Aptamer Microarrays—Current Status and Future Prospects
Abstract
:1. Introduction
2. Aptamer Microarrays
2.1. Conventional Microarrays vs. Aptamer Microarrays
2.2. Strategies for Aptamer Microarray Fabrication
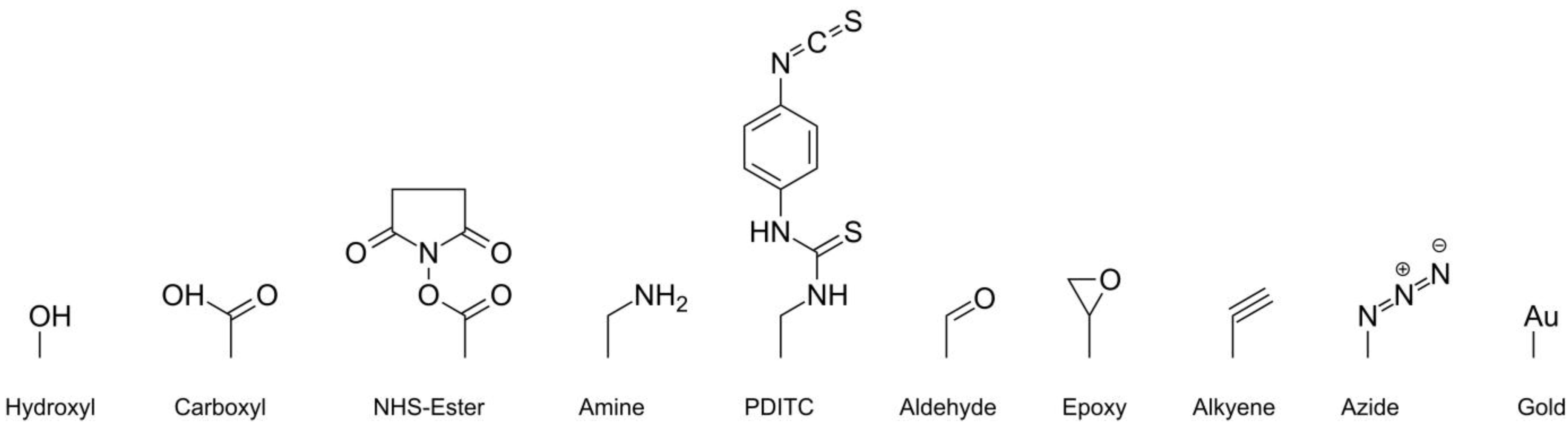
2.3. Attachment Chemistries for Post Synthesis Immobilization
2.4. Steric Requirements of Aptamers
2.4.1. Surface Density
2.4.2. Surface Charge
2.4.3. Proximity to Surface and Spacers
| Spacer type | Abbreviation | Length/unit [angstrom] | Hydrophobicity |
|---|---|---|---|
| Aliphatic carbon chain | C | 1.57 [32] | Hydrophobic |
| Polyethylene glycol | PEG | 3.51 [33] | Hydrophilic |
| Polyethylene imine | PEI | 2.9–3.5 [34] | Hydrophilic |
| Poly thymine | poly(dT) | 3.4 [35] | Hydrophilic |
2.5. Assay Formats
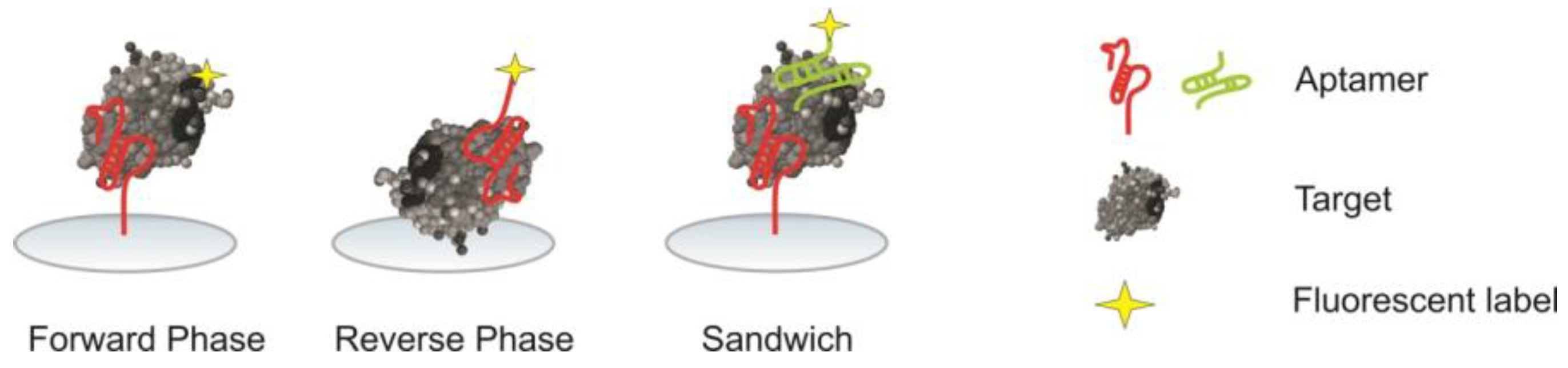
2.5.1. Forward Phase
2.5.2. Reverse Phase
2.5.3. Sandwich
2.6. Detection Methods
2.6.1. Fluorescent Dye-Based
2.6.2. SPRi
2.6.3. Other
2.7. Applications of Aptamer Microarrays
| Target Name | Assay-Format | Multiplexing | Detection technique | Limit of detection | Author + Year |
|---|---|---|---|---|---|
| Toll-like receptor 2 | forward | fluorescence scanner | Chang 2009 [54] | ||
| Thrombin/ E. coli total protein | forward | 12 | fluorescence scanner | Chen 2013 [38] | |
| Human Thrombin/VEGF | forward | 2 | SPR | Chen 2012 [60] | |
| 4 different proteins | forward | 2 × 2 | fluorescence scanner | pM–nM | Cho 2006 [10] |
| Human angiopoietin-2 | forward | 15.000 | fluorescence scanner | Cho 2013 [61] | |
| Lysozyme; IgE | forward | 4 | fluorescence scanner | 70 fM; 5.2 fM | Collett 2005 [62] |
| Lysozyme | forward | 26 | fluorescence scanner | 70 fM | Collett 2005 [39] |
| Thrombin | forward | SPR | 100 pM | Daniel 2013 [63] | |
| IgE | forward | 15.000 | fluorescence scanner | Fischer 2008 [55] | |
| Streptavidin | forward | yes (?) | fluorescence scanner | Franssen-van Hal 2013 [9] | |
| Thrombin | forward | 2 | fluorescence scanner | 30–50 pM | Lao 2009 [35] |
| human fIXa | forward | 5 | SPR | 10 nM | Li 2006 [64] |
| 4 different proteins | forward | 4 | fluorescence polarization | McCauley2003 [65] | |
| Thrombin | forward | 4.6 × 104 | fluorescence scanner | Platt 2009 [57] | |
| PFEI-His | forward | fluorescence scanner | Sinitsyna 2012 [28] | ||
| BGL-His + streptavidin | forward | fluorescence scanner | Walter 2008 [30] | ||
| IgE; IgG | forward | SPRi | 2 nM | Wang 2007 [66] | |
| PFEI-His | forward | fluorescence scanner | Zhu 2011 [36] | ||
| 17 different proteins | forward/heterogenic sandwich | 17 | fluorescence scanner | pM–nM | Bock 2004 [41] |
| Thrombin | forward/sandwich | resonance waveguide diffraction; fluorescence reader | Edwards 2010 [27] | ||
| IgE | heterogenic sandwich | SPR | 1 fM | Kim 2010 [67] | |
| Thrombin / VEGF | heterogenic sandwich | 4 + 1 Control | SPR | 1 pM (VEGF) | Li 2007 [68] |
| PFEI-His | reverse | fluorescence scanner | 30 nM | Lübbecke 2012 [8] | |
| HCVNS3 protein | reverse | confocal laser scanning microscope | 73 pM | Roh 2010 [69] | |
| Yeast TBP (TATA Binding Protein) | sandwich | fluorescence scanner | nM–µM | Ahn 2010 [47] | |
| Thrombin | sandwich | fluorescence scanner | 64 pM | Edwards 2010 [70] | |
| Thrombin | sandwich | fluorescence scanner | 0.17/0.75 nM | Meneghello 2012 [71] | |
| 8 different proteins | sandwich | 8 | dual laser flow system; fluorescence | 1–100 pM | Ochsner 2014 [72] |
| C-reactive protein (CRP) | sandwich | fluorescence scanner | 43 pM | Pultar 2009 [29] | |
| Thrombin + VEGF | sandwich | 2 | fluorescence scanner | 50 nM | Sosic 2013 [73] |
| Thrombin | sandwich | fluorescence scanner | Sosic 2011 [74] | ||
| Thrombin | sandwich | fluorescence microscope | 0.27 nM | Tennico 2010 [75] | |
| Ethanolamine | pseudo-sandwich (TID) | fluorescence scanner | 10 pM | Heilkenbrinker 2014 [76] |
2.8. Current Limitations and Future Prospects
3. Conclusion and Outlook
Author Contributions
Conflicts of Interest
References
- Patel, D.J.; Suri, A.K.; Jiang, F.; Jiang, L.; Fan, P.; Kumar, R.A.; Nonin, S. Structure, recognition and adaptive binding in RNA aptamer complexes. J. Mol. Biol. 1997, 272, 645–664. [Google Scholar] [CrossRef] [PubMed]
- Hermann, T.; Patel, D.J. Adaptive recognition by nucleic acid aptamers. Science 2000, 287, 820–825. [Google Scholar] [CrossRef] [PubMed]
- Stoltenburg, R.; Reinemann, C.; Strehlitz, B. SELEX—A (r)evolutionary method to generate high-affinity nucleic acid ligands. Biomol. Eng. 2007, 24, 381–403. [Google Scholar] [CrossRef] [PubMed]
- Ozer, A.; Pagano, J.M.; Lis, J.T. New Technologies Provide Quantum Changes in the Scale, Speed, and Success of SELEX Methods and Aptamer Characterization. Mol. Ther. Nucleic Acids 2014, 3, e183. [Google Scholar] [CrossRef] [PubMed]
- Walter, J.-G.; Stahl, F.; Scheper, T. Aptamers as affinity ligands for downstream processing. Eng. Life Sci. 2012, 12, 496–506. [Google Scholar] [CrossRef]
- Walter, J.-G.; Heilkenbrinker, A.; Austerjost, J.; Timur, S.; Stahl, F.; Scheper, T. Aptasensors for Small Molecule Detection. Zeitschrift für Naturforsch. B 2012, 67b, 976–986. [Google Scholar] [CrossRef]
- Lönne, M.; Zhu, G.; Stahl, F.; Walter, J.G. Aptamer-modified nanoparticles as biosensors. Adv. Biochem. Eng. Biotechnol. 2014, 140, 121–154. [Google Scholar] [PubMed]
- Lübbecke, M.; Walter, J.-G.; Stahl, F.; Scheper, T. Aptamers as detection molecules on reverse phase protein microarrays for the analysis of cell lysates. Eng. Life Sci. 2012, 12, 144–151. [Google Scholar] [CrossRef]
- Franssen-van Hal, N.L.; van der Putte, P.; Hellmuth, K.; Matysiak, S.; Kretschy, N.; Somoza, M.M. Optimized Light-Directed Synthesis of Aptamer Microarrays. Anal. Chem. 2013, 85, 5950–5957. [Google Scholar] [CrossRef] [PubMed]
- Cho, E.J.; Collett, J.R.; Szafranska, A.E.; Ellington, A.D. Optimization of aptamer microarray technology for multiple protein targets. Anal. Chim. Acta 2006, 564, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Krishnan, S.; Weinman, C.J.; Ober, C.K. Advances in polymers for anti-biofouling surfaces. J. Mater. Chem. 2008, 18, 3405. [Google Scholar] [CrossRef]
- Soler, M.; Estevez, M.C.; Alvarez, M.; Otte, M.A.; Sepulveda, B.; Lechuga, L.M. Direct detection of protein biomarkers in human fluids using site-specific antibody immobilization strategies. Sensors 2014, 14, 2239–2258. [Google Scholar] [CrossRef] [PubMed]
- Kwiatkowski, M.; Nilsson, M.; Landegren, U. Synthesis of full-length oligonucleotides: Cleavage of apurinic molecules on a novel support. Nucleic Acids Res. 1996, 24, 4632–4638. [Google Scholar] [CrossRef] [PubMed]
- Gao, X.; Gulari, E.; Zhou, X. In Situ Synthesis of Oligonucleotide Microarrays. Biopolymers 2004, 73, 579–596. [Google Scholar] [CrossRef] [PubMed]
- Piran, U.; Riordan, W.J. Dissociation rate constant of the biotin-streptavidin complex. J. Immunol. Methods 1990, 133, 141–143. [Google Scholar] [CrossRef] [PubMed]
- Balamurugan, S.; Obubuafo, A.; Soper, S.A.; Spivak, D. A Surface immobilization methods for aptamer diagnostic applications. Anal. Bioanal. Chem. 2008, 390, 1009–1021. [Google Scholar] [CrossRef] [PubMed]
- Potyrailo, R.A.; Conrad, R.C.; Ellington, A.D.; Hieftje, G.M. Adapting selected nucleic acid ligands (aptamers) to biosensors. Anal. Chem. 1998, 70, 3419–3425. [Google Scholar] [CrossRef] [PubMed]
- Sehgal, D.; Vijay, I.K. A method for the high efficiency of water-soluble carbodiimide-mediated amidation. Anal. Biochem. 1994, 218, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Gong, P.; Grainger, D.W. Comparison of DNA immobilization efficiency on new and regenerated commercial amine-reactive polymer microarray surfaces. Surf. Sci. 2004, 570, 67–77. [Google Scholar] [CrossRef]
- Lee, H.; Rho, J.; Messersmith, P.B. Facile conjugation of biomolecules onto surfaces via mussel adhesive protein inspired coatings. Adv. Mater. 2009, 21, 431–434. [Google Scholar] [CrossRef] [PubMed]
- Kalia, J.; Raines, R.T. Catalysis of imido-group hydrolysis in a maleimide conjugate. Bioorg. Med. Chem. Lett. 2007, 17, 6286–6289. [Google Scholar] [CrossRef] [PubMed]
- Miller, A.W.; Robyt, J.F. Sodium cyanoborohydride in the immobilization of proteins to glutaraldehyde-activated aminoalkyl silica. Biotechnol. Bioeng. 1983, 25, 2795–2800. [Google Scholar] [CrossRef] [PubMed]
- Uszczyńska, B.; Ratajczak, T.; Frydrych, E.; Maciejewski, H.; Figlerowicz, M.; Markiewicz, W.T.; Chmielewski, M.K. Application of click chemistry to the production of DNA microarrays. Lab Chip 2012, 12, 1151–1156. [Google Scholar] [CrossRef] [PubMed]
- Escorihuela, J.; Bañuls, M.J.; Grijalvo, S.; Eritja, R.; Puchades, R.; Maquieira, Á. Direct covalent attachment of DNA microarrays by rapid thiol-ene “click” chemistry. Bioconjug. Chem. 2014, 25, 618–627. [Google Scholar] [CrossRef] [PubMed]
- Piana, S.; Bilic, A. The nature of the adsorption of nucleobases on the gold [111] surface. J. Phys. Chem. B 2006, 110, 23467–23471. [Google Scholar] [CrossRef] [PubMed]
- Zammatteo, N.; Jeanmart, L.; Hamels, S.; Courtois, S.; Louette, P.; Hevesi, L.; Remacle, J. Comparison between different strategies of covalent attachment of DNA to glass surfaces to build DNA microarrays. Anal. Biochem. 2000, 280, 143–150. [Google Scholar] [CrossRef] [PubMed]
- Edwards, K.A.; Baeumner, A.J. Aptamer sandwich assays: Label-free and fluorescence investigations of heterogeneous binding events. Anal. Bioanal. Chem. 2010, 398, 2635–2644. [Google Scholar] [CrossRef] [PubMed]
- Sinitsyna, E.S.; Walter, J.G.; Vlakh, E.G.; Stahl, F.; Kasper, C.; Tennikova, T.B. Macroporous methacrylate-based monoliths as platforms for DNA microarrays. Talanta 2012, 93, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Pultar, J.; Sauer, U.; Domnanich, P.; Preininger, C. Aptamer-antibody on-chip sandwich immunoassay for detection of CRP in spiked serum. Biosens. Bioelectron. 2009, 24, 1456–1461. [Google Scholar] [CrossRef] [PubMed]
- Walter, J.-G.; Kökpinar, O.; Friehs, K.; Stahl, F.; Scheper, T. Systematic investigation of optimal aptamer immobilization for protein-microarray applications. Anal. Chem. 2008, 80, 7372–7378. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Zhang, K.; Libera, J.A.; Olvera de la Cruz, M.; Bedzyk, M.J. Polynucleotide adsorption to negatively charged surfaces in divalent salt solutions. Biophys. J. 2006, 90, 1164–1174. [Google Scholar] [CrossRef] [PubMed]
- Allen, F.H.; Kennard, O.; Watson, D.G.; Brammer, L.; Orpen, A.G.; Taylor, R. Tables of bond lengths determined by X-ray and neutron diffraction. Part 1. Bond lengths in organic compounds. J. Chem. Soc. Perkin Trans. 1987, 2, S1–S19. [Google Scholar] [CrossRef]
- Ham, A.S.; Klibanov, A.L.; Lawrence, M.B. Action at a distance: Lengthening adhesion bonds with Poly(ethylene glycol) spacers enhances mechanically stressed affinity for improved vascular targeting of microparticles. Langmuir 2009, 25, 10038–10044. [Google Scholar] [CrossRef] [PubMed]
- Ziebarth, J.D.; Wang, Y. Understanding the protonation behavior of linear polyethylenimine in solutions through Monte Carlo simulations. Biomacromolecules 2010, 11, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Lao, Y.; Peck, K.; Chen, L. Enhancement of Aptamer Microarray Sensitivity through Spacer Optimization and Avidity Effect. Anal. Chem. 2009, 1747–1754. [Google Scholar] [CrossRef]
- Zhu, G.; Lübbecke, M.; Walter, J.-G.; Stahl, F.; Scheper, T. Characterization of Optimal Aptamer-Microarray Binding Chemistry and Spacer Design. Chem. Eng. Technol. 2011, 34, 2022–2028. [Google Scholar] [CrossRef]
- Brut, M.; Trapaidze, A.; Estève, A.; Bancaud, A.; Estève, D.; Landa, G.; Djafari-Rouhani, M. Bringing aptamers into technologies: Impact of spacer terminations. Appl. Phys. Lett. 2012, 163702, 3–6. [Google Scholar]
- Chen, L.-C.; Tzeng, S.-C.; Peck, K. Aptamer microarray as a novel bioassay for protein-protein interaction discovery and analysis. Biosens. Bioelectron. 2013, 42, 248–255. [Google Scholar] [CrossRef] [PubMed]
- Collett, J.R.; Cho, E.J.; Lee, J.F.; Levy, M.; Hood, A.J.; Wan, C.; Ellington, A.D. Functional RNA microarrays for high-throughput screening of antiprotein aptamers. Anal. Biochem. 2005, 338, 113–123. [Google Scholar] [CrossRef] [PubMed]
- Ramakrishnan, R.; Dorris, D.; Lublinsky, A.; Nguyen, A.; Domanus, M.; Prokhorova, A.; Gieser, L.; Touma, E.; Lockner, R.; Tata, M.; et al. An assessment of Motorola CodeLink microarray performance for gene expression profiling applications. Nucleic Acids Res. 2002, 30, e30. [Google Scholar] [CrossRef] [PubMed]
- Bock, C.; Coleman, M.; Collins, B.; Davis, J.; Foulds, G.; Gold, L.; Greef, C.; Heil, J.; Heilig, J.S.; Hicke, B.; et al. Photoaptamer arrays applied to multiplexed proteomic analysis. Proteomics 2004, 4, 609–618. [Google Scholar] [CrossRef] [PubMed]
- Lyng, H.; Badiee, A.; Svendsrud, D.H.; Hovig, E.; Myklebost, O.; Stokke, T. Profound influence of microarray scanner characteristics on gene expression ratios: Analysis and procedure for correction. BMC Genomics 2004, 5, 10. [Google Scholar] [CrossRef] [PubMed]
- Wear, M.A.; Patterson, A.; Malone, K.; Dunsmore, C.; Turner, N.J.; Walkinshaw, M.D. A surface plasmon resonance-based assay for small molecule inhibitors of human cyclophilin A. Anal. Biochem. 2005, 345, 214–226. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Wilson, W.D. Quantitative analysis of small molecule-nucleic acid interactions with a biosensor surface and surface plasmon resonance detection. Methods Mol. Biol. 2010, 613, 1–23. [Google Scholar] [PubMed]
- Ghindilis, A.L.; Smith, M.W.; Schwarzkopf, K.R.; Roth, K.M.; Peyvan, K.; Munro, S.B.; Lodes, M.J.; Stöver, A.G.; Bernards, K.; Dill, K.; et al. CombiMatrix oligonucleotide arrays: Genotyping and gene expression assays employing electrochemical detection. Biosens. Bioelectron. 2007, 22, 1853–1860. [Google Scholar] [CrossRef] [PubMed]
- Cole, J.R.; Dick, L.W.; Morgan, E.J.; McGown, L.B. Affinity capture and detection of immunoglobulin E in human serum using an aptamer-modified surface in matrix-assisted laser desorption/ionization mass spectrometry. Anal. Chem. 2007, 79, 273–279. [Google Scholar] [CrossRef] [PubMed]
- Ahn, J.-Y.Y.; Lee, S.W.; Kang, H.S.; Jo, M.; Lee, D.K.; Laurell, T.; Kim, S. Aptamer Microarray Mediated Capture and Mass Spectrometry Identification of Biomarker in Serum Samples. J. Proteome Res. 2010, 9, 5568–5573. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, P. A perspective on protein microarrays. Nat. Biotechnol. 2002, 20, 225–229. [Google Scholar] [CrossRef] [PubMed]
- Urmann, K.; Walter, J.-G.; Scheper, T.; Segal, E. Label-Free Optical Biosensors Based on Aptamer-Functionalized Porous Silicon Scaffolds. Anal. Chem. 2015, 87, 1999–2006. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Mehta, G.; Srivastava, S. Label-free detection techniques for protein microarrays: Prospects, merits and challenges. Proteomics 2010, 10, 731–748. [Google Scholar] [CrossRef] [PubMed]
- Ostroff, R.M.; Bigbee, W.L.; Franklin, W.; Gold, L.; Mehan, M.; Miller, Y.E.; Pass, H.I.; Rom, W.N.; Siegfried, J.M.; Stewart, A.; et al. Unlocking biomarker discovery: Large scale application of aptamer proteomic technology for early detection of lung cancer. PLoS One 2010, 5, e15003. [Google Scholar] [CrossRef] [PubMed]
- Webber, J.; Stone, T.C.; Katilius, E.; Smith, B.C.; Gordon, B.; Mason, M.D.; Tabi, Z.; Brewis, I.A.; Clayton, A. Proteomics analysis of cancer exosomes using a novel modified aptamer-based array (SOMAscanTM) platform. Mol. Cell. Proteomics 2014, 13, 1050–1064. [Google Scholar] [CrossRef] [PubMed]
- Straw, S.; Ferrigno, P.K.; Song, Q.; Tomlinson, D.; Galdo, F. Del Proof of concept study to identify candidate biomarkers of fibrosis using high throughput peptide aptamer microarray and validate by enzyme linked immunosorbant assay. J. Biomed. Sci. Eng. 2013, 6, 32–42. [Google Scholar] [CrossRef]
- Chang, Y.-C.; Kao, W.-C.; Wang, W.-Y.; Wang, W.-Y.; Yang, R.-B.; Peck, K. Identification and characterization of oligonucleotides that inhibit Toll-like receptor 2-associated immune responses. FASEB J. 2009, 23, 3078–3088. [Google Scholar] [CrossRef] [PubMed]
- Fischer, N.O.; Tok, J.B.-H.; Tarasow, T.M. Massively parallel interrogation of aptamer sequence, structure and function. PLoS One 2008, 3, e2720. [Google Scholar] [CrossRef] [PubMed]
- Katilius, E.; Flores, C.; Woodbury, N.W. Exploring the sequence space of a DNA aptamer using microarrays. Nucleic Acids Res. 2007, 35, 7626–7635. [Google Scholar] [CrossRef] [PubMed]
- Platt, M.; Rowe, W.; Knowles, J.; Day, P.J.; Kell, D.B. Analysis of aptamer sequence activity relationships. Integr. Biol. (Camb). 2009, 1, 116–122. [Google Scholar] [CrossRef] [PubMed]
- Knight, C.G.; Platt, M.; Rowe, W.; Wedge, D.C.; Khan, F.; Day, P.J.R.; Mcshea, A.; Knowles, J.; Kell, D.B. Array-based evolution of DNA aptamers allows modelling of an explicit sequence-fitness landscape. Nucleic Acids Res. 2009, 37, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Platt, M.; Rowe, W.; Wedge, D.C.; Kell, D.B.; Knowles, J.; Day, P.J.R. Aptamer evolution for array-based diagnostics. Anal. Biochem. 2009, 390, 203–205. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Nakamoto, K.; Niwa, O.; Corn, R.M. On-Chip Synthesis of RNA Aptamer Microarrays for Multiplexed Protein Biosensing with SPR Imaging Measurements. Langmuir 2012, 28, 8281–8285. [Google Scholar] [CrossRef] [PubMed]
- Cho, M.; Soo, O.S.; Nie, J.; Stewart, R.; Eisenstein, M.; Chambers, J.; Marth, J.D.; Walker, F.; Thomson, J.A.; Soh, H.T. Quantitative selection and parallel characterization of aptamers. Proc. Natl. Acad. Sci. USA 2013, 110, 18460–18465. [Google Scholar] [CrossRef] [PubMed]
- Collett, J.R.; Cho, E.J.; Ellington, A.D. Production and processing of aptamer microarrays. Methods 2005, 37, 4–15. [Google Scholar] [CrossRef] [PubMed]
- Daniel, C.; Roupioz, Y.; Gasparutto, D.; Livache, T.; Buhot, A. Solution-phase vs. surface-phase aptamer-protein affinity from a label-free kinetic biosensor. PLoS One 2013, 8, e75419. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lee, H.J.; Corn, R.M. Fabrication and characterization of RNA aptamer microarrays for the study of protein-aptamer interactions with SPR imaging. Nucleic Acids Res. 2006, 34, 6416–6424. [Google Scholar] [CrossRef] [PubMed]
- McCauley, T.G.; Hamaguchi, N.; Stanton, M. Aptamer-based biosensor arrays for detection and quantification of biological macromolecules. Anal. Biochem. 2003, 319, 244–250. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wilkop, T.; Xu, D.; Dong, Y.; Ma, G.; Cheng, Q. Surface plasmon resonance imaging for affinity analysis of aptamer-protein interactions with PDMS microfluidic chips. Anal. Bioanal. Chem. 2007, 389, 819–825. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Lee, J.; Lee, S.J.; Lee, H.J. Ultra-sensitive detection of IgE using biofunctionalized nanoparticle-enhanced SPR. Talanta 2010, 81, 1755–1759. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lee, H.J.; Corn, R.M. Detection of Protein Biomarkers Using RNA Aptamer Microarrays and Enzymatically Amplified Surface Plasmon Resonance Imaging. Anal. Chem. 2007, 79, 1082–1088. [Google Scholar] [CrossRef] [PubMed]
- Roh, C.; Lee, H.-Y.; Kim, S.-E.; Jo, S.-K. Quantum-dots-based detection of hepatitis C virus (HCV) NS3 using RNA aptamer on chip. J. Chem. Technol. Biotechnol. 2010, 85, 1130–1134. [Google Scholar] [CrossRef]
- Edwards, K.A.; Wang, Y.; Baeumner, A.J. Aptamer sandwich assays: Human α-thrombin detection using liposome enhancement. Anal. Bioanal. Chem. 2010, 398, 2645–2654. [Google Scholar] [CrossRef] [PubMed]
- Meneghello, A.; Sosic, A.; Antognoli, A.; Cretaio, E.; Gatto, B. Development and Optimization of a Thrombin Sandwich Aptamer Microarray. Microarrays 2012, 1, 95–106. [Google Scholar] [CrossRef]
- Rohloff, J.C.; Gelinas, A.D.; Jarvis, T.C.; Ochsner, U.A.; Schneider, D.J.; Gold, L.; Janjic, N. Nucleic Acid Ligands With Protein-like Side Chains: Modified Aptamers and Their Use as Diagnostic and Therapeutic Agents. Mol. Ther. Nucleic Acids 2014, 3, e201. [Google Scholar]
- Sosic, A.; Meneghello, A.; Antognoli, A.; Cretaio, E.; Gatto, B. Development of a multiplex sandwich aptamer microarray for the detection of VEGF165 and thrombin. Sensors 2013, 13, 13425–13438. [Google Scholar]
- Sosic, A.; Meneghello, A.; Cretaio, E.; Gatto, B. Human thrombin detection through a sandwich aptamer microarray: Interaction analysis in solution and in solid phase. Sensors 2011, 11, 9426–9441. [Google Scholar]
- Tennico, Y.H.; Hutanu, D.; Koesdjojo, M.T.; Bartel, C.M.; Remcho, V.T. On-chip aptamer-based sandwich assay for thrombin detection employing magnetic beads and quantum dots. Anal. Chem. 2010, 82, 5591–5597. [Google Scholar] [PubMed]
- Heilkenbrinker, A.; Reinemann, C.; Stoltenburg, R.; Walter, J.-G.; Jochums, A.; Stahl, F.; Zimmermann, S.; Strehlitz, B.; Scheper, T. Identification of the Target Binding Site of Ethanolamine-Binding Aptamers and Its Exploitation for Ethanolamine Detection. Anal. Chem. 2014, 87, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Kökpinar, O.; Walter, J.-G.; Shoham, Y.; Stahl, F.; Scheper, T. Aptamer-based downstream processing of his-tagged proteins utilizing magnetic beads. Biotechnol. Bioeng. 2011, 108, 2371–2379. [Google Scholar] [CrossRef] [PubMed]
- Jayasena, S.D. Aptamers: An emerging class of molecules that rival antibodies in diagnostics. Clin. Chem. 1999, 45, 1628–1650. [Google Scholar] [PubMed]
- Ochsner, U.A.; Green, L.S.; Gold, L.; Janjic, N. Systematic selection of modified aptamer pairs for diagnostic sandwich assays. Biotechniques 2014, 56, 125–128, 130, 132–133. [Google Scholar] [PubMed]
- McKeague, M.; Derosa, M.C. Challenges and opportunities for small molecule aptamer development. J. Nucleic Acids 2012, 2012, 748913. [Google Scholar] [CrossRef] [PubMed]
- Nikolaus, N.; Strehlitz, B. DNA-aptamers binding aminoglycoside antibiotics. Sensors 2014, 14, 3737–3755. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Witt, M.; Walter, J.-G.; Stahl, F. Aptamer Microarrays—Current Status and Future Prospects. Microarrays 2015, 4, 115-132. https://doi.org/10.3390/microarrays4020115
Witt M, Walter J-G, Stahl F. Aptamer Microarrays—Current Status and Future Prospects. Microarrays. 2015; 4(2):115-132. https://doi.org/10.3390/microarrays4020115
Chicago/Turabian StyleWitt, Martin, Johanna-Gabriela Walter, and Frank Stahl. 2015. "Aptamer Microarrays—Current Status and Future Prospects" Microarrays 4, no. 2: 115-132. https://doi.org/10.3390/microarrays4020115
APA StyleWitt, M., Walter, J.-G., & Stahl, F. (2015). Aptamer Microarrays—Current Status and Future Prospects. Microarrays, 4(2), 115-132. https://doi.org/10.3390/microarrays4020115






