Phenotypic MicroRNA Microarrays
Abstract
:1. Introduction
2. Experimental Methods
2.1. Chemicals and Cell Culture
2.2. miRNA and siRNA Transient Transfection
2.3. Phenotypic NF-κB Assay and Data Analysis
2.4. RNA Extraction and Real-Time PCR
2.5. siRNA and miRNA Printing
2.6. Microarray-Based Phenotypic Assay and Data Analysis
3. Results and Discussion
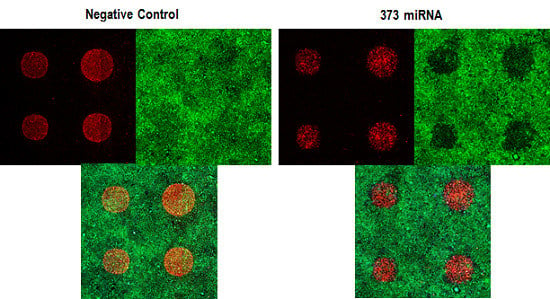
3.1. Phenotypic Assay of siRNA and miRNA Knock-down Effect in Cellular Model


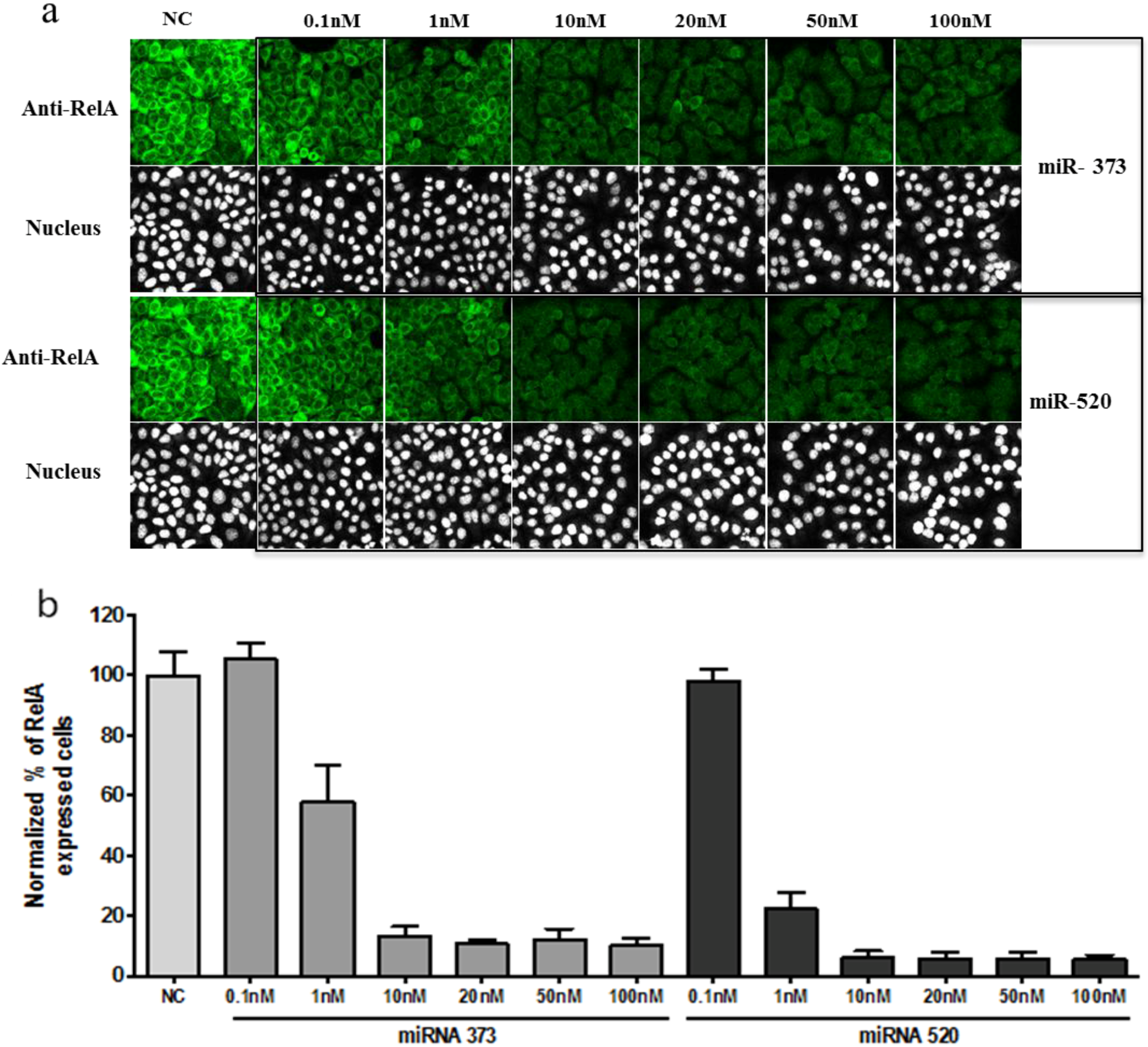
3.2. RNA Interference Microarrays
3.2.1. High-Density Spot Microarrays for Reverse Transfection

| Component of mixture for reverse transfection | |||||
|---|---|---|---|---|---|
| in wells | on array | ||||
| 1 | lipofectamine | 0.1 μL | 1 | lipofectamine | 3 μL |
| DMEM | 9.9 uL | ||||
| 2 | SiRNA (1 μM) | 5 μL (1 μM) | 2 | SiRNA (20 μM) | 2 μL (20 μM) |
| DMEM | 5 μL | 3 | 0.3 M Sucrose in OptiMEM | 2 μL (0.3 M) | |
| RNAse free water | 2 μL | ||||
| Mixed and incubate for 20 min | Mixed and incubate for 20 min | ||||
| 3 | OptiMEM | 80 μL | 4 | siGLO (20 μM) | 2 μL |
| 5 | 0.2% gelatine | 5 μL | |||
| add 100 μL to cells in 1 well/96 well plate or 20 μL in 1 well/384 well plate | Printed 3–6 μL/300 μm spot | ||||
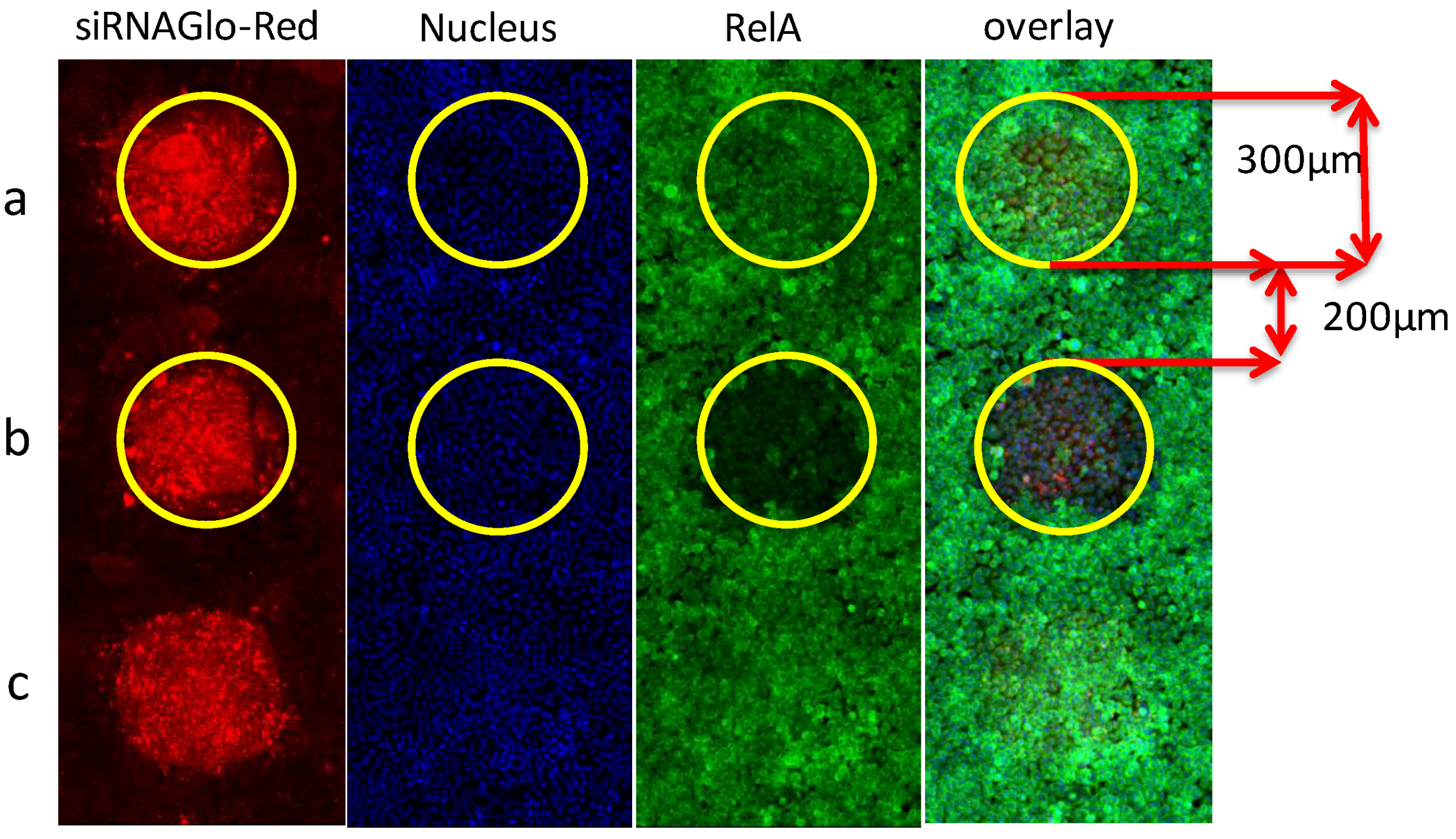

3.2.2. Optimization of the Transfection Reaction for siRNA and miRNA on Microarrays
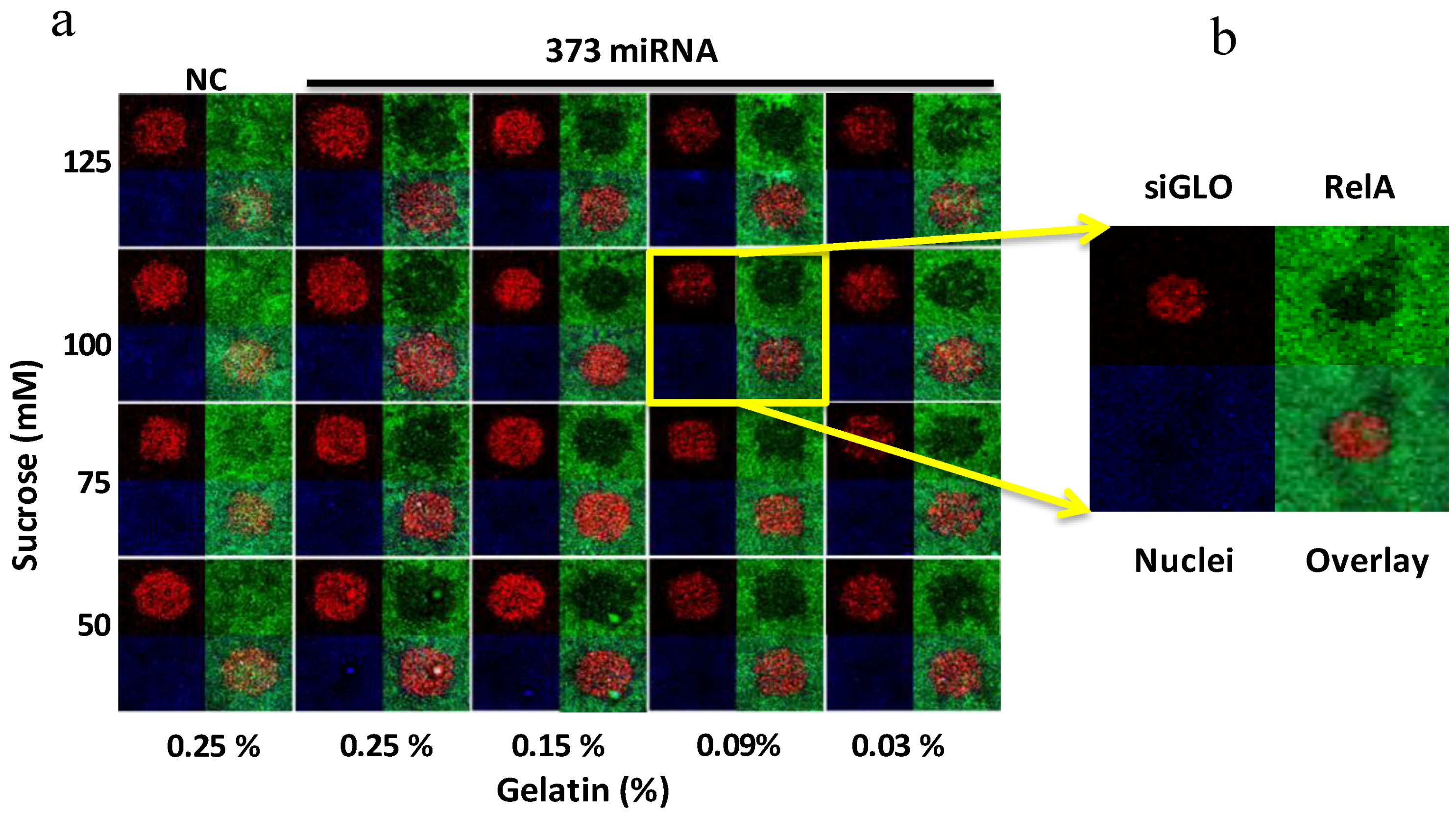

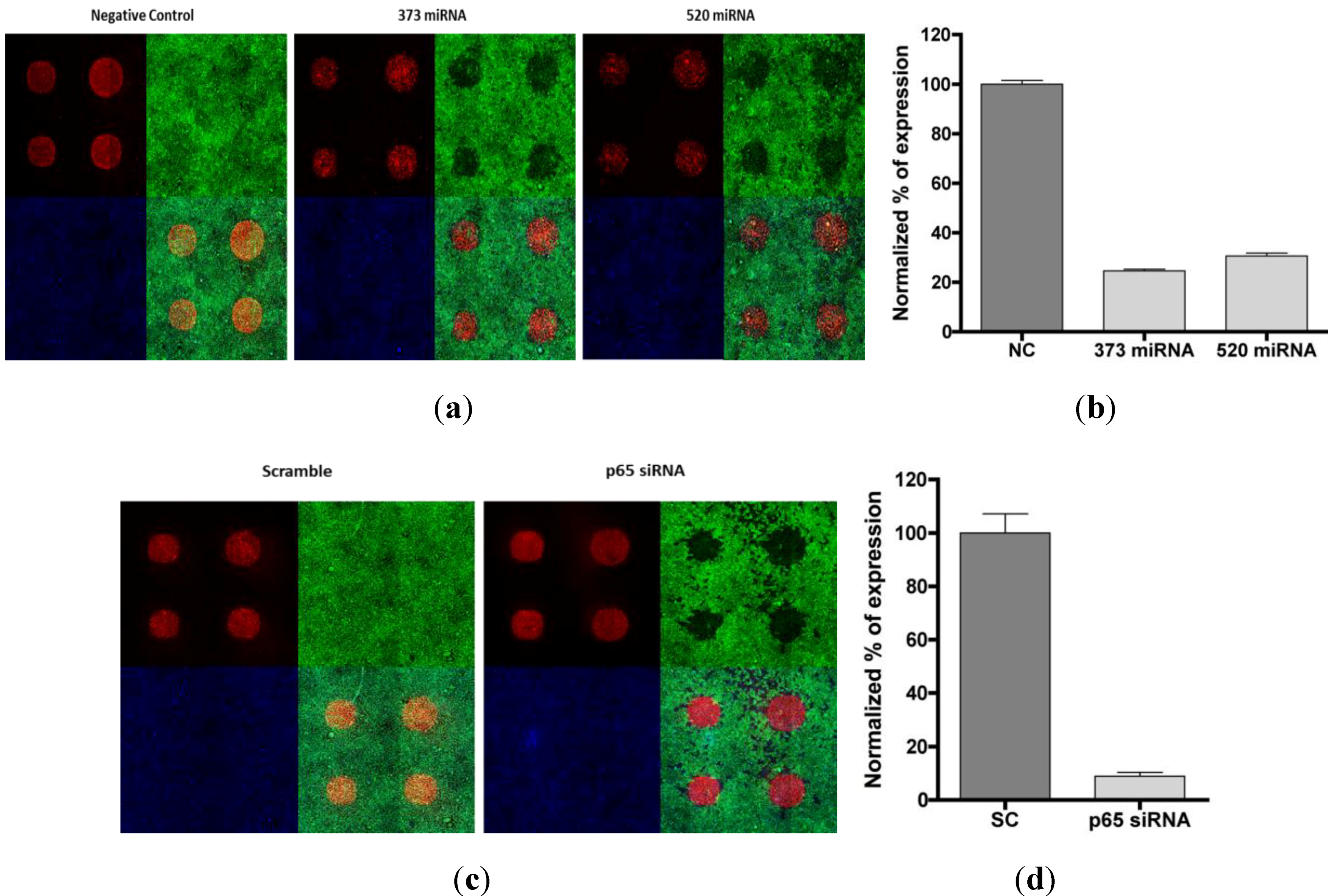
3.3. Phenotypic Assay for Cellular Monolayer Microarrays with miRNA and siRNA
3.4. Discussion
4. Conclusions
Acknowledgments
References
- Fire, A. RNA-triggered gene silencing. Trend Genet 1999, 15, 358–363. [Google Scholar] [CrossRef]
- Sharp, P.A. RNA interference—2001. Genes Dev. 2001, 15, 485–490. [Google Scholar] [CrossRef]
- Elbashir, S.M.; Harborth, J.; Lendeckel, W.; Yalch, A.; Weber, K.; Tuschl, T. Duplex of 21-nucleotide RNAs mediates RNA interference in cultured mammalian cells. Nature 2001, 411, 494–498. [Google Scholar]
- Wianny, F.; Zrnicka-Goetz, M. Specific interference with gene function by double-stranded RNA in early mouse development. Nature Cell Biol. 2000, 2, 70–75. [Google Scholar] [CrossRef]
- Sharma, S.; Rao, A. RNAi screening: Tips and techniques. Nat. Immunol. 2009, 10, 799–804. [Google Scholar] [CrossRef]
- Seyhan, A.A.; Ryan, T.E. RNAi screening for the discovery of novel modulators of human diseases. Curr. Pharmaceut. Biotechnol. 2010, 11, 735–756. [Google Scholar] [CrossRef]
- Mohr, S.; Bakal, C.; Perrimon, N. Genomic screening with RNAi: Results and challenges. Ann. Rev. Biochem. 2010, 79, 37–64. [Google Scholar] [CrossRef]
- Sood, P.; Krek, A.; Zavolan, M.; Macino, G.; Rajewsky, N. Cell-type-specific signatures of microRNAs on target mRNA expression. Proc. Natl. Acad. Sci. USA 2006, 103, 2746–2751. [Google Scholar]
- Baek, D.; Villen, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of microRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar]
- Guo, H.; Ingolia, N.T. Mammalian microRNAs predominantly act to decrease target mRNA level. Nature 2010, 466, 835–840. [Google Scholar]
- Pasquinelli, A.E.; Hunter, S.; Bracht, J. MicroRNAs: A developing story. Curr. Opin. Genet. Dev. 2005, 15, 200–205. [Google Scholar] [CrossRef]
- Janas, M.M.; Wang, E.; Love, T.; Harris, A.S.; Stevenson, K.; Semmelmann, K.; Shaffer, J.M.; Chen, P.H.; Novina, C.D. Reduced expression of ribosomal proteins relieves micro-RNA-mediate repression. Mol. Cell 2012, 46, 171–186. [Google Scholar]
- Wang, Z. The guideline of the design and validation of miRNA mimics. Methods Mol. Biol. 2011, 676, 211–223. [Google Scholar] [CrossRef]
- Sokilde, R.; Barken, K.B.; Mouritzen, P.; Moller, S.; Litman, T. MicroRNA expression analysis by LNA enhanced microarrays. In MicroRNA Profiling in Cancer; Gusev, Y., Ed.; Pan Stanford Publisher: Singapore, 2010. [Google Scholar]
- Pandey, P.; Brors, B.; Srivastava, P.K.; Bott, A.; Boehn, S.N.E.; Groene, H.J.; Gretz, N. Microarrays-based approach identifies microRNAs and their target functional patterns in polycystic kidney disease. BMC Genomics 2008. [Google Scholar] [CrossRef] [Green Version]
- Keklikoglou, I.; Koerner, C.; Schmidt, C.; Zhang, J.D.; Heckmann, D.; Shavinskaya, A.; Allgayer, H.; Guckel, B.; Fehm, T.; Schneewewiss, A.; Sahin, O.; Wiemann, S.; Tschulena, U. MicroRNA-520/373 family functions as a tumor suppressor in estrogen receptor negative breast cancer by targeting NF-κB and TGF-β signaling pathways. Oncogene 2011. [Google Scholar] [CrossRef]
- Zhang, Y.; Fan, K.J.; Sun, Q.; Chen, A.Z.; Shen, W.L.; Zhao, Z.H.; Zheng, X.F.; Yang, X. Functional screening for miRNAs targeting Smad4 identified miR-199a as a negative regulator of TGF-β signaling pathway. Nucl. Acids Res. 2012, 40. [Google Scholar] [CrossRef]
- Eulalio, A.; Mank, M.; Ferro, M.D.; Zentilin, L.; Sinagra, G.; Zacchigna, S.; Giacca, M. Functional screening identifies miRNAs inducing cardiac regeneration. Nature 2012, 492, 376–381. [Google Scholar]
- Schena, M.; Shalon, D.; Davis, R.W.; Brown, P.O. Quantitative monitoring of gene expression patterns with a complementary DNA microarray. Science 1995, 270, 467–470. [Google Scholar]
- Cheung, V.G.; Morley, M.; Aguilar, F.; Massimi, A.; Kucherlapati, R.; Childs, G. Making and Reading Microarrays. Available online: http://www.rose-hulman.edu/~ahmed/making%20and%20reading%20cdna%20microarrays.pdf (accessed on 8 February 2013).
- Barbulovic-Nad, I.; Lucente, M.; Sun, Y.; Zhang, M.; Wheeler, A.R.; Bussmann, M. Bio-microarray fabrication techniques—A review. Crit. Rev. Biotechnol. 2006, 26, 237–259. [Google Scholar] [CrossRef]
- Ziauddin, J.; Sabatini, D.M. Micro-array of cells expressing defined cDNAs. Nature 2001, 411, 107–110. [Google Scholar] [CrossRef]
- Silva, J.M.; Mizuno, H.; Brady, A.; Lucito, R.; Hannon, G.J. RNA interference microarrays: High-throughput loss-of-function genetics in mammalian cells. PNAS 2004, 101, 6548–6552. [Google Scholar]
- Mousses, S.; Caplen, N.J.; Cornelison, R.; Weaver, D.; Basik, M.; Hautaniemi, S.; Elkahloun, A.G.; Lotufo, R.A.; Choudary, A.; Dougherty, E.R.; Suh, E.; Kallioniemi, O. RNAi microarrays analysis in cultured mammalian cells. Genome Res. 2007. [Google Scholar] [CrossRef]
- Karin, M.; Greten, F.R. NF-κB: Linking inflammation and immunity to cancer development and progression. Nat. Rev. Immunol. 2005, 5, 739–759. [Google Scholar]
- Karin, M. Nuclear factor-κB in cancer development and progression. Nature 2006 441, 431–436.
- Hoffmann, A.; Baltimore, D. Circuitry of nuclear factor κB signaling. Immunol. Rev. 2006, 210, 171–186. [Google Scholar] [CrossRef]
- Oeckinghaus, A.; Ghosh, S. The NF-κB family of transcription factors and its regulation. Cold Spring Harb. Perspct. Biol. 2009. [Google Scholar] [CrossRef]
- O’Donnell, K.A.; Wentzel, E.A. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 2005, 435, 839–843. [Google Scholar]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; Hammond, S.M. A microRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar] [CrossRef]
- Mraz, M.; Pospisilova, S.; Malinova, K.; Slapak, I.; Mayer, J. MicroRNAs in chronic lymphocytic leukemia pathogenesis and disease subtypes. Leuk. Lymphoma 2009, 50, 506–509. [Google Scholar] [CrossRef]
- Cordes, K.; Srivastava, D. MicroRNA regulation of cardiovascular development. Circ. Res. 2009. [Google Scholar] [CrossRef]
- Ma, X.; Becker Buscaglia, L.E; Barker, J.R.; Li, Y. MicroRNAs in NF-κB signaling. J. Mol. Cell Biol. 2011. [Google Scholar] [CrossRef]
- Liu, P.; Wilson, M.J. MiR-520c and miR373 upregulate MMP9 expression by targeting mTOR and SIRT1, and activate the Ras/Raf/MEK/Erk signaling pathway and NF-κB factor in Human fibrosarcoma cells. J. Cell. Physiol. 2012, 277, 867–876. [Google Scholar]
- Wang, L.; Kang, F.; Shan, B.; Liu, L.; Sang, M. Targeting NF-κB p65 with an artificial microRNA suppress growth of MDA-MB-231 human triple-negative breast cancer cell line. Gene Ther. Mol. Biol. 2012, 14, 30–41. [Google Scholar]
- Huang, S.; Robinson, J.B.; Deguzman, A.; Bucana, C.D.; Fidler, I.J. Blockade of nuclear factor-κB signaling inhibits angiogenesis and tumorigenecity of human ovarian cancer cells by suppressing expression of vascular endothelial growth factor and interleukin 8. Cancer Res. 2000, 60, 5334–5339. [Google Scholar]
- Huang, Q.; Gumireddy, K.; Schrier, M.; le Sage, C.; Nagel, R.; Nair, S.; Egan, D.A.; Li, A.; Huang, G.; Pure, E.; Agami, R. The microRNAs miR-373 and miR-520c promote tumor invasion and metastasis. Nat. Cell Biol. 2008, 10, 202–210. [Google Scholar] [CrossRef]
- Erfle, H.; Neumann, B.; Liebel, U.; Rogers, P.; Held, M.; Walter, T.; Ellenberg, J.; Pepperkok, R. Reverse transfection on cell arreys for high content screening microscopy. Nat. Protocol. 2007, 2, 392–399. [Google Scholar]
- Erfle, H.; Neumann, B.; Rogers, P.; Bulkescher, J.; Ellenberg, J.; Pepperkok, R. Work flow for multiplexing siRNA assays by solid-pahse reverse trasnfection in multiwell plates. J. Biomol. Screen. 2008, 13, 575–580. [Google Scholar] [CrossRef]
- Genovesio, A.; Giardini, M.A.; Kwon, Y.J.; Dossin, F.D.M.; Choi, S.Y.; Kim, N.Y.; Kim, H.C.; Jung, S.Y.; Schenkman, S.; Almeida, I.C.; Emans, N.; Freitas-Junior, L.H.F. Visual genome-wide RNAi screening to identify human hostfactors requered for Trypanosoma cruzi infection. PLoS One 2011, 6, e19733. [Google Scholar] [CrossRef] [Green Version]
- Genovesio, A.; Kwon, Y.J.; Windisch, M.P.; Kim, N.Y.; Choi, S.Y.; Kim, H.C.; Jung, S.; Mammano, F.; Perrin, V.; Boese, A.S.; Casartelli, N.; Swartz, O.; Nehrbass, U.; Emans, N. Automated genome-wide visual profiling of cellular proteins involved in HIV infection. J. Biomol. Screen. 2011, 16, 945–958. [Google Scholar] [CrossRef]
- Ikeda, S.; Kong, S.W.; Lu, J.; Bisping, E.; Zhang, H.; Allen, P.D.; Golub, T.R.; Pieske, B.; Pu, W.T. Altered microRNA expression in human heart disease. Physiol. Genom. 2007, 31, 367–373. [Google Scholar] [CrossRef]
- Zhu, S.; Pan, W.; Sing, X.; Liu, Y.; Tang, Y.; Liang, D.; He, D.; Wang, H.; Liu, W.; Shi, Y.; Harley, J.B.; Shen, N.; Qian, Y. The microRNA mir-23b suppress IL-17-associated autoimmune inflammation by Targeting TAB2, TAB3 and IKK-α. Nat. Med. 2012, 18. [Google Scholar] [CrossRef]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; Downing, J.R.; Jacks, T.; Horvitz, H.R.; Golub, T.R. MicroRNA expression profiles classify human cancers. Nature 2005, 435. [Google Scholar] [CrossRef]
- Tavazoie, S.F.; Alarcon, C.; Oskarsson, T.; Padua, D.; Wang, Q.; Bos, P.D.; Gerald, W.L.; Massague, J. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 2008, 451, 147–152. [Google Scholar]
- Dai, R.; Zhang, Y.; Khan, D.; Heid, B.; Caudell, D.; Crasta, O.; Ahmed, S.A. Identification of a common lupus disease-associated microRNA expression pattern in three different murine models of Lupus. PLoS One 2010, 5, e14302. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Kwon, Y.-J.; Heo, J.Y.; Kim, H.C.; Kim, J.Y.; Liuzzi, M.; Soloveva, V. Phenotypic MicroRNA Microarrays. Microarrays 2013, 2, 63-80. https://doi.org/10.3390/microarrays2020063
Kwon Y-J, Heo JY, Kim HC, Kim JY, Liuzzi M, Soloveva V. Phenotypic MicroRNA Microarrays. Microarrays. 2013; 2(2):63-80. https://doi.org/10.3390/microarrays2020063
Chicago/Turabian StyleKwon, Yong-Jun, Jin Yeong Heo, Hi Chul Kim, Jin Yeop Kim, Michel Liuzzi, and Veronica Soloveva. 2013. "Phenotypic MicroRNA Microarrays" Microarrays 2, no. 2: 63-80. https://doi.org/10.3390/microarrays2020063
APA StyleKwon, Y.-J., Heo, J. Y., Kim, H. C., Kim, J. Y., Liuzzi, M., & Soloveva, V. (2013). Phenotypic MicroRNA Microarrays. Microarrays, 2(2), 63-80. https://doi.org/10.3390/microarrays2020063