Avian and Mammalian Facilitative Glucose Transporters
Abstract
:1. Introduction
2. Glucose Transport
3. GLUT Transporter Classes
3.1. Class I GLUTs
3.2. Class II GLUTs
3.3. Class III GLUTs
4. GLUT Annotation
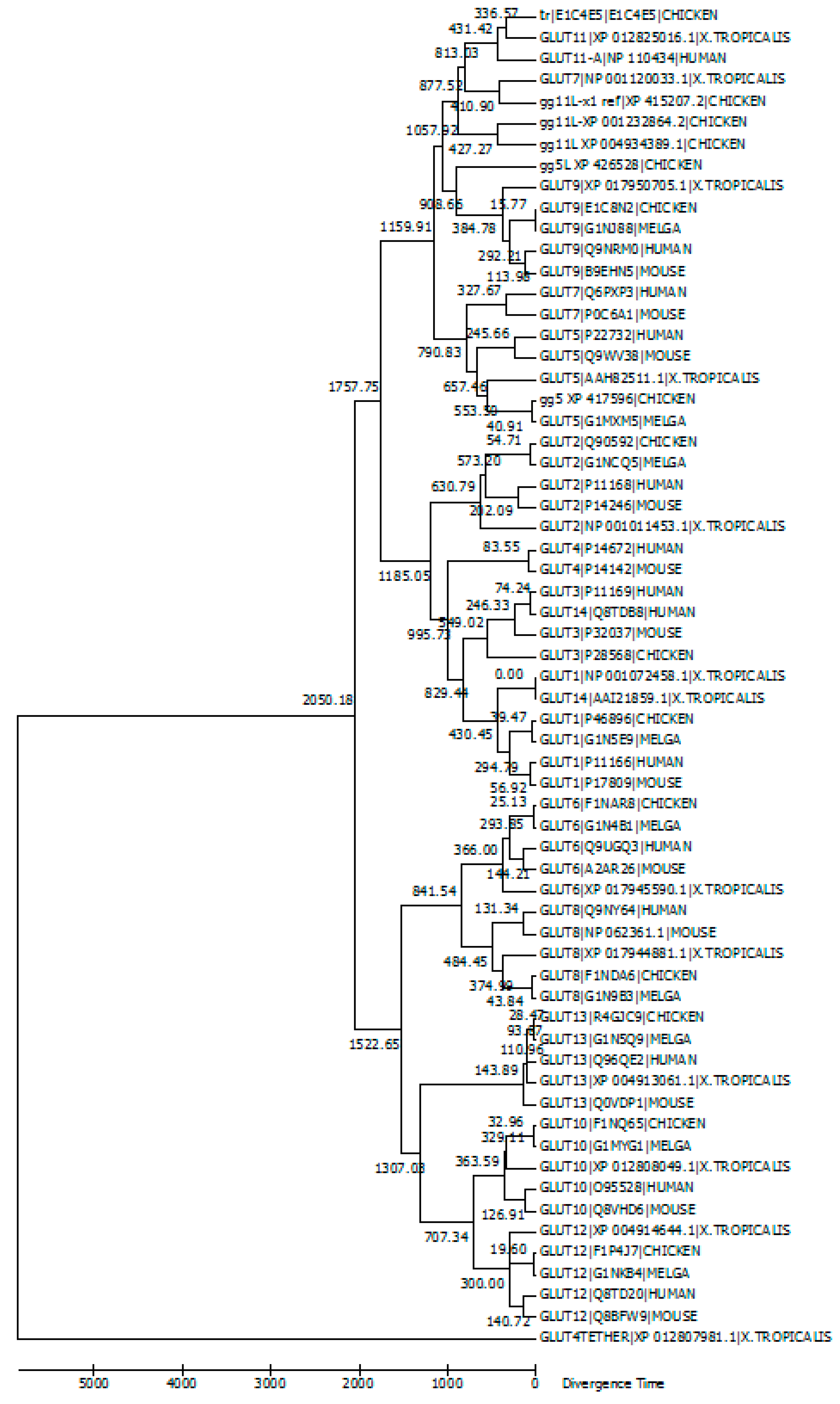
5. Evolutionary Relationships among GLUT Members
6. Chicken GLUT Members
7. Prospects and Conclusions
Acknowledgments
Author Contribution
Conflicts of Interest
References
- Kono, T.; Nishida, M.; Nishiki, Y.; Seki, Y.; Sato, K.; Akiba, Y. Characterisation of glucose transporter (GLUT) gene expression in broiler chickens. Br. Poult. Sci. 2005, 46, 510–515. [Google Scholar] [CrossRef] [PubMed]
- Seki, Y.; Sato, K.; Kono, T.; Abe, H.; Akiba, Y. Broiler chickens (Ross strain) lack insulin-responsive glucose transporter GLUT4 and have GLUT8 cDNA. Gen. Comp. Endocrinol. 2003, 133, 80–87. [Google Scholar] [CrossRef]
- Gibbs, E.M.; Calderhead, D.M.; Holman, G.D.; Gould, G.W. Phorbol ester only partially mimics the effects of insulin on glucose transport and glucose-transporter distribution in 3T3-L1 adipocytes. Biochem. J. 1991, 275 Pt 1, 145–150. [Google Scholar] [CrossRef] [PubMed]
- Wood, I.S.; Hunter, L.; Trayhurn, P. Expression of Class III facilitative glucose transporter genes (GLUT-10 and GLUT-12) in mouse and human adipose tissues. Biochem. Biophys. Res. Commun. 2003, 308, 43–49. [Google Scholar] [CrossRef]
- Mueckler, M.; Thorens, B. The SLC2 (GLUT) family of membrane transporters. Mol. Aspects Med. 2013, 34, 121–138. [Google Scholar] [CrossRef] [PubMed]
- Wu, X.; Freeze, H.H. GLUT14, a duplicon of GLUT3, is specifically expressed in testis as alternative splice forms. Genomics 2002, 80, 553–557. [Google Scholar] [CrossRef] [PubMed]
- Thorens, B.; Mueckler, M. Glucose transporters in the 21st Century. Am. J. Physiol. Endocrinol. Metab. 2010, 298, E141–E145. [Google Scholar] [CrossRef] [PubMed]
- McQuade, D.T.; Plutschack, M.B.; Seeberger, P.H. Passive fructose transporters in disease: A molecular overview of their structural specificity. Org. Biomol. Chem. 2013, 11, 4909–4920. [Google Scholar] [CrossRef] [PubMed]
- Ibberson, M.; Uldry, M.; Thorens, B. GLUTX1, a novel mammalian glucose transporter expressed in the central nervous system and insulin-sensitive tissues. J. Biol. Chem. 2000, 275, 4607–4612. [Google Scholar] [CrossRef] [PubMed]
- Patel, C.; Sugimoto, K.; Douard, V.; Shah, A.; Inui, H.; Yamanouchi, T.; Ferraris, R.P. Effect of dietary fructose on portal and systemic serum fructose levels in rats and in KHK-/- and GLUT5-/- mice. Am. J. Physiol. Gastrointest. Liver Physiol. 2015, 309, G779–G790. [Google Scholar] [CrossRef] [PubMed]
- Wright, E.M.; Loo, D.D.; Hirayama, B.A.; Turk, E. Surprising versatility of Na+-glucose cotransporters: SLC5. Physiology (Bethesda) 2004, 19, 370–376. [Google Scholar] [CrossRef] [PubMed]
- Kellett, G.L.; Brot-Laroche, E. Apical GLUT2: A major pathway of intestinal sugar absorption. Diabetes 2005, 54, 3056–3062. [Google Scholar] [CrossRef]
- Debnam, E.S.; Levin, R.J. An experimental method of identifying and quantifying the active transfer electrogenic component from the diffusive component during sugar absorption measured in vivo. J. Physiol. 1975, 246, 181–196. [Google Scholar] [CrossRef] [PubMed]
- Kellett, G.L. The facilitated component of intestinal glucose absorption. J. Physiol. 2001, 531, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Maher, F.; Vannucci, S.J.; Simpson, I.A. Glucose transporter proteins in brain. FASEB J. 1994, 8, 1003–1011. [Google Scholar] [PubMed]
- Mueckler, M.; Makepeace, C. Model of the exofacial substrate-binding site and helical folding of the human Glut1 glucose transporter based on scanning mutagenesis. Biochemistry 2009, 48, 5934–5942. [Google Scholar] [CrossRef] [PubMed]
- Mueckler, M.; Makepeace, C. Transmembrane segment 6 of the Glut1 glucose transporter is an outer helix and contains amino acid side chains essential for transport activity. J. Biol. Chem. 2008, 283, 11550–11555. [Google Scholar] [CrossRef] [PubMed]
- Klepper, J.; Wang, D.; Fischbarg, J.; Vera, J.C.; Jarjour, I.T.; O’Driscoll, K.R.; De Vivo, D.C. Defective glucose transport across brain tissue barriers: A newly recognized neurological syndrome. Neurochem. Res. 1999, 24, 587–594. [Google Scholar] [CrossRef] [PubMed]
- Pascual, J.M.; Van Heertum, R.L.; Wang, D.; Engelstad, K.; De Vivo, D.C. Imaging the metabolic footprint of Glut1 deficiency on the brain. Ann. Neurol. 2002, 52, 458–464. [Google Scholar] [CrossRef] [PubMed]
- Klepper, J.; Willemsen, M.; Verrips, A.; Guertsen, E.; Herrmann, R.; Kutzick, C.; Florcken, A.; Voit, T. Autosomal dominant transmission of GLUT1 deficiency. Hum. Mol. Genet. 2001, 10, 63–68. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; Kranz-Eble, P.; De Vivo, D.C. Mutational analysis of GLUT1 (SLC2A1) in Glut-1 deficiency syndrome. Hum. Mutat. 2000, 16, 224–231. [Google Scholar] [CrossRef]
- Brockmann, K.; Wang, D.; Korenke, C.G.; von Moers, A.; Ho, Y.Y.; Pascual, J.M.; Kuang, K.; Yang, H.; Ma, L.; Kranz-Eble, P.; et al. Autosomal dominant glut-1 deficiency syndrome and familial epilepsy. Ann. Neurol. 2001, 50, 476–485. [Google Scholar] [CrossRef] [PubMed]
- Leen, W.G.; Klepper, J.; Verbeek, M.M.; Leferink, M.; Hofste, T.; van Engelen, B.G.; Wevers, R.A.; Arthur, T.; Bahi-Buisson, N.; Ballhausen, D.; et al. Glucose transporter-1 deficiency syndrome: The expanding clinical and genetic spectrum of a treatable disorder. Brain 2010, 133, 655–670. [Google Scholar] [CrossRef] [PubMed]
- Mohammad, S.S.; Coman, D.; Calvert, S. Glucose transporter 1 deficiency syndrome and hemiplegic migraines as a dominant presenting clinical feature. J. Paediatr. Child Health 2014, 50, 1025–1026. [Google Scholar]
- Pellegrin, S.; Cantalupo, G.; Opri, R.; Dalla Bernardina, B.; Darra, F. EEG findings during “paroxysmal hemiplegia” in a patient with GLUT1-deficiency. Eur. J. Paediatr. Neurol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Mullen, S.A.; Suls, A.; De Jonghe, P.; Berkovic, S.F.; Scheffer, I.E. Absence epilepsies with widely variable onset are a key feature of familial GLUT1 deficiency. Neurology 2010, 75, 432–440. [Google Scholar] [CrossRef] [PubMed]
- Deng, D.; Xu, C.; Sun, P.; Wu, J.; Yan, C.; Hu, M.; Yan, N. Crystal structure of the human glucose transporter GLUT1. Nature 2014, 510, 121–125. [Google Scholar] [CrossRef] [PubMed]
- Du, M.J.; Shen, Q.; Yin, H.; Rao, Q.; Zhou, M.X. Diagnostic roles of MUC1 and GLUT1 in differentiating thymic carcinoma from type B3 thymoma. Pathol. Res. Pract. 2016, 212, 1048–1051. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.M.; Lee, G.; Oh, T.I.; Kim, B.M.; Shim, D.W.; Lee, K.H.; Kim, Y.J.; Lim, B.O.; Lim, J.H. Inhibition of glutamine utilization sensitizes lung cancer cells to apigenin-induced apoptosis resulting from metabolic and oxidative stress. Int. J. Oncol. 2016, 48, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Rancourt, R.C.; Schellong, K.; Ott, R.; Bogatyrev, S.; Tzschentke, B.; Plagemann, A. Acquired alterations of hypothalamic gene expression of insulin and leptin receptors and glucose transporters in prenatally high-glucose exposed three-week old chickens do not coincide with aberrant promoter DNA methylation. PLoS ONE 2015, 10, e0119213. [Google Scholar] [CrossRef]
- Zhao, J.P.; Bao, J.; Wang, X.J.; Jiao, H.C.; Song, Z.G.; Lin, H. Altered gene and protein expression of glucose transporter1 underlies dexamethasone inhibition of insulin-stimulated glucose uptake in chicken muscles. J. Anim. Sci. 2012, 90, 4337–4345. [Google Scholar] [CrossRef] [PubMed]
- Murphy, M.; Brown, G.; Wallin, C.; Tatusova, T.; Pruitt, K.; Murphy, T.; Maglott, D. Gene Help: Integrated Access to Genes of Genomes in the Reference Sequence Collection. 2006. Available online: https://www.ncbi.nlm.nih.gov/books/NBK3841/ (accessed on 10 January 2016).
- Zhang, W.; Sumners, L.H.; Siegel, P.B.; Cline, M.A.; Gilbert, E.R. Quantity of glucose transporter and appetite-associated factor mRNA in various tissues after insulin injection in chickens selected for low or high body weight. Physiol. Genom. 2013, 45, 1084–1094. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.Y.; Tsai, M.Y.; Wang, C. Identification of chicken liver glucose transporter. Arch. Biochem. Biophys. 1994, 310, 172–179. [Google Scholar] [CrossRef] [PubMed]
- Borglykke, A.; Grarup, N.; Sparso, T.; Linneberg, A.; Fenger, M.; Jeppesen, J.; Hansen, T.; Pedersen, O.; Jorgensen, T. Genetic variant SLC2A2 [corrected] Is associated with risk of cardiovascular disease—Assessing the individual and cumulative effect of 46 type 2 diabetes related genetic variants. PLoS ONE 2012, 7, e50418. [Google Scholar] [CrossRef] [PubMed]
- Seatter, M.J.; De la Rue, S.A.; Porter, L.M.; Gould, G.W. QLS motif in transmembrane helix VII of the glucose transporter family interacts with the C-1 position of d-glucose and is involved in substrate selection at the exofacial binding site. Biochemistry 1998, 37, 1322–1326. [Google Scholar] [CrossRef] [PubMed]
- Coordinators, National Center for Biotechnology Inforamtion online Database: Gene. 2016. Available online: https://www.ncbi.nlm.nih.gov/ (accessed on 12 January 2016).
- Deng, D.; Sun, P.; Yan, C.; Ke, M.; Jiang, X.; Xiong, L.; Ren, W.; Hirata, K.; Yamamoto, M.; Fan, S.; et al. Molecular basis of ligand recognition and transport by glucose transporters. Nature 2015, 526, 391–396. [Google Scholar] [CrossRef] [PubMed]
- Vittori, A.; Breda, C.; Repici, M.; Orth, M.; Roos, R.A.; Outeiro, T.F.; Giorgini, F.; Hollox, E.J.; REGISTRY investigators of the European Huntington’s Disease Network. Copy-number variation of the neuronal glucose transporter gene SLC2A3 and age of onset in Huntington’s disease. Hum. Mol. Genet. 2014, 23, 3129–3137. [Google Scholar] [CrossRef] [PubMed]
- Ban, Y.; Rizzolo, L.J. Regulation of glucose transporters during development of the retinal pigment epithelium. Brain Res. Dev. Brain Res. 2000, 121, 89–95. [Google Scholar] [CrossRef]
- Hussey, S.E.; McGee, S.L.; Garnham, A.; McConell, G.K.; Hargreaves, M. Exercise increases skeletal muscle GLUT4 gene expression in patients with type 2 diabetes. Diabetes Obes. Metab. 2012, 14, 768–771. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.T.; Li, C.L.; Tian, H.; Li, J.; Pei, Y.; Liu, Y.; Gong, Y.P.; Fang, F.S.; Sun, B.R. MiR-199a is overexpressed in plasma of type 2 diabetes patients which contributes to type 2 diabetes by targeting GLUT4. Mol. Cell Biochem. 2014, 397, 45–51. [Google Scholar] [CrossRef] [PubMed]
- Tokushima, Y.; Sulistiyanto, B.; Takahashi, K.; Akiba, Y. Insulin-glucose interactions characterised in newly hatched broiler chicks. Br. Poult. Sci. 2003, 44, 746–751. [Google Scholar] [CrossRef] [PubMed]
- Dupont, J.; Metayer-Coustard, S.; Ji, B.; Rame, C.; Gespach, C.; Voy, B.; Simon, J. Characterization of major elements of insulin signaling cascade in chicken adipose tissue: Apparent insulin refractoriness. Gen. Comp. Endocrinol. 2012, 176, 86–93. [Google Scholar] [CrossRef] [PubMed]
- Dupont, J.; Dagou, C.; Derouet, M.; Simon, J.; Taouis, M. Early steps of insulin receptor signaling in chicken and rat: Apparent refractoriness in chicken muscle. Domest. Anim. Endocrinol. 2004, 26, 127–142. [Google Scholar] [CrossRef] [PubMed]
- NCBI Resource Coordinators. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 2016, 44, D7–D19. [Google Scholar]
- Wang, W.; Yu, J.T.; Zhang, W.; Cui, W.Z.; Wu, Z.C.; Zhang, Q.; Tan, L. Genetic association of SLC2A14 polymorphism with Alzheimer’s disease in a Han Chinese population. J. Mol. Neurosci. 2012, 47, 481–484. [Google Scholar] [CrossRef] [PubMed]
- Berlth, F.; Monig, S.; Pinther, B.; Grimminger, P.; Maus, M.; Schlosser, H.; Plum, P.; Warnecke-Eberz, U.; Harismendy, O.; Drebber, U.; et al. Both GLUT-1 and GLUT-14 are Independent Prognostic Factors in Gastric Adenocarcinoma. Ann. Surg. Oncol. 2015, 22 (Suppl. 3), S822–S831. [Google Scholar] [CrossRef] [PubMed]
- Burant, C.F.; Takeda, J.; Brot-Laroche, E.; Bell, G.I.; Davidson, N.O. Fructose transporter in human spermatozoa and small intestine is GLUT5. J. Biol. Chem. 1992, 267, 14523–14526. [Google Scholar] [PubMed]
- Rand, E.B.; Depaoli, A.M.; Davidson, N.O.; Bell, G.I.; Burant, C.F. Sequence, tissue distribution, and functional characterization of the rat fructose transporter GLUT5. Am. J. Physiol. 1993, 264, G1169–G1176. [Google Scholar] [PubMed]
- Fagerberg, L.; Hallstrom, B.M.; Oksvold, P.; Kampf, C.; Djureinovic, D.; Odeberg, J.; Habuka, M.; Tahmasebpoor, S.; Danielsson, A.; Edlund, K.; et al. Analysis of the human tissue-specific expression by genome-wide integration of transcriptomics and antibody-based proteomics. Mol. Cell Proteom. 2014, 13, 397–406. [Google Scholar] [CrossRef] [PubMed]
- Medina Villaamil, V.; Aparicio Gallego, G.; Valbuena Rubira, L.; Garcia Campelo, R.; Valladares-Ayerbes, M.; Grande Pulido, E.; Victoria Bolos, M.; Santamarina Cainzos, I.; Anton Aparicio, L.M. Fructose transporter GLUT5 expression in clear renal cell carcinoma. Oncol. Rep. 2011, 25, 315–323. [Google Scholar] [CrossRef] [PubMed]
- Garriga, C.; Barfull, A.; Planas, J.M. Kinetic characterization of apical D-fructose transport in chicken jejunum. J. Membr. Biol. 2004, 197, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimi, R.; Jahromi, M.F.; Liang, J.B.; Farjam, A.S.; Shokryazdan, P.; Idrus, Z. Effect of Dietary Lead on Intestinal Nutrient Transporters mRNA Expression in Broiler Chickens. Biomed. Res. Int. 2015, 2015, 149745. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Manolescu, A.; Ritzel, M.; Yao, S.; Slugoski, M.; Young, J.D.; Chen, X.Z.; Cheeseman, C.I. Cloning and functional characterization of the human GLUT7 isoform SLC2A7 from the small intestine. Am. J. Physiol. Gastrointest. Liver Physiol. 2004, 287, G236–G242. [Google Scholar] [CrossRef] [PubMed]
- Boutet, E.; Lieberherr, D.; Tognolli, M.; Schneider, M.; Bansal, P.; Bridge, A.J.; Poux, S.; Bougueleret, L.; Xenarios, I. UniProtKB/Swiss-Prot, the Manually Annotated Section of the UniProt KnowledgeBase: How to Use the Entry View. Methods Mol. Biol. 2016, 1374, 23–54. [Google Scholar] [PubMed]
- Augustin, R.; Carayannopoulos, M.O.; Dowd, L.O.; Phay, J.E.; Moley, J.F.; Moley, K.H. Identification and characterization of human glucose transporter-like protein-9 (GLUT9): Alternative splicing alters trafficking. J. Biol. Chem. 2004, 279, 16229–16236. [Google Scholar] [CrossRef] [PubMed]
- Vitart, V.; Rudan, I.; Hayward, C.; Gray, N.K.; Floyd, J.; Palmer, C.N.; Knott, S.A.; Kolcic, I.; Polasek, O.; Graessler, J.; et al. SLC2A9 is a newly identified urate transporter influencing serum urate concentration, urate excretion and gout. Nat. Genet. 2008, 40, 437–442. [Google Scholar] [CrossRef] [PubMed]
- Urano, W.; Taniguchi, A.; Anzai, N.; Inoue, E.; Sekita, C.; Endou, H.; Kamatani, N.; Yamanaka, H. Association between GLUT9 and gout in Japanese men. Ann. Rheum. Dis. 2010, 69, 932–933. [Google Scholar] [CrossRef] [PubMed]
- Doege, H.; Bocianski, A.; Scheepers, A.; Axer, H.; Eckel, J.; Joost, H.G.; Schurmann, A. Characterization of human glucose transporter (GLUT) 11 (encoded by SLC2A11), a novel sugar-transport facilitator specifically expressed in heart and skeletal muscle. Biochem. J. 2001, 359, 443–449. [Google Scholar] [CrossRef] [PubMed]
- Sasaki, T.; Minoshima, S.; Shiohama, A.; Shintani, A.; Shimizu, A.; Asakawa, S.; Kawasaki, K.; Shimizu, N. Molecular cloning of a member of the facilitative glucose transporter gene family GLUT11 (SLC2A11) and identification of transcription variants. Biochem. Biophys. Res. Commun. 2001, 289, 1218–1224. [Google Scholar] [CrossRef] [PubMed]
- Scheepers, A.; Schmidt, S.; Manolescu, A.; Cheeseman, C.I.; Bell, A.; Zahn, C.; Joost, H.G.; Schurmann, A. Characterization of the human SLC2A11 (GLUT11) gene: Alternative promoter usage, function, expression, and subcellular distribution of three isoforms, and lack of mouse orthologue. Mol. Membr. Biol. 2005, 22, 339–351. [Google Scholar] [CrossRef] [PubMed]
- Doege, H.; Bocianski, A.; Joost, H.G.; Schurmann, A. Activity and genomic organization of human glucose transporter 9 (GLUT9), a novel member of the family of sugar-transport facilitators predominantly expressed in brain and leucocytes. Biochem. J. 2000, 350, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Byrne, F.L.; Poon, I.K.; Modesitt, S.C.; Tomsig, J.L.; Chow, J.D.; Healy, M.E.; Baker, W.D.; Atkins, K.A.; Lancaster, J.M.; Marchion, D.C.; et al. Metabolic vulnerabilities in endometrial cancer. Cancer Res. 2014, 74, 5832–5845. [Google Scholar] [CrossRef] [PubMed]
- Maria, Z.; Campolo, A.R.; Lacombe, V.A. Diabetes Alters the Expression and Translocation of the Insulin-Sensitive Glucose Transporters 4 and 8 in the Atria. PLoS ONE 2015, 10, e0146033. [Google Scholar] [CrossRef] [PubMed]
- Dawson, P.A.; Mychaleckyj, J.C.; Fossey, S.C.; Mihic, S.J.; Craddock, A.L.; Bowden, D.W. Sequence and functional analysis of GLUT10: A glucose transporter in the Type 2 diabetes-linked region of chromosome 20q12-13.1. Mol. Genet. Metab. 2001, 74, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Nemeth, C.E.; Marcolongo, P.; Gamberucci, A.; Fulceri, R.; Benedetti, A.; Zoppi, N.; Ritelli, M.; Chiarelli, N.; Colombi, M.; Willaert, A.; et al. Glucose transporter type 10-lacking in arterial tortuosity syndrome-facilitates dehydroascorbic acid transport. FEBS Lett. 2016, 590, 1630–1640. [Google Scholar] [CrossRef] [PubMed]
- Pujol-Gimenez, J.; Perez, A.; Reyes, A.M.; Loo, D.D.; Lostao, M.P. Functional characterization of the human facilitative glucose transporter 12 (GLUT12) by electrophysiological methods. Am. J. Physiol. Cell Physiol. 2015, 308, C1008–C1022. [Google Scholar] [CrossRef] [PubMed]
- Rogers, S.; Macheda, M.L.; Docherty, S.E.; Carty, M.D.; Henderson, M.A.; Soeller, W.C.; Gibbs, E.M.; James, D.E.; Best, J.D. Identification of a novel glucose transporter-like protein-GLUT-12. Am. J. Physiol. Endocrinol. Metab. 2002, 282, E733–E738. [Google Scholar] [PubMed]
- Pujol-Gimenez, J.; Martisova, E.; Perez-Mediavilla, A.; Lostao, M.P.; Ramirez, M.J. Expression of the glucose transporter GLUT12 in Alzheimer’s disease patients. J. Alzheimers Dis. 2014, 42, 97–101. [Google Scholar] [PubMed]
- Linden, K.C.; DeHaan, C.L.; Zhang, Y.; Glowacka, S.; Cox, A.J.; Kelly, D.J.; Rogers, S. Renal expression and localization of the facilitative glucose transporters GLUT1 and GLUT12 in animal models of hypertension and diabetic nephropathy. Am. J. Physiol. Renal Physiol. 2006, 290, F205–F213. [Google Scholar] [CrossRef] [PubMed]
- Coudert, E.; Pascal, G.; Dupont, J.; Simon, J.; Cailleau-Audouin, E.; Crochet, S.; Duclos, M.J.; Tesseraud, S.; Metayer-Coustard, S. Phylogenesis and Biological Characterization of a New Glucose Transporter in the Chicken (Gallus gallus), GLUT12. PLoS ONE 2015, 10, e0139517. [Google Scholar] [CrossRef] [PubMed]
- Uldry, M.; Ibberson, M.; Horisberger, J.D.; Chatton, J.Y.; Riederer, B.M.; Thorens, B. Identification of a mammalian H(+)-myo-inositol symporter expressed predominantly in the brain. EMBO J. 2001, 20, 4467–4477. [Google Scholar] [CrossRef] [PubMed]
- Di Daniel, E.; Mok, M.H.; Mead, E.; Mutinelli, C.; Zambello, E.; Caberlotto, L.L.; Pell, T.J.; Langmead, C.J.; Shah, A.J.; Duddy, G.; et al. Evaluation of expression and function of the H+/myo-inositol transporter HMIT. BMC Cell Biol. 2009, 10, 54. [Google Scholar] [CrossRef] [PubMed]
- Bankovic, J.; Stojsic, J.; Jovanovic, D.; Andjelkovic, T.; Milinkovic, V.; Ruzdijic, S.; Tanic, N. Identification of genes associated with non-small-cell lung cancer promotion and progression. Lung Cancer 2010, 67, 151–159. [Google Scholar] [CrossRef] [PubMed]
- Satake, W.; Nakabayashi, Y.; Mizuta, I.; Hirota, Y.; Ito, C.; Kubo, M.; Kawaguchi, T.; Tsunoda, T.; Watanabe, M.; Takeda, A.; et al. Genome-wide association study identifies common variants at four loci as genetic risk factors for Parkinson’s disease. Nat. Genet. 2009, 41, 1303–1307. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- International Chicken Genome Sequencing Consortium. Sequence and comparative analysis of the chicken genome provide unique perspectives on vertebrate evolution. Nature 2004, 432, 695–716. [Google Scholar]
- Wagstaff, P.; Kang, H.Y.; Mylott, D.; Robbins, P.J.; White, M.K. Characterization of the Avian Glut1 Glucose-Transporter - Differential Regulation of Glut1 and Glut3 in Chicken-Embryo Fibroblasts. Mol. Biol. Cell 1995, 6, 1575–1589. [Google Scholar] [CrossRef] [PubMed]
- Carver, F.M.; Shibley, I.A., Jr.; Pennington, J.S.; Pennington, S.N. Differential expression of glucose transporters during chick embryogenesis. Cell. Mol. Life Sci. 2001, 58, 645–652. [Google Scholar] [CrossRef] [PubMed]
- Carayannopoulos, M.O.; Chi, M.M.; Cui, Y.; Pingsterhaus, J.M.; McKnight, R.A.; Mueckler, M.; Devaskar, S.U.; Moley, K.H. GLUT8 is a glucose transporter responsible for insulin-stimulated glucose uptake in the blastocyst. Proc. Natl. Acad. Sci. USA 2000, 97, 7313–7318. [Google Scholar] [CrossRef] [PubMed]
- Rice, B.B.; Zhang, W.; Bai, S.; Siegel, P.B.; Cline, M.A.; Gilbert, E.R. Insulin-induced hypoglycemia associations with gene expression changes in liver and hypothalamus of chickens from lines selected for low or high body weight. Gen. Comp. Endocrinol. 2014, 208, 1–4. [Google Scholar] [CrossRef] [PubMed]
| Gene | Alias | Accession Number | Species | Chromosome | Exons | A.A. | Start | End | Span |
|---|---|---|---|---|---|---|---|---|---|
| slc2a6 | 6, 9 | NP_060055.2 | Human | 9 | 11 | 507 | 133,472,024 | 133,479,059 | 7036 |
| slc2a9 | NP_064425.2 | Human | 4 | 24 | 540 | 9,826,400 | 10,021,429 | 195,030 | |
| slc2a6 | 6, 9 | AAI41169.1 | Mouse | 2 | 10 | 443 | 27,021,917 | 27,027,905 | 5989 |
| slc2a9 | AAI38214.1 | Mouse | 5 | 20 | 523 | 38,351,086 | 38,483,364 | 132,279 | |
| slc2a6 | 6, 9 | XP_017945590.1 | X. tropicalis | Unknown | 10 | 504 | 95,667 | 104,372 | 8706 |
| slc2a9 | XP_017950705.1 | X. tropicalis | 1 | 15 | 527 | 195,610,477 | 195,628,747 | 18,271 | |
| slc2a10 | NP_110404.1 | Human | 20 | 8 | 541 | 5,931,524 | 5,933,981 | 2458 | |
| slc2a11 | 10, 11 | NP_110434.3 | Human | 15 | 14 | 503 | 8,026,649 | 8,027,108 | 460 |
| slc2a8 | NP_055395.2 | Human | 9 | 11 | 477 | 127,397,231 | 127,407,246 | 10,016 | |
| slc2a12 | 8, 12 | EAW47994.1 | Human | 6 | 7 | 617 | 133,991,158 | 134,052,480 | 61,323 |
| Gene | Orthologs | Human | Chicken |
|---|---|---|---|
| glut1/slc2a1 | Conserved in human, chicken, chimpanzee, cow, mouse, rat, Rhesus monkey, zebrafish, Eremothecium gossypii. 122 organisms contain orthologs with slc2a1 [46]. | Blood-brain barrier [15]. Receptor for T-cell leukemia virus I and II [46]. Microcephaly, childhood epilepsy [18,19] , hypoglycorrhachia [20,21] , cryohydrocytosis [22], choreathetosis [23], ataxia [22], migraines [24,25] , spasticity, dyskinesia [26], indicator for cancer [27], thymic carcinoma [28]. | Hypothalamus, basal glucose uptake, ubiquitous [31,34]. |
| glut2/slc2a2 | Conserved in human, chicken, dog, chimpanzee, cow, Rhesus monkey, rat, frog and zebrafish. 168 organisms have orthologs of slc2a2 [46]. | Glycoprotein, bidirectional transport in liver, islet beta cells, intestine, kidney, glucose sensor, gene mutations associated with susceptibility to disease, noninsulin-dependent diabetes, Fanconi–Bickel syndrome. Alternative splicing results in multiple transcript variants of this gene [33,34]. | Fructose, galactose, liver, pancreas, small intestine, kidneys [1,33,57], insulin dependent [33]. |
| glut3/slc2a3 | Conserved in dog, cow, frog, chimpanzee, Rhesus monkey, mouse, rat, chicken, zebrafish, fruit fly, mosquito, Caenorhabditis elegans, Saccharomyces cerevisiae, Kluyveromyces lactis, Magnaporthe oryzae, Neurosporra crassa, Arabidopsis thaliana and rice. 70 organisms have orthologs of slc2a3 [46] | Mediates uptake of glucose, 2-deoxyglucose, galactose, mannose, xylose, fucose and other monosaccharides across the cell membrane. Gene mutation associated with Huntington’s disease [39,40] . | Neurons [1,30], insulin dependent [33]. |
| glut4/slc2a4 | Conserved in chimpanzee, Rhesus monkey, dog, cow, mouse and rat. 114 organisms have orthologs of slc2a4 [46]. | Insulin-regulated. Upon insulin stimulation, GLUT4 translocates to cell surface to transport glucose across the cell membrane. Gene mutations are associated with noninsulin-dependent diabetes mellitus [41]. | Not exist in chickens [2,80]. |
| glut5/slc2a5 | Conserved in chicken, dog, mouse, rat, chimpanzee, Rhesus monkey, cow and frog. 123 organisms have orthologs of slc2a5 [46]. | Thought to be cytochalasin β-sensitive carrier, expression in small intestine [49], adipose tissue, skeletal muscle [50], duodenum, bone marrow, kidney [51], renal cell carcinoma [52]. | Fructose, small intestine [55,57]. |
| glut6/slc2a6 | Conserved in chicken, dog, cow, chimpanzee, mouse, Rhesus monkey, zebrafish, fruit fly, mosquito and frog. 169 organisms have orthologs of slc2a6 [46]. | GLUT6/GLUT9 [46], hexose transport [63], endometrial cancer [64]. | Uncharacterized protein [78]. |
| glut7/slc2a7 | Conserved in mouse, rat, chimpanzee and Rhesus monkey. Orthologs found in 55 organisms [46]. | Glucose, fructose transport, expression in small intestine and colon, lower levels in testis and prostate [55]. | Not found in chickens. |
| glut8/slc2a8 | Conserved in chicken, dog, mouse, rat, chimpanzee, Rhesus monkey, cow, zebrafish, fruit fly, rice, A. thaliana and frog. Orthologs found in156 organisms [46]. | Insulin-regulated, binds cytochalasin β in glucose-inhibitable manner, may be dual-specific, as it is inhibitable by fructose [69]. | Ubiquitous, especially in adipose tissue, kidneys, insulin response [1,2]. |
| glut9/slc2a9 | Conserved in chicken, dog, cow, chimpanzee, mouse, rat and frog. Orthologs found in 153 organisms [46]. | Fructose, urate transport, and glucose at a low rate, urate reabsorption by proximal tubules, regulatory role in development and survival of chondrocytes [59]. | Liver [33]. |
| glut10/slc2a10 | Conserved in chicken, dog, mouse, rat, chimpanzee, Rhesus monkey, cow, frog and zebrafish. Orthologs found in 166 organisms [46]. | Liver and pancreas [66], glucose regulation, gene mutations are associated with arterial tortuosity syndrome [46]. | Uncharacterized [78]. |
| glut11/slc2a11 | Conserved in chicken, dog, cow, chimpanzee, frog, Rhesus monkey and zebrafish and frog. Orthologs found in 111 organisms [46]. | Glucose, fructose. 11-A: skeletal muscle, heart, kidney. 11-B: adipose tissue, kidney, placenta. 11-C: skeletal muscle, heart, adipose tissue, pancreas [62], 11-D [46]. | Uncharacterized [78]. |
| glut12/slc2a12 | Conserved in chicken, dog, mouse, rat, chimpanzee, Rhesus monkey, cow, frog, zebrafish, rice and A. thaliana. Orthologs found in 177 organisms [46]. | GLUT8/GLUT12 [46], skeletal muscle, heart, prostate, lower levels in brain, placenta, kidneys [69], wide variety of hexoses [68], Alzheimer’s, hypertension, diabetic neuropathy [71]. | Insulin-sensitive. May act as GLUT4 in skeletal and cardiac muscle [72]. |
| glut13/slc2a13 | Conserved in chicken, dog, cow, chimpanzee, rice, Rhesus monkey, mouse, rat, frog, zebrafish, C. elegans, S. cerevisiae, K. lactis, E. gossypii, Schizosaccharomyces pombe and A. thaliana. Orthologs found in 151 organisms [46]. | Glial cells and neurons [73], myo-inositol and related stereoisomers [74], non-small-cell lung cancer [75], Parkinson’s [76]. | Uncharacterized. |
| glut14/slc2a14 | 2 organisms have orthologs of human slc2a14 [46]. | Spermatogenesis [46], Alzheimer’s disease [47], gastric adenocarcinoma [48]. | N/A |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Byers, M.S.; Howard, C.; Wang, X. Avian and Mammalian Facilitative Glucose Transporters. Microarrays 2017, 6, 7. https://doi.org/10.3390/microarrays6020007
Byers MS, Howard C, Wang X. Avian and Mammalian Facilitative Glucose Transporters. Microarrays. 2017; 6(2):7. https://doi.org/10.3390/microarrays6020007
Chicago/Turabian StyleByers, Mary Shannon, Christianna Howard, and Xiaofei Wang. 2017. "Avian and Mammalian Facilitative Glucose Transporters" Microarrays 6, no. 2: 7. https://doi.org/10.3390/microarrays6020007
APA StyleByers, M. S., Howard, C., & Wang, X. (2017). Avian and Mammalian Facilitative Glucose Transporters. Microarrays, 6(2), 7. https://doi.org/10.3390/microarrays6020007






