both ND1 and ND6 encode subunits of complex I critical for cardiac contractility [119]. Recent advances in detection methods have identified N6-methyldeoxyadenine (6mA)—a methylation mark once thought restricted to bacteria—in eukaryotes [120]. Its regulatory role in mitochondrial stress has been reported [120,121]. Methyltransferase-like protein 4 (METTL4), a putative mammalian methyltransferase, catalyzes mtDNA 6mA methylation, leading to repression of transcription and reduction in mtDNA copy-number [121].
Although the prevalence and functional significance of mtDNA methylation remain debated, numerous studies have documented correlations between mtDNA methylation (and hydroxymethylation) patterns and cell type, differentiation state, age, and disease status [4,41]. In metabolically active organs such as the heart, altered mtDNA methylation impacts bioenergetics and can promote systemic inflammation, vascular endothelial dysfunction, and myocardial injury [116]. The methylation and downregulation of mitochondrial gene COX2, which encodes COII, are biomarkers of aging in heart mesenchymal stem cells [122]. Moreover, in platelets from patients with cardiovascular disease, increased methylation of mtDNA encoding cytochrome c oxidase and tRNA leucine 1 has been reported [123]. Mitochondrial-targeted overexpression of DNMT1 also impairs mitochondrial gene expression, compromises mitochondrial function, and reduces VSMC contractility [118]. Finally, mtDNA methylation appears to influence clinical presentation in coronary artery disease: patients with acute coronary syndrome exhibit higher mtDNA methylation and lower mtDNA copy-number compared to those with stable coronary artery disease [124].
3.5. MtDNA Release
Under stress, mitochondria can release mtDNA into the cytosol, where it acts as a damage-associated molecular pattern (DAMP) and triggers inflammation [52,125]. During apoptosis, pro-apoptotic proteins BAX and BAK promote mitochondrial outer membrane permeabilization (MOMP), allowing release of intermembrane-space proteins—such as cytochrome c—and subsequent activation of caspases, culminating in cell death [126,127]. However, cell death can also proceed via caspase-independent pathways, which are accompanied by mtDNA leakage [126]. When caspases are inhibited, BAX/BAK-mediated MOMP drives herniation of the inner mitochondrial membrane into the cytosol, carrying matrix components, including mtDNA [126,128,129]. Rather than preventing herniation, apoptotic caspases function to clear dying cells and suppress mtDNA-induced innate immune signaling via the cGAS–STING pathway, thereby maintaining immunological silence during apoptosis [129].
Large mitochondrial membrane pores can also form independently of BAX/BAK. Under oxidative stress, voltage-dependent anion channel 1 (VDAC1)—the most abundant outer-membrane protein, which regulates Ca2+ influx, metabolism, inflammasome activation, and cell death—oligomerizes to create large pores that facilitate mtDNA release [130]. It has been proposed that VDAC1 oligomers mediate mtDNA efflux under moderate stress, whereas BAX/BAK macropores predominate during severe stress or apoptosis [130]. Another debated mechanism involves the mitochondrial permeability transition pore (MPTP), a nonspecific 2–3 nm inner-membrane channel formed in response to Ca2+ overload and other stimuli [66,131]. The MPTP permits passage of molecules smaller than ~1500 Da and may allow egress of mtDNA fragments [66], although its precise role remains controversial and is sometimes conflated with VDAC1 function [131,132].
Mitochondria-derived vesicles have also been suggested as carriers of mtDNA; these small vesicles bud from mitochondria carrying proteins and lipids while preserving membrane integrity [131,133]. However, evidence for mtDNA incorporation into mitochondria-derived vesicles and its subsequent release into the cytosol to activate cGAS remains limited [131]. Finally, excessive mitochondrial fission can disrupt membrane integrity, causing rupture and release of mtDNA into the cytosol [18].
Myocardial mitochondrial dysfunction and consequent mtDNA release are crucial pathophysiological hallmarks or mediating factors for the pathogenesis of myocarditis [134], post-MI cardiac dysfunction [18], myocardial ischemia/reperfusion injury (MI/RI) [23], and thrombosis [135]. Perfusion of the coronary circulation with mtDNA fragments during heart ischemia increases infarct size, underscoring mtDNA’s cardiotoxic potential [26]. In an atherosclerosis mouse model, oxidized mtDNA is released via VDAC1-dependent MPTP opening and activates the STING–PERK pathway [132]. Similar interactions between released mtDNA and inflammation have been observed in animal models of hypertension [15] and doxorubicin-induced cardiotoxicity [136]. Circulating mtDNA has also been proposed as a blood-based biomarker for atrial fibrillation [137].
The lipotoxicity-mediated mtDNA release has also been reported. Upon cellular entry, fatty acids may be esterified into triglycerides for storage in lipid droplets [138,139]. Upon demand, stored triacylglycerols are released and hydrolyzed into fatty acids, which are subsequently converted to fatty acyl-CoA [139]. While fatty acyl-CoA normally undergoes β-oxidation in mitochondria to support energy production, cardiac cells may also divert it toward the synthesis of harmful lipids such as ceramides, diacylglycerols, and triglycerides [140]. The resulting lipid accumulation causes lipotoxicity, promoting endoplasmic reticulum stress, apoptosis, and inflammation [138]. Importantly, lipotoxicity can trigger mtDNA release, activating cGAS–STING signaling and inflammatory pathways, ultimately leading to cardiac cell death and fibrosis [29,30].
3.6. Autophagy
Autophagy serves as another line of defense when insults overwhelm the protective antioxidant, repair pathways, and inhibition of mtDNA release. This homeostatic mechanism sequesters harmful cytosolic components within double-membrane autophagosomes, which subsequently fuse with lysosomes to degrade their contents [125,141]. Under low-energy conditions, AMP-activated protein kinase (AMPK) is activated and phosphorylates ULK1 (unc-51–like kinase 1), initiating phagophore formation. This curved double membrane structure expands via autophagy-related (ATG) proteins into a mature autophagosome [127]. Microtubule-associated protein 1 light chain 3 beta (LC3B), a member of the ATG8 family, integrates into the growing phagophore and drives membrane elongation, of which LC3B-I turns into lipidated form termed LC3B-II that binds to the phagophore [127].
Mitophagy, the selective removal of damaged mitochondria, begins when PTEN-induced kinase 1 (PINK1) accumulates on the outer mitochondrial membrane and recruits Parkin. Parkin ubiquitinates mitochondrial surface proteins, facilitating recruitment of LC3-binding adaptors and targeting mitochondria for autophagic degradation [142]. Upon mitochondrial stress, PINK1 directly phosphorylates and activates Parkin on Ser65 in its ubiquitin-like domain, which leads to ubiquitination of substrates on damaged mitochondria that function as autophagy-mediated degradation signals [143]. Mitophagy mediated by this signaling pathway attenuates pathological cardiac alternation via inhibiting the mtDNA release [144,145]. On the other hand, although PINK1 and Parkin are not encoded by mitochondrial genes, the 3733G>C mutation in mtDNA downregulated PINK1/Parkin-mediated mitophagy pathway [146]. In selective autophagy, autophagy adaptors act as selective takers, which contain both a ubiquitin-binding domain for recognizing ubiquitin chains and an LC3-interacting region for recruiting phagophore membranes [143]. During mitophagy, all known autophagy adaptors are recruited to damaged mitochondria in a PINK1/Parkin-dependent manner [147].
In addition to mitochondrial proteins functioning as substrates of PINK1/Parkin-mediated ubiquitination, mitophagy also relies on various mitochondrial cargo receptors, including BCL2-interacting protein 3 (BNIP3), BNIP3-like protein (BNIP3L, known as NIX), FUN14 domain containing 1(FUNDC1), and others [148,149]. BNIP3 and NIX function primarily under stress, which integrate into the outer mitochondrial membrane via their carboxy terminal transmembrane domains, with the remaining bulk of both proteins facing into the cytosol, where a conserved LC3 interaction region near the amino-terminal end interacts with processed LC3/GABARAP [148]. NIX undergoes ubiquitylation by Parkin to facilitate recruitment of the autophagy adaptor NBR1, which promotes formation of autophagosomes surrounding mitochondria by simultaneously binding to ubiquitin and LC3/GABARAP, while BNIP3 interacts with and stabilizes PINK1 on the outer mitochondrial membrane, leading to Parkin translocation to mitochondria [143]. Cardiac progenitor cells harboring mtDNA mutations fail to efficiently activate the mitophagy in response to differentiation, while this adverse situation is fortunately improved by NIX-and FUNDC1-mediated mitophagy [149].
Autophagosomes carrying impaired mitochondria fuse with the lysosomes and deoxyribonuclease (DNase) degrades mtDNA in the autophagy system to eliminate damaged mtDNA [141,150,151]. However, mtDNA that escapes autophagic clearance elicits inflammatory responses mediated by toll-like receptor (TLR) 9, the NLRP3 inflammasome, or the cGAS-STING pathway [16,125,141,152]. TLR9 recognizes unmethylated CpG motifs in mtDNA delivered to lysosomes by autophagosomes [153,154]; intriguingly, TLR9 activation via this route also contributes to mitochondrial biogenesis, suggesting context-dependent roles based on the source of the TLR9 ligand and how or where the receptor/ligand interaction takes place [154]. Meanwhile, autophagy proteins regulate innate immune responses by limiting NLRP3 inflammasome-mediated mtDNA release [125].
Apoptosis-induced autophagy can occur independently of caspase activation: BAX/BAK-mediated mitochondrial permeabilization rapidly activates AMPK and ULK1, driving autophagic flux that degrades mitochondria and reduces mtDNA release and downstream innate immune signaling [127]. However, it is suspected that cGAS/STING activation occurs too quickly for autophagic clearance to inhibit this early step effectively. For this question, the authors speculated that BAX/BAK-stimulated autophagy causes the cytokine to be sequestered rather than released, but it remains to be determined [127]. Moreover, STING itself can induce autophagy from the ER-Golgi intermediate compartment, as evidenced by LC3 lipidation and autophagosome formation, although the precise mechanism remains unclear [128]. We hypothesize that STING activation at the ER-Golgi intermediate compartment stimulates the local production of autophagy-specific phosphoinositides or facilitates the transfer of lipids to create a lipid microenvironment conducive to the elongation and curvature of phagophores. Because phosphatidylinositol 4-phosphate, the most abundant cellular phosphoinositide with the highest abundance at the trans-Golgi network, is an energy source for lipid-transfer [155]. Lipid transfer plays a critical role in membrane remodeling and is directly implicated in autophagy [156].
Physiologically, mitophagy maintains mitochondrial quality by removing damaged organelles. In some metabolic disorders and cardiovascular diseases, mitophagy is often impaired, leading to the accumulation of dysfunctional mitochondria, elevated mtDNA fragments, and chronic inflammation [157]. For instance, mtDNA, if escaping from autophagy, can provoke inflammation in cardiomyocytes, even myocarditis, and dilated cardiomyopathy [141]. In atherosclerotic plaques, endogenous mtDNA complexed with LL-37 evades DNase II and autophagic degradation, persistently activating TLR9 and driving chemokine and cytokine production that exacerbates lesion development [16]. Similarly, impaired mitophagy facilitates NLRP3 inflammasome activation in atherosclerosis [158], and in silica nanoparticle exposure-induced cardiac adverse events, defective autophagic flux permits cytosolic mtDNA accumulation and cGAS-STING-mediated pyroptosis [152]. During myocardial ischemia/reperfusion or hypoxia/reoxygenation, autophagy deficiency promotes mitochondrial fission, mtDNA release, and sterile inflammation, i.e., the activation of the STING/IRF3 axis [24].
3.7. Inflammation
Because mtDNA contains unmethylated CpG motifs similar to bacterial DNA, it elicits stronger inflammatory responses than nDNA [16]. When mtDNA enters the cytosol, extracellular space, or circulation, it activates multiple pattern-recognition receptors, triggering pro-inflammatory and type I interferon responses [2]. Different mtDNA topology affects different inflammatory responses of cardiomyocytes [159]. As a prototypical DAMP, mtDNA–driven sterile inflammation plays a central role in the pathogenesis of diverse cardiovascular diseases [160,161].
Endolysosomal TLR9 recognizes unmethylated CpG motifs in mtDNA released from injured cells. Upon binding mtDNA, TLR9 initiates a signaling cascade that activates nuclear factor-κB (NFκB) and induces expression of pro-inflammatory cytokines such as TNF-α, IL-1β, and IL-6, as well as assembly of the NLRP3 inflammasome [125,141,160,162,163,164,165]. In the injured heart, these cytokines elevate intracellular ROS, which further compromise mitochondrial membrane integrity and exacerbate cell death [134]. NLRP3 activation itself appears to depend on hexokinase dissociation from mitochondria, thereby promoting the oligomerization of voltage-dependent anion channel (VDAC) on the outer membrane; together with cytosolic mtDNA fragments, oligomerized VDAC recruits NLRP3 to the mitochondrial surface for inflammasome assembly [166]. Oxidized mtDNA released during apoptosis can also directly bind and activate NLRP3 in primed macrophages, driving IL-1β secretion [167].
The cGAS-STING signaling pathway senses pathogenic DNA and initiates a tightly regulated signaling cascade to trigger various inflammatory effectors [30,168]. Cytosolic mtDNA is sensed by cGAS, which catalyzes synthesis of 2′,3′-cyclic GMP-AMP (2′,3′-cGAMP) from ATP and GTP upon DNA binding [128,169]. cGAMP then functions as a secondary messenger activating STING, leading to phosphorylation of interferon regulatory factor 3 (IRF3) and NFκB via the TANK-binding kinase 1 (TBK1) and I-kappa-B kinase (IKK), respectively [30,128,168]. Phosphorylated IRF3 dimerizes and translocates to the nucleus to drive type I interferon transcription [33], while NFκB initiates the transcription of pro-inflammatory cytokines implicated in atherosclerosis, pulmonary arterial hypertension, and myocarditis [132,134,170]. In addition, it may also be involved in metabolic regulation by rewiring the metabolic circuitry, such as ornithine decarboxylase-putrescine metabolic flux [171]. Beyond these canonical pathways, STING can also activate PERK in endothelial cells [132], engage MAPKs, stimulate NLRP3, trigger autophagy, and even induce lytic cell death, underscoring its multifunctional role in inflammation and cell fate [128,157,168].
Clinically, robust myocardial infarction (MI)-mediated inflammation contributes to adverse remodeling and dysfunction across cardiovascular contexts. In this process, cGAS-STING-driven macrophage polarization toward an M1 phenotype exacerbates tissue damage and remodeling; conversely, inhibiting this axis enhances wound healing, neovascularization, and survival post-MI [19,20]. Additionally, NFκB activated by the STING, transactivates ornithine decarboxylase-putrescine metabolic flux to prompt the pathogenesis of cardiac hypertrophy [171]. In hypertension, elevated circulating mtDNA coupled with impaired DNase activity activates TLR9, raising arterial pressure and promoting vascular dysfunction [172]. In atherogenesis, ROS-induced mtDNA damage and release fuel vascular inflammation and VSMC phenotypic switching [17,77], while VCAM1-mediated aberrant mtDNA synthesis in macrophages triggers STING-dependent inflammation and lesion progression [173]. Similarly, STING signaling drives VSMC senescence and fibrous-cap thinning in chronic-kidney-disease-associated atherosclerosis [174]. Finally, activated NLRP3 in cardiomyocytes promotes collagen deposition, left ventricular hypertrophy, fibrosis, and diastolic dysfunction [30].
3.8. Pyroptosis
Pyroptosis is an inflammatory form of programmed cell death characterized by caspase-1-dependent formation of membrane pores, resulting in the release of pro-inflammatory cytokines IL-1β and IL-18 [21]. This process is closely linked to activation of the NLRP3 inflammasome, a multimeric complex composed of NLRP3, the adaptor protein ASC (apoptosis-associated speck-like protein containing a CARD), and caspase-1 [21]. Upon assembly, the inflammasome cleaves pro–caspase-1 into active caspase-1, which in turn processes pro-IL-1β and pro–IL-18 into their mature forms and simultaneously cleaves gasdermin D (GSDMD) to generate the pore-forming N-terminal fragment (GSDMD-N). GSDMD-N mediates secretion of IL-1β and IL-18 and disrupts plasma membrane integrity, culminating in pyroptotic cell death [30,175].
In cardiovascular pathology, NLRP3 inflammasome activation and elevated IL-1β/IL-18 levels drive recruitment of macrophages to aortic lesions, promoting foam-cell formation and plaque progression [176]. Within the myocardium, NLRP3 inflammasome activation in cardiac fibroblasts stimulates IL-1β release and cardiomyocyte pyroptosis, exacerbating cardiac inflammation and adverse remodeling [158]. Additionally, this inflammatory cascade and pyroptosis have been observed in myocardial ischemia [21].
Metabolic stress further accentuates pyroptosis in the heart. In diabetic cardiomyopathy, free fatty acid–induced mtDNA release into the cytosol activates cGAS–STING, triggering pyroptosis and inflammatory responses that drive cardiac hypertrophy [30]. Similarly, exposure to silica nanoparticles disrupts autophagy, leading to cytosolic mtDNA accumulation and cGAS–STING–dependent pyroptosis, as described above [152]. Intriguingly, recent data suggest that cGAS–STING–NLRP3–mediated pyroptosis may enhance the efficacy of radiotherapy in hypertrophic cardiomyopathy, highlighting context-dependent roles for this pathway [177].

Table 1.
Representative examples of cardiovascular pathologies driven by mtDNA abnormalities.
Table 1.
Representative examples of cardiovascular pathologies driven by mtDNA abnormalities.
| Clinical Condition | Mechanism | Evidence/Example | Reference(s) |
|---|---|---|---|
| Hypertension | MtDNA mutations; released mtDNA activates innate immune responses, contributing to vascular dysfunction | m.A14696G and m.A14693G mutations lead to failure in tRNAGlu metabolism | [12] |
| m.15992A>G and m.15077G>A mutations may disturb mitochondrial function | [13] | ||
| m.T10410C and m.T10454C mutations affect the structure and function of tRNAArg | [14] | ||
| Released mtDNA mediates inflammation | [15] | ||
| Elevated circulating mtDNA and impaired DNase activity activate TLR9 | [172] | ||
| Atherosclerosis | Damaged mtDNA escapes autophagy and promotes vascular inflammation and VSMC senescence | MtDNA promotes atherosclerosis through mitochondrial dysfunction | [71] |
| MtDNA4977 deletion is a hallmark oxidative lesion, accumulating in vascular aging and atherosclerosis | [72,73] | ||
| Atherosclerosis can result from ROS-independent mtDNA damage in smooth muscle cells and monocytes | [75] | ||
| Oxidized mtDNA is released and activates the STING–PERK pathway | [132] | ||
| MtDNA complexed with LL-37 evades autophagic degradation | [16] | ||
| Impaired mitophagy facilitates NLRP3 inflammasome activation | [158] | ||
| ROS-induced mtDNA damage and release promote vascular inflammation and VSMC phenotypic switching | [11,17,77] | ||
| Aberrant mtDNA synthesis in macrophages exacerbates STING-dependent inflammation and atherosclerosis | [173] | ||
| STING signaling drives VSMC senescence and fibrous-cap thinning | [174] | ||
| NLRP3 inflammasome activation and elevated IL-1β/IL-18 levels drive recruitment of macrophages to aortic lesions, promoting foam-cell formation and plaque progression | [176] | ||
| MtDNA repair can regulate NLRP3 inflammasome and prevent atherosclerosis | [178] | ||
| Myocardial infarction | Damaged mtDNA is released and triggers severe inflammatory responses, exacerbating tissue damage | Myocardial mitochondrial impairment, mtDNA release, and subsequently STING/p65 activation mediate post-MI cardiac dysfunction | [18] |
| cGAS functions as a cytosolic DNA receptor, promoting macrophage polarization and governing myocardial ischemic injury | [19,20] | ||
| PCSK9 initiates mtDNA damage, and induces pyroptosis | [21] | ||
| Protection of mitochondria and mtDNA can ameliorate isoproterenol-induced MI | [22] | ||
| Myocardial ischemia/reperfusion injury | DNA methylation, mtDNA damage, and release amplify sterile inflammation | Age-associated DNA methylation augments cardiac sensitivity towards MI/RI | [119] |
| Increased mtDNA release enhances pro-inflammatory cytokines | [23] | ||
| Autophagy deficiency promotes mitochondrial fission, mtDNA release, and sterile inflammation | [24] | ||
| ROS can initiate DNA single-strand breakage and severe lesions subsequently | [25] | ||
| Perfusion of coronary circulation with free mtDNA fragments aggravates infarct | [26] | ||
| Heart Failure | MtDNA lesions, mutations, and depletion impairing OXPHOS and ATP production | Oxidative mtDNA damage mediates cardiac dysfunction | [9] |
| ROS increase can lead to a catastrophic cycle of mtDNA damage and cellular injury in heart failure | [63] | ||
| MtDNA toxicity can impair dynamics and function of mitochondria, leading to heart failure | [78] | ||
| Angiotensin II-mediated mtDNA oxidative damage, homozygous POLG mutations, or prolonged zidovudine treatment contribute to cardiac hypertrophy, fibrosis, and failure | [79] | ||
| Accumulation of mtDNA mutations induces premature aging, dilated cardiac hypertrophy, and fatal congestive heart failure | [93] | ||
| MtDNA copy number depletion is an independent risk factor for heart failure | [108] | ||
| Reduced mtDNA replication and depletion of mtDNA impair mitochondrial biogenesis | [10] | ||
| Patients with heart failure exhibit reduced mtDNA content and mtDNA-encoded proteins | [111] | ||
| TFAM overexpression ameliorates mitochondrial deficiencies, increases mtDNA copy-number, and improves cardiac failure | [27,28,179] | ||
| Diabetic Cardiomyopathy | Hyperglycemia induces mtDNA oxidative damage and release, activating cGAS-STING-mediated pyroptosis | Leukocyte levels of 8-hydroxy-2′-deoxyguanosine predict coronary artery disease risk in type 2 diabetes | [69] |
| Oxidized mtDNA release and triggering cGAS-STING-mediated pyroptosis | [30] | ||
| Diabetic cardiomyopathy reduces mtDNA replication and transcription, together with impairing mitochondrial ultrastructure | [180] | ||
| Greater mtDNA damage is found in patients with diabetes mellitus and clinical atherosclerosis | [76] | ||
| Cardiac Hypertrophy | MtDNA mutations, mtDNA damage and chronic inflammation promote fibrosis and hypertrophy | Mutations in tRNAIle including m.4277T>C, m.4295A>G, m.4300A>G, m.4320C>T | [91] |
| Oxidative stress-derived mtDNA damage and deletion are partly linked to cardiac hypertrophy | [72] | ||
| MtDNA lesions cause a vicious cycle with decreased cardiac bioenergetics and ROS accumulation | [80] | ||
| NFκB activated by the STING, prompts cardiac hypertrophy | [171] | ||
| Dilated cardiomyopathy | MtDNA mutations and impaired autophagic flux | Pathological mtDNA mutations leading to abnormal mitochondria | [89] |
| A homoplasmic tRNAIle mutation accompanied by profound mtDNA depletion | [91] | ||
| MtDNA that escapes from autophagy provokes myocarditis and dilated cardiomyopathy | [141] | ||
| Arrhythmia | MtDNA oxidative lesions, mutations, and copy-number variations | Oxidative lesions and mtDNA deletions in cardiomyocytes are increased in atrial fibrillation | [31] |
| Kearns–Sayre syndrome, caused by large-scale mtDNA rearrangements, leads to progressive conduction system degeneration. | [7,94] | ||
| MtDNA mutations are detected in patients with chronic atrial fibrillation | [32] | ||
| MtDNA copy-number is a risk factor for atrial fibrillation | [107] | ||
| Others | MtDNA mutations | m.3243A>G mutation of MTTL1 in sudden cardiac death | [90] |
| m.4269A>G mutation causes encephalocardiomyopathy and m.4317A>G leads to fatal infantile cardiomyopathy | [91] | ||
| m.3243A>G mutation in tRNALeuUUR commonly produces MELAS syndrome | [7,92] | ||
| MtDNA copy-number variations | In Chagasic cardiomyopathy, poly (ADP-ribose) polymerase 1 impairs mtDNA maintenance | [105] | |
| MtDNA methylation abnormality | The methylation and downregulation of COX2 are biomarkers of aging in heart mesenchymal stem cells | [122] | |
| Altered mtDNA methylation can promote systemic inflammation, vascular endothelial dysfunction, and myocardial injury | [116] | ||
| Overexpression of DNMT1 impairs mitochondrial gene expression, compromises mitochondrial function, and reduces VSMC contractility | [118] | ||
| MtDNA release | Impaired mitochondria and released mtDNA mediate myocarditis | [134] | |
| Released mtDNA induces platelet activation, leading to thrombosis | [135] | ||
| Impaired autophagy, mtDNA release, and pyroptosis | In cardiomyocytes exposed to silica nanoparticles, defective autophagic flux permits cytosolic mtDNA accumulation and cGAS-STING-mediated pyroptosis | [152] | |
| Pyroptosis | NLRP3 activation in cardiac fibroblasts stimulates cardiomyocyte pyroptosis, exacerbating cardiac inflammation and adverse remodeling | [158] |
4. Therapeutic Strategies Targeting mtDNA and Its Related Pathways in Cardiovascular Diseases
Maintaining mtDNA integrity and function, and interrupting its downstream inflammatory and cell-death cascades, are crucial for preserving mitochondrial health and preventing cardiovascular disease. Table 2 summarizes current mtDNA-targeted interventions and their mechanisms.
4.1. Repairs of mtDNA Oxidative Damage
The circular structure and proximity of mtDNA to the respiratory chain render it highly susceptible to ROS-mediated damage. In the context of MI/RI, a burst of ROS leads to extensive mtDNA lesions that contribute to cardiomyocyte dysfunction and death [9,11,25,63,181]. Ecklonia cava extract [182] and oleoylethanolamide [183] have been shown to mitigate mtDNA oxidative damage: the former attenuates vascular calcification in hypertensive models, while the latter counteracts STING1-accelerated VSMC senescence-associated vascular calcification. In heterozygous mitochondrial superoxide dismutase knockout mice, overexpression of the mitochondrial helicase Twinkle enhances repair of oxidative mtDNA lesions and protects against cardiomyopathy [184]. Additionally, in spontaneously hypertensive rats, miRNA-21 was found to upregulate translation of mtDNA-encoded cytochrome b, reducing ROS production and ameliorating hypertension-related cardiac pathology [185].
ROS attack both bases and the sugar-phosphate backbone of mtDNA, resulting in mutations, deletions, and strand breaks [26,66]. To counteract these insults, mitochondria rely on a robust base-excision repair (BER) pathway that mitigates oxidative and other minor base damage. BER is initiated by lesion-specific DNA glycosylases possessing apurinic/apyrimidinic (AP) lyase activity, which recognize and remove oxidized or otherwise aberrant bases [2,26,181,186]. The resultant abasic site is processed by AP endonucleases and phosphodiesterases to remove the deoxyribose moiety; POLγ then fills the gap, and DNA ligase III seals the nick [181,186]. Guanine is particularly prone to oxidation, forming 8-oxo-guanine, which is excised by the glycosylase OGG1 [66,181,187]. Enhanced OGG1 activity confers significant protection across multiple cardiac injury models, including MI/RI [26], diabetic cardiomyopathy [186], atherogenesis [178], and cardiac failure [188].
In addition to OGG1, endonuclease III (Endo III) is another mitochondria-targeted DNA glycosylase/AP lyase peptide that preferentially targets oxidized pyrimidines. This fusion protein, Exscien1-III, provided pronounced cardioprotection in mouse models of MI/RI and transverse aortic constriction–induced heart failure [26,189]. However, some studies suggest that BER alone may not suffice to preserve cardiac function in vivo following MI/RI, potentially due to limitations in functional assessment techniques, observation time windows, or the overwhelming ROS burden that exceeds repair capacity [26,181,190,191]. To address this, combined therapy with Endo III and DNase I has been employed: Endo III minimizes DAMP generation by repairing mtDNA lesions, while DNase I degrades irreversibly damaged mtDNA fragments that escape repair [26]. Therefore, such combination therapy targeting both mtDNA damage and its immunostimulatory properties, alongside other therapeutic strategies, is expected to exhibit broad applicability in cardiovascular protection. For instance, Ecklonia cava extract [182], RU.521 [192], and Hirudin [193] collectively ameliorate mitochondrial dysfunction-induced hypertension or hypertensive cardiac injury by targeting multiple stages of the pathological cascade: inhibiting mtDNA oxidative damage, suppressing cGAS activation, and blocking NLRP3 inflammasome assembly, respectively.
4.2. MtDNA Quality Control
Given the central role of TFAM in maintaining mtDNA integrity, therapies that restore TFAM levels confer cardioprotection. In MI/RI rat models, lycopene treatment [194] and the traditional Chinese formulation Huoxue Huatan Decoction [195] preserve cardiomyocyte viability by upregulating TFAM and reducing mtDNA oxidative damage. In angiotensin II-induced hypertensive mice, voluntary exercise enhances endothelial mitochondrial remodeling and lowers blood pressure by promoting p53–TFAM binding, thereby improving mtDNA integrity [196]. Diabetic hearts exhibit diminished TFAM expression and activity, correlated with impaired ETC function; these defects are reversed by resveratrol [54]. Moreover, TFAM overexpression [27,28,179] and TFAM acetylation [197] mitigate mtDNA depletion and mitochondrial dysfunction in models of MI, volume overload, and transverse aortic constriction-induced heart failure. Carvedilol—a third-generation, nonselective β-blocker used in heart failure-stimulates mitochondrial biogenesis via the PGC-1α/TFAM axis in human umbilical vein endothelial cells [198]. In cardiomyocyte hypertrophy models, recombinant human TFAM protein increases mtDNA copy number and inhibits nuclear factor of activated T cells, exerting further cardioprotective effects [199].
Beyond TFAM, mitochondrial quality control encompasses biogenesis, dynamics, and autophagic flux, all of which can be modulated pharmacologically. Agents such as Tat-Beclin 1 [200], sappanone A [201], β-hydroxybutyrate [202], suberoylanilide hydroxamic acid [203], ablating lysocardiolipin acyltransferase 1 [204], astragaloside IV [205], resveratrol [206], and oleuropein [207] reduce mtDNA damage, elevate mtDNA copy number, and improve mitochondrial function in cardiovascular diseases. Activation of peroxisome proliferator–activated receptors (PPARs) also regulates mitochondrial homeostasis [208]: PPARα activation [209] and PPAR 1α deacetylation [210], protect against hyperlipidemia—induced vascular calcification and atherogenesis, respectively. Additionally, high-density lipoprotein (HDL) facilitates cholesterol efflux, maintains the integrity and functionality of mitochondria [211,212], and exhibits antioxidant, anti-inflammatory, and other beneficial properties, contributing to its cardiovascular protective effects [213,214]. The beneficial properties of HDL may interact positively with mtDNA, as suggested by the consistent levels observed between HDL (or HDL-C) and mtDNA [215,216]. Additional interventions—Dl-3-n-butylphthalide [217], the SIRT1 activator RT1720 [218], and Twinkle overexpression [219]—augment mtDNA content and biogenesis in H9c2 cardiomyocytes after H2O2 insult and in MI models. In age-related cardiac hypertrophy, enalapril confers protection by attenuating oxidative damage, increasing mitochondrial mass, and preserving quality-control mechanisms [72].
4.3. MtDNA Content Maintenance
As MtDNA copy-number correlates positively with mitochondrial gene transcription and reflects organelle functionality [80]. Calcitriol supplementation [220] and chronic aerobic exercise [221] increase mtDNA copy-number and improve mitochondrial function, ameliorating age-related hypertension and aortic sclerosis with endothelial dysfunction, respectively. Overexpression of Twinkle, the nuclear-encoded mtDNA helicase, enhances mtDNA replication, raising copy number and conferring protection in volume overload-induced [179] and pressure overload-induced [222] heart failure. Similarly, mitochondrial topoisomerase I relieves mtDNA topological stress and entanglements generated during replication and transcription; its activity shields against doxorubicin—induced cardiotoxicity by maintaining mtDNA topology [223]. Exercise training [180] and recombinant human glucagon-like peptide-1 [224] preserve mtDNA replication and transcription, prevent copy-number decline, and ameliorate mitochondrial ultrastructural defects in diabetic cardiomyopathy and intermittent hypoxia-induced cardiac injury. In addition, omega-3 fatty acids, having been reported as an inducer of mitochondrial biogenesis and function, and their appropriate supplementation exerts their neuro-cardio protective role [225].
Paradoxically, mtDNA–depleted J774A.1 murine macrophages—generated by ethidium bromide treatment—exhibit resistance to oxidized low-density-lipoprotein-induced pyroptosis [226]. This apparent contradiction likely reflects reduced release of mtDNA as a DAMP, thereby attenuating inflammatory cell death.
4.4. Regulation of mtDNA Methylation
DNMT1 is the principal enzyme catalyzing D-loop methylation of mtDNA. Inhibition of DNMT1 by 5-azacytidine attenuates mtDNA hypermethylation and confers protection against MI/RI [117,118,227]. One key target of mtDNA methylation is the COX2 gene, which encodes subunit II of cytochrome c oxidase (Complex IV). In human cardiac mesenchymal stem cells, age-associated senescence is accompanied by COX2 hypermethylation and downregulation; treatment with the DNA methyltransferase inhibitor 5-aza-2′-deoxycytidine restores COX2 expression and improves mitochondrial function [122].
Beyond cytosine methylation, excessive N6-methyladenine (6mA) in mtDNA—mediated by methyltransferase-like protein 4 (METTL4)—impairs mitochondrial gene expression and contributes to heart failure. Genetic or pharmacologic strategies that normalize METTL4 activity and reduce mtDNA 6mA levels alleviate cardiac dysfunction in experimental models [40]. However, developing therapies that selectively target mitochondrial epigenetics remains challenging due to systemic effects on nuclear DNA. For example, systemic administration of 5-aza-2′-deoxycytidine induces genomic hypomethylation, which can lead to genomic instability and tumorigenesis [4]. Tailored delivery systems or mitochondria-specific inhibitors will be essential to exploit mtDNA methylation as a therapeutic avenue without off-target genomic consequences [4].
4.5. Inhibition of mtDNA Release
Beyond repairing oxidative lesions, strategies that prevent mtDNA efflux from damaged mitochondria have shown therapeutic promise. Epigallocatechin-3-gallate attenuates mtDNA release under stress conditions, thereby reducing downstream inflammation [228]. Similarly, blockade of cell-surface nucleolin-using midkine or the aptamer AS1411 suppresses mtDNA uptake by cardiomyocytes during hypoxia-reoxygenation, mitigating inflammatory injury [229].
In the context of acute myocardial infarction, mitochondrial impairment and mtDNA release drive STING-p65 signaling and cardiomyocyte dysfunction [18]. Genetic deletion of large tumor suppressor kinase 2 (LATS2) prevents mtDNA liberation into the cytosol, thereby inhibiting STING–p65 activation and attenuating post-infarction injury [18]. Pharmacologic agents such as astaxanthin [230], thymoquinone [231], and octreotide [22] similarly reduce isoproterenol-induced myocardial damage by decreasing oxidative stress, preserving mitochondrial morphology and function, and restoring mtDNA copy-number.
Metabolic derangements also exacerbate mtDNA release: hyperglycemia impairs mitochondrial integrity via the SIRT1–AMPK–PGC1α axis, promoting secretion of mtDNA-enriched vesicles in diabetic cardiomyopathy. Both exogenous and endogenous interleukin-37 counteract this process, preserving mitochondrial homeostasis and limiting inflammation [232]. Glycerol-3-phosphate acyltransferase 4 (GPAT4), an endoplasmic reticulum–anchored enzyme, supports cardiac development by maintaining ER–mitochondrial communication and preventing mtDNA escape [233]. In human cardiac progenitor–like cells, argon preconditioning enhances cell membrane integrity and suppresses mtDNA release, protecting against oxygen–glucose deprivation–induced cell death [234].
Once in the cytosol or extracellular space, mtDNA can itself be targeted. Z-DNA binding protein 1 (ZBP1) binds released mtDNA to inhibit myocardial inflammation and adverse remodeling after MI, acting as an endogenous brake on DAMP signaling [160]. Intriguingly, in doxorubicin-treated cardiomyocytes, ZBP1 cooperates with cGAS to sustain type I interferon responses, illustrating its context-dependent dual roles in sensing mtDNA and modulating inflammation [235]. These findings underscore ZBP1’s multifunctionality, with its protective or pro-inflammatory effects determined by cell type and stimulus [160].
4.6. Regulation of Autophagy
When oxidative damage exceeds mitochondrial repair capacity, damaged mitochondria are selectively removed via autophagy, preventing accumulation of DAMPs such as mtDNA that would otherwise trigger sterile inflammation and release of pro-fibrotic cytokines [236,237]. In a heart-failure model induced by transverse aortic constriction, DNase II within autolysosomes degrades mtDNA fragments, protecting the myocardium from hemodynamic-stress-induced inflammation [141]. In ischemic myocardial injury, metformin suppresses inflammasome activation by enhancing macrophage autophagy, thereby degrading DAMPs and limiting myocardial injury [237]. Crizotinib—an ALK inhibitor used for metastatic non-small cell lung cancer—interrupts autophagosome-lysosome fusion, contributing to its cardiotoxicity; co-treatment with metformin restores autophagy and rescues crizotinib-induced cardiac injury [238]. Administration of β-hydroxybutyrate at the onset of reperfusion increases autophagic flux and maintains mitochondrial homeostasis by inhibiting the mTOR pathway [202]. Similarly, restoration of mitophagy with the inhibitor of MTORC1 can mitigate mitochondrial dysfunction and cardiomyopathy in Barth syndrome [239].
The PINK1/Parkin pathway is well-known as a mediator of mitophagy [240] and reasonably serves as a target for the treatment of cardiovascular diseases. PINK1 overexpression [144], omega-3 fatty acids [241], and vericiguat [145] protect the heart through promoting PINK1-mediated mitophagy and inhibiting the mtDNA release. On the other hand, astragaloside IV promotes the autophagy of angiotensin II-induced impaired mitochondria by facilitating the expression of Parkin and Drp1 [205].
Components of the cGAS-STING pathway, including cGAMP, cGAS, and TBK1, directly or indirectly induce autophagy and are essential for clearance of cytosolic DNA after genotoxic stress [157]. In Alzheimer’s disease–associated cardiac dysfunction, amyloid-β–induced reductions in melatonin impair mitochondrial aldehyde dehydrogenase activity, leading to mitochondrial damage and mtDNA release. Chronic cytosolic mtDNA accumulation then desensitizes cGAS-STING-TBK1 signaling, suppressing autophagy and further exacerbating amyloid-β and mitochondrial clearance deficits; melatonin supplementation reverses these effects by restoring aldehyde dehydrogenase activity, mitochondrial integrity, cGAS-STING-TBK1 signaling, and autophagic flux [242]. Paradoxically, in hypertension, mtDNA released from microglial mitochondria in the paraventricular nucleus activates cGAS, which in this context blocks autophagic flux and promotes neuroinflammation, sympathetic overdrive, and hypertensive cardiac injury [192]. We hypothesize that mtDNA-triggered, STING-dependent suppression of mitophagy forms a positive feedback loop in hypertensive heart disease, wherein STING activation impairs mitochondrial clearance, leading to further mtDNA release and sustained inflammation. Intracisternal infusion of the cGAS inhibitor RU.521 reverses these effects, underscoring the context-dependent interplay between mtDNA sensing and autophagy [192].
4.7. Ameliorating mtDNA-Triggered Inflammation
MtDNA released into the cytosol potently activates the cGAS–STING pathway, driving inflammatory cascades implicated in cardiovascular pathology. Consequently, inhibition of this axis has emerged as a promising therapeutic strategy. In murine models of high-fat diet–induced and aging-related atherosclerosis, the iridoid glycoside aucubin [243] and the phosphodiesterase-3 inhibitor cilostazol [244], respectively, attenuate lesion progression by suppressing STING signaling. STING deficiency likewise protects against diabetic cardiomyopathy and coxsackievirus B3–induced myocarditis [30,245]. In pulmonary arterial hypertension, cGAS–STING–NFκB activation by cytosolic mtDNA contributes to vascular remodeling; this is mitigated by calcitonin gene–related peptide administration [170]. In neuroinflammation-driven hypertensive cardiac injury, intracisternal infusion of the cGAS inhibitor RU.521 prevents mtDNA-mediated damage [192]. More recently, targeted disruption of cardiomyocyte–macrophage crosstalk has been achieved using a hydrogel patch delivering Rb1/PDA nanoparticles into the infarct border zone; these nanoparticles inhibit macrophage STING activation by cardiomyocyte-derived mtDNA, improving post-MI remodeling [20,246]. However, chronic blockade of cytosolic DNA sensing raises concerns about increased susceptibility to infection [247].
Released mtDNA also engages endosomal Toll-like receptors, particularly TLR9, initiating NFκB–dependent pro-inflammatory responses that promote NLRP3 inflammasome assembly and cytokine release [160,165]. In a SARS-CoV-2 myocarditis model, extracellular vesicles from human umbilical vein endothelial cells inhibit TLR-mediated NFκB activation, reducing myocardial inflammation and mitochondrial damage, and preserving cardiac function [134]. Similarly, TLR4 knockdown protects against circulating mtDNA–mediated inflammation in experimental autoimmune myocarditis [248].
Because the NLRP3 inflammasome mediates mtDNA-driven sterile inflammation, macrophage recruitment, and pyroptosis, its inhibition offers further therapeutic opportunities. The small-molecule inhibitor MCC950 [175], melatonin [249], and metformin [250] each suppress NLRP3 activation in macrophages or endothelial cells, attenuating atherosclerotic lesion development [162]. Electroacupuncture preconditioning decreases circulating mtDNA, prevents NLRP3 overactivation, and expedites the transition from pro-inflammatory to reparative phases after MI/RI [251]. In cardiac hypertrophy, the thrombin inhibitor hirudin reduces ROS, lowers cytosolic mtDNA, and inhibits NLRP3 assembly [193]. Upstream of inflammasome activation, enhancement of mitochondrial aldehyde dehydrogenase activity inhibits the caspase-1/GSDMD pyroptotic pathway, improving survival and preventing septic shock–associated cardiac dysfunction [252].
4.8. Others
Beyond the pathways detailed above, additional interventions targeting mtDNA-mediated mechanisms have demonstrated efficacy:
- •
- PCSK9 Inhibition. Proprotein convertase subtilisin/kexin type 9 (PCSK9) has been implicated in pyroptosis via mtDNA damage during chronic myocardial ischemia. Pharmacologic inhibition of PCSK9 reduces mtDNA lesions, attenuates NLRP3 activation, and confers cardioprotection in ischemic models [21,253]. Furthermore, PCSK9 inhibition exerts a lipid-lowering effect [254,255] and overcomes the limitations of statins in interfering with the attachment of mtDNA to the inner mitochondrial membrane [256], mtDNA depletion [257], suppressing coenzyme Q10 synthesis and inducing mtDNA oxidative damage [258,259], especially the lipophilic statins due to their non-selective diffusion to extrahepatic tissues [260].
- •
- Kearns–Sayre Syndrome Management. Kearns–Sayre syndrome, caused by large-scale mtDNA rearrangements, leads to progressive conduction system degeneration. Prophylactic implantation of a permanent pacemaker is often life-saving, preventing bradyarrhythmias and sudden cardiac death in affected patients [7,261].
- •
- Mitochondrial–Nuclear Exchange Models. In mitochondrial-nuclear exchange mice—where the mtDNA haplotype from C3H/HeN mice is placed on a C57BL/6J nuclear background and vice versa—mtDNA variations modulate nuclear gene expression, mitochondrial morphology, and function. Notably, the C3H mtDNA haplotype attenuates adverse remodeling in volume-overload heart failure, highlighting the protective role of specific mitochondrial genomes in cardiac stress responses [262].

Table 2.
Summary of mtDNA-targeted therapeutic strategies and mechanisms.
Table 2.
Summary of mtDNA-targeted therapeutic strategies and mechanisms.
| Diseases | Treatments | Mechanisms |
|---|---|---|
| Hypertension | Calcitonin gene-related peptide [170] | Alleviating mitochondrial damage, mtDNA release, and cGAS-STING-NFκB activation |
| RU.521 [192] | Inhibiting cGAS | |
| Ecklonia cava extract [182] | Decreasing mtDNA damage and mtROS generation | |
| Voluntary exercise [196] | Improving mtDNA integrity | |
| Oleuropein [207], Calcitriol [220] | Increasing mtDNA copy-number and regulating mitochondrial function | |
| MiRNA-21 [185] | Enhancing translation of mtDNA-encoded cytochrome b | |
| Atherosclerosis | Chronic aerobic exercise [221] | Preserving mitochondrial function |
| Ethidium bromide treatment [226] | MtDNA-depletion | |
| SRT1720 [210] | Improving mtDNA damage and mitochondrial function | |
| OGG1 [178] | DNA repair | |
| Aucubin [243], Cilostazol [132] | Suppressing STING signaling | |
| MCC950 [175], Melatonin [249], Metformin [250] | NLRP3 inhibition | |
| Myocardial infarction | Astaxanthin [230], Octreotide [22] | Reducing oxidative damage, protecting mitochondria, and mtDNA |
| Thymoquinone [231] | Inhibiting mtDNA loss, oxidative stress, inflammation, and apoptosis | |
| Dl-3-n-butylphthalide [217], SRT1720 [218], Twinkle overexpression [219] | Regulating mitochondrial function and biogenesis | |
| Metformin [237] | Alleviating autophagy-ROS-NLRP3 axis-mediated inflammatory response | |
| Rb1/PDA NPs-loaded hydrogel [246] | MtDNA-STING crosstalk modulation | |
| Deleting large tumor suppressor kinase 2 [18] | Preventing mtDNA release | |
| Myocardial ischemia/reperfusion injury | Lycopene [194], Huoxue Huatan Decoction [195] | Restoring TFAM |
| 5-azacytidine [227] | Inhibiting DNMT1 | |
| Tat-Beclin 1 [200], β-hydroxybutyrate [202] | Increasing autophagic flux | |
| Sappanone A [201] | Mitochondrial quality control | |
| Suberoylanilide hydroxamic acid [203] | Inducing autophagy and mitochondrial biogenesis | |
| Elevating OGG1 activity [26], Exscien1-III [189] | Base-excision repair | |
| Combination of Endo III with DNase I [26] | Repairing mtDNA and removing destroyed mtDNA fragments | |
| Epigallocatechin-3-gallate [228], Argon preconditioning [234] | Inhibition of mtDNA release | |
| Midkine, AS1411 [229] | Preventing mtDNA uptake | |
| Electroacupuncture preconditioning [251] | Reducing plasma mtDNA and modulating dynamic inflammatory response | |
| Suppressing proprotein convertase subtilisin/Kexin type 9 [21,253] | Partly through inhibition of pyroptosis via suppressing mtDNA damage | |
| Diabetic cardiomyopathy | STING deficiency [30] | Inhibiting cardiomyocyte pyroptosis and inflammatory response |
| Resveratrol [54] | Reducing TFAM acetylation | |
| IL-37 administration or IL-37 transgenosis [232] | Regulating mtDNA-enriched vesicle release | |
| Exercise training [180] | Reversing reduced mtDNA replication and transcription, and impaired mitochondrial ultrastructure | |
| A dominant negative O-GlcNAc transferase mutant F460A [186] | Restoring OGG1 enzymatic activity | |
| Heart failure | TFAM overexpression [27,28,179] | Ameliorating decreased mtDNA copy-number and mitochondrial deficiencies [27,179] Inhibiting MMP9 protease expression and pathological cardiac remodeling [28] |
| General control of amino acid synthesis 5-like 1 [197] | TFAM acetylation | |
| Twinkle overexpression [179,222] | Increasing mtDNA copy-number | |
| Mitochondrial topoisomerase 1 [223] | Maintaining mtDNA homeostasis | |
| Rectifying N6-methyladenine excess [40] | Inhibition of mtDNA methylation | |
| OGG1 overexpression [188], Exscien1-III [189] | Base-excision repair | |
| DNase II [141] | Digests mtDNA in the autophagy system | |
| Z-DNA binding protein 1 [160] | Suppressing mtDNA-induced inflammation | |
| Myocarditis | Extracellular vesicles derived from human umbilical endothelial cells [134] | Inhibiting TLR-mediated NFκB activation |
| STING deficiency [245] | Resist cardiac inflammation | |
| TLR4 knockdown [248] | Protecting against circulating mtDNA-mediated cardiac inflammation and injury | |
| Cardiac hypertrophy | Recombinant human TFAM protein [199] | Increasing mtDNA and inhibiting nuclear factor of activated T cells |
| Hirudin [193] | Inhibiting NLRP3 inflammasome formation | |
| PINK1 overexpression [144] | Ameliorating autophagy disturbance | |
| Radiotherapy [177] | Inducing oxidative stress, which causes mtDNA leakage and cGAS/STING/NLRP3-mediated pyroptosis | |
| Enalapril [72], Ablating lysocardiolipin acyltransferase 1 [204] | Attenuating mtDNA oxidative damage [72], and maintaining mitochondrial quality control [72,204] | |
| Others | Recombinant human glucagon-like peptide-1 [224] | Preserving mtDNA content and mitochondrial biogenesis |
| Prophylactic placement of a pacemaker [261] | Preserving heart conduction in Kearns-Sayre syndrome | |
| Twinkle [184], Astragaloside IV [205], omega-3 fatty acids [241] | Attenuating mtDNA oxidative damage [184,205], mediating autophagy [205,241] | |
| Oleoylethanolamide [209] | Attenuating mtDNA stress by activating PPARα | |
| 5-aza-2′-deoxycytidine [122] | Inhibiting COX2 gene methylation and downregulation | |
| C3Hmt haplotype [262] | Modulating genome expression and mitochondrial structure/function | |
| Metformin [238], Resveratrol [206], Vericiguat [145], Rapamycin [239] | Ameliorating mitophagy disturbance | |
| Melatonin [242] | Boosting ALDH2 activity | |
| Aldehyde dehydrogenase [252] | Inhibiting mitochondrion-NLRP3 pathway | |
| Glycerol-3-phosphate acyltransferase 4 [233] | Preventing mtDNA escape |
5. Conclusions and Perspectives
In conclusion, mtDNA dysfunction constitutes a self-perpetuating vicious cycle at cardiovascular pathogenesis, mechanistically linking bioenergetic collapse to sterile inflammation. This cycle acts both as an instigator in susceptible individuals and an amplifier of injury in established disease. Disrupting this deleterious cycle requires a two-pronged therapeutic approach: alleviating mtDNA pathological characteristics and suppressing innate immune activation.
Despite these promising strategies, key questions remain unresolved. In some settings, mtDNA defects may be downstream consequences rather than primary causes. The heteroplasmy thresholds at which mutant mtDNA becomes pathogenic are poorly defined and most proposed therapies have not advanced beyond preclinical models. Rigorous mechanistic investigations and well-designed clinical trials will be essential to establish causality, identify at-risk patient populations, and translate this unified pathogenic framework into effective treatments.
Ultimately, future work must integrate mtDNA-focused diagnostics with targeted therapies to break this pathogenic cycle and improve cardiovascular outcomes.
Author Contributions
M.X. and M.Y.: Data curation, Writing-original draft, Investigation, Conceptualization. X.O., A.S. and S.L.: Methodology, Writing-review & editing, Conceptualization, Validation. L.Z. and D.H.: Writing-review & editing, Validation, Supervision, Resources, Project administration, Funding acquisition, Conceptualization. All authors have read and agreed to the published version of the manuscript.
Funding
This work was supported by the National Natural Science Foundations of China (Nos. 82474287, 82274317, 82070136).
Conflicts of Interest
The authors confirmed that there is no conflict of interest.
Abbreviations
ROS, reactive oxygen species; mtDNA, mitochondrial deoxyribonucleic acid; tRNA, transfer ribonucleic acid; rRNA, ribosomal ribonucleic acid; H-strand, heavy strand; L-strand, light strand; ATP, adenosine triphosphate; nDNA, nuclear DNA; D-loop, displacement loop; TFAM, mitochondrial transcription factor A; TFB2M, mitochondrial transcription factor B2; POLRMT, mitochondrial RNA polymerase; POLγ, polymerase γ; LSP, light strand promoter; HSP, heavy strand promoter; OXPHOS, oxidative phosphorylation; ETC, electron transport chain; DNMT1, DNA methyltransferase 1; VSMC, vascular smooth muscle cell; cGAS, cyclic GMP-AMP synthase; STING, stimulator of interferon genes; GTP, guanosine triphosphate; cGAMP, cyclic GMP-AMP; MPTP, mitochondrial permeability transition pore; MI, myocardial infarction; MI/RI, myocardial ischemia/reperfusion injury; AMPK, AMP-activated protein kinase; LC3, microtubule-associated protein 1 light chain 3; DNase, deoxyribonuclease; TLR, toll-like receptor; NLRP3, nucleotide-binding domain and leucine-rich-repeat family pyrin domain containing 3; DAMP, damage-associated molecular pattern; IRF3, interferon regulatory factor 3; NFκB, nuclear factor kappa B; TBK1, TANK-binding kinase 1; IKK, I-kappa-B kinase; ASC, apoptosis-associated speck-like protein containing a CARD; GSDMD, gasdermin D; BER, base-excision repair; OGG1, 8-oxoguanine glycosylase; SIRT, sirtuin; Endo III, endonuclease III; NADH, nicotinamide adenine dinucleotide; FADH2, flavin adenine dinucleotide; PPAR, peroxisome proliferator activated receptor; COX, cytochrome c oxidase; LC3B, microtubule-associated protein 1 light chain 3 beta. PINK1, PTEN-induced kinase 1; BNIP3, BCL2 interacting protein 3; BNIP3L, BNIP3-like protein/Nip3-like protein X (NIX); FUNDC1, FUN14 domain containing 1.
References
- Memme, J.M.; Erlich, A.T.; Phukan, G.; Hood, D.A. Exercise and mitochondrial health. J. Physiol. 2021, 599, 803–817. [Google Scholar] [CrossRef]
- West, A.P.; Shadel, G.S. Mitochondrial DNA in innate immune responses and inflammatory pathology. Nat. Rev. Immunol. 2017, 17, 363–375. [Google Scholar] [CrossRef]
- Chinnery, P.F.; Hudson, G. Mitochondrial genetics. Br. Med. Bull. 2013, 106, 135–159. [Google Scholar] [CrossRef]
- Mongelli, A.; Mengozzi, A.; Geiger, M.; Gorica, E.; Mohammed, S.A.; Paneni, F.; Ruschitzka, F.; Costantino, S. Mitochondrial epigenetics in aging and cardiovascular diseases. Front. Cardiovasc. Med. 2023, 10, 1204483. [Google Scholar] [CrossRef]
- Copeland, W.C. Defects in mitochondrial DNA replication and human disease. Crit. Rev. Biochem. Mol. Biol. 2012, 47, 64–74. [Google Scholar] [CrossRef]
- Liao, T.; Xiong, L.; Wang, X.; Yang, S.; Liang, Z. Mitochondrial disorders as a mechanism for the development of obese Sarcopenia. Diabetol. Metab. Syndr. 2023, 15, 224. [Google Scholar] [CrossRef] [PubMed]
- Santorelli, F.M.; Tessa, A.; D’Amati, G.; Casali, C. The emerging concept of mitochondrial cardiomyopathies. Am. Heart J. 2001, 141, 5A–12A. [Google Scholar] [CrossRef]
- Matsushima, S.; Ide, T.; Yamato, M.; Matsusaka, H.; Hattori, F.; Ikeuchi, M.; Kubota, T.; Sunagawa, K.; Hasegawa, Y.; Kurihara, T.; et al. Overexpression of mitochondrial peroxiredoxin-3 prevents left ventricular remodeling and failure after myocardial infarction in mice. Circulation 2006, 113, 1779–1786. [Google Scholar] [CrossRef] [PubMed]
- Suematsu, N.; Tsutsui, H.; Wen, J.; Kang, D.; Ikeuchi, M.; Ide, T.; Hayashidani, S.; Shiomi, T.; Kubota, T.; Hamasaki, N.; et al. Oxidative stress mediates tumor necrosis factor-alpha-induced mitochondrial DNA damage and dysfunction in cardiac myocytes. Circulation 2003, 107, 1418–1423. [Google Scholar] [CrossRef] [PubMed]
- Karamanlidis, G.; Nascimben, L.; Couper, G.S.; Shekar, P.S.; del Monte, F.; Tian, R. Defective DNA replication impairs mitochondrial biogenesis in human failing hearts. Circ. Res. 2010, 106, 1541–1548. [Google Scholar] [CrossRef]
- Ballinger, S.W.; Patterson, C.; Knight-Lozano, C.A.; Burow, D.L.; Conklin, C.A.; Hu, Z.; Reuf, J.; Horaist, C.; Lebovitz, R.; Hunter, G.C.; et al. Mitochondrial integrity and function in atherogenesis. Circulation 2002, 106, 544–549. [Google Scholar] [CrossRef]
- Wang, C.; Deng, X.; Li, L.; Li, M. Maternally Inherited Essential Hypertension May Be Associated with the Mutations in Mitochondrial tRNA(Glu) Gene. Pharmacogenomics Pers. Med. 2024, 17, 13–26. [Google Scholar] [CrossRef]
- Guo, H.; Guo, L.; Yuan, Y.; Liang, X.Y.; Bi, R. Co-occurrence of m.15992A>G and m.15077G>A Is Associated With a High Penetrance of Maternally Inherited Hypertension in a Chinese Pedigree. Am. J. Hypertens. 2022, 35, 96–102. [Google Scholar] [CrossRef]
- Ding, Y.; Yu, J.; Guo, Q.; Gao, B.; Huang, J. Molecular characterization of two Chinese pedigrees with maternally inherited hypertension. J. Gene Med. 2021, 23, e3328. [Google Scholar] [CrossRef] [PubMed]
- Dikalova, A.E.; Pandey, A.; Xiao, L.; Arslanbaeva, L.; Sidorova, T.; Lopez, M.G.; Billings, F.T.T.; Verdin, E.; Auwerx, J.; Harrison, D.G.; et al. Mitochondrial Deacetylase Sirt3 Reduces Vascular Dysfunction and Hypertension While Sirt3 Depletion in Essential Hypertension Is Linked to Vascular Inflammation and Oxidative Stress. Circ. Res. 2020, 126, 439–452. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Meng, P.; Han, Y.; Shen, C.; Li, B.; Hakim, M.A.; Zhang, X.; Lu, Q.; Rong, M.; Lai, R. Mitochondrial DNA-LL-37 Complex Promotes Atherosclerosis by Escaping from Autophagic Recognition. Immunity 2015, 43, 1137–1147. [Google Scholar] [CrossRef]
- Ding, Z.; Liu, S.; Wang, X.; Dai, Y.; Khaidakov, M.; Deng, X.; Fan, Y.; Xiang, D.; Mehta, J.L. LOX-1, mtDNA damage, and NLRP3 inflammasome activation in macrophages: Implications in atherogenesis. Cardiovasc. Res. 2014, 103, 619–628. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Huang, S.; Du, Y.; Zhou, H.; Zhang, K.; He, J. Lats2 deficiency protects the heart against myocardial infarction by reducing inflammation and inhibiting mitochondrial fission and STING/p65 signaling. Int. J. Biol. Sci. 2023, 19, 3428–3440. [Google Scholar] [CrossRef]
- Frangogiannis, N.G. The inflammatory response in myocardial injury, repair, and remodelling. Nat. Rev. Cardiol. 2014, 11, 255–265. [Google Scholar] [CrossRef]
- Cao, D.J.; Schiattarella, G.G.; Villalobos, E.; Jiang, N.; May, H.I.; Li, T.; Chen, Z.J.; Gillette, T.G.; Hill, J.A. Cytosolic DNA Sensing Promotes Macrophage Transformation and Governs Myocardial Ischemic Injury. Circulation 2018, 137, 2613–2634. [Google Scholar] [CrossRef]
- Wang, X.; Li, X.; Liu, S.; Brickell, A.N.; Zhang, J.; Wu, Z.; Zhou, S.; Ding, Z. PCSK9 regulates pyroptosis via mtDNA damage in chronic myocardial ischemia. Basic Res. Cardiol. 2020, 115, 66. [Google Scholar] [CrossRef]
- Khalifa, A.A.; El Sokkary, N.H.; Elblehi, S.S.; Diab, M.A.; Ali, M.A. Potential cardioprotective effect of octreotide via NOXs mitigation, mitochondrial biogenesis and MAPK/Erk1/2/STAT3/NF-kβ pathway attenuation in isoproterenol-induced myocardial infarction in rats. Eur. J. Pharmacol. 2022, 925, 174978. [Google Scholar] [CrossRef]
- Feng, Y.; Aliagan, A.I.; Tombo, N.; Bopassa, J.C. Mitofilin Heterozygote Mice Display an Increase in Myocardial Injury and Inflammation after Ischemia/Reperfusion. Antioxidants 2023, 12, 921. [Google Scholar] [CrossRef]
- Tam, E.; Song, E.; Noskovicova, N.; Hinz, B.; Xu, A.; Sweeney, G. Autophagy deficiency exacerbated hypoxia-reoxygenation induced inflammation and cell death via a mitochondrial DNA/STING/IRF3 pathway. Life Sci. 2024, 358, 123173. [Google Scholar] [CrossRef] [PubMed]
- Cuzzocrea, S.; Riley, D.P.; Caputi, A.P.; Salvemini, D. Antioxidant therapy: A new pharmacological approach in shock, inflammation, and ischemia/reperfusion injury. Pharmacol. Rev. 2001, 53, 135–159. [Google Scholar] [CrossRef]
- Yang, X.M.; Cui, L.; White, J.; Kuck, J.; Ruchko, M.V.; Wilson, G.L.; Alexeyev, M.; Gillespie, M.N.; Downey, J.M.; Cohen, M.V. Mitochondrially targeted Endonuclease III has a powerful anti-infarct effect in an in vivo rat model of myocardial ischemia/reperfusion. Basic Res. Cardiol. 2015, 110, 3. [Google Scholar] [CrossRef]
- Ikeuchi, M.; Matsusaka, H.; Kang, D.; Matsushima, S.; Ide, T.; Kubota, T.; Fujiwara, T.; Hamasaki, N.; Takeshita, A.; Sunagawa, K.; et al. Overexpression of mitochondrial transcription factor a ameliorates mitochondrial deficiencies and cardiac failure after myocardial infarction. Circulation 2005, 112, 683–690. [Google Scholar] [CrossRef]
- Kunkel, G.H.; Kunkel, C.J.; Ozuna, H.; Miralda, I.; Tyagi, S.C. TFAM overexpression reduces pathological cardiac remodeling. Mol. Cell. Biochem. 2019, 454, 139–152. [Google Scholar] [CrossRef] [PubMed]
- Ma, X.M.; Geng, K.; Law, B.Y.; Wang, P.; Pu, Y.L.; Chen, Q.; Xu, H.W.; Tan, X.Z.; Jiang, Z.Z.; Xu, Y. Lipotoxicity-induced mtDNA release promotes diabetic cardiomyopathy by activating the cGAS-STING pathway in obesity-related diabetes. Cell Biol. Toxicol. 2023, 39, 277–299. [Google Scholar] [CrossRef] [PubMed]
- Yan, M.; Li, Y.; Luo, Q.; Zeng, W.; Shao, X.; Li, L.; Wang, Q.; Wang, D.; Zhang, Y.; Diao, H.; et al. Mitochondrial damage and activation of the cytosolic DNA sensor cGAS-STING pathway lead to cardiac pyroptosis and hypertrophy in diabetic cardiomyopathy mice. Cell Death Discov. 2022, 8, 258. [Google Scholar] [CrossRef]
- Lin, P.H.; Lee, S.H.; Su, C.P.; Wei, Y.H. Oxidative damage to mitochondrial DNA in atrial muscle of patients with atrial fibrillation. Free Radic. Biol. Med. 2003, 35, 1310–1318. [Google Scholar] [CrossRef]
- Park, H.W.; Ahn, Y.; Jeong, M.H.; Cho, J.G.; Park, J.C.; Kang, J.C.; Shin, M.G.; Shin, J.H.; Suh, S.P.; Ryang, D.W.; et al. Chronic atrial fibrillation associated with somatic mitochondrial DNA mutations in human atrial tissue. J. Clin. Pathol. 2007, 60, 948–950. [Google Scholar] [CrossRef]
- Heilig, R.; Lee, J.; Tait, S.W.G. Mitochondrial DNA in cell death and inflammation. Biochem. Soc. Trans. 2023, 51, 457–472. [Google Scholar] [CrossRef]
- Hu, H.; Lin, Y.; Xu, X.; Lin, S.; Chen, X.; Wang, S. The alterations of mitochondrial DNA in coronary heart disease. Exp. Mol. Pathol. 2020, 114, 104412. [Google Scholar] [CrossRef] [PubMed]
- West, A.P.; Khoury-Hanold, W.; Staron, M.; Tal, M.C.; Pineda, C.M.; Lang, S.M.; Bestwick, M.; Duguay, B.A.; Raimundo, N.; MacDuff, D.A.; et al. Mitochondrial DNA stress primes the antiviral innate immune response. Nature 2015, 520, 553–557. [Google Scholar] [CrossRef]
- Wang, K.Z.; Zhu, J.; Dagda, R.K.; Uechi, G.; Cherra, S.J., 3rd; Gusdon, A.M.; Balasubramani, M.; Chu, C.T. ERK-mediated phosphorylation of TFAM downregulates mitochondrial transcription: Implications for Parkinson’s disease. Mitochondrion 2014, 17, 132–140. [Google Scholar] [CrossRef] [PubMed]
- Ngo, H.B.; Kaiser, J.T.; Chan, D.C. The mitochondrial transcription and packaging factor Tfam imposes a U-turn on mitochondrial DNA. Nat. Struct. Mol. Biol. 2011, 18, 1290–1296. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Liu, H.; Zhang, F.; Xu, H. The initiation of mitochondrial DNA replication. Biochem. Soc. Trans. 2024, 52, 1243–1251. [Google Scholar] [CrossRef]
- Falkenberg, M.; Larsson, N.G.; Gustafsson, C.M. Replication and Transcription of Human Mitochondrial DNA. Annu. Rev. Biochem. 2024, 93, 47–77. [Google Scholar] [CrossRef]
- Zhang, F.; Zhang, L.; Hu, G.; Chen, X.; Liu, H.; Li, C.; Guo, X.; Huang, C.; Sun, F.; Li, T.; et al. Rectifying METTL4-Mediated N(6)-Methyladenine Excess in Mitochondrial DNA Alleviates Heart Failure. Circulation 2024. [CrossRef]
- van der Wijst, M.G.; Rots, M.G. Mitochondrial epigenetics: An overlooked layer of regulation? Trends Genet. 2015, 31, 353–356. [Google Scholar] [CrossRef]
- Rubio-Cosials, A.; Sidow, J.F.; Jiménez-Menéndez, N.; Fernández-Millán, P.; Montoya, J.; Jacobs, H.T.; Coll, M.; Bernadó, P.; Solà, M. Human mitochondrial transcription factor A induces a U-turn structure in the light strand promoter. Nat. Struct. Mol. Biol. 2011, 18, 1281–1289. [Google Scholar] [CrossRef]
- Ott, M.; Amunts, A.; Brown, A. Organization and Regulation of Mitochondrial Protein Synthesis. Annu. Rev. Biochem. 2016, 85, 77–101. [Google Scholar] [CrossRef]
- Kummer, E.; Schubert, K.N.; Schoenhut, T.; Scaiola, A.; Ban, N. Structural basis of translation termination, rescue, and recycling in mammalian mitochondria. Mol. Cell 2021, 81, 2566–2582.e6. [Google Scholar] [CrossRef]
- Kummer, E.; Ban, N. Mechanisms and regulation of protein synthesis in mitochondria. Nat. Rev. Mol. Cell Biol. 2021, 22, 307–325. [Google Scholar] [CrossRef]
- Schwartzbach, C.J.; Spremulli, L.L. Bovine mitochondrial protein synthesis elongation factors. Identification and initial characterization of an elongation factor Tu-elongation factor Ts complex. J. Biol. Chem. 1989, 264, 19125–19131. [Google Scholar] [CrossRef] [PubMed]
- Tsuboi, M.; Morita, H.; Nozaki, Y.; Akama, K.; Ueda, T.; Ito, K.; Nierhaus, K.H.; Takeuchi, N. EF-G2mt is an exclusive recycling factor in mammalian mitochondrial protein synthesis. Mol. Cell 2009, 35, 502–510. [Google Scholar] [CrossRef] [PubMed]
- Bhargava, K.; Templeton, P.; Spremulli, L.L. Expression and characterization of isoform 1 of human mitochondrial elongation factor G. Protein Expr. Purif. 2004, 37, 368–376. [Google Scholar] [CrossRef]
- Chung, H.K.; Spremulli, L.L. Purification and characterization of elongation factor G from bovine liver mitochondria. J. Biol. Chem. 1990, 265, 21000–21004. [Google Scholar] [CrossRef] [PubMed]
- Eberly, S.L.; Locklear, V.; Spremulli, L.L. Bovine mitochondrial ribosomes. Elongation factor specificity. J. Biol. Chem. 1985, 260, 8721–8725. [Google Scholar] [CrossRef]
- Kremer, L.S. Rehling, Coordinating mitochondrial translation with assembly of the OXPHOS complexes. Hum. Mol. Genet. 2024, 33, R47–R52. [Google Scholar] [CrossRef] [PubMed]
- Elorza, A.A.; Soffia, J.P. mtDNA Heteroplasmy at the Core of Aging-Associated Heart Failure. An Integrative View of OXPHOS and Mitochondrial Life Cycle in Cardiac Mitochondrial Physiology. Front. Cell Dev. Biol. 2021, 9, 625020. [Google Scholar] [CrossRef]
- Fernandez-Vizarra, E.; Zeviani, M. Mitochondrial disorders of the OXPHOS system. FEBS Lett. 2021, 595, 1062–1106. [Google Scholar] [CrossRef]
- Bagul, P.K.; Katare, P.B.; Bugga, P.; Dinda, A.K.; Banerjee, S.K. SIRT-3 Modulation by Resveratrol Improves Mitochondrial Oxidative Phosphorylation in Diabetic Heart through Deacetylation of TFAM. Cells 2018, 7, 235. [Google Scholar] [CrossRef]
- Hirst, J. Why does mitochondrial complex I have so many subunits? Biochem. J. 2011, 437, e1–e3. [Google Scholar] [CrossRef] [PubMed]
- Angerer, H.; Zwicker, K.; Wumaier, Z.; Sokolova, L.; Heide, H.; Steger, M.; Kaiser, S.; Nübel, E.; Brutschy, B.; Radermacher, M.; et al. A scaffold of accessory subunits links the peripheral arm and the distal proton-pumping module of mitochondrial complex I. Biochem. J. 2011, 437, 279–288. [Google Scholar] [CrossRef]
- Rai, N.K.; Venugopal, H.; Rajesh, R.; Ancha, P.; Venkatesh, S. Mitochondrial complex-1 as a therapeutic target for cardiac diseases. Mol. Cell. Biochem. 2024, 480, 869–890. [Google Scholar] [CrossRef]
- Cañadas-Garre, M.; Maqueda, J.J.; Baños-Jaime, B.; Hill, C.; Skelly, R.; Cappa, R.; Brennan, E.; Doyle, R.; Godson, C.; Maxwell, A.P.; et al. Mitochondrial related variants associated with cardiovascular traits. Front. Physiol. 2024, 15, 1395371. [Google Scholar] [CrossRef]
- Ballinger, S.W.; Patterson, C.; Yan, C.N.; Doan, R.; Burow, D.L.; Young, C.G.; Yakes, F.M.; Van Houten, B.; Ballinger, C.A.; Freeman, B.A.; et al. Hydrogen peroxide- and peroxynitrite-induced mitochondrial DNA damage and dysfunction in vascular endothelial and smooth muscle cells. Circ. Res. 2000, 86, 960–966. [Google Scholar] [CrossRef]
- Liu, J.; Yan, P.; Liu, X.; Long, Z.; Bing, T.; Zhang, N.; Shangguan, D. Heptamethine Cyanine-Based Molecule Release Triggered by Mitochondrial ROS. ACS Appl. Bio Mater. 2023, 7, 362–368. [Google Scholar] [CrossRef] [PubMed]
- Bandy, B.; Davison, A.J. Mitochondrial mutations may increase oxidative stress: Implications for carcinogenesis and aging? Free Radic. Biol. Med. 1990, 8, 523–539. [Google Scholar] [CrossRef]
- Gredilla, R.; Bohr, V.A.; Stevnsner, T. Mitochondrial DNA repair and association with aging—An update. Exp. Gerontol. 2010, 45, 478–488. [Google Scholar]
- Tsutsui, H.; Kinugawa, S.; Matsushima, S. Oxidative stress and heart failure. Am. J. Physiol. Heart Circ. Physiol. 2011, 301, H2181–H2190. [Google Scholar] [CrossRef] [PubMed]
- Muftuoglu, M.; Mori, M.P.; de Souza-Pinto, N.C. Formation and repair of oxidative damage in the mitochondrial DNA. Mitochondrion 2014, 17, 164–181. [Google Scholar] [CrossRef]
- Hayashi, G.; Cortopassi, G. Oxidative stress in inherited mitochondrial diseases. Free Radic. Biol. Med. 2015, 88, 10–17. [Google Scholar] [CrossRef]
- García, N.; Chávez, E. Mitochondrial DNA fragments released through the permeability transition pore correspond to specific gene size. Life Sci. 2007, 81, 1160–1166. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Xu, T.; Wu, Z.; Li, N.; Liang, Q. Rebalancing redox homeostasis: A pivotal regulator of the cGAS-STING pathway in autoimmune diseases. Autoimmun. Rev. 2025, 24, 103823. [Google Scholar] [CrossRef] [PubMed]
- Cooke, M.S.; Evans, M.D.; Dizdaroglu, M.; Lunec, J. Oxidative DNA damage: Mechanisms, mutation, and disease. FASEB J. 2003, 17, 1195–1214. [Google Scholar] [CrossRef]
- Wang, X.B.; Cui, N.H.; Liu, X.; Liu, X. Mitochondrial 8-hydroxy-2′-deoxyguanosine and coronary artery disease in patients with type 2 diabetes mellitus. Cardiovasc. Diabetol. 2020, 19, 22. [Google Scholar] [CrossRef]
- Corral-Debrinski, M.; Stepien, G.; Shoffner, J.M.; Lott, M.T.; Kanter, K.; Wallace, D.C. Hypoxemia is associated with mitochondrial DNA damage and gene induction. Implications for cardiac disease. JAMA 1991, 266, 1812–1816. [Google Scholar] [CrossRef]
- Yu, E.P.; Bennett, M.R. The role of mitochondrial DNA damage in the development of atherosclerosis. Free Radic. Biol. Med. 2016, 100, 223–230. [Google Scholar] [CrossRef]
- Picca, A.; Sirago, G.; Pesce, V.; Lezza, A.M.S.; Calvani, R.; Bossola, M.; Villani, E.R.; Landi, F.; Leeuwenburgh, C.; Bernabei, R.; et al. Administration of Enalapril Started Late in Life Attenuates Hypertrophy and Oxidative Stress Burden, Increases Mitochondrial Mass, and Modulates Mitochondrial Quality Control Signaling in the Rat Heart. Biomolecules 2018, 8, 177. [Google Scholar] [CrossRef]
- Vecoli, C.; Borghini, A.; Andreassi, M.G. The molecular biomarkers of vascular aging and atherosclerosis: Telomere length and mitochondrial DNA(4977) common deletion. Mutat. Res. Rev. Mutat. Res. 2020, 784, 108309. [Google Scholar] [CrossRef] [PubMed]
- Yu, T.; Robotham, J.L.; Yoon, Y. Increased production of reactive oxygen species in hyperglycemic conditions requires dynamic change of mitochondrial morphology. Proc. Natl. Acad. Sci. USA 2006, 103, 2653–2658. [Google Scholar] [CrossRef]
- Yu, E.; Calvert, P.A.; Mercer, J.R.; Harrison, J.; Baker, L.; Figg, N.L.; Kumar, S.; Wang, J.C.; Hurst, L.A.; Obaid, D.R.; et al. Mitochondrial DNA damage can promote atherosclerosis independently of reactive oxygen species through effects on smooth muscle cells and monocytes and correlates with higher-risk plaques in humans. Circulation 2013, 128, 702–712. [Google Scholar] [CrossRef]
- Fetterman, J.L.; Holbrook, M.; Westbrook, D.G.; Brown, J.A.; Feeley, K.P.; Bretón-Romero, R.; Linder, E.A.; Berk, B.D.; Weisbrod, R.M.; Widlansky, M.E.; et al. Mitochondrial DNA damage and vascular function in patients with diabetes mellitus and atherosclerotic cardiovascular disease. Cardiovasc. Diabetol. 2016, 15, 53. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Li, X.; Song, S.; Sun, Z.; Li, Y.; Yang, L.; Xie, Z.; Cai, Y.; Zhao, Y. Mitochondria spatially and temporally modulate VSMC phenotypes via interacting with cytoskeleton in cardiovascular diseases. Redox Biol. 2023, 64, 102778. [Google Scholar] [CrossRef]
- Lauritzen, K.H.; Kleppa, L.; Aronsen, J.M.; Eide, L.; Carlsen, H.; Haugen, Ø.P.; Sjaastad, I.; Klungland, A.; Rasmussen, L.J.; Attramadal, H.; et al. Impaired dynamics and function of mitochondria caused by mtDNA toxicity leads to heart failure. Am. J. Physiol. Heart Circ. Physiol. 2015, 309, H434–H449. [Google Scholar] [CrossRef]
- Dai, D.F.; Johnson, S.C.; Villarin, J.J.; Chin, M.T.; Nieves-Cintrón, M.; Chen, T.; Marcinek, D.J.; Dorn, G.W., 2nd; Kang, Y.J.; Prolla, T.A.; et al. Mitochondrial oxidative stress mediates angiotensin II-induced cardiac hypertrophy and Galphaq overexpression-induced heart failure. Circ. Res. 2011, 108, 837–846. [Google Scholar] [CrossRef]
- Wang, X.; Yu, Q.; Liao, X.; Fan, M.; Liu, X.; Liu, Q.; Wang, M.; Wu, X.; Huang, C.K.; Tan, R.; et al. Mitochondrial Dysfunction in Arrhythmia and Cardiac Hypertrophy. Rev. Cardiovasc. Med. 2023, 24, 364. [Google Scholar] [CrossRef] [PubMed]
- Schon, E.A.; DiMauro, S.; Hirano, M. Human mitochondrial DNA: Roles of inherited and somatic mutations. Nat. Rev. Genet. 2012, 13, 878–890. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.W.; Giordano, C.; Davidson, M.M.; d’Amati, G.; Bain, H.; Hayes, C.M.; Leonard, H.; Barron, M.J.; Casali, C.; Santorelli, F.M.; et al. A homoplasmic mitochondrial transfer ribonucleic acid mutation as a cause of maternally inherited hypertrophic cardiomyopathy. J. Am. Coll. Cardiol. 2003, 41, 1786–1796. [Google Scholar] [CrossRef] [PubMed]
- Dabravolski, S.A.; Khotina, V.A.; Sukhorukov, V.N.; Kalmykov, V.A.; Mikhaleva, L.M.; Orekhov, A.N. The Role of Mitochondrial DNA Mutations in Cardiovascular Diseases. Int. J. Mol. Sci. 2022, 23, 952. [Google Scholar] [CrossRef] [PubMed]
- Salnikova, D.; Orekhova, V.; Grechko, A.; Starodubova, A.; Bezsonov, E.; Popkova, T.; Orekhov, A. Mitochondrial Dysfunction in Vascular Wall Cells and Its Role in Atherosclerosis. Int. J. Mol. Sci. 2021, 22, 8990. [Google Scholar] [CrossRef]
- Bates, M.G.; Bourke, J.P.; Giordano, C.; d’Amati, G.; Turnbull, D.M.; Taylor, R.W. Cardiac involvement in mitochondrial DNA disease: Clinical spectrum, diagnosis, and management. Eur. Heart J. 2012, 33, 3023–3033. [Google Scholar] [CrossRef]
- Viscomi, C.; Zeviani, M. MtDNA-maintenance defects: Syndromes and genes. J. Inherit. Metab. Dis. 2017, 40, 587–599. [Google Scholar] [CrossRef]
- Chen, H.; Vermulst, M.; Wang, Y.E.; Chomyn, A.; Prolla, T.A.; McCaffery, J.M.; Chan, D.C. Mitochondrial fusion is required for mtDNA stability in skeletal muscle and tolerance of mtDNA mutations. Cell 2010, 141, 280–289. [Google Scholar] [CrossRef]
- Jackson, C.B.; Turnbull, D.M.; Minczuk, M.; Gammage, P.A. Therapeutic Manipulation of mtDNA Heteroplasmy: A Shifting Perspective. Trends Mol. Med. 2020, 26, 698–709. [Google Scholar] [CrossRef]
- Arbustini, E.; Diegoli, M.; Fasani, R.; Grasso, M.; Morbini, P.; Banchieri, N.; Bellini, O.; Bello, B.D.; Pilotto, A.; Magrini, G.; et al. Mitochondrial DNA mutations and mitochondrial abnormalities in dilated cardiomyopathy. Am. J. Pathol. 1998, 153, 1501–1510. [Google Scholar] [CrossRef]
- Majamaa-Voltti, K.; Turkka, J.; Kortelainen, M.L.; Huikuri, H.; Majamaa, K. Causes of death in pedigrees with the 3243A>G mutation in mitochondrial DNA. J. Neurol. Neurosurg. Psychiatry 2008, 79, 209–211. [Google Scholar] [CrossRef]
- Alila-Fersi, O.; Tabebi, M.; Maalej, M.; Belguith, N.; Keskes, L.; Mkaouar-Rebai, E.; Fakhfakh, F. First description of a novel mitochondrial mutation in the MT-TI gene associated with multiple mitochondrial DNA deletion and depletion in family with severe dilated mitochondrial cardiomyopathy. Biochem. Biophys. Res. Commun. 2018, 497, 1049–1054. [Google Scholar] [CrossRef]
- Pavlakis, S.G.; Phillips, P.C.; DiMauro, S.; De Vivo, D.C.; Rowland, L.P. Mitochondrial myopathy, encephalopathy, lactic acidosis, and strokelike episodes: A distinctive clinical syndrome. Ann. Neurol. 1984, 16, 481–488. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Mott, J.L.; Chang, S.W.; Denniger, G.; Feng, Z.; Zassenhaus, H.P. Construction of transgenic mice with tissue-specific acceleration of mitochondrial DNA mutagenesis. Genomics 2000, 69, 151–161. [Google Scholar] [CrossRef]
- Yazdani, M. Cellular and Molecular Responses to Mitochondrial DNA Deletions in Kearns-Sayre Syndrome: Some Underlying Mechanisms. Mol. Neurobiol. 2024, 61, 5665–5679. [Google Scholar] [CrossRef]
- Bai, R.K.; Wong, L.J. Simultaneous detection and quantification of mitochondrial DNA deletion(s), depletion, and over-replication in patients with mitochondrial disease. J. Mol. Diagn. 2005, 7, 613–622. [Google Scholar] [CrossRef] [PubMed]
- Ashar, F.N.; Zhang, Y.; Longchamps, R.J.; Lane, J.; Moes, A.; Grove, M.L.; Mychaleckyj, J.C.; Taylor, K.D.; Coresh, J.; Rotter, J.I.; et al. Association of Mitochondrial DNA Copy Number With Cardiovascular Disease. JAMA Cardiol. 2017, 2, 1247–1255. [Google Scholar] [CrossRef]
- Misic, J.; Milenkovic, D.; Al-Behadili, A.; Xie, X.; Jiang, M.; Jiang, S.; Filograna, R.; Koolmeister, C.; Siira, S.J.; Jenninger, L.; et al. Mammalian RNase H1 directs RNA primer formation for mtDNA replication initiation and is also necessary for mtDNA replication completion. Nucleic Acids Res. 2022, 50, 8749–8766. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Sun, R.; Eriksson, S. Basic biochemical characterization of cytosolic enzymes in thymidine nucleotide synthesis in adult rat tissues: Implications for tissue specific mitochondrial DNA depletion and deoxynucleoside-based therapy for TK2-deficiency. BMC Mol. Cell Biol. 2020, 21, 33. [Google Scholar] [CrossRef]
- Silva Ramos, E.; Motori, E.; Brüser, C.; Kühl, I.; Yeroslaviz, A.; Ruzzenente, B.; Kauppila, J.H.K.; Busch, J.D.; Hultenby, K.; Habermann, B.H.; et al. Mitochondrial fusion is required for regulation of mitochondrial DNA replication. PLoS Genet. 2019, 15, e1008085. [Google Scholar] [CrossRef]
- L’Ecuyer, T.; Sanjeev, S.; Thomas, R.; Novak, R.; Das, L.; Campbell, W.; Heide, R.V. DNA damage is an early event in doxorubicin-induced cardiac myocyte death. Am. J. Physiol. Heart Circ. Physiol. 2006, 291, H1273–H1280. [Google Scholar] [CrossRef]
- Lebrecht, D.; Walker, U.A. Role of mtDNA lesions in anthracycline cardiotoxicity. Cardiovasc. Toxicol. 2007, 7, 108–113. [Google Scholar] [CrossRef] [PubMed]
- Cui, L.; Guo, J.; Zhang, Q.; Yin, J.; Li, J.; Zhou, W.; Zhang, T.; Yuan, H.; Zhao, J.; Zhang, L.; et al. Erythropoietin activates SIRT1 to protect human cardiomyocytes against doxorubicin-induced mitochondrial dysfunction and toxicity. Toxicol. Lett. 2017, 275, 28–38. [Google Scholar] [CrossRef]
- Gilkerson, R.W.; Margineantu, D.H.; Capaldi, R.A.; Selker, J.M. Mitochondrial DNA depletion causes morphological changes in the mitochondrial reticulum of cultured human cells. FEBS Lett. 2000, 474, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Picca, A.; Guerra, F.; Calvani, R.; Coelho-Júnior, H.J.; Leeuwenburgh, C.; Bucci, C.; Marzetti, E. The contribution of mitochondrial DNA alterations to aging, cancer, and neurodegeneration. Exp. Gerontol. 2023, 178, 112203. [Google Scholar] [CrossRef] [PubMed]
- Wen, J.J.; Yin, Y.W.; Garg, N.J. PARP1 depletion improves mitochondrial and heart function in Chagas disease: Effects on POLG dependent mtDNA maintenance. PLoS Pathog. 2018, 14, e1007065. [Google Scholar] [CrossRef]
- Pisano, A.; Cerbelli, B.; Perli, E.; Pelullo, M.; Bargelli, V.; Preziuso, C.; Mancini, M.; He, L.; Bates, M.G.; Lucena, J.R.; et al. Impaired mitochondrial biogenesis is a common feature to myocardial hypertrophy and end-stage ischemic heart failure. Cardiovasc. Pathol. 2016, 25, 103–112. [Google Scholar] [CrossRef]
- Zhao, D.; Bartz, T.M.; Sotoodehnia, N.; Post, W.S.; Heckbert, S.R.; Alonso, A.; Longchamps, R.J.; Castellani, C.A.; Hong, Y.S.; Rotter, J.I.; et al. Mitochondrial DNA copy number and incident atrial fibrillation. BMC Med. 2020, 18, 246. [Google Scholar] [CrossRef]
- Huang, J.; Tan, L.; Shen, R.; Zhang, L.; Zuo, H.; Wang, D.W. Decreased Peripheral Mitochondrial DNA Copy Number is Associated with the Risk of Heart Failure and Long-term Outcomes. Medicine 2016, 95, e3323. [Google Scholar] [CrossRef]
- Lewis, W. Defective mitochondrial DNA replication and NRTIs: Pathophysiological implications in AIDS cardiomyopathy. Am. J. Physiol. Heart Circ. Physiol. 2003, 284, H1–H9. [Google Scholar] [CrossRef][Green Version]
- Poirier, M.C.; Gibbons, A.T.; Rugeles, M.T.; Andre-Schmutz, I.; Blanche, S. Fetal consequences of maternal antiretroviral nucleoside reverse transcriptase inhibitor use in human and nonhuman primate pregnancy. Curr. Opin. Pediatr. 2015, 27, 233–239. [Google Scholar] [CrossRef]
- Sabbah, H.N. Targeting the Mitochondria in Heart Failure: A Translational Perspective. JACC Basic Transl. Sci. 2020, 5, 88–106. [Google Scholar] [CrossRef] [PubMed]
- Berger, S.L.; Kouzarides, T.; Shiekhattar, R.; Shilatifard, A. An operational definition of epigenetics. Genes Dev. 2009, 23, 781–783. [Google Scholar] [CrossRef] [PubMed]
- Yasumizu, Y.; Hagiwara, M.; Umezu, Y.; Fuji, H.; Iwaisako, K.; Asagiri, M.; Uemoto, S.; Nakamura, Y.; Thul, S.; Ueyama, A.; et al. Neural-net-based cell deconvolution from DNA methylation reveals tumor microenvironment associated with cancer prognosis. NAR Cancer 2024, 6, zcae022. [Google Scholar] [CrossRef]
- Bellizzi, D.; D’Aquila, P.; Scafone, T.; Giordano, M.; Riso, V.; Riccio, A.; Passarino, G. The control region of mitochondrial DNA shows an unusual CpG and non-CpG methylation pattern. DNA Res. 2013, 20, 537–547. [Google Scholar] [CrossRef]
- Coppedè, F. One-carbon epigenetics and redox biology of neurodegeneration. Free Radic. Biol. Med. 2021, 170, 19–33. [Google Scholar] [CrossRef]
- Rehman, A.; Kumari, R.; Kamthan, A.; Tiwari, R.; Srivastava, R.K.; van der Westhuizen, F.H.; Mishra, P.K. Cell-free circulating mitochondrial DNA: An emerging biomarker for airborne particulate matter associated with cardiovascular diseases. Free Radic. Biol. Med. 2023, 195, 103–120. [Google Scholar] [CrossRef]
- Shock, L.S.; Thakkar, P.V.; Peterson, E.J.; Moran, R.G.; Taylor, S.M. DNA methyltransferase; cytosine methylation, and cytosine hydroxymethylation in mammalian mitochondria. Proc. Natl. Acad. Sci. USA 2011, 108, 3630–3635. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.F.; Zhu, J.J.; Tian, X.Y.; Liu, H.; Zhang, T.; Zhang, Y.P.; Xie, S.A.; Zheng, M.; Kong, W.; Yao, W.J.; et al. Hypermethylation of mitochondrial DNA in vascular smooth muscle cells impairs cell contractility. Cell Death Dis. 2020, 11, 35. [Google Scholar] [CrossRef]
- Boovarahan, S.R.; Ali, N.; AlAsmari, A.F.; Alameen, A.A.; Khan, R.; Kurian, G.A. Age-associated global DNA hypermethylation augments the sensitivity of hearts towards ischemia-reperfusion injury. Front. Genet. 2022, 13, 995887. [Google Scholar] [CrossRef]
- Ma, C.; Niu, R.; Huang, T.; Shao, L.W.; Peng, Y.; Ding, W.; Wang, Y.; Jia, G.; He, C.; Li, C.Y.; et al. N6-methyldeoxyadenine is a transgenerational epigenetic signal for mitochondrial stress adaptation. Nat. Cell Biol. 2019, 21, 319–327. [Google Scholar] [CrossRef]
- Hao, Z.; Wu, T.; Cui, X.; Zhu, P.; Tan, C.; Dou, X.; Hsu, K.W.; Lin, Y.T.; Peng, P.H.; Zhang, L.S.; et al. N(6)-Deoxyadenosine Methylation in Mammalian Mitochondrial DNA. Mol. Cell 2020, 78, 382–395.e8. [Google Scholar] [CrossRef]
- Sun, X.; Wang, Z.; Cong, X.; Lv, Y.; Li, Z.; Rong, L.; Yang, T.; Yu, D. Mitochondrial gene COX2 methylation and downregulation is a biomarker of aging in heart mesenchymal stem cells. Int. J. Mol. Med. 2021, 47, 161–170. [Google Scholar] [CrossRef]
- Baccarelli, A.A.; Byun, H.M. Platelet mitochondrial DNA methylation: A potential new marker of cardiovascular disease. Clin. Epigenetics. 2015, 7, 44. [Google Scholar] [CrossRef]
- Park, S.H.; Lee, S.Y.; Kim, S.A. Mitochondrial DNA Methylation Is Higher in Acute Coronary Syndrome Than in Stable Coronary Artery Disease. In Vivo 2021, 35, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Nakahira, K.; Haspel, J.A.; Rathinam, V.A.; Lee, S.J.; Dolinay, T.; Lam, H.C.; Englert, J.A.; Rabinovitch, M.; Cernadas, M.; Kim, H.P.; et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 2011, 12, 222–230. [Google Scholar] [CrossRef]
- Riley, J.S.; Quarato, G.; Cloix, C.; Lopez, J.; O’Prey, J.; Pearson, M.; Chapman, J.; Sesaki, H.; Carlin, L.M.; Passos, J.F.; et al. Mitochondrial inner membrane permeabilisation enables mtDNA release during apoptosis. EMBO J. 2018, 37, e99238. [Google Scholar] [CrossRef]
- Lindqvist, L.M.; Frank, D.; McArthur, K.; Dite, T.A.; Lazarou, M.; Oakhill, J.S.; Kile, B.T.; Vaux, D.L. Autophagy induced during apoptosis degrades mitochondria and inhibits type I interferon secretion. Cell Death Differ. 2018, 25, 784–796. [Google Scholar] [CrossRef]
- Hopfner, K.P.; Hornung, V. Molecular mechanisms and cellular functions of cGAS-STING signalling. Nat. Rev. Mol. Cell Biol. 2020, 21, 501–521. [Google Scholar] [CrossRef] [PubMed]
- McArthur, K.; Whitehead, L.W.; Heddleston, J.M.; Li, L.; Padman, B.S.; Oorschot, V.; Geoghegan, N.D.; Chappaz, S.; Davidson, S.; Chin, H.S.; et al. BAK/BAX macropores facilitate mitochondrial herniation and mtDNA efflux during apoptosis. Science 2018, 359, eaao6047. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Gupta, R.; Blanco, L.P.; Yang, S.; Shteinfer-Kuzmine, A.; Wang, K.; Zhu, J.; Yoon, H.E.; Wang, X.; Kerkhofs, M.; et al. VDAC oligomers form mitochondrial pores to release mtDNA fragments and promote lupus-like disease. Science 2019, 366, 1531–1536. [Google Scholar] [CrossRef]
- Kanneganti, T.D.; Kundu, M.; Green, D.R. Innate immune recognition of mtDNA--an undercover signal? Cell Metab. 2015, 21, 793–794. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Li, X.; Chen, X.; Zheng, L.; Chen, M.; Zhang, Y.; Zhu, R.; Chen, J.; Gu, J.; Yin, Q.; Jiang, H.; et al. Non-canonical STING-PERK pathway dependent epigenetic regulation of vascular endothelial dysfunction via integrating IRF3 and NF-κB in inflammatory response. Acta Pharm. Sin. B 2023, 13, 4765–4784. [Google Scholar] [CrossRef]
- Zecchini, V.; Paupe, V.; Herranz-Montoya, I.; Janssen, J.; Wortel, I.M.N.; Morris, J.L.; Ferguson, A.; Chowdury, S.R.; Segarra-Mondejar, M.; Costa, A.S.H.; et al. Fumarate induces vesicular release of mtDNA to drive innate immunity. Nature 2023, 615, 499–506. [Google Scholar] [CrossRef]
- Lu, R.X.Z.; Rafatian, N.; Zhao, Y.; Wagner, K.T.; Beroncal, E.L.; Li, B.; Lee, C.; Chen, J.; Churcher, E.; Vosoughi, D.; et al. Cardiac tissue model of immune-induced dysfunction reveals the role of free mitochondrial DNA and the therapeutic effects of exosomes. Sci. Adv. 2024, 10, eadk0164. [Google Scholar] [CrossRef] [PubMed]
- Xin, G.; Wei, Z.; Ji, C.; Zheng, H.; Gu, J.; Ma, L.; Huang, W.; Morris-Natschke, S.L.; Yeh, J.L.; Zhang, R.; et al. Metformin Uniquely Prevents Thrombosis by Inhibiting Platelet Activation and mtDNA Release. Sci. Rep. 2016, 6, 36222. [Google Scholar] [CrossRef]
- Xiong, W.; Li, B.; Pan, J.; Li, D.; Yuan, H.; Wan, X.; Zhang, Y.; Fu, L.; Zhang, J.; Lei, M.; et al. Mitochondrial Amount Determines Doxorubicin-Induced Cardiotoxicity in Cardiomyocytes. Adv. Sci. 2025, 12, e2412017. [Google Scholar] [CrossRef]
- Wiersma, M.; van Marion, D.M.S.; Bouman, E.J.; Li, J.; Zhang, D.; Ramos, K.S.; Lanters, E.A.H.; de Groot, N.M.S.; Brundel, B. Cell-Free Circulating Mitochondrial DNA: A Potential Blood-Based Marker for Atrial Fibrillation. Cells 2020, 9, 1159. [Google Scholar] [CrossRef]
- Liu, K.; Qiu, D.; Liang, X.; Huang, Y.; Wang, Y.; Jia, X.; Li, K.; Zhao, J.; Du, C.; Qiu, X.; et al. Lipotoxicity-induced STING1 activation stimulates MTORC1 and restricts hepatic lipophagy. Autophagy 2022, 18, 860–876. [Google Scholar] [CrossRef]
- Boone, C.; Lewis, S.C. Bridging lipid metabolism and mitochondrial genome maintenance. J. Biol. Chem. 2024, 300, 107498. [Google Scholar] [CrossRef]
- Lee, S.R.; Heo, J.H.; Jo, S.L.; Kim, G.; Kim, S.J.; Yoo, H.J.; Lee, K.P.; Kwun, H.J.; Shin, H.J.; Baek, I.J.; et al. Progesterone receptor membrane component 1 reduces cardiac steatosis and lipotoxicity via activation of fatty acid oxidation and mitochondrial respiration. Sci. Rep. 2021, 11, 8781. [Google Scholar] [CrossRef] [PubMed]
- Oka, T.; Hikoso, S.; Yamaguchi, O.; Taneike, M.; Takeda, T.; Tamai, T.; Oyabu, J.; Murakawa, T.; Nakayama, H.; Nishida, K.; et al. Mitochondrial DNA that escapes from autophagy causes inflammation and heart failure. Nature 2012, 485, 251–255. [Google Scholar] [CrossRef]
- Li, L.; Huang, T.; Yang, J.; Yang, P.; Lan, H.; Liang, J.; Cai, D.; Zhong, H.; Jiao, W.; Song, Y. PINK1/Parkin pathway-mediated mitophagy by AS-IV to explore the molecular mechanism of muscle cell damage. Biomed. Pharmacother. 2023, 161, 114533. [Google Scholar] [CrossRef]
- Onishi, M.; Yamano, K.; Sato, M.; Matsuda, N.; Okamoto, K. Molecular mechanisms and physiological functions of mitophagy. EMBO j. 2021, 40, e104705. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Wang, X.; Xu, T.; Gan, D.; Ma, Z.; Zhang, H.; Zhang, J.; Zeng, Q.; Xu, D. PINK1-mediated mitophagy attenuates pathological cardiac hypertrophy by suppressing the mtDNA release-activated cGAS-STING pathway. Cardiovasc. Res. 2025, 121, 128–142. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Zhang, H.; Xu, T.; Mei, X.; Wang, X.; Yang, Q.; Luo, Z.; Zeng, Q.; Xu, D.; Ren, H. Vericiguat attenuates doxorubicin-induced cardiotoxicity through the PRKG1/PINK1/STING axis. Transl. Res. 2024, 273, 90–103. [Google Scholar] [CrossRef]
- Yasheng, M.; Ji, Y.; He, Y.; Yi, Q.; Zhang, H.; Shan, W.; Wang, K.; Zhang, J.; Li, Y.; Meng, F.; et al. Leber’s hereditary optic neuropathy-associated ND1 3733G>C mutation ameliorates the mitochondrial quality control and cellular homeostasis. J. Biol. Chem. 2025, 301, 110464. [Google Scholar] [CrossRef]
- Lazarou, M.; Sliter, D.A.; Kane, L.A.; Sarraf, S.A.; Wang, C.; Burman, J.L.; Sideris, D.P.; Fogel, A.I.; Youle, R.J. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature 2015, 524, 309–314. [Google Scholar] [CrossRef] [PubMed]
- Poole, L.P.; Macleod, K.F. Mitophagy in tumorigenesis and metastasis. Cell. Mol. Life Sci. 2021, 78, 3817–3851. [Google Scholar] [CrossRef]
- Lampert, M.A.; Orogo, A.M.; Najor, R.H.; Hammerling, B.C.; Leon, L.J.; Wang, B.J.; Kim, T.; Sussman, M.A.; Gustafsson, Å.B. BNIP3L/NIX and FUNDC1-mediated mitophagy is required for mitochondrial network remodeling during cardiac progenitor cell differentiation. Autophagy 2019, 15, 1182–1198. [Google Scholar] [CrossRef]
- Mizushima, N.; Komatsu, M. Autophagy: Renovation of cells and tissues. Cell 2011, 147, 728–741. [Google Scholar] [CrossRef]
- Ravikumar, B.; Sarkar, S.; Davies, J.E.; Futter, M.; Garcia-Arencibia, M.; Green-Thompson, Z.W.; Jimenez-Sanchez, M.; Korolchuk, V.I.; Lichtenberg, M.; Luo, S.; et al. Regulation of mammalian autophagy in physiology and pathophysiology. Physiol. Rev. 2010, 90, 1383–1435. [Google Scholar] [CrossRef]
- Han, D.; Xu, T.; Lyu, X.; Li, K.; Xu, S. Smaller-Sized Silica Nanoparticles Exacerbated Cardiomyocyte Pyroptosis by Impairing Mitophagy to Activate mtDNA-cGAS-STING Signaling. J. Agric. Food Chem. 2025, 73, 9359–9369. [Google Scholar] [CrossRef]
- De Leo, M.G.; Staiano, L.; Vicinanza, M.; Luciani, A.; Carissimo, A.; Mutarelli, M.; Di Campli, A.; Polishchuk, E.; Di Tullio, G.; Morra, V.; et al. Autophagosome-lysosome fusion triggers a lysosomal response mediated by TLR9 and controlled by OCRL. Nat. Cell Biol. 2016, 18, 839–850. [Google Scholar] [CrossRef]
- Carchman, E.H.; Whelan, S.; Loughran, P.; Mollen, K.; Stratamirovic, S.; Shiva, S.; Rosengart, M.R.; Zuckerbraun, B.S. Experimental sepsis-induced mitochondrial biogenesis is dependent on autophagy, TLR4, and TLR9 signaling in liver. FASEB j. 2013, 27, 4703–4711. [Google Scholar] [CrossRef]
- Fang, R.; Jiang, Q.; Jia, X.; Jiang, Z. ARMH3-mediated recruitment of PI4KB directs Golgi-to-endosome trafficking and activation of the antiviral effector STING. Immunity 2023, 56, 500–515.e6. [Google Scholar] [CrossRef]
- Varela, Y.R.; Aguirre, C.C.; Iriondo, M.N.; Ballesteros, U.; Collado, M.I.; Etxaniz, A.; Montes, L.R.; Goñi, F.M.; Alonso, A. Ceramide and the membrane-fusion activity of LC3/GABARAP autophagy proteins. Cell. Mol. Life Sci. 2025, 82, 283. [Google Scholar] [CrossRef]
- Oduro, P.K.; Zheng, X.; Wei, J.; Yang, Y.; Wang, Y.; Zhang, H.; Liu, E.; Gao, X.; Du, M.; Wang, Q. The cGAS-STING signaling in cardiovascular and metabolic diseases: Future novel target option for pharmacotherapy. Acta Pharm. Sin. B 2022, 12, 50–75. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Zeng, X.; Li, X.; Mehta, J.L.; Wang, X. Role of NLRP3 inflammasome in the pathogenesis of cardiovascular diseases. Basic Res. Cardiol. 2018, 113, 5. [Google Scholar] [CrossRef]
- Yu, F.; Yang, L.; Zhang, R.; Hu, F.; Yuan, Y.; Wang, Z.; Yang, W. Low levels of supercoiled mitochondrial DNA are involved in heart failure induced by transverse aortic constriction in mice via an inflammatory response mediated by ZBP1. Exp. Cell Res. 2024, 442, 114187. [Google Scholar] [CrossRef] [PubMed]
- Enzan, N.; Matsushima, S.; Ikeda, S.; Okabe, K.; Ishikita, A.; Yamamoto, T.; Sada, M.; Miyake, R.; Tsutsui, Y.; Nishimura, R.; et al. ZBP1 Protects Against mtDNA-Induced Myocardial Inflammation in Failing Hearts. Circ. Res. 2023, 132, 1110–1126. [Google Scholar] [CrossRef] [PubMed]
- Bliksøen, M.; Mariero, L.H.; Torp, M.K.; Baysa, A.; Ytrehus, K.; Haugen, F.; Seljeflot, I.; Vaage, J.; Valen, G.; Stensløkken, K.O. Extracellular mtDNA activates NF-κB via toll-like receptor 9 and induces cell death in cardiomyocytes. Basic Res. Cardiol. 2016, 111, 42. [Google Scholar] [CrossRef]
- Takeuchi, O.; Akira, S. Pattern recognition receptors and inflammation. Cell 2010, 140, 805–820. [Google Scholar] [CrossRef]
- De Gaetano, A.; Solodka, K.; Zanini, G.; Selleri, V.; Mattioli, A.V.; Nasi, M.; Pinti, M. Molecular Mechanisms of mtDNA-Mediated Inflammation. Cells 2021, 1, e000100. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Raoof, M.; Chen, Y.; Sumi, Y.; Sursal, T.; Junger, W.; Brohi, K.; Itagaki, K.; Hauser, C.J. Circulating mitochondrial DAMPs cause inflammatory responses to injury. Nature 2010, 464, 104–107. [Google Scholar] [CrossRef] [PubMed]
- King, K.R.; Aguirre, A.D.; Ye, Y.X.; Sun, Y.; Roh, J.D.; Ng, R.P., Jr.; Kohler, R.H.; Arlauckas, S.P.; Iwamoto, Y.; Savol, A.; et al. IRF3 and type I interferons fuel a fatal response to myocardial infarction. Nat. Med. 2017, 23, 1481–1487. [Google Scholar] [CrossRef]
- Baik, S.H.; Ramanujan, V.K.; Becker, C.; Fett, S.; Underhill, D.M.; Wolf, A.J. Hexokinase dissociation from mitochondria promotes oligomerization of VDAC that facilitates NLRP3 inflammasome assembly and activation. Sci. Immunol. 2023, 8, eade7652. [Google Scholar] [CrossRef]
- Shimada, K.; Crother, T.R.; Karlin, J.; Dagvadorj, J.; Chiba, N.; Chen, S.; Ramanujan, V.K.; Wolf, A.J.; Vergnes, L.; Ojcius, D.M.; et al. Oxidized mitochondrial DNA activates the NLRP3 inflammasome during apoptosis. Immunity 2012, 36, 401–414. [Google Scholar] [CrossRef] [PubMed]
- Ablasser, A.; Chen, Z.J. cGAS in action: Expanding roles in immunity and inflammation. Science 2019, 363, eaat8657. [Google Scholar] [CrossRef]
- Ishikawa, H.; Barber, G.N. STING is an endoplasmic reticulum adaptor that facilitates innate immune signalling. Nature 2008, 455, 674–678. [Google Scholar] [CrossRef]
- Yan, X.; Huang, J.; Zeng, Y.; Zhong, X.; Fu, Y.; Xiao, H.; Wang, X.; Lian, H.; Luo, H.; Li, D.; et al. CGRP attenuates pulmonary vascular remodeling by inhibiting the cGAS-STING-NFκB pathway in pulmonary arterial hypertension. Biochem. Pharmacol. 2024, 222, 116093. [Google Scholar] [CrossRef]
- Han, W.; Du, C.; Zhu, Y.; Ran, L.; Wang, Y.; Xiong, J.; Wu, Y.; Lan, Q.; Wang, Y.; Wang, L.; et al. Targeting Myocardial Mitochondria-STING-Polyamine Axis Prevents Cardiac Hypertrophy in Chronic Kidney Disease. JACC Basic Transl. Sci. 2022, 7, 820–840. [Google Scholar] [CrossRef]
- McCarthy, C.G.; Wenceslau, C.F.; Goulopoulou, S.; Ogbi, S.; Baban, B.; Sullivan, J.C.; Matsumoto, T.; Webb, R.C. Circulating mitochondrial DNA and Toll-like receptor 9 are associated with vascular dysfunction in spontaneously hypertensive rats. Cardiovasc. Res. 2015, 107, 119–130. [Google Scholar] [CrossRef] [PubMed]
- Natarajan, N.; Florentin, J.; Johny, E.; Xiao, H.; O’Neil, S.P.; Lei, L.; Shen, J.; Ohayon, L.; Johnson, A.R.; Rao, K.; et al. Aberrant mitochondrial DNA synthesis in macrophages exacerbates inflammation and atherosclerosis. Nat. Commun. 2024, 15, 7337. [Google Scholar] [CrossRef]
- Bi, X.; Du, C.; Wang, X.; Wang, X.Y.; Han, W.; Wang, Y.; Qiao, Y.; Zhu, Y.; Ran, L.; Liu, Y.; et al. Mitochondrial Damage-Induced Innate Immune Activation in Vascular Smooth Muscle Cells Promotes Chronic Kidney Disease-Associated Plaque Vulnerability. Adv. Sci. 2021, 8, 2002738. [Google Scholar] [CrossRef]
- Zeng, W.; Wu, D.; Sun, Y.; Suo, Y.; Yu, Q.; Zeng, M.; Gao, Q.; Yu, B.; Jiang, X.; Wang, Y. The selective NLRP3 inhibitor MCC950 hinders atherosclerosis development by attenuating inflammation and pyroptosis in macrophages. Sci. Rep. 2021, 11, 19305. [Google Scholar] [CrossRef]
- Westerterp, M.; Fotakis, P.; Ouimet, M.; Bochem, A.E.; Zhang, H.; Molusky, M.M.; Wang, W.; Abramowicz, S.; la Bastide-van Gemert, S.; Wang, N.; et al. Cholesterol Efflux Pathways Suppress Inflammasome Activation, NETosis, and Atherogenesis. Circulation 2018, 138, 898–912. [Google Scholar] [CrossRef]
- Li, H.; Wang, X.; Luo, X.; Shi, H.; Li, J. cGAS/STING/NLRP3 Signaling Pathway-Mediated Pyroptosis in Hypertrophic Cardiomyopathy Radiotherapy. Front. Biosci. 2025, 30, 26084. [Google Scholar] [CrossRef]
- Tumurkhuu, G.; Shimada, K.; Dagvadorj, J.; Crother, T.R.; Zhang, W.; Luthringer, D.; Gottlieb, R.A.; Chen, S.; Arditi, M. Ogg1-Dependent DNA Repair Regulates NLRP3 Inflammasome and Prevents Atherosclerosis. Circ. Res. 2016, 119, e76–e90. [Google Scholar] [CrossRef]
- Ikeda, M.; Ide, T.; Fujino, T.; Arai, S.; Saku, K.; Kakino, T.; Tyynismaa, H.; Yamasaki, T.; Yamada, K.; Kang, D.; et al. Overexpression of TFAM or twinkle increases mtDNA copy number and facilitates cardioprotection associated with limited mitochondrial oxidative stress. PLoS ONE 2015, 10, e0119687. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Bei, Y.; Lu, Y.; Sun, W.; Liu, Q.; Wang, Y.; Cao, Y.; Chen, P.; Xiao, J.; Kong, X. Exercise Prevents Cardiac Injury and Improves Mitochondrial Biogenesis in Advanced Diabetic Cardiomyopathy with PGC-1α and Akt Activation. Cell. Physiol. Biochem. 2015, 35, 2159–2168. [Google Scholar] [CrossRef] [PubMed]
- Bliksøen, M.; Baysa, A.; Eide, L.; Bjørås, M.; Suganthan, R.; Vaage, J.; Stensløkken, K.O.; Valen, G. Mitochondrial DNA damage and repair during ischemia-reperfusion injury of the heart. J. Mol. Cell. Cardiol. 2015, 78, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Byun, K.A.; Oh, S.; Yang, J.Y.; Lee, S.Y.; Son, K.H.; Byun, K. Ecklonia cava extracts decrease hypertension-related vascular calcification by modulating PGC-1α and SOD2. Biomed. Pharmacother. 2022, 153, 113283. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Li, X.; Sun, X.; Xiao, S.; Chen, T.; Ren, L.; Liu, N. STING1-accelerated vascular smooth muscle cell senescence-associated vascular calcification in diabetes is ameliorated by oleoylethanolamide via improved mitochondrial DNA oxidative damage. Free Radic. Biol. Med. 2024, 222, 437–455. [Google Scholar] [CrossRef]
- Pohjoismäki, J.L.; Williams, S.L.; Boettger, T.; Goffart, S.; Kim, J.; Suomalainen, A.; Moraes, C.T.; Braun, T. Overexpression of Twinkle-helicase protects cardiomyocytes from genotoxic stress caused by reactive oxygen species. Proc. Natl. Acad. Sci. USA 2013, 110, 19408–19413. [Google Scholar] [CrossRef]
- Li, H.; Zhang, X.; Wang, F.; Zhou, L.; Yin, Z.; Fan, J.; Nie, X.; Wang, P.; Fu, X.D.; Chen, C.; et al. MicroRNA-21 Lowers Blood Pressure in Spontaneous Hypertensive Rats by Upregulating Mitochondrial Translation. Circulation 2016, 134, 734–751. [Google Scholar] [CrossRef]
- Cividini, F.; Scott, B.T.; Dai, A.; Han, W.; Suarez, J.; Diaz-Juarez, J.; Diemer, T.; Casteel, D.E.; Dillmann, W.H. O-GlcNAcylation of 8-Oxoguanine DNA Glycosylase (Ogg1) Impairs Oxidative Mitochondrial DNA Lesion Repair in Diabetic Hearts. J. Biol. Chem. 2016, 291, 26515–26528. [Google Scholar] [CrossRef]
- Dobson, A.W.; Xu, Y.; Kelley, M.R.; LeDoux, S.P.; Wilson, G.L. Enhanced mitochondrial DNA repair and cellular survival after oxidative stress by targeting the human 8-oxoguanine glycosylase repair enzyme to mitochondria. J. Biol. Chem. 2000, 275, 37518–37523. [Google Scholar] [CrossRef]
- Wang, J.; Wang, Q.; Watson, L.J.; Jones, S.P.; Epstein, P.N. Cardiac overexpression of 8-oxoguanine DNA glycosylase 1 protects mitochondrial DNA and reduces cardiac fibrosis following transaortic constriction. Am. J. Physiol. Heart Circ. Physiol. 2011, 301, H2073–H2080. [Google Scholar] [CrossRef]
- Bradley, J.M.; Li, Z.; Organ, C.L.; Polhemus, D.J.; Otsuka, H.; Islam, K.N.; Bhushan, S.; Gorodnya, O.M.; Ruchko, M.V.; Gillespie, M.N.; et al. A novel mtDNA repair fusion protein attenuates maladaptive remodeling and preserves cardiac function in heart failure. Am. J. Physiol. Heart Circ. Physiol. 2018, 314, H311–H321. [Google Scholar] [CrossRef]
- Croteau, D.L.; Stierum, R.H.; Bohr, V.A. Mitochondrial DNA repair pathways. Mutat. Res. 1999, 434, 137–148. [Google Scholar] [CrossRef] [PubMed]
- Van Houten, B.; Woshner, V.; Santos, J.H. Role of mitochondrial DNA in toxic responses to oxidative stress. DNA Repair 2006, 5, 145–152. [Google Scholar] [CrossRef]
- Han, C.; Qian, X.; Ren, X.; Zhang, S.; Hu, L.; Li, J.; Huang, Y.; Huang, R.; Ooi, K.; Lin, H.; et al. Inhibition of cGAS in Paraventricular Nucleus Attenuates Hypertensive Heart Injury Via Regulating Microglial Autophagy. Mol. Neurobiol. 2022, 59, 7006–7024. [Google Scholar] [CrossRef]
- Luo, G.; Chen, L.; Chen, M.; Mao, L.; Zeng, Q.; Zou, Y.; Xue, J.; Liu, P.; Wu, Q.; Yang, S.; et al. Hirudin inhibit the formation of NLRP3 inflammasome in cardiomyocytes via suppressing oxidative stress and activating mitophagy. Heliyon 2024, 10, e23077. [Google Scholar] [CrossRef] [PubMed]
- Yue, R.; Xia, X.; Jiang, J.; Yang, D.; Han, Y.; Chen, X.; Cai, Y.; Li, L.; Wang, W.E.; Zeng, C. Mitochondrial DNA oxidative damage contributes to cardiomyocyte ischemia/reperfusion-injury in rats: Cardioprotective role of lycopene. J. Cell. Physiol. 2015, 230, 2128–2141. [Google Scholar] [CrossRef]
- Lin, F.; Tan, Y.Q.; He, X.H.; Guo, L.L.; Wei, B.J.; Li, J.P.; Chen, Z.; Chen, H.W.; Wang, J. Huoxue Huatan Decoction Ameliorates Myocardial Ischemia/Reperfusion Injury in Hyperlipidemic Rats via PGC-1α-PPARα and PGC-1α-NRF1-mtTFA Pathways. Front. Pharmacol. 2020, 11, 546825. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.; Hong, S.G.; Choi, S.Y.; Rath, M.E.; Saredy, J.; Jovin, D.G.; Sayoc, J.; Park, H.S.; Eguchi, S.; Rizzo, V.; et al. Flow-induced endothelial mitochondrial remodeling mitigates mitochondrial reactive oxygen species production and promotes mitochondrial DNA integrity in a p53-dependent manner. Redox Biol. 2022, 50, 102252. [Google Scholar] [CrossRef]
- Zhang, M.; Feng, N.; Peng, Z.; Thapa, D.; Stoner, M.W.; Manning, J.R.; McTiernan, C.F.; Yang, X.; Jurczak, M.J.; Guimaraes, D.; et al. Reduced acetylation of TFAM promotes bioenergetic dysfunction in the failing heart. iScience 2023, 26, 106942. [Google Scholar] [CrossRef] [PubMed]
- Yao, K.; Zhang, W.W.; Yao, L.; Yang, S.; Nie, W.; Huang, F. Carvedilol promotes mitochondrial biogenesis by regulating the PGC-1/TFAM pathway in human umbilical vein endothelial cells (HUVECs). Biochem. Biophys. Res. Commun. 2016, 470, 961–966. [Google Scholar] [CrossRef] [PubMed]
- Fujino, T.; Ide, T.; Yoshida, M.; Onitsuka, K.; Tanaka, A.; Hata, Y.; Nishida, M.; Takehara, T.; Kanemaru, T.; Kitajima, N.; et al. Recombinant mitochondrial transcription factor A protein inhibits nuclear factor of activated T cells signaling and attenuates pathological hypertrophy of cardiac myocytes. Mitochondrion 2012, 12, 449–458. [Google Scholar] [CrossRef]
- He, L.; Chu, Y.; Yang, J.; He, J.; Hua, Y.; Chen, Y.; Benavides, G.; Rowe, G.C.; Zhou, L.; Ballinger, S.; et al. Activation of Autophagic Flux Maintains Mitochondrial Homeostasis during Cardiac Ischemia/Reperfusion Injury. Cells 2022, 11, 2111. [Google Scholar] [CrossRef]
- Shi, X.; Li, Y.; Wang, Y.; Ding, T.; Zhang, X.; Wu, N. Pharmacological postconditioning with sappanone A ameliorates myocardial ischemia reperfusion injury and mitochondrial dysfunction via AMPK-mediated mitochondrial quality control. Toxicol. Appl. Pharmacol. 2021, 427, 115668. [Google Scholar] [CrossRef] [PubMed]
- Chu, Y.; Hua, Y.; He, L.; He, J.; Chen, Y.; Yang, J.; Mahmoud, I.; Zeng, F.; Zeng, X.; Benavides, G.A.; et al. β-hydroxybutyrate administered at reperfusion reduces infarct size and preserves cardiac function by improving mitochondrial function through autophagy in male mice. J. Mol. Cell. Cardiol. 2024, 186, 31–44. [Google Scholar] [CrossRef]
- Yang, J.; He, J.; Ismail, M.; Tweeten, S.; Zeng, F.; Gao, L.; Ballinger, S.; Young, M.; Prabhu, S.D.; Rowe, G.C.; et al. HDAC inhibition induces autophagy and mitochondrial biogenesis to maintain mitochondrial homeostasis during cardiac ischemia/reperfusion injury. J. Mol. Cell. Cardiol. 2019, 130, 36–48. [Google Scholar] [CrossRef]
- Liu, X.; Ye, B.; Miller, S.; Yuan, H.; Zhang, H.; Tian, L.; Nie, J.; Imae, R.; Arai, H.; Li, Y.; et al. Ablation of ALCAT1 mitigates hypertrophic cardiomyopathy through effects on oxidative stress and mitophagy. Mol. Cell. Biol. 2012, 32, 4493–4504. [Google Scholar] [CrossRef]
- Lu, Y.; Li, S.; Wu, H.; Bian, Z.; Xu, J.; Gu, C.; Chen, X.; Yang, D. Beneficial effects of astragaloside IV against angiotensin II-induced mitochondrial dysfunction in rat vascular smooth muscle cells. Int. J. Mol. Med. 2015, 36, 1223–1232. [Google Scholar] [CrossRef] [PubMed]
- Kuno, A.; Hosoda, R.; Sebori, R.; Hayashi, T.; Sakuragi, H.; Tanabe, M.; Horio, Y. Resveratrol Ameliorates Mitophagy Disturbance and Improves Cardiac Pathophysiology of Dystrophin-deficient mdx Mice. Sci. Rep. 2018, 8, 15555. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Wang, X.; Hou, C.; Yang, L.; Li, H.; Guo, J.; Huo, C.; Wang, M.; Miao, Y.; Liu, J.; et al. Oleuropein improves mitochondrial function to attenuate oxidative stress by activating the Nrf2 pathway in the hypothalamic paraventricular nucleus of spontaneously hypertensive rats. Neuropharmacology 2017, 113, 556–566. [Google Scholar] [CrossRef]
- Chang, M.M.; Chu, D.T.; Lin, S.C.; Lee, J.S.; Vu, T.D.; Vu, H.T.; Ramasamy, T.S.; Lin, S.P.; Wu, C.C. Enhanced mitochondrial function and delivery from adipose-derived stem cell spheres via the EZH2-H3K27me3-PPARγ pathway for advanced therapy. Stem Cell Res. Ther. 2025, 16, 129. [Google Scholar] [CrossRef]
- Chen, Z.; Sun, X.; Li, X.; Liu, N. Oleoylethanolamide alleviates hyperlipidaemia-mediated vascular calcification via attenuating mitochondrial DNA stress triggered autophagy-dependent ferroptosis by activating PPARα. Biochem. Pharmacol. 2023, 208, 115379. [Google Scholar] [CrossRef]
- Sung, J.Y.; Kim, S.G.; Kang, Y.J.; Park, S.Y.; Choi, H.C. SIRT1-dependent PGC-1α deacetylation by SRT1720 rescues progression of atherosclerosis by enhancing mitochondrial function. Biochim. Biophys. Acta Mol. Cell Biol. Lipids 2024, 1869, 159453. [Google Scholar] [CrossRef]
- Zhang, H.; Komatsu, T.; Nakashima, S.; Abe, S.; Kawachi, E.; Imaizumi, S.; Saku, K.; Uehara, Y. HDL Mimetics Enhances Mitochondrial Function via Stimulation of PGC1-α. Eur. Cardiol. 2023, 18, e27. [Google Scholar] [CrossRef]
- Sposito, A.C.; de Lima-Junior, J.C.; Moura, F.A.; Barreto, J.; Bonilha, I.; Santana, M.; Virginio, V.W.; Sun, L.; Carvalho, L.S.F.; Soares, A.A.S.; et al. Reciprocal Multifaceted Interaction Between HDL (High-Density Lipoprotein) and Myocardial Infarction. Arter. Thromb. Vasc. Biol. 2019, 39, 1550–1564. [Google Scholar] [CrossRef]
- Zhao, Y.; Wang, Y.; Zheng, H.; Xu, Q.; Zhou, K.; Liu, H.; Xia, Y.; Wei, D.H.; Jiang, M.; Tang, Z.H.; et al. Hydrogen sulfide upregulates SIRT1 to inhibit ox-HDL-induced endothelial cell damage and mitochondrial dysfunction. Nitric Oxide 2024, 152, 78–89. [Google Scholar] [CrossRef]
- Huang, J.; Tao, H.; Yancey, P.G.; Leuthner, Z.; May-Zhang, L.S.; Jung, J.Y.; Zhang, Y.; Ding, L.; Amarnath, V.; Liu, D.; et al. Scavenging dicarbonyls with 5’-O-pentyl-pyridoxamine increases HDL net cholesterol efflux capacity and attenuates atherosclerosis and insulin resistance. Mol. Metab. 2023, 67, 101651. [Google Scholar] [CrossRef]
- Jenkins, A.J.; Carroll, L.M.; Huang, M.L.H.; Wen-Loh, Y.; Mangani, A.; O’Neal, D.N.; Januszewski, A.S. Mitochondrial DNA copy number in adults with and without Type 1 diabetes. Diabetes Res. Clin. Pract. 2023, 203, 110877. [Google Scholar] [CrossRef] [PubMed]
- Agius, R.; Pace, N.P.; Fava, S. Reduced leukocyte mitochondrial copy number in metabolic syndrome and metabolically healthy obesity. Front. Endocrinol. 2022, 13, 886957. [Google Scholar] [CrossRef]
- Tian, X.; He, W.; Yang, R.; Liu, Y. Dl-3-n-butylphthalide protects the heart against ischemic injury and H9c2 cardiomyoblasts against oxidative stress: Involvement of mitochondrial function and biogenesis. J. Biomed. Sci. 2017, 24, 38. [Google Scholar] [CrossRef]
- Zeng, Z.; Yu, K.; Hu, W.; Cheng, S.; Gao, C.; Liu, F.; Chen, J.; Kong, M.; Zhang, F.; Liu, X.; et al. SRT1720 Pretreatment Promotes Mitochondrial Biogenesis of Aged Human Mesenchymal Stem Cells and Improves Their Engraftment in Postinfarct Nonhuman Primate Hearts. Stem Cells Dev. 2021, 30, 386–398. [Google Scholar] [CrossRef]
- Inoue, T.; Ikeda, M.; Ide, T.; Fujino, T.; Matsuo, Y.; Arai, S.; Saku, K.; Sunagawa, K. Twinkle overexpression prevents cardiac rupture after myocardial infarction by alleviating impaired mitochondrial biogenesis. Am. J. Physiol. Heart Circ. Physiol. 2016, 311, H509–H519. [Google Scholar] [CrossRef] [PubMed]
- Hua, R.; Liu, B.; He, W.; Zhang, H.; Liu, Y.; Xie, Q.; Zhou, L.; Pei, F. Calcitriol reverses age-related hypertension via downregulating renal AP1/AT(1)R pathway through regulating mitochondrial function. Clin. Exp. Hypertens. 2023, 45, 2277653. [Google Scholar] [CrossRef] [PubMed]
- Gu, Q.; Wang, B.; Zhang, X.F.; Ma, Y.P.; Liu, J.D.; Wang, X.Z. Chronic aerobic exercise training attenuates aortic stiffening and endothelial dysfunction through preserving aortic mitochondrial function in aged rats. Exp. Gerontol. 2014, 56, 37–44. [Google Scholar] [CrossRef]
- Tanaka, A.; Ide, T.; Fujino, T.; Onitsuka, K.; Ikeda, M.; Takehara, T.; Hata, Y.; Ylikallio, E.; Tyynismaa, H.; Suomalainen, A.; et al. The overexpression of Twinkle helicase ameliorates the progression of cardiac fibrosis and heart failure in pressure overload model in mice. PLoS ONE 2013, 8, e67642. [Google Scholar] [CrossRef]
- Khiati, S.; Rosa, I.D.; Sourbier, C.; Ma, X.; Rao, V.A.; Neckers, L.M.; Zhang, H.; Pommier, Y. Mitochondrial topoisomerase I (top1mt) is a novel limiting factor of doxorubicin cardiotoxicity. Clin. Cancer Res. 2014, 20, 4873–4881. [Google Scholar] [CrossRef] [PubMed]
- Tao, L.; Wang, L.; Yang, X.; Jiang, X.; Hua, F. Recombinant human glucagon-like peptide-1 protects against chronic intermittent hypoxia by improving myocardial energy metabolism and mitochondrial biogenesis. Mol. Cell. Endocrinol. 2019, 481, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Dantham, S.; Srivastava, A.K.; Gulati, S.; Rajeswari, M.R. Plasma circulating cell-free mitochondrial DNA in the assessment of Friedreich’s ataxia. J. Neurol. Sci. 2016, 365, 82–88. [Google Scholar] [CrossRef]
- Yan, H.; Li, Y.; Peng, X.; Huang, D.; Gui, L.; Huang, B. Resistance of mitochondrial DNA-depleted cells against oxidized low-density lipoprotein-induced macrophage pyroptosis. Mol. Med. Rep. 2016, 13, 4393–4399. [Google Scholar] [CrossRef]
- Boovarahan, S.R.; Kurian, G.A. Investigating the role of DNMT1 gene expression on myocardial ischemia reperfusion injury in rat and associated changes in mitochondria. Biochim. Biophys. Acta Bioenerg. 2022, 1863, 148566. [Google Scholar] [CrossRef]
- Qin, C.Y.; Zhang, H.W.; Gu, J.; Xu, F.; Liang, H.M.; Fan, K.J.; Shen, J.Y.; Xiao, Z.H.; Zhang, E.Y.; Hu, J. Mitochondrial DNA-induced inflammatory damage contributes to myocardial ischemia reperfusion injury in rats: Cardioprotective role of epigallocatechin. Mol. Med. Rep. 2017, 16, 7569–7576. [Google Scholar] [CrossRef] [PubMed]
- Mariero, L.H.; Torp, M.K.; Heiestad, C.M.; Baysa, A.; Li, Y.; Valen, G.; Vaage, J.; Stensløkken, K.O. Inhibiting nucleolin reduces inflammation induced by mitochondrial DNA in cardiomyocytes exposed to hypoxia and reoxygenation. Br. J. Pharmacol. 2019, 176, 4360–4372. [Google Scholar] [CrossRef]
- Mahmoud, D.S.E.; Kamel, M.A.; El-Sayed, I.E.; Binsuwaidan, R.; Elmongy, E.I.; Razzaq, M.K.; Eldaim, M.A.A.; Ahmed, E.; Shaker, S.A. Astaxanthin ameliorated isoproterenol induced myocardial infarction via improving the mitochondrial function and antioxidant activity in rats. J. Biochem. Mol. Toxicol. 2024, 38, e23804. [Google Scholar] [CrossRef]
- Khalifa, A.A.; Rashad, R.M.; El-Hadidy, W.F. Thymoquinone protects against cardiac mitochondrial DNA loss, oxidative stress, inflammation and apoptosis in isoproterenol-induced myocardial infarction in rats. Heliyon 2021, 7, e07561. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Chen, T.; Li, J.; Wang, Y.; Shi, H.; Yu, Y.; Ji, Q.; Shen, X.; Sun, T.; Shi, H.; et al. IL-37 ameliorates myocardial fibrosis by regulating mtDNA-enriched vesicle release in diabetic cardiomyopathy mice. J. Transl. Med. 2024, 22, 494. [Google Scholar] [CrossRef]
- Zhao, T.; Jin, K.; Wang, X.; Su, X.; Wang, Y.; Gao, M.; Luo, W.; Yang, H.; Yang, Z. GPAT4 sustains endoplasmic reticulum homeostasis in endocardial cells and safeguards heart development. Nat. Commun. 2025, 16, 3345. [Google Scholar] [CrossRef]
- Qi, H.; Soto-Gonzalez, L.; Krychtiuk, K.A.; Ruhittel, S.; Kaun, C.; Speidl, W.S.; Kiss, A.; Podesser, B.K.; Yao, S.; Markstaller, K.; et al. Pretreatment With Argon Protects Human Cardiac Myocyte-Like Progenitor Cells from Oxygen Glucose Deprivation-Induced Cell Death by Activation of AKT and Differential Regulation of Mapkinases. Shock 2018, 49, 556–563. [Google Scholar] [CrossRef]
- Lei, Y.; VanPortfliet, J.J.; Chen, Y.F.; Bryant, J.D.; Li, Y.; Fails, D.; Torres-Odio, S.; Ragan, K.B.; Deng, J.; Mohan, A.; et al. Cooperative sensing of mitochondrial DNA by ZBP1 and cGAS promotes cardiotoxicity. Cell 2023, 186, 3013–3032.e22. [Google Scholar] [CrossRef]
- Moya, G.E.; Rivera, P.D.; Dittenhafer-Reed, K.E. Evidence for the Role of Mitochondrial DNA Release in the Inflammatory Response in Neurological Disorders. Int. J. Mol. Sci. 2021, 22, 7030. [Google Scholar] [CrossRef] [PubMed]
- Fei, Q.; Ma, H.; Zou, J.; Wang, W.; Zhu, L.; Deng, H.; Meng, M.; Tan, S.; Zhang, H.; Xiao, X.; et al. Metformin protects against ischaemic myocardial injury by alleviating autophagy-ROS-NLRP3-mediated inflammatory response in macrophages. J. Mol. Cell. Cardiol. 2020, 145, 1–13. [Google Scholar] [CrossRef]
- Xu, Z.; Pan, Z.; Jin, Y.; Gao, Z.; Jiang, F.; Fu, H.; Chen, X.; Zhang, X.; Yan, H.; Yang, X.; et al. Inhibition of PRKAA/AMPK (Ser485/491) phosphorylation by crizotinib induces cardiotoxicity via perturbing autophagosome-lysosome fusion. Autophagy 2024, 20, 416–436. [Google Scholar] [CrossRef]
- Zhang, J.; Liu, X.; Nie, J.; Shi, Y. Restoration of mitophagy ameliorates cardiomyopathy in Barth syndrome. Autophagy 2022, 18, 2134–2149. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Liao, W.; Wang, Y.; Liao, J.; Chen, N.; Jia, C.; Zeng, L. Human adipose-derived stem cell exosomes reduce mitochondrial DNA common deletion through PINK1/Parkin-mediated mitophagy to improve skin photoaging. Stem Cell Res. Ther. 2025, 16, 365. [Google Scholar] [CrossRef]
- Choi, D.H.; Lee, S.M.; Park, B.N.; Lee, M.H.; Yang, D.E.; Son, Y.K.; Kim, S.E.; An, W.S. Omega-3 Fatty Acids Modify Drp1 Expression and Activate the PINK1-Dependent Mitophagy Pathway in the Kidney and Heart of Adenine-Induced Uremic Rats. Biomedicines 2024, 12, 2107. [Google Scholar] [CrossRef]
- Wang, S.; Wang, L.; Qin, X.; Turdi, S.; Sun, D.; Culver, B.; Reiter, R.J.; Wang, X.; Zhou, H.; Ren, J. ALDH2 contributes to melatonin-induced protection against APP/PS1 mutation-prompted cardiac anomalies through cGAS-STING-TBK1-mediated regulation of mitophagy. Signal Transduct. Target. Ther. 2020, 5, 119. [Google Scholar] [CrossRef]
- Liu, Y.; Zhang, Y.; Zhu, H.; Shen, W.; Chen, Z.; Bai, J.; Shuang, T.; Chen, Q. Aucubin administration suppresses STING signaling and mitigated high-fat diet-induced atherosclerosis and steatohepatosis in LDL receptor deficient mice. Food Chem. Toxicol. 2022, 169, 113422. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Z.H.; Wang, J.J.; Lin, J.G.; Ye, W.L.; Zou, J.M.; Liang, L.Y.; Yang, P.L.; Qiu, W.L.; Li, Y.Y.; Yang, S.J.; et al. Cytosolic DNA initiates a vicious circle of aging-related endothelial inflammation and mitochondrial dysfunction via STING: The inhibitory effect of Cilostazol. Acta Pharmacol. Sin. 2024, 45, 1879–1897. [Google Scholar] [CrossRef]
- Qin, A.; Wen, Z.; Xiong, S. Myocardial Mitochondrial DNA Drives Macrophage Inflammatory Response through STING Signaling in Coxsackievirus B3-Induced Viral Myocarditis. Cells 2023, 12, 2555. [Google Scholar] [CrossRef]
- Zheng, Z.; Sun, J.; Wang, J.; He, S.; Liu, Z.; Xie, J.; Yu, C.Y.; Wei, H. Enhancing myocardial infarction treatment through bionic hydrogel-mediated spatial combination therapy via mtDNA-STING crosstalk modulation. J. Control Release 2024, 371, 570–587. [Google Scholar] [CrossRef]
- Jiménez-Loygorri, J.I.; Villarejo-Zori, B.; Viedma-Poyatos, Á.; Zapata-Muñoz, J.; Benítez-Fernández, R.; Frutos-Lisón, M.D.; Tomás-Barberán, F.A.; Espín, J.C.; Area-Gómez, E.; Gomez-Duran, A.; et al. Mitophagy curtails cytosolic mtDNA-dependent activation of cGAS/STING inflammation during aging. Nat. Commun. 2024, 15, 830. [Google Scholar] [CrossRef]
- Wu, B.; Ni, H.; Li, J.; Zhuang, X.; Zhang, J.; Qi, Z.; Chen, Q.; Wen, Z.; Shi, H.; Luo, X.; et al. The Impact of Circulating Mitochondrial DNA on Cardiomyocyte Apoptosis and Myocardial Injury After TLR4 Activation in Experimental Autoimmune Myocarditis. Cell Physiol. Biochem. 2017, 42, 713–728. [Google Scholar] [CrossRef]
- Zhang, Y.; Liu, X.; Bai, X.; Lin, Y.; Li, Z.; Fu, J.; Li, M.; Zhao, T.; Yang, H.; Xu, R.; et al. Melatonin prevents endothelial cell pyroptosis via regulation of long noncoding RNA MEG3/miR-223/NLRP3 axis. J. Pineal Res. 2018, 64, e12449. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Lu, L.; Zhong, X.; Yue, Y.; Hong, Y.; Li, Y.; Li, Y. Metformin reduced NLRP3 inflammasome activity in Ox-LDL stimulated macrophages through adenosine monophosphate activated protein kinase and protein phosphatase 2A. Eur. J. Pharmacol. 2019, 852, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Bai, H.; Xu, S.L.; Shi, J.J.; Ding, Y.P.; Liu, Q.Q.; Jiang, C.H.; He, L.L.; Zhang, H.R.; Lu, S.F.; Gu, Y.H. Electroacupuncture preconditioning protects against myocardial ischemia-reperfusion injury by modulating dynamic inflammatory response. Heliyon 2023, 9, e19396. [Google Scholar] [CrossRef]
- Zhang, Y.; Lv, Y.; Zhang, Q.; Wang, X.; Han, Q.; Liang, Y.; He, S.; Yuan, Q.; Zheng, J.; Xu, C.; et al. ALDH2 attenuates myocardial pyroptosis through breaking down Mitochondrion-NLRP3 inflammasome pathway in septic shock. Front. Pharmacol. 2023, 14, 1125866. [Google Scholar] [CrossRef]
- Wu, Z.M.; Kan, J.; Ye, F.; You, W.; Wu, X.Q.; Tian, N.L.; Lin, S.; Ge, Z.; Liu, Z.Z.; Li, X.B.; et al. PCSK9 inhibitor added to high-intensity statin therapy to prevent cardiovascular events in patients with acute coronary syndrome after percutaneous coronary intervention: A randomized, double- blind, placebo-controlled, multicenter SHAWN study. Am. Heart J. 2024, 277, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.H.; Lee, Y.Y.; Jung, J.S.; Hyun, S.H.; Lee, J.; Park, J.H.; Park, S.K.; Lee, S.H. Cholesterol-modulating effects of Korean red ginseng (KRG) targeting PCSK9 in hyperlipidemia. Sci. Rep. 2025, 15, 30637. [Google Scholar] [CrossRef]
- Liu, G.; Zhao, B.; Chen, Q.; Li, X.; Zhu, X.; Duan, M.; Zhang, M.; Liu, Z.; Wen, X.; Guo, J.; et al. HIIT and MICT mitigate endothelial dysfunction in early atherosclerotic mice via PCSK9 inhibition. Sci. Rep. 2025, 15, 30411. [Google Scholar] [CrossRef] [PubMed]
- Cirigliano, A.; Amelina, A.; Biferali, B.; Macone, A.; Mozzetta, C.; Bianchi, M.M.; Mori, M.; Botta, B.; Pick, E.; Negri, R.; et al. Statins interfere with the attachment of S. cerevisiae mtDNA to the inner mitochondrial membrane. J. Enzym. Inhib. Med. Chem. 2020, 35, 129–137. [Google Scholar] [CrossRef]
- Stringer, H.A.; Sohi, G.K.; Maguire, J.A.; Côté, H.C. Decreased skeletal muscle mitochondrial DNA in patients with statin-induced myopathy. J. Neurol. Sci. 2013, 325, 142–147. [Google Scholar] [CrossRef] [PubMed]
- Tavintharan, S.; Ong, C.N.; Jeyaseelan, K.; Sivakumar, M.; Lim, S.C.; Sum, C.F. Reduced mitochondrial coenzyme Q10 levels in HepG2 cells treated with high-dose simvastatin: A possible role in statin-induced hepatotoxicity? Toxicol. Appl. Pharmacol. 2007, 223, 173–179. [Google Scholar] [CrossRef]
- Ruiz-Pesini, E.; Bayona-Bafaluy, M.P.; Sanclemente, T.; Puzo, J.; Montoya, J.; Pacheu-Grau, D. Mitochondrial Genetic Background May Impact Statins Side Effects and Atherosclerosis Development in Familial Hypercholesterolemia. Int. J. Mol. Sci. 2022, 24, 471. [Google Scholar] [CrossRef]
- Bortolin, R.H.; de Souza Leite, F.; Luchessi, A.D.; Esposito, J.; Barbosa, I.N.; Freitas, R.C.C.; Sonawane, A.R.; Singh, S.A.; Aikawa, E.; Telles-Silva, K.A.; et al. Translational insights into statin-induced myotoxicity: Differential impact of lipophilic and hydrophilic statins on iPSC-derived skeletal muscle cells from patients with familial hypercholesterolemia. Toxicology 2025, 515, 154159. [Google Scholar] [CrossRef]
- Anan, R.; Nakagawa, M.; Miyata, M.; Higuchi, I.; Nakao, S.; Suehara, M.; Osame, M.; Tanaka, H. Cardiac involvement in mitochondrial diseases. A study on 17 patients with documented mitochondrial DNA defects. Circulation 1995, 91, 955–961. [Google Scholar] [CrossRef] [PubMed]
- Guichard, J.L.; Kane, M.S.; Grenett, M.; Sandel, M.; Benavides, G.A.; Bradley, W.E.; Powell, P.C.; Darley-Usmar, V.; Ballinger, S.W.; Dell, L.J. Mitochondrial haplotype modulates genome expression and mitochondrial structure/function in cardiomyocytes following volume overload. Am. J. Physiol. Heart Circ. Physiol. 2023, 324, H484–H493. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).