A New Distribution Family for Microarray Data
Abstract
:1. Introduction
2. Methods
2.1. Gpower-normal Distribution
2.2. Relationship between Gpower-normal Models and Pseudo-dispersion Models
2.3. Quantiles and Moments
2.4. Parameter Estimation
- Given a data set represented by vector , , …, , to obtain a profile likelihood for the power p, we consider a grid of values , , …, .
- For each , the transformed data are calculated as
- Then, for each , the corresponding and are estimated, maximizing the likelihood function of the truncated normal variable.
- Then, , and are used to obtain the log-likelihood function of whose density was given by (2):
- Finally, p is chosen as the one that maximizes the log-likelihood in the grid:
3. Results
3.1. Some Examples of Gpower-normal Densities
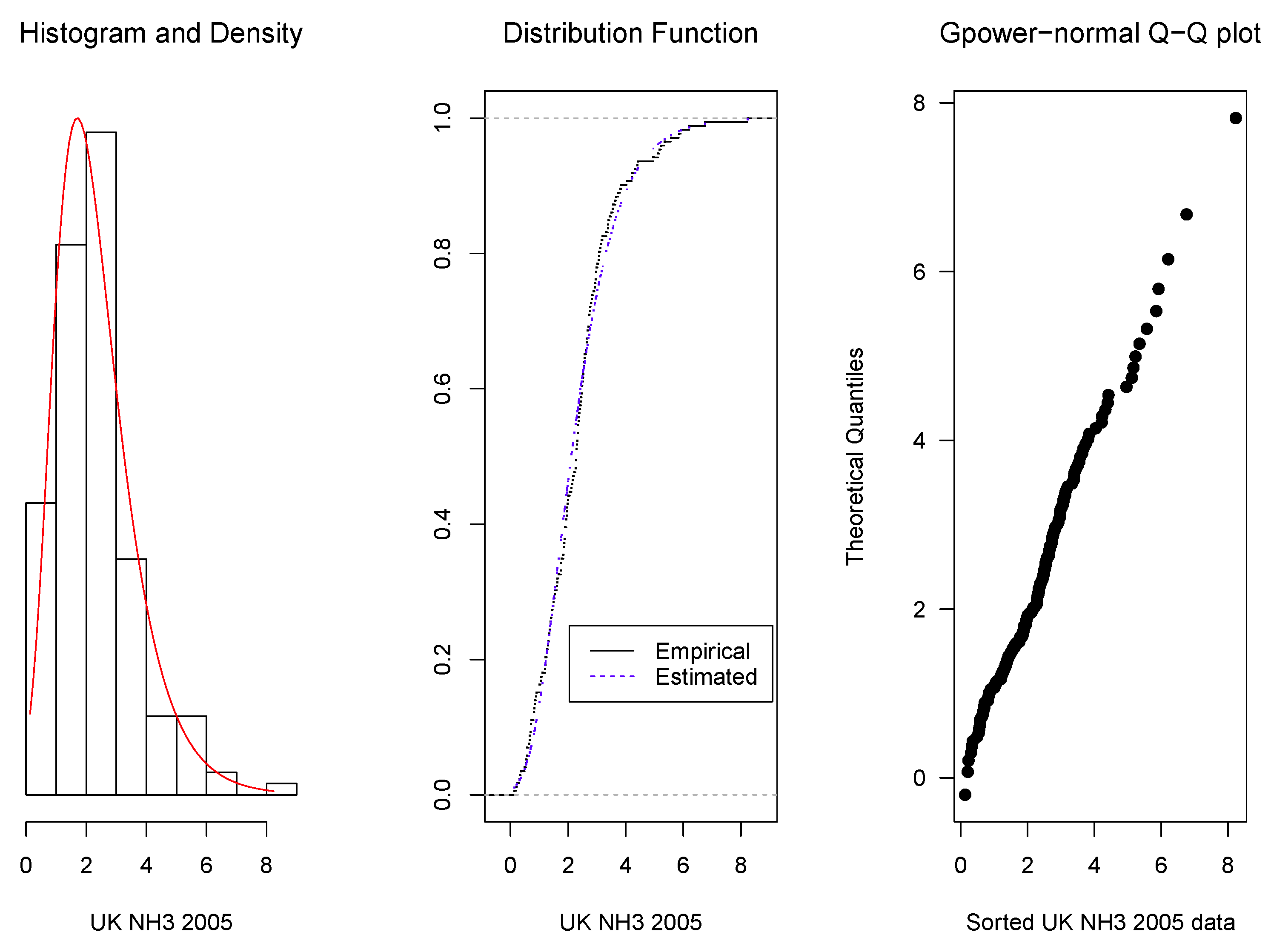
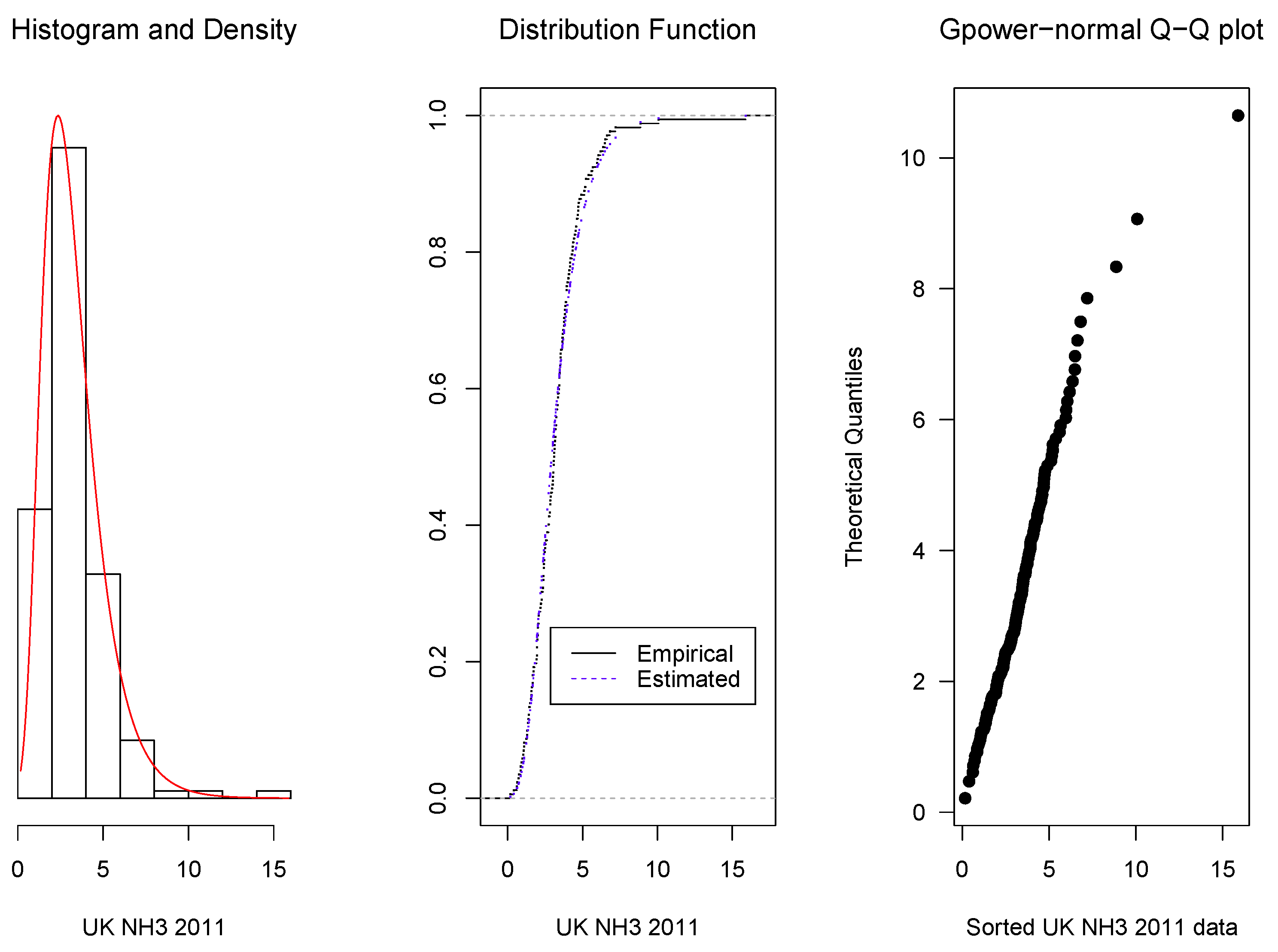
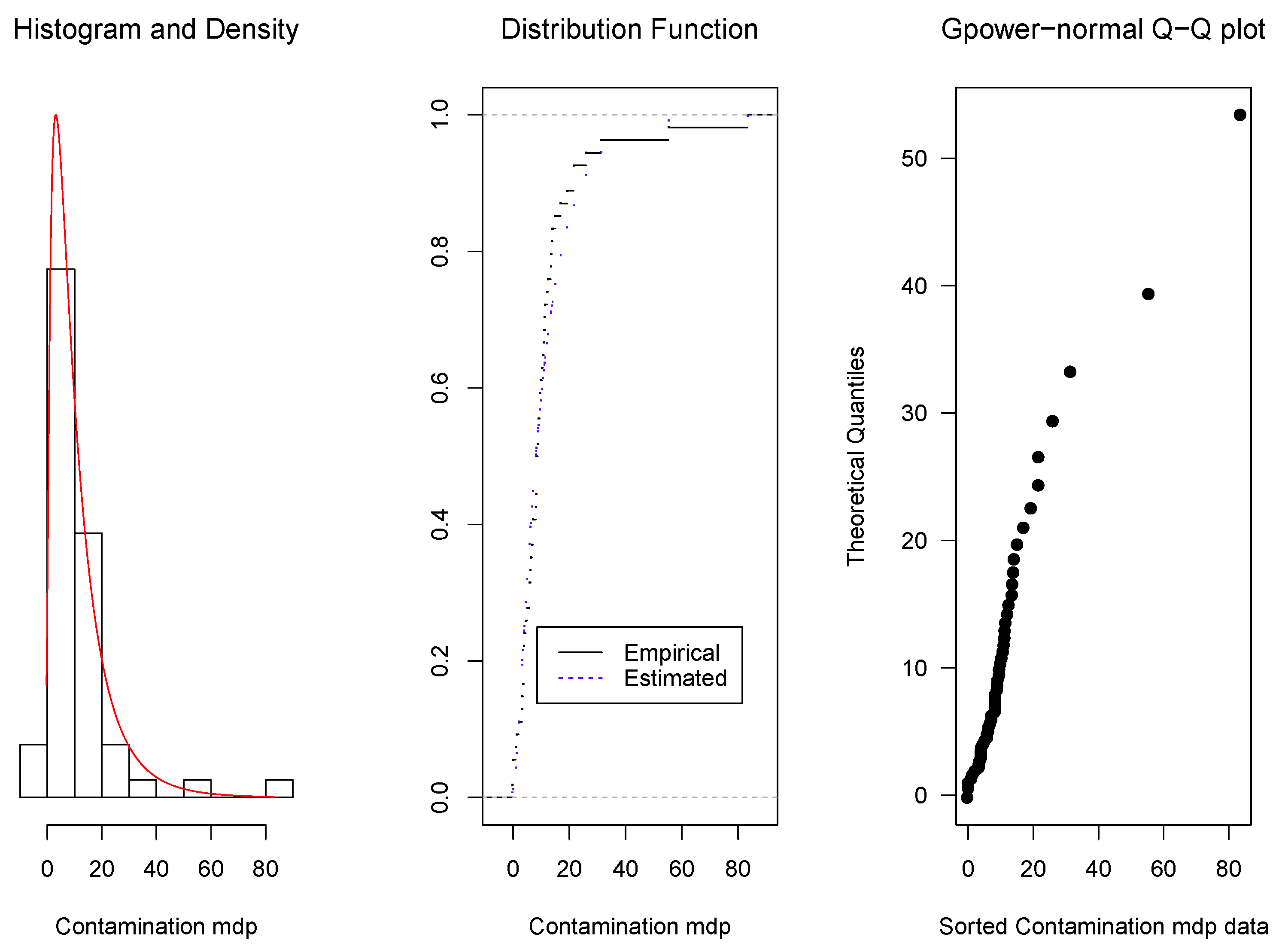
3.2. Real Data Applications
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Appendix A. Proof of Theorem 1
Appendix B. Relationship between Gpower-normal Models and Pseudo-dispersion Models
Appendix C. Moments of Gpower-normal Family
Appendix D. Data used in Examples
Example 1
Example 2
- 2005:
- 3.16 6.21 5.17 5.57 4.96 2.58 1.01 1.43 1.27 1.87 0.71 0.63 0.67 3.06 0.49 1.85 1.94 0.33 1.34 2.73 2.45 2.71 3.57 2.30 1.39 2.52 2.17 2.54 2.46 1.89 2.06 2.50 2.26 2.97 3.50 1.89 1.97 3.07 3.39 3.08 4.33 8.23 5.35 5.85 4.04 3.81 3.44 5.23 5.92 4.22 2.67 1.55 1.50 1.95 1.43 3.40 2.38 2.96 1.36 0.13 0.21 0.69 1.96 0.23 0.55 2.13 3.55 4.40 2.73 5.12 3.64 3.85 2.12 2.49 2.33 2.53 2.64 4.42 3.67 3.13 2.64 3.02 2.92 2.73 2.97 2.36 3.74 2.16 2.76 3.12 3.33 1.84 2.29 2.63 2.54 2.41 2.98 2.47 2.34 2.29 2.57 2.67 2.32 2.28 1.40 1.76 2.79 2.34 2.55 2.82 2.29 2.22 1.63 0.54 0.31 1.01 1.62 0.71 1.94 1.47 0.36 0.57 1.61 0.81 2.43 2.65 1.20 1.38 1.26 1.85 1.54 1.31 0.89 0.83 0.85 1.30 1.75 1.34 1.76 2.83 2.00 1.22 1.51 1.88 2.37 2.88 3.20 3.38 4.23 2.01 1.06 1.05 1.77 1.22 0.90 2.77 1.21 1.89 0.84 0.72 1.11 0.81 0.66 2.28 2.33 2.49 2.93 1.67 1.98 0.58 6.76 0.58.
- 2011:
- 3.25 6.82 7.20 6.49 5.40 4.09 2.84 2.28 2.08 3.25 1.57 1.06 0.75 3.44 0.86 2.37 2.45 0.63 1.67 3.43 3.45 4.29 4.47 2.77 1.95 3.12 2.77 3.47 4.19 2.73 3.76 3.92 4.17 6.19 4.37 2.46 2.82 3.65 4.71 3.50 6.50 15.89 8.87 10.08 5.97 6.36 4.61 5.99 6.64 4.69 3.06 1.66 2.02 1.73 1.35 3.20 2.02 2.73 1.43 0.17 0.40 1.43 2.36 0.87 1.24 2.88 4.15 4.78 4.74 6.05 3.92 3.98 3.67 2.83 3.14 3.54 3.01 5.22 4.56 3.94 3.27 3.81 3.82 3.92 4.33 3.45 5.12 3.64 4.42 5.16 5.67 2.90 5.22 4.70 4.32 3.70 3.88 3.35 3.02 3.06 3.18 3.55 3.10 3.22 1.93 2.19 3.52 3.15 3.33 3.85 2.83 2.97 2.49 1.55 0.61 1.59 2.26 1.97 4.11 2.55 1.67 2.38 2.05 2.22 3.05 3.75 3.09 1.96 2.09 2.64 2.40 2.01 1.50 1.35 1.41 1.96 2.46 1.97 2.50 3.52 2.64 1.80 2.08 2.37 3.77 4.05 4.61 4.91 5.61 3.20 1.38 3.36 3.35 1.60 1.09 3.48 1.71 2.40 0.99 1.04 1.30 1.10 0.93 3.00 3.08 3.27 3.66 2.28 2.42 0.72 4.54 1.29.
Example 3
References
- Speed, T. Statistical Analysis of Gene Expression Data; Chapman and Hall: London, UK, 2003. [Google Scholar]
- Smyth, G.; Yang, Y.; Speed, T. Statistical Issues in cDNA Microarray Data Analysis. Methods Mol. Biol. 2003, 224, 111–136. [Google Scholar] [PubMed]
- Durbin, B.; Hardin, J.; Hawkins, D.; Rocke, D. A variance-stabilizing transformation for gene-expression microarray data. Bioinformatics 2002, 18, 247–252. [Google Scholar] [CrossRef]
- Huber, W.H.; Sueltmann, H.A.; Poustka, A.; Vingron, M. Parameter estimation for the calibration and variance stabilization of microarray data. Stat. Appl. Genet. Mol. Biol. 2003, 2. [Google Scholar] [CrossRef]
- Kelmansky, D.M.; Martínez, E.J.; Leiva, V. A new variance stabilizing transformation for gene expression data. Stat. Appl. Genet. Mol. Biol. 2013, 12, 653–666. [Google Scholar] [CrossRef] [PubMed]
- Box, G.E.P.; Cox, D.R. An Analysis of Transformations. J. R. Stat. Soc. Ser. B (Meth.) 1964, 26, 211–252. [Google Scholar]
- Yang, Y.; Dudoit, S.; Luu, P.; Lin, D.; Peng, V.; Ngai, J.; Speed, T. Normalization for cDNA microarray data: A robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 2002, 30, e15. [Google Scholar] [CrossRef] [PubMed]
- Allison, D.B.; Cui, X.; Page, G.P.; Sabripour, M. Microarray data analysis: From disarray to consolidation and consensus. Nat. Rev. Genet. 2006, 7, 55–65. [Google Scholar] [CrossRef] [PubMed]
- Dabney, A.R.; Storey, J.D. Normalization of two-channel microarrays accounting for experimental design and intensity-dependent relationships. Genome Biol. 2007, 8, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson, H.; Hössjer, O. Methodological study of affine transformations of gene expression data with proposed robust non-parametric multi-dimensional normalization method. BMC Bioinform. 2006, 7, 1–18. [Google Scholar]
- Leiva, V.; Sanhueza, A.; Kelmansky, D.; Martinez, E. On the glog-normal distribution and its association with the gene expression problem. Comput. Stat. Data Anal. 2009, 53, 1613–1621. [Google Scholar] [CrossRef]
- Freeman, J.; Modarres, S. Inverse Box-Cox: The power-normal distribution. Stat. Probab. Lett. 2006, 76, S105–S110. [Google Scholar] [CrossRef]
- Dhrymes, P.J. Moments of Truncated (Normal) Distributions 2005. Available online: http://www.columbia.edu/lpjd1/l (accessed on 15 May 2012).
- Jørgensen, B. The Theory of Dispersion Models; Chapman and Hall: London, UK, 1997. [Google Scholar]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013. [Google Scholar]
- MicroArray Quality Control (MAQC) Consortium. Available online: ftp://ftp.ncbi.nlm.nih.gov/geo/series/GSE5nnn/GSE5350/suppl/GSE5350_MAQC_H25K_2_30GPRs.zip (accessed on 2 February 2007).
- Natural Environment Research Council (NERC), UK. Available online: http://www.pollutantdeposition.ceh.ac.uk/data (accessed on 5 June 2013).
- Chaparro, M.A.; Miranda, A.C.; Chaparro, D.M.; Gargiulo, J.L.; Bohnel, H. Biomonitoreo Magnético de Polvos Antropogénicos en Árboles de Mar del Plata (Argentina). In Proceedings of the Reunión Anual 2016 Unión Geofísica Mexicana, Puerto Vallarta, Jalisco, México, 30 October–4 November 2016.
- Morris, C.N. Natural exponential families with quadratic variance functions. Ann. Stat. 1982, 10, 65–80. [Google Scholar] [CrossRef]


| p = 0.05 | p = 0.50 | |||||
|---|---|---|---|---|---|---|
| α | σ = 1 | σ = 5 | σ = 10 | σ = 1 | σ = 5 | σ = 10 |
| 0.01 | −11.78 | −3.65 × 7 | −1.63 × 22 | −166.37 | −508.68 | −182.81 |
| 0.05 | −5.39 | −3.98 × 104 | −2.73 × 1011 | −13.84 | −20.83 | −7.25 |
| 0.10 | −3.49 | −2.26 × 103 | −4.64 × 107 | −5.67 | −5.17 | −1.33 |
| 0.50 | 0.00 | 3.97 × 10−4 | 0.57 | 0.06 | 4.25 | 14.16 |
| 0.90 | 3.17 | 259.00 | 2.17 × 104 | 2.34 | 22.75 | 78.52 |
| 0.95 | 4.65 | 981.00 | 1.74 × 105 | 3.04 | 31.37 | 109.85 |
| 0.99 | 8.92 | 9590.00 | 5.24 × 106 | 4.48 | 51.85 | 185.57 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Kelmansky, D.M.; Ricci, L. A New Distribution Family for Microarray Data. Microarrays 2017, 6, 5. https://doi.org/10.3390/microarrays6010005
Kelmansky DM, Ricci L. A New Distribution Family for Microarray Data. Microarrays. 2017; 6(1):5. https://doi.org/10.3390/microarrays6010005
Chicago/Turabian StyleKelmansky, Diana Mabel, and Lila Ricci. 2017. "A New Distribution Family for Microarray Data" Microarrays 6, no. 1: 5. https://doi.org/10.3390/microarrays6010005
APA StyleKelmansky, D. M., & Ricci, L. (2017). A New Distribution Family for Microarray Data. Microarrays, 6(1), 5. https://doi.org/10.3390/microarrays6010005




