Protein Engineering of Electron Transfer Components from Electroactive Geobacter Bacteria
Abstract
1. Introduction
2. Thermodynamic Characterization of MCs
2.1. Redox Titrations Followed by Visible Spectroscopy
2.2. Redox Titrations Followed by NMR Spectroscopy
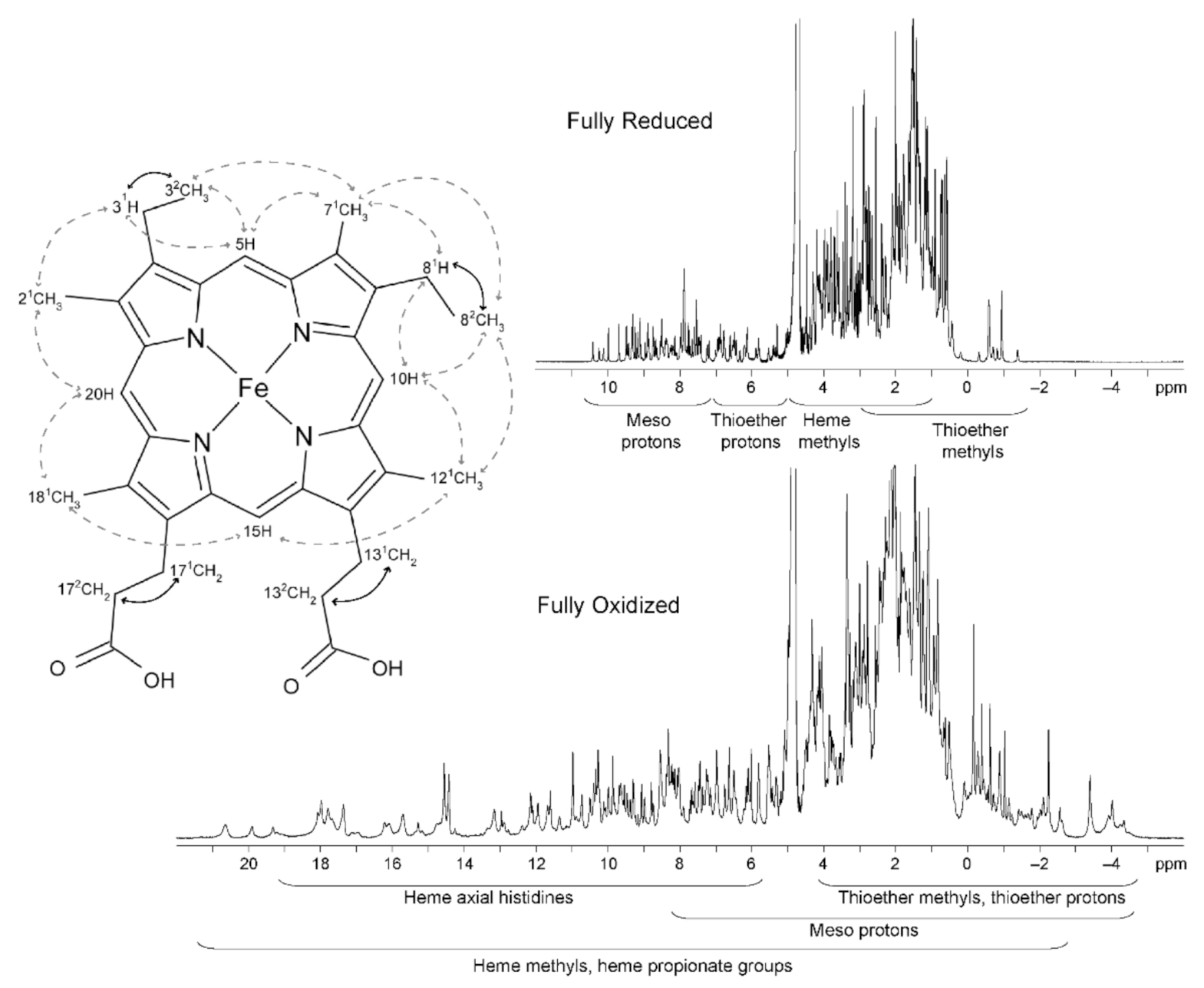
2.2.1. NMR Spectral Features of c-Type Cytochromes
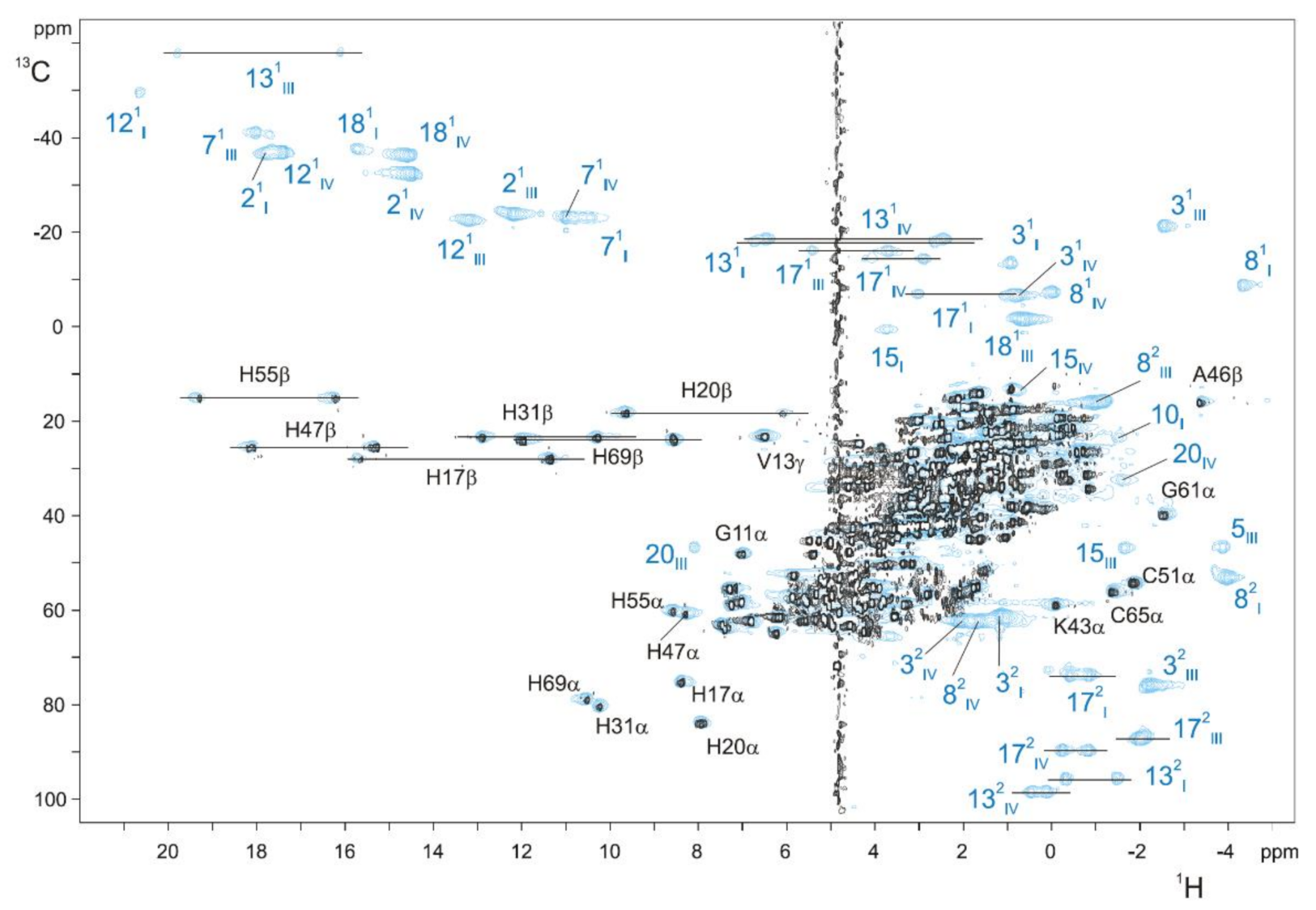
2.2.2. Heme Substituents Assignment
2.2.3. Probing the Heme Oxidation Profiles by EXSY NMR
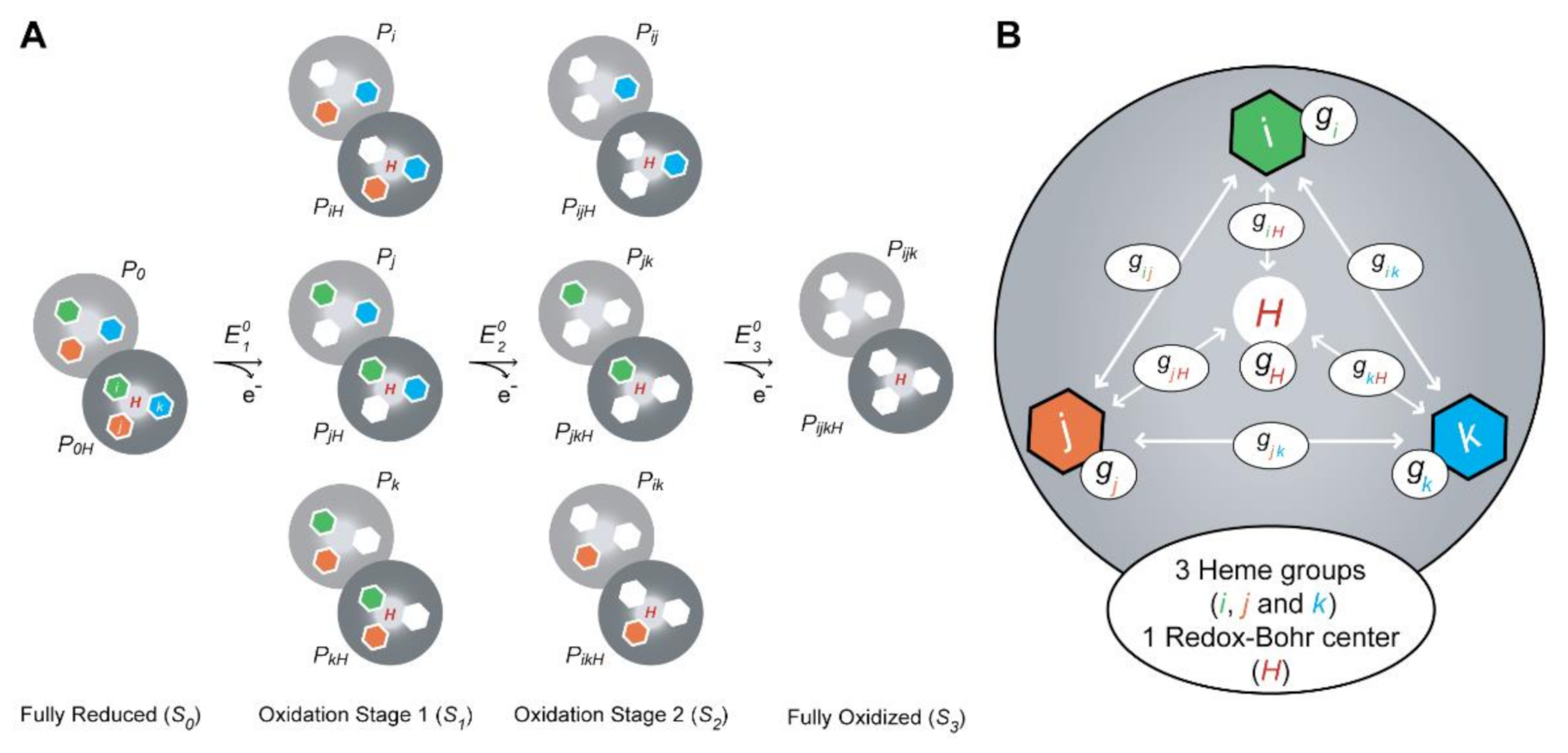
2.3. Redox Network
2.4. The Example of the Triheme Cytochrome PpcA
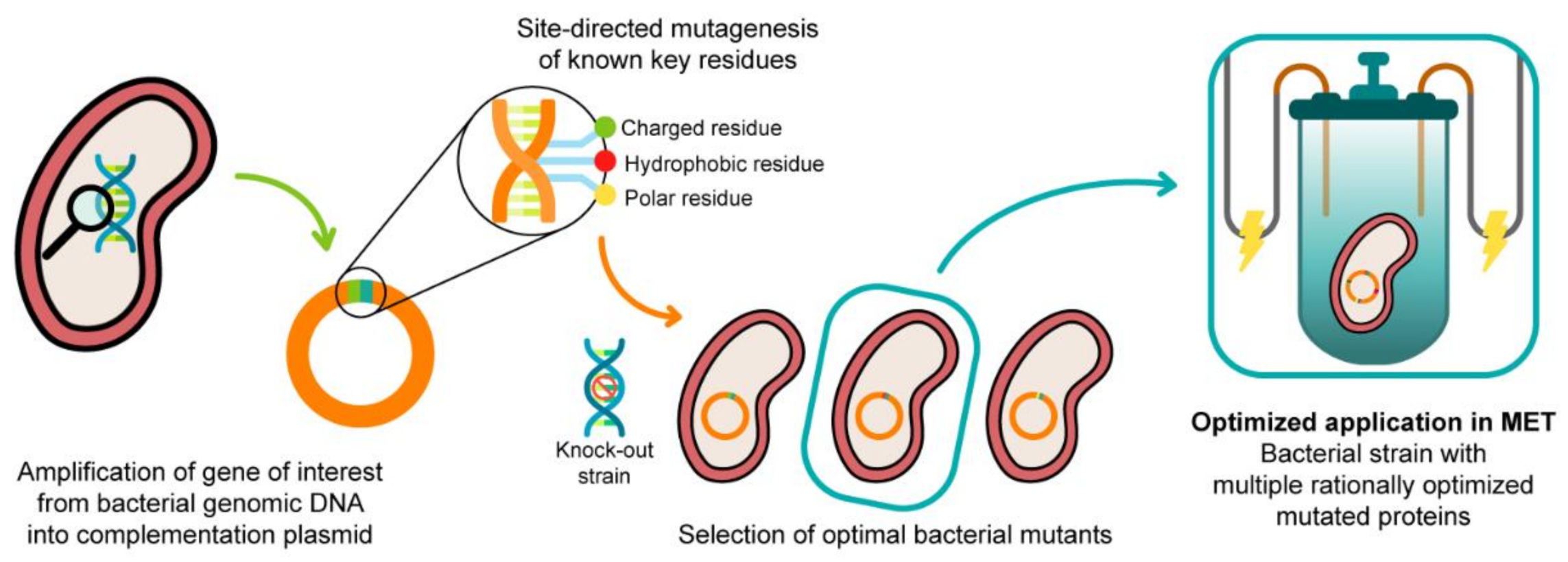
3. Modulation of the Redox Properties of MC for Optimized Geobacter Strains
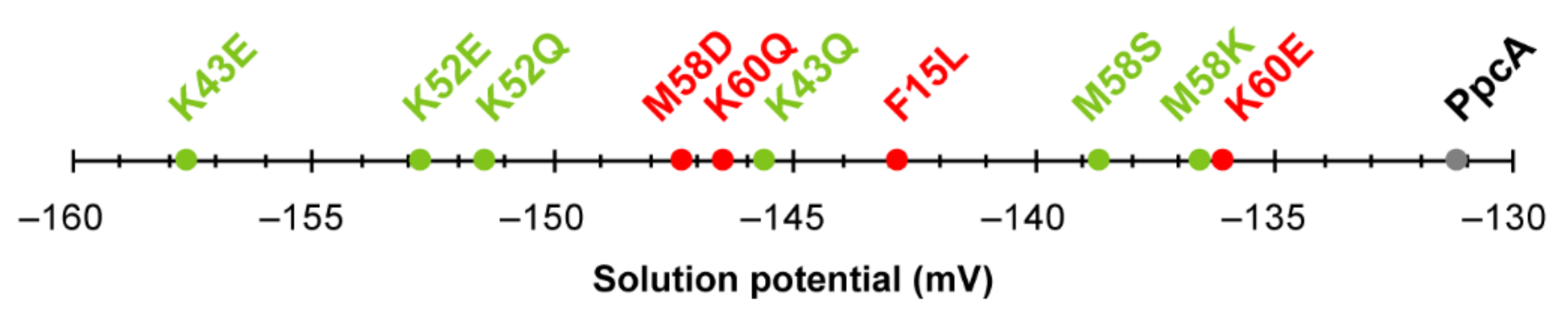
3.1. Enginnering of MCs—The Example of PpcA Mutants
3.2. In Vivo Testing of the Impact of the Selected Mutations of PpcA
3.3. Ongoing Biotic Strategies to Improve Extracellular Electron Transfer
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| EET | Extracellular electron transfer |
| METs | Microbial electrochemical technologies |
| MC(s) | Multiheme cytochrome(s) |
References
- Potter, M.C. Electrical effects accompanying the decomposition of organic compounds. Proc. R. Soc. Lond. Ser. B Contain. Pap. Biol. Character 1911, 84, 260–276. [Google Scholar]
- Logan, B.E.; Rossi, R.; Saikaly, P.E. Electroactive microorganisms in bioelectrochemical systems. Nat. Rev. Microbiol. 2019, 17, 307–319. [Google Scholar] [CrossRef] [PubMed]
- Wang, X.; Aulenta, F.; Puig, S.; Esteve-Núñez, A.; He, Y.; Mu, Y.; Rabaey, K. Microbial electrochemistry for bioremediation. Environ. Sci. Ecotechnol. 2020, 1, 100013. [Google Scholar] [CrossRef]
- Arends, J.B.A.; Verstraete, W. 100 years of microbial electricity production: Three concepts for the future. Microb. Biotechnol. 2012, 5, 333–346. [Google Scholar] [CrossRef]
- Lovley, D.R. Microbial fuel cells: Novel microbial physiologies and engineering approaches. Curr. Opin. Biotechnol. 2006, 17, 327–332. [Google Scholar] [CrossRef]
- Lovley, D.R. Cleaning up with genomics: Applying molecular biology to bioremediation. Nat. Rev. Microbiol. 2003, 1, 35–44. [Google Scholar] [CrossRef]
- Ing, N.L.; El-Naggar, M.Y.; Hochbaum, A.I. Going the distance: Long-range conductivity in protein and peptide bioelectronic materials. J. Phys. Chem. B 2018, 122, 10403–10423. [Google Scholar] [CrossRef]
- Lovley, D.R. e-Biologics: Fabrication of sustainable electronics with “green” biological materials. mBio 2017, 8, e00695-17. [Google Scholar] [CrossRef]
- Smith, A.F.; Liu, X.; Woodard, T.L.; Fu, T.; Emrick, T.; Jiménez, J.M.; Lovley, D.R.; Yao, J. Bioelectronic protein nanowire sensors for ammonia detection. Nano Res. 2020, 13, 1479–1484. [Google Scholar] [CrossRef]
- Mohan, Y.; Kumar, S.M.M.; Das, D. Electricity generation using microbial fuel cells. Int. J. Hydrogen Energy 2008, 33, 423–426. [Google Scholar] [CrossRef]
- Rani, N.; Sangwan, P.; Joshi, M.; Sagar, A.; Bala, K. Chapter 5—Microbes: A key player in industrial wastewater treatment. In Microbial Wastewater Treatment; Elsevier: Amsterdam, The Netherlands, 2019; pp. 83–102. [Google Scholar]
- Saeed, H.M.; Husseini, G.A.; Yousef, S.; Saif, J.; Al-Asheh, S.; Fara, A.A.; Azzam, S.; Khawaga, R.; Aidan, A. Microbial desalination cell technology: A review and a case study. Desalination 2015, 359, 1–13. [Google Scholar] [CrossRef]
- Nandi, R.; Sengupta, S. Microbial production of hydrogen: An overview. Crit. Rev. Microbiol. 1998, 24, 61–84. [Google Scholar] [CrossRef]
- Schnürer, A. Biogas Production: Microbiology and technology, Anaerobes in Biotechnology; Springer International Publishing: Cham, Switzerland, 2016; pp. 195–234. [Google Scholar]
- Liao, J.C.; Mi, L.; Pontrelli, S.; Luo, S. Fuelling the future: Microbial engineering for the production of sustainable biofuels. Nat. Rev. Microbiol. 2016, 14, 288–304. [Google Scholar] [CrossRef]
- Wu, S.; Li, H.; Zhou, X.; Liang, P.; Zhang, X.; Jiang, Y.; Huang, X. A novel pilot-scale stacked microbial fuel cell for efficient electricity generation and wastewater treatment. Water Res. 2016, 98, 396–403. [Google Scholar] [CrossRef]
- Anderson, R.T.; Vrionis, H.A.; Ortiz-Bernad, I.; Resch, C.T.; Long, P.E.; Dayvault, R.; Karp, K.; Marutzky, S.; Metzler, D.R.; Peacock, A.; et al. Stimulating the in situ activity of Geobacter species to remove uranium from the groundwater of a uranium-contaminated aquifer. Appl. Environ. Microbiol. 2003, 69, 5884–5891. [Google Scholar] [CrossRef]
- Mathuriya, A.S.; Jadhav, D.A.; Ghangrekar, M.M. Architectural adaptations of microbial fuel cells. Appl. Microbiol. Biot. 2018, 102, 9419–9432. [Google Scholar] [CrossRef]
- Zhou, M.; Chi, M.; Luo, J.; He, H.; Jin, T. An overview of electrode materials in microbial fuel cells. J. Power Sources 2011, 196, 4427–4435. [Google Scholar] [CrossRef]
- Jiang, D.; Li, B.; Jia, W.; Lei, Y. Effect of inoculum types on bacterial adhesion and power production in microbial fuel cells. Appl. Biochem. Biotechnol. 2009, 160, 182. [Google Scholar] [CrossRef]
- Li, F.; Sharma, Y.; Lei, Y.; Li, B.; Zhou, Q. Microbial fuel cells: The effects of configurations, electrolyte solutions, and electrode materials on power generation. Appl. Biochem. Biotechnol. 2009, 160, 168. [Google Scholar] [CrossRef]
- Schaetzle, O.; Barrière, F.; Baronian, K. Bacteria and yeasts as catalysts in microbial fuel cells: Electron transfer from micro-organisms to electrodes for green electricity. Energy Environ. Sci. 2008, 1, 607–620. [Google Scholar] [CrossRef]
- Yi, H.; Nevin, K.P.; Kim, B.-C.; Franks, A.E.; Klimes, A.; Tender, L.M.; Lovley, D.R. Selection of a variant of Geobacter sulfurreducens with enhanced capacity for current production in microbial fuel cells. Biosens. Bioelectron. 2009, 24, 3498–3503. [Google Scholar] [CrossRef]
- Hirose, A.; Kasai, T.; Koga, R.; Suzuki, Y.; Kouzuma, A.; Watanabe, K. Understanding and engineering electrochemically active bacteria for sustainable biotechnology. Bioresour. Bioprocess. 2019, 6, 10. [Google Scholar] [CrossRef]
- Zhao, J.; Li, F.; Cao, Y.; Zhang, X.; Chen, T.; Song, H.; Wang, Z. Microbial extracellular electron transfer and strategies for engineering electroactive microorganisms. Biotechnol. Adv. 2020, 107682. [Google Scholar] [CrossRef]
- Lin, W.C.; Coppi, M.V.; Lovley, D.R. Geobacter sulfurreducens can grow with oxygen as a terminal electron acceptor. Appl. Environ. Microbiol. 2004, 70, 2525–2528. [Google Scholar] [CrossRef]
- Lovley, D.R. Dissimilatory metal reduction. Annu. Rev. Microbiol. 1993, 47, 263–290. [Google Scholar] [CrossRef]
- Williams, K.H.; Bargar, J.R.; Lloyd, J.R.; Lovley, D.R. Bioremediation of uranium-contaminated groundwater: A systems approach to subsurface biogeochemistry. Curr. Opin. Biotechnol. 2013, 24, 489–497. [Google Scholar] [CrossRef]
- Methé, B.A.; Nelson, K.E.; Eisen, J.A.; Paulsen, I.T.; Nelson, W.; Heidelberg, J.F.; Wu, D.; Wu, M.; Ward, N.; Beanan, M.J.; et al. Genome of Geobacter sulfurreducens: Metal reduction in subsurface environments. Science 2003, 302, 1967–1969. [Google Scholar] [CrossRef]
- Aklujkar, M.; Coppi, M.V.; Leang, C.; Kim, B.C.; Chavan, M.A.; Perpetua, L.A.; Giloteaux, L.; Liu, A.; Holmes, D.E. Proteins involved in electron transfer to Fe(III) and Mn(IV) oxides by Geobacter sulfurreducens and Geobacter uraniireducens. Microbiology 2013, 159, 515–535. [Google Scholar] [CrossRef]
- Ding, Y.H.; Hixson, K.K.; Aklujkar, M.A.; Lipton, M.S.; Smith, R.D.; Lovley, D.R.; Mester, T. Proteome of Geobacter sulfurreducens grown with Fe(III) oxide or Fe(III) citrate as the electron acceptor. BBA Proteins Proteom. 2008, 1784, 1935–1941. [Google Scholar] [CrossRef]
- Ding, Y.H.; Hixson, K.K.; Giometti, C.S.; Stanley, A.; Esteve-Núñez, A.; Khare, T.; Tollaksen, S.L.; Zhu, W.; Adkins, J.N.; Lipton, M.S.; et al. The proteome of dissimilatory metal-reducing microorganism Geobacter sulfurreducens under various growth conditions. Biochim. Biophys. Acta 2006, 1764, 1198–1206. [Google Scholar] [CrossRef]
- Hernández-Eligio, A.; Pat-Espadas, A.M.; Vega-Alvarado, L.; Huerta-Amparán, M.; Cervantes, F.J.; Juárez, K. Global transcriptional analysis of Geobacter sulfurreducens under palladium reducing conditions reveals new key cytochromes involved. Appl. Microbiol. Biot. 2020, 104, 4059–4069. [Google Scholar] [CrossRef] [PubMed]
- Holmes, D.E.; Chaudhuri, S.K.; Nevin, K.P.; Mehta, T.; Methé, B.A.; Liu, A.; Ward, J.E.; Woodard, T.L.; Webster, J.; Lovley, D.R. Microarray and genetic analysis of electron transfer to electrodes in Geobacter sulfurreducens. Environ. Microbiol. 2006, 8, 1805–1815. [Google Scholar] [CrossRef]
- Otero, F.J.; Chan, C.H.; Bond, D.R. Identification of different putative outer membrane electron conduits necessary for Fe(III) citrate, Fe(III) oxide, Mn(IV) oxide, or electrode reduction by Geobacter sulfurreducens. J. Bacteriol. 2018, 200, e00347-18. [Google Scholar]
- Kim, B.C.; Leang, C.; Ding, Y.H.; Glaven, R.H.; Coppi, M.V.; Lovley, D.R. OmcF, a putative c-type monoheme outer membrane cytochrome required for the expression of other outer membrane cytochromes in Geobacter sulfurreducens. J. Bacteriol. 2005, 187, 4505–4513. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.C.; Lovley, D.R. Investigation of direct vs indirect involvement of the c-type cytochrome MacA in Fe(III) reduction by Geobacter sulfurreducens. FEMS Microbiol. Lett. 2008, 286, 39–44. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.C.; Postier, B.L.; DiDonato, R.J.; Chaudhuri, S.K.; Nevin, K.P.; Lovley, D.R. Insights into genes involved in electricity generation in Geobacter sulfurreducens via whole genome microarray analysis of the OmcF-deficient mutant. Bioelectrochemistry 2008, 73, 70–75. [Google Scholar] [CrossRef]
- Kim, B.-C.; Qian, X.; Leang, C.; Coppi, M.V.; Lovley, D.R. Two putative c-type multiheme cytochromes required for the expression of OmcB, an outer membrane protein essential for optimal Fe(III) reduction in Geobacter sulfurreducens. J. Bacteriol. 2006, 188, 3138. [Google Scholar] [CrossRef]
- Leang, C.; Coppi, M.V.; Lovley, D.R. OmcB, a c-type polyheme cytochrome, involved in Fe(III) reduction in Geobacter sulfurreducens. J. Bacteriol. 2003, 185, 2096–2103. [Google Scholar] [CrossRef]
- Levar, C.E.; Chan, C.H.; Mehta-Kolte, M.G.; Bond, D.R. An inner membrane cytochrome required only for reduction of high redox potential extracellular electron acceptors. mBio 2014, 5, e02034. [Google Scholar] [CrossRef]
- Liu, Y.; Fredrickson, J.K.; Zachara, J.M.; Shi, L. Direct involvement of ombB, omaB, and omcB genes in extracellular reduction of Fe(III) by Geobacter sulfurreducens PCA. Front. Microbiol. 2015, 6, 1075. [Google Scholar] [CrossRef]
- Mehta, T.; Coppi, M.V.; Childers, S.E.; Lovley, D.R. Outer membrane c-type cytochromes required for Fe(III) and Mn(IV) oxide reduction in Geobacter sulfurreducens. Appl. Environ. Microbiol. 2005, 71, 8634–8641. [Google Scholar] [CrossRef]
- Mollaei, M.; Timmers, P.H.A.; Suarez-Diez, M.; Boeren, S.; van Gelder, A.H.; Stams, A.J.M.; Plugge, C.M. Comparative proteomics of Geobacter sulfurreducens PCAT in response to acetate, formate and/or hydrogen as electron donor. Environ. Microbiol. 2021, 23, 299–315. [Google Scholar] [CrossRef]
- Nevin, K.P.; Kim, B.C.; Glaven, R.H.; Johnson, J.P.; Woodard, T.L.; Methe, B.A.; Didonato, R.J.; Covalla, S.F.; Franks, A.E.; Liu, A.; et al. Anode biofilm transcriptomics reveals outer surface components essential for high density current production in Geobacter sulfurreducens fuel cells. PLoS ONE 2009, 4, e5628. [Google Scholar] [CrossRef]
- Orellana, R.; Hixson, K.K.; Murphy, S.; Mester, T.; Sharma, M.L.; Lipton, M.S.; Lovley, D.R. Proteome of Geobacter sulfurreducens in the presence of U(VI). Microbiology 2014, 160, 2607–2617. [Google Scholar] [CrossRef]
- Rollefson, J.B.; Levar, C.E.; Bond, D.R. Identification of genes involved in biofilm formation and respiration via mini-Himar transposon mutagenesis of Geobacter sulfurreducens. J. Bacteriol. 2009, 191, 4207–4217. [Google Scholar] [CrossRef]
- Shelobolina, E.S.; Coppi, M.V.; Korenevsky, A.A.; DiDonato, L.N.; Sullivan, S.A.; Konishi, H.; Xu, H.; Leang, C.; Butler, J.E.; Kim, B.C.; et al. Importance of c-type cytochromes for U(VI) reduction by Geobacter sulfurreducens. BMC Microbiol. 2007, 7, 16. [Google Scholar] [CrossRef]
- Tremblay, P.L.; Summers, Z.M.; Glaven, R.H.; Nevin, K.P.; Zengler, K.; Barrett, C.L.; Qiu, Y.; Palsson, B.O.; Lovley, D.R. A c-type cytochrome and a transcriptional regulator responsible for enhanced extracellular electron transfer in Geobacter sulfurreducens revealed by adaptive evolution. Environ. Microbiol. 2011, 13, 13–23. [Google Scholar] [CrossRef]
- Voordeckers, J.W.; Kim, B.C.; Izallalen, M.; Lovley, D.R. Role of Geobacter sulfurreducens outer surface c-type cytochromes in reduction of soil humic acid and anthraquinone-2,6-disulfonate. Appl. Environ. Microbiol. 2010, 76, 2371–2375. [Google Scholar] [CrossRef]
- Zacharoff, L.; Chan, C.H.; Bond, D.R. Reduction of low potential electron acceptors requires the CbcL inner membrane cytochrome of Geobacter sulfurreducens. Bioelectrochemistry 2016, 107, 7–13. [Google Scholar] [CrossRef]
- Bowman, S.E.; Bren, K.L. The chemistry and biochemistry of heme c: Functional bases for covalent attachment. Nat. Prod. Rep. 2008, 25, 1118–1130. [Google Scholar] [CrossRef]
- Moore, G.R.; Pettigrew, G.W. Cytochromes c: Evolutionary, structural and physicochemical aspects. In Molecular Biology; Springer: Berlin/Heidelberg, Germany, 1990. [Google Scholar]
- Esteve-Núñez, A.; Sosnik, J.; Visconti, P.; Lovley, D.R. Fluorescent properties of c-type cytochromes reveal their potential role as an extracytoplasmic electron sink in Geobacter sulfurreducens. Environ. Microbiol. 2008, 10, 497–505. [Google Scholar] [CrossRef] [PubMed]
- Pokkuluri, P.R.; Londer, Y.Y.; Wood, S.J.; Duke, N.E.; Morgado, L.; Salgueiro, C.A.; Schiffer, M. Outer membrane cytochrome c, OmcF, from Geobacter sulfurreducens: High structural similarity to an algal cytochrome c6. Proteins 2009, 74, 266–270. [Google Scholar] [CrossRef] [PubMed]
- Morgado, L.; Bruix, M.; Pokkuluri, P.R.; Salgueiro, C.A.; Turner, D.L. Redox- and pH-linked conformational changes in triheme cytochrome PpcA from Geobacter sulfurreducens. Biochem. J. 2017, 474, 231–246. [Google Scholar] [CrossRef] [PubMed]
- Pokkuluri, P.R.; Londer, Y.Y.; Duke, N.E.; Pessanha, M.; Yang, X.; Orshonsky, V.; Orshonsky, L.; Erickson, J.; Zagyansky, Y.; Salgueiro, C.A.; et al. Structure of a novel dodecaheme cytochrome c from Geobacter sulfurreducens reveals an extended 12 nm protein with interacting hemes. J. Struct. Biol. 2011, 174, 223–233. [Google Scholar] [CrossRef]
- Rogers, N.K.; Moore, G.R. On the energetics of conformational changes and pH dependent redox behaviour of electron transfer proteins. FEBS Lett. 1988, 228, 69–73. [Google Scholar] [CrossRef]
- Xavier, A.V. Energy transduction coupling mechanisms in multiredox center proteins. J. Inorg. Biochem. 1986, 28, 239–243. [Google Scholar] [CrossRef]
- Page, C.C.; Moser, C.C.; Chen, X.; Dutton, P.L. Natural engineering principles of electron tunnelling in biological oxidation-reduction. Nature 1999, 402, 47–52. [Google Scholar] [CrossRef]
- Armstrong, F.A. Evaluations of reduction potential data in relation to coupling, kinetics and function. J. Biol. Inorg. Chem. 1997, 2, 139–142. [Google Scholar] [CrossRef]
- Das, D.K.; Medhi, O.K. The role of heme propionate in controlling the redox potential of heme: Square wave voltammetry of protoporphyrinato IX iron (III) in aqueous surfactant micelles. J. Inorg. Biochem. 1998, 70, 83–90. [Google Scholar] [CrossRef]
- de Lavalette, A.d.; Barucq, L.; Alric, J.; Rappaport, F.; Zito, F. Is the redox state of the ci heme of the cytochrome b6f complex dependent on the occupation and structure of the Qi site and vice-versa? J. Biol. Chem. 2009, 284, 20822–20829. [Google Scholar] [CrossRef]
- Gunner, M.R.; Alexov, E.; Torres, E.; Lipovaca, S. The importance of the protein in controlling the electrochemistry of heme metalloproteins: Methods of calculation and analysis. J. Biol. Inorg. Chem. 1997, 2, 126–134. [Google Scholar] [CrossRef]
- Liu, Y.; Moenne-Loccoz, P.; Hildebrand, D.P.; Wilks, A.; Loehr, T.M.; Mauk, A.G.; de Montellano, P.R.O. Replacement of the proximal histidine iron ligand by a cysteine or tyrosine converts heme oxygenase to an oxidase. Biochemistry 1999, 38, 3733–3743. [Google Scholar] [CrossRef]
- Ma, J.G.; Zhang, J.; Franco, R.; Jia, S.L.; Moura, I.; Moura, J.J.; Kroneck, P.M.; Shelnutt, J.A. The structural origin of nonplanar heme distortions in tetraheme ferricytochromes c3. Biochemistry 1998, 37, 12431–12442. [Google Scholar] [CrossRef]
- Sarma, S.; DiGate, R.J.; Goodin, D.B.; Miller, C.J.; Guiles, R.D. Effect of axial ligand plane reorientation on electronic and electrochemical properties observed in the A67V mutant of rat cytochrome b5. Biochemistry 1997, 36, 5658–5668. [Google Scholar] [CrossRef]
- Tezcan, F.A.; Winkler, J.R.; Gray, H.B. Effects of ligation and folding on reduction potentials of heme proteins. J. Am. Chem. Soc. 1998, 120, 13383–13388. [Google Scholar] [CrossRef]
- Assfalg, M.; Banci, L.; Bertini, I.; Bruschi, M.; Turano, P. 800 MHz 1H NMR solution structure refinement of oxidized cytochrome c7 from Desulfuromonas acetoxidans. Eur. J. Biochem. 1998, 256, 261–270. [Google Scholar] [CrossRef]
- Fernandes, A.P.; Couto, I.; Morgado, L.; Londer, Y.Y.; Salgueiro, C.A. Isotopic labeling of c-type multiheme cytochromes overexpressed in E. coli. Prot. Expr. Purif. 2008, 59, 182–188. [Google Scholar] [CrossRef]
- Messias, A.C.; Kastrau, D.H.; Costa, H.S.; LeGall, J.; Turner, D.L.; Santos, H.; Xavier, A.V. Solution structure of Desulfovibrio vulgaris (Hildenborough) ferrocytochrome c3: Structural basis for functional cooperativity. J. Mol. Biol. 1998, 281, 719–739. [Google Scholar] [CrossRef]
- Morgado, L.; Fernandes, A.P.; Londer, Y.Y.; Bruix, M.; Salgueiro, C.A. One simple step in the identification of the cofactors signals, one giant leap for the solution structure determination of multiheme proteins. Biochem. Biophys. Res. Commun. 2010, 393, 466–470. [Google Scholar] [CrossRef]
- Santos, H.; Turner, D.L.; Xavier, A.V.; le Gall, J. Two-dimensional NMR studies of electron transfer in cytochrome c3. J. Magn. Reson. 1984, 59, 177–180. [Google Scholar] [CrossRef]
- Turner, D.L.; Salgueiro, C.A.; Schenkels, P.; le Gall, J.; Xavier, A.V. Carbon-13 NMR studies of the influence of axial ligand orientation on haem electronic structure. Biochim. Biophys. Acta 1995, 1246, 24–28. [Google Scholar] [CrossRef]
- Morgado, L.; Bruix, M.; Pessanha, M.; Londer, Y.Y.; Salgueiro, C.A. Thermodynamic characterization of a triheme cytochrome family from Geobacter sulfurreducens reveals mechanistic and functional diversity. Biophys. J. 2010, 99, 293–301. [Google Scholar] [CrossRef]
- Wilson, G.S. Determination of Oxidation-Reduction Potentials, Methods in Enzymology; Academic Press: Cambridge, MA, USA, 1978; pp. 396–410. [Google Scholar]
- Heitmann, D.; Einsle, O. Structural and biochemical characterization of DHC2, a novel diheme cytochrome c from Geobacter sulfurreducens. Biochemistry 2005, 44, 12411–12419. [Google Scholar] [CrossRef] [PubMed]
- Santos, H.; Moura, J.J.; Moura, I.; le Gall, J.; Xavier, A.V. NMR studies of electron transfer mechanisms in a protein with interacting redox centres: Desulfovibrio gigas cytochrome c3. Eur. J. Biochem. 1984, 141, 283–296. [Google Scholar] [CrossRef] [PubMed]
- Jeener, J.; Meier, B.H.; Bachmann, P.; Ernst, R.R. Investigation of exchange processes by two-dimensional NMR spectroscopy. J. Chem. Phys. 1979, 71, 4546–4553. [Google Scholar] [CrossRef]
- Dantas, J.M.; Morgado, L.; Aklujkar, M.; Bruix, M.; Londer, Y.Y.; Schiffer, M.; Pokkuluri, P.R.; Salgueiro, C.A. Rational engineering of Geobacter sulfurreducens electron transfer components: A foundation for building improved Geobacter-based bioelectrochemical technologies. Front. Microbiol. 2015, 6, 752. [Google Scholar] [CrossRef] [PubMed]
- Perrin, C.L.; Dwyer, T.J. Application of two-dimensional NMR to kinetics of chemical exchange. Chem. Rev. 1990, 90, 935–967. [Google Scholar] [CrossRef]
- Johnson, C.E.; Bovey, F.A. Calculation of nuclear magnetic resonance spectra of aromatic hydrocarbons. J. Chem. Phys. 1958, 29, 1012–1014. [Google Scholar] [CrossRef]
- Moss, G.P. Nomenclature of tetrapyrroles. Recommendations 1986 IUPAC-IUB joint commission on biochemical nomenclature (JCBN). Eur. J. Biochem. 1988, 178, 277–328. [Google Scholar] [CrossRef]
- Rule, G.S.; Hitchens, T.K. Fundamentals of Protein NMR Spectroscopy, 1st ed.; Springer: Dordrecht, The Netherlands, 2006. [Google Scholar]
- Turner, D.L.; Salgueiro, C.A.; le Gall, J.; Xavier, A.V. Structural studies of Desulfovibrio vulgaris ferrocytochrome c3 by two-dimensional NMR. Eur. J. Biochem. 1992, 210, 931–936. [Google Scholar] [CrossRef]
- Keller, R.M.; Wüthrich, K. Assignment of the heme c resonances in the 360 MHz 1H NMR spectra of cytochrome c. Biochim. Biophys. Acta Protein Struct. 1978, 533, 195–208. [Google Scholar] [CrossRef]
- Turner, D.L.; Costa, H.S.; Coutinho, I.B.; Legall, J.; Xavier, A.V. Assignment of the ligand geometry and redox potentials of the trihaem ferricytochrome c3 from Desulfuromonas acetoxidans. Eur. J. Biochem. 1997, 243, 474–481. [Google Scholar] [CrossRef]
- Salgueiro, C.A.; Turner, D.L.; Xavier, A.V. Use of paramagnetic NMR probes for structural analysis in cytochrome c3 from Desulfovibrio vulgaris. Eur. J. Biochem. 1997, 244, 721–734. [Google Scholar] [CrossRef]
- Fernandes, T.M.; Morgado, L.; Salgueiro, C.A. Thermodynamic and functional characterization of the periplasmic triheme cytochrome PpcA from Geobacter metallireducens. Biochem. J. 2018, 475, 2861–2875. [Google Scholar] [CrossRef]
- Xavier, A.V. Thermodynamic and choreographic constraints for energy transduction by cytochrome c oxidase. BBA Bioenergy 2004, 1658, 23–30. [Google Scholar] [CrossRef][Green Version]
- Marquardt, D.W. An algorithm for least-squares estimation of nonlinear parameters. J. Soc. Ind. Appl. Math. 1963, 11, 431–441. [Google Scholar] [CrossRef]
- Pessanha, M.; Morgado, L.; Louro, R.O.; Londer, Y.Y.; Pokkuluri, P.R.; Schiffer, M.; Salgueiro, C.A. Thermodynamic characterization of triheme cytochrome PpcA from Geobacter sulfurreducens: Evidence for a role played in e−/H+ energy transduction. Biochemistry 2006, 45, 13910–13917. [Google Scholar] [CrossRef]
- Morgado, L.; Bruix, M.; Orshonsky, V.; Londer, Y.Y.; Duke, N.E.; Yang, X.; Pokkuluri, P.R.; Schiffer, M.; Salgueiro, C.A. Structural insights into the modulation of the redox properties of two Geobacter sulfurreducens homologous triheme cytochromes. BBA Bioenergy 2008, 1777, 1157–1165. [Google Scholar] [CrossRef]
- Mahadevan, R.; Bond, D.R.; Butler, J.E.; Esteve-Núñez, A.; Coppi, M.V.; Palsson, B.O.; Schilling, C.H.; Lovley, D.R. Characterization of metabolism in the Fe(III)-reducing organism Geobacter sulfurreducens by constraint-based modeling. Appl. Environ. Microbiol. 2006, 72, 1558–1568. [Google Scholar] [CrossRef]
- Dantas, J.M.; Portela, P.C.; Fernandes, A.P.; Londer, Y.Y.; Yang, X.; Duke, N.E.C.; Schiffer, M.; Pokkuluri, P.R.; Salgueiro, C.A. Structural and functional relevance of the conserved residue V13 in the triheme cytochrome PpcA from Geobacter sulfurreducens. J. Phys. Chem. B 2019, 123, 3050–3060. [Google Scholar] [CrossRef]
- Dantas, J.M.; Morgado, L.; Londer, Y.Y.; Fernandes, A.P.; Louro, R.O.; Pokkuluri, P.R.; Schiffer, M.; Salgueiro, C.A. Pivotal role of the strictly conserved aromatic residue F15 in the cytochrome c7 family. J. Biol. Inorg. Chem. 2012, 17, 11–24. [Google Scholar] [CrossRef] [PubMed]
- Dantas, J.M.; Simões, T.; Morgado, L.; Caciones, C.; Fernandes, A.P.; Silva, M.A.; Bruix, M.; Pokkuluri, P.R.; Salgueiro, C.A. Unveiling the structural basis that regulates the energy transduction properties within a family of triheme cytochromes from Geobacter sulfurreducens. J. Phys. Chem. B 2016, 120, 10221–10233. [Google Scholar] [CrossRef] [PubMed]
- Morgado, L.; Dantas, J.M.; Simões, T.; Londer, Y.Y.; Pokkuluri, P.R.; Salgueiro, C.A. Role of Met58 in the regulation of electron/proton transfer in trihaem cytochrome PpcA from Geobacter sulfurreducens. Biosci. Rep. 2013, 33, e00002. [Google Scholar] [CrossRef] [PubMed]
- Morgado, L.; Lourenço, S.; Londer, Y.Y.; Schiffer, M.; Pokkuluri, P.R.; Salgueiro, C.A. Dissecting the functional role of key residues in triheme cytochrome PpcA: A path to rational design of G. sulfurreducens strains with enhanced electron transfer capabilities. PLoS ONE 2014, 9, e105566. [Google Scholar] [CrossRef] [PubMed]
- Lloyd, J.R.; Leang, C.; Myerson, A.L.H.; Coppi, M.V.; Cuifo, S.; Methe, B.; Sandler, S.J.; Lovley, D.R. Biochemical and genetic characterization of PpcA, a periplasmic c-type cytochrome in Geobacter sulfurreducens. Biochem. J. 2003, 369, 153–161. [Google Scholar] [CrossRef] [PubMed]
- Ueki, T.; DiDonato, L.N.; Lovley, D.R. Toward establishing minimum requirements for extracellular electron transfer in Geobacter sulfurreducens. FEMS Microbiol. Lett. 2017, 364, fnx093. [Google Scholar] [CrossRef] [PubMed]
- Coppi, M.V.; Leang, C.; Sandler, S.J.; Lovley, D.R. Development of a genetic system for Geobacter sulfurreducens. Appl. Environ. Microbiol. 2001, 67, 3180–3187. [Google Scholar] [CrossRef]
- Jensen, H.M.; Albers, A.E.; Malley, K.R.; Londer, Y.Y.; Cohen, B.E.; Helms, B.A.; Weigele, P.; Groves, J.T.; Ajo-Franklin, C.M. Engineering of a synthetic electron conduit in living cells. Proc. Natl. Acad. Sci. USA 2010, 107, 19213. [Google Scholar] [CrossRef]
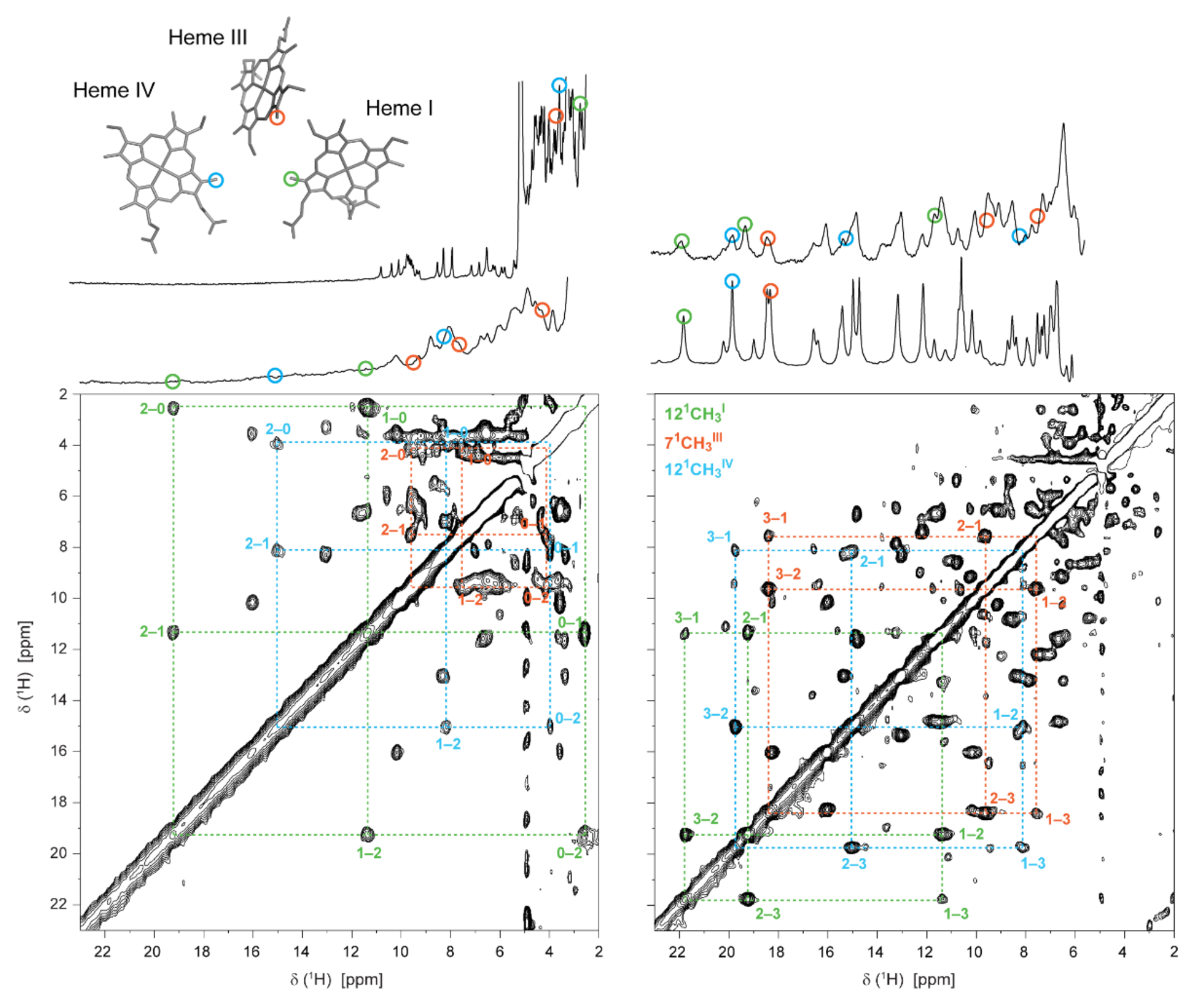
| pH 8 | Chemical Shift (ppm) | xi | ∑xi | ||||
|---|---|---|---|---|---|---|---|
| S | 121CH3I | 71CH3III | 121CH3IV | 121CH3I | 71CH3III | 121CH3IV | |
| 0 | 2.55 | 4.14 | 3.95 | 0.00 | 0.00 | 0.00 | 0.00 |
| 1 | 11.22 | 7.38 | 8.44 | 0.45 | 0.23 | 0.28 | 0.96 |
| 2 | 19.25 | 9.40 | 15.31 | 0.87 | 0.37 | 0.72 | 1.95 |
| 3 | 21.79 | 18.42 | 19.81 | 1.00 | 1.00 | 1.00 | 3.00 |
| pH 6 | Chemical Shift (ppm) | xi | ∑xi | ||||
| S | 121CH3I | 71CH3III | 121CH3IV | 121CH3I | 71CH3III | 121CH3IV | |
| 0 | 2.55 | 4.14 | 3.95 | 0.00 | 0.00 | 0.00 | 0.00 |
| 1 | 12.70 | 8.13 | 5.92 | 0.53 | 0.28 | 0.14 | 0.95 |
| 2 | 19.78 | 11.61 | 11.19 | 0.90 | 0.52 | 0.53 | 1.95 |
| 3 | 21.62 | 18.42 | 17.69 | 1.00 | 1.00 | 1.00 | 3.00 |
| Energy (meV) | ||||
|---|---|---|---|---|
| Heme I | Heme III | Heme IV | Redox-Bohr Center | |
| Heme I | −154 (5) | 27 (2) | 16 (3) | −32 (4) |
| Heme III | −138 (5) | 41 (3) | −31 (4) | |
| Heme IV | −125 (5) | −58 (4) | ||
| Redox-Bohr center | 495 (8) | |||
| Protein | eapp (mV) | Order of Heme Oxidation | Electron Transfer Pathway | ||
|---|---|---|---|---|---|
| Heme I | Heme III | Heme IV | |||
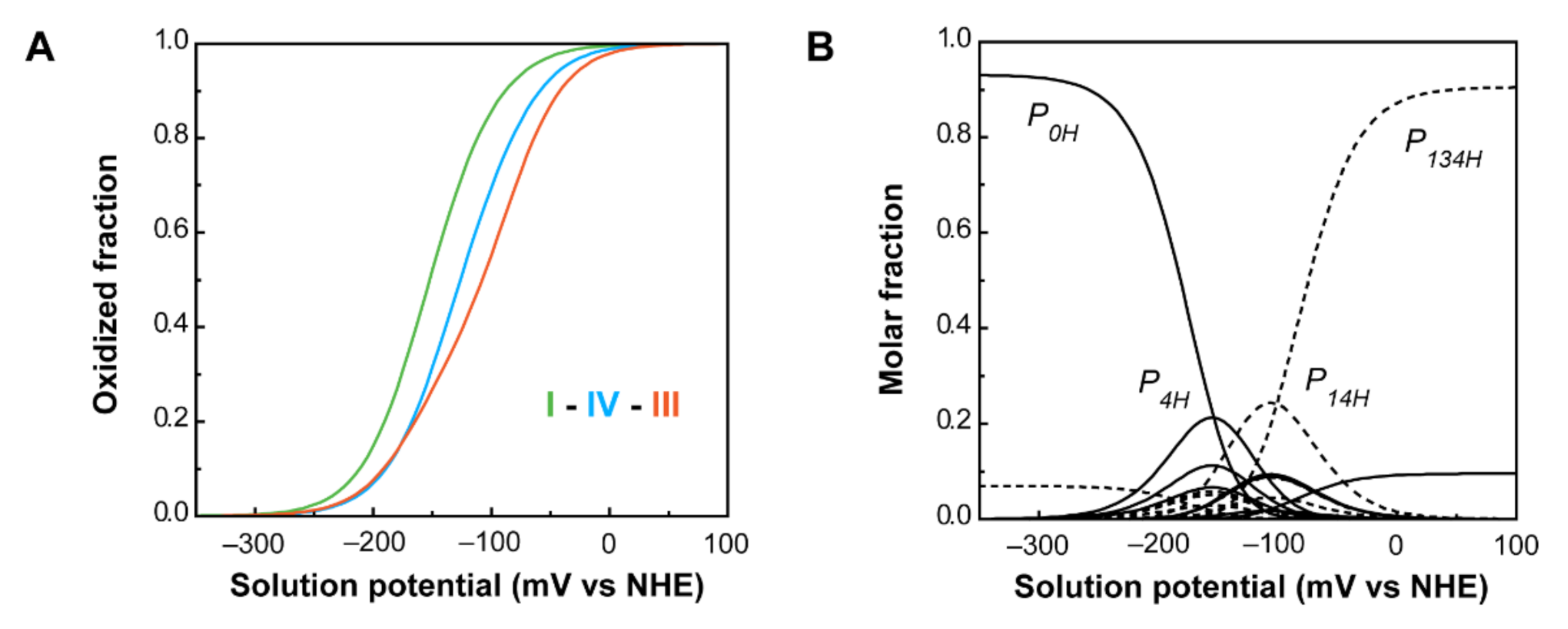
| PpcA | −152 | −108 | −126 | I-IV-III | P0H→P1H→P14→P134 |
| F15L | −155 | −146 | −125 | I-III-IV | No preferential pathway |
| M58D | −160 | −139 | −140 | I-(III,IV) | No preferential pathway |
| M58K | −159 | −91 | −146 | I-IV-III | P0H→P14→P134 |
| M58S | −159 | −110 | −139 | I-IV-III | P0H→P1H→P14→P134 |
| K43E | −165 | −117 | −180 | IV-I-III | P0H→P14→P134 |
| K43Q | −162 | −117 | −150 | I-IV-III | P0H→P1H→P14→P134 |
| K52E | −156 | −116 | −177 | IV-I-III | P0H→P14→P134 |
| K52Q | −157 | −111 | −175 | IV-I-III | P0H→P14→P134 |
| K60E | −145 | −146 | −119 | (III,I)-IV | No preferential pathway |
| K60Q | −161 | −143 | −134 | I-III-IV | No preferential pathway |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fernandes, T.M.; Morgado, L.; Turner, D.L.; Salgueiro, C.A. Protein Engineering of Electron Transfer Components from Electroactive Geobacter Bacteria. Antioxidants 2021, 10, 844. https://doi.org/10.3390/antiox10060844
Fernandes TM, Morgado L, Turner DL, Salgueiro CA. Protein Engineering of Electron Transfer Components from Electroactive Geobacter Bacteria. Antioxidants. 2021; 10(6):844. https://doi.org/10.3390/antiox10060844
Chicago/Turabian StyleFernandes, Tomás M., Leonor Morgado, David L. Turner, and Carlos A. Salgueiro. 2021. "Protein Engineering of Electron Transfer Components from Electroactive Geobacter Bacteria" Antioxidants 10, no. 6: 844. https://doi.org/10.3390/antiox10060844
APA StyleFernandes, T. M., Morgado, L., Turner, D. L., & Salgueiro, C. A. (2021). Protein Engineering of Electron Transfer Components from Electroactive Geobacter Bacteria. Antioxidants, 10(6), 844. https://doi.org/10.3390/antiox10060844







