Microarray for Identification of the Chiropteran Host Species of Rabies Virus in Canada
Abstract
:1. Introduction
2. Experimental Section
2.1. Bat Specimens
| Species | Abbreviation | Scientific Name | Numbers Tested |
|---|---|---|---|
| Little brown bat | LBB | Myotis lucifugus | 24 |
| Northern long-eared bat | NLB | Myotis septentrionalis | 13 |
| California bat | CLB | Myotis californicus | 7 |
| Long-legged bat | LLB | Myotis volans | 5 |
| Western long-eared bat | LEB | Myotis evotis | 12 |
| Yuma bat | YUB | Myotis yumanensis | 8 |
| Keen’s bat | KEB | Myotis keenii | 14 |
| Big brown bat | BBB | Eptesicus fuscus | 25 |
| Townsend’s big-eared bat | WEB | Corynorhinus townsendii | 9 |
| Hoary bat | HRB | Lasiurus cinereus | 10 |
| Eastern red bat | REB | Lasiurus borealis | 9 |
| Silver-haired bat | SHB | Lasionycteris noctivagans | 12 |
| Tricolored bat * | EPB | Perimyotis subflavus | 3 |
| Pallid bat | PAB | Antrozous pallidus | 2 |
| Total | 153 |
2.2. Primer and Probe Design
2.3. DNA Extraction
2.4. Slot Blot Analysis
2.5. PCR Amplification for Microarray Analysis
2.6. Microarray
3. Results and Discussion
3.1. Development of a Universal PCR Primer Pair
3.2. PCR for Partial Amplification of the Cytochrome c Oxidase Subunit I (COI) Gene of Canadian Bats
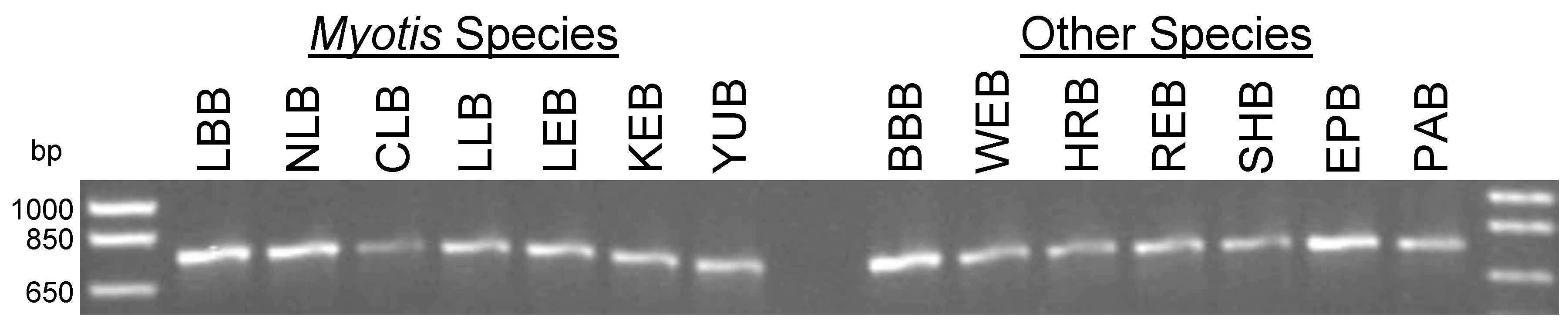
| Name | Specific Target | Sequence (5′-3′) | Location 1 |
|---|---|---|---|
| BB1 2 | BBB | CTGCCCTGAGTCTGCT | 130–145 |
| BB2 2 | BBB | GTGGACCTGACCATT | 465–479 |
| BB-E 2 | BBB (eastern) | CTTTTATTCGGCGCTTGA | 96–113 |
| BB-W 2 | BBB (western) | TTCTGTTCGGCGCCTGA | 97–113 |
| EP1 | EPB | CGCACACGCCTTTG | 215–228 |
| EP2 2 | EPB | CTTTCTTTTACTCCTAGCAT | 362–381 |
| LAS | HRB/REB | TATTATTGGCATCWTC | 370–385 |
| HR1 2 | HRB | AGTACCACTGATGATTGG | 284–301 |
| HR2 2 | HRB | CCACCTGCCTTGTCC | 555–569 |
| RE1 2 | REB | TCTATAACGTTATCGTGAC | 196–214 |
| RE2 2 | REB | GGTACCCCTTATRATCG | 284–300 |
| SH1 | SHB | GGGCTCTACTTGGAGAT | 172–188 |
| SH2 2 | SHB | CAACTGGTTAGTTCCTCTG | 275–293 |
| WE1 2 | WEB | TGATTAATCCCACTAATGAT | 279–298 |
| WE2 2 | WEB | CTTACATCTGGCTGG | 485–497 |
| CL1 2 | CLB | CTGGGAGACGATCAAATT | 180–197 |
| CL2 2 | CLB | TTTTATAGTTATGCCAATCAT | 239–259 |
| LB1 2 | LBB | CGCTGAGCTAGGTCAA | 152–167 |
| LB2 2 | LBB | TTTCTATTACTGCTGGC | 363–379 |
| LE1 | LEB | TGACATAGCCTTTCC | 308–322 |
| LE2 2 | LEB | CTGTCTTACTCCTTCTC | 613–629 |
| LL1 | LLB | TGTTGGGGGACGATCAGA | 178–195 |
| LL2 2 | LLB | TCTTCTCTCTGCACTTA | 478–494 |
| NL1 | NLB | AACTGGGCCAGCCA | 157–170 |
| NL2 2 | NLB | ATTCGTTTGGTCCGT | 587–601 |
| NL3 2 | NLB | ATTCGTTTGGTCTGT | 587–601 |
| KE1 2 | KEB | TCATAGTTATGCCCATTA | 241–258 |
| KE2 | KEB | TATGCCCATTATAATTGG | 248–265 |
| YU1 | YUB | CCCTTTTAGGGGATG | 175–189 |
| YU2 2 | YUB | TATAGTAATGCCGATTATAATC | 242–263 |
| PA 2 | PAB | TAATGTAATTGTCACAGCA | 200–218 |
3.3. Evaluation of Probes by Slot Blot Analysis
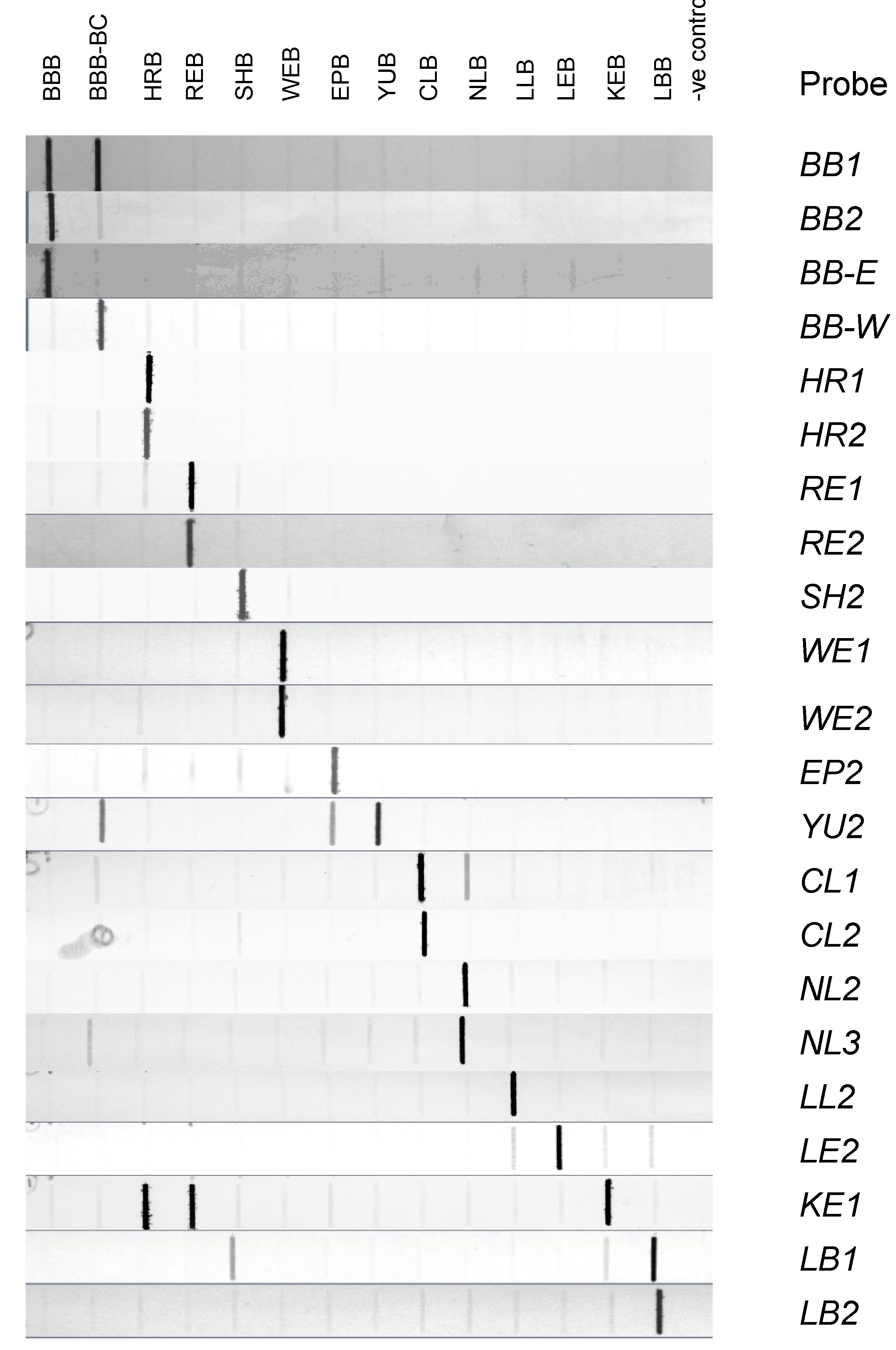
3.4. Evaluation and Validation of Probes by Microarray
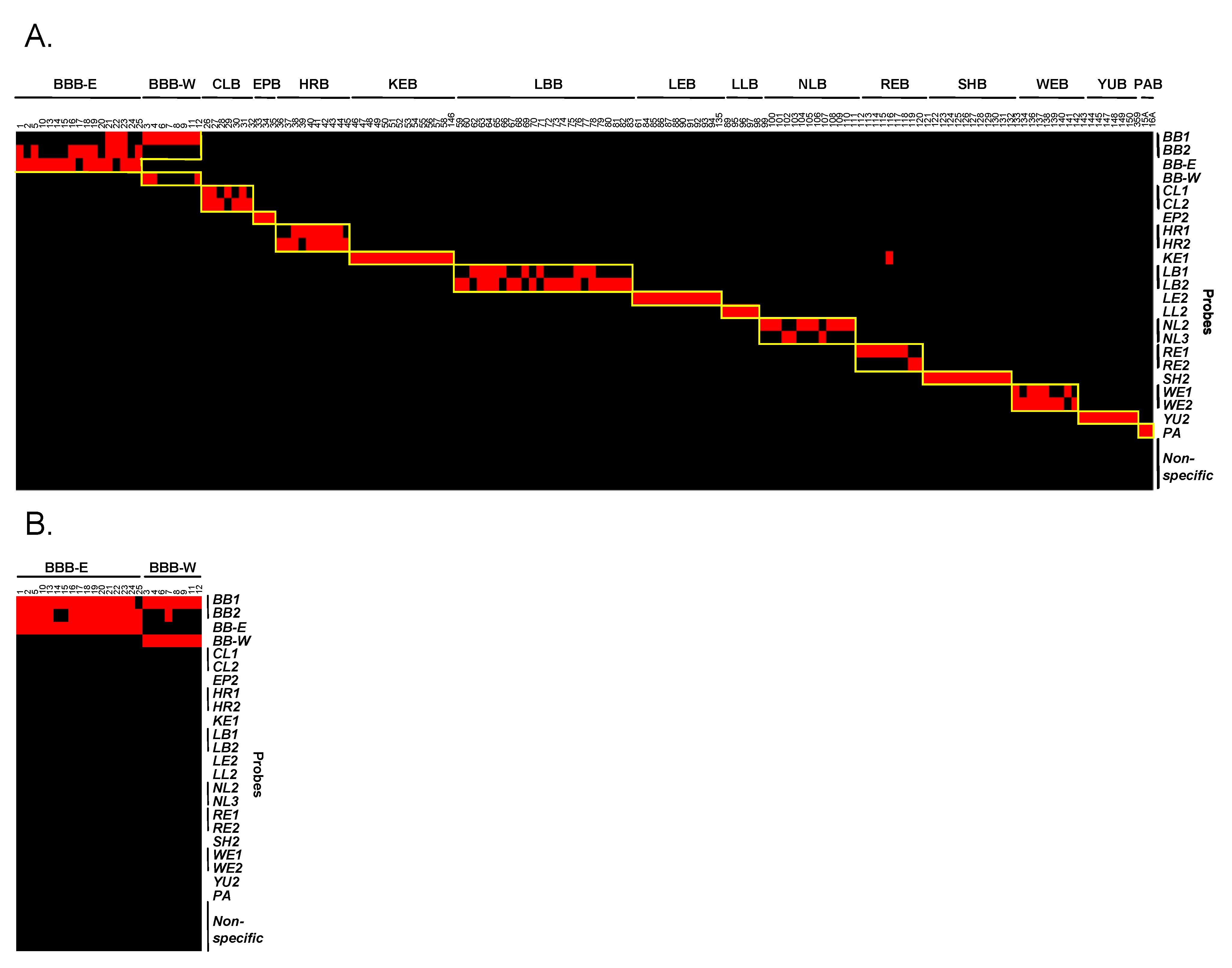
| Sample/Probe | Sequence (5′-3′) |
|---|---|
| REB #116 | TTA TAG TTA TGC CCA TTA 1 |
| REB #115 | TTA TAG TCA TGC CCA TTA 1 |
| KE1 Probe | TCA TAG TTA TGC CCA TTA 1,2 |
| KEB #51 | TCA TAG TTA TGC CCA TTA |
4. Conclusions
Acknowledgments
Conflict of Interest
References
- Arnason, U.; Adegoke, J.A.; Bodin, K.; Born, E.W.; Esa, Y.B.; Gullberg, A.; Nilsson, M.; Short, R.V.; Xu, X.; Janke, A. Mammalian mitogenomic relationships and the root of the eutherian tree. Proc. Natl. Acad. Sci. USA 2002, 99, 8151–8156. [Google Scholar] [CrossRef]
- Frézal, L.; Leblois, R. Four years of DNA barcoding: Current advances and prospects. Infect. Genet. Evol. 2008, 8, 727–736. [Google Scholar] [CrossRef]
- Hajibabaei, M.; Singer, G.A.C.; Hebert, P.D.N.; Hickey, D.A. DNA barcoding: How it complements taxonomy, molecular phylogenetics and population genetics. Trends Genet. 2007, 23, 167–172. [Google Scholar] [CrossRef]
- Tobe, S.S.; Kitchener, A.C.; Linacre, A.M.T. Reconstructing mammalian phylogenies: A detailed comparison of the cytochrome b and cytochrome oxidase subunit I mitochondrial genes. PLoS ONE 2010, 5, e14156. [Google Scholar] [CrossRef] [Green Version]
- Hebert, P.D.N.; Cywinska, A.; Ball, S.L.; de Waard, J.R. Biological identification through DNA barcodes. Proc. R. Soc. Lond. B 2003, 270, 313–321. [Google Scholar] [CrossRef]
- Hebert, P.D.N.; Ratnasingham, S.; de Waard, J.R. Barcoding animal life: Cytochrome c oxidase subunit 1 divergences among closely related species. Proc. R. Soc. Lond. B 2003, 270, S96–S99. [Google Scholar] [CrossRef]
- Ratnasingham, S.; Hebert, P.D.N. BOLD: The barcode of life data system. Mol. Ecol. Notes 2007, 7, 355–364. [Google Scholar]
- Clare, E.I.; Lim, B.K.; Engstrom, M.D.; Eger, J.I.; Hebert, P.D.N. DNA barcoding of Neotropical bats: Species identification and discovery within Guyana. Mol. Ecol. Notes 2007, 7, 184–190. [Google Scholar] [CrossRef]
- Mayer, F.; Dietz, C.; Kiefer, A. Molecular species identification boosts bat diversity. Front. Zool. 2007, 4, 4. [Google Scholar] [CrossRef]
- Jones, K.E.; Bininda-Emonds, O.; Gittleman, J. Bats, clocks, and rocks: Diversification patterns in chiroptera. Evolution 2005, 59, 2243–2255. [Google Scholar]
- Teeling, E.C.; Springer, M.S.; Madsen, O.; Bates, P.; O’Brien, S.J.; Murphy, W.J. A molecular phylogeny for bats illuminates biogeography and the fossil record. Science 2005, 307, 580–584. [Google Scholar] [CrossRef]
- Tudge, C. The Variety of Life: A Survey and a Celebration of All the Creatures that Have Ever Lived; Oxford University Press: Oxford, UK, 2000. [Google Scholar]
- Calisher, C.H.; Childs, J.E.; Field, H.E.; Holmes, K.V.; Schountz, T. Bats: Important reservoir hosts of emerging viruses. Clin. Microbiol. Rev. 2006, 19, 531–545. [Google Scholar] [CrossRef]
- Wibbelt, G.; Moore, M.S.; Schountz, T.; Voigt, C.C. Emerging diseases in Chiroptera. Why bats? Biol. Lett. 2010, 6, 438–440. [Google Scholar] [CrossRef]
- Banyard, A.C.; Hayman, D.; Johnson, N.; McElhinney, L.; Fooks, A.R. Bats and lyssaviruses. Adv. Virus Res. 2011, 79, 239–289. [Google Scholar] [CrossRef]
- Wong, S.; Lau, S.; Woo, P.; Yuen, K.Y. Bats as a continuing source of emerging infections in humans. Rev. Med. Virol. 2007, 17, 67–91. [Google Scholar] [CrossRef]
- Kuzmin, I.V.; Rupprecht, C.E. Bat rabies. In Rabies, 2nd ed.; Jackson, A.C., Wunner, W.H., Eds.; Academic Press: San Diego, CA, USA, 2007; pp. 259–307. [Google Scholar]
- Hanlon, C.A.; Niezgoda, M.; Rupprecht, C.E. Rabies in terrestrial animals. In Rabies, 2nd ed.; Jackson, A.C., Wunner, W.H., Eds.; Academic Press: San Diego, CA, USA, 2007; pp. 201–258. [Google Scholar]
- Nadin-Davis, S.A.; Fehlner-Gardiner, C. Lyssaviruses: Current trends. Adv. Virus Res. 2008, 71, 207–250. [Google Scholar] [CrossRef]
- Nadin-Davis, S.A.; Huang, W.; Armstrong, J.; Casey, G.A.; Bahloul, C.; Tordo, N.; Wandeler, A.I. Antigenic and genetic divergence of rabies viruses from bat species indigenous to Canada. Virus Res. 2001, 74, 139–156. [Google Scholar] [CrossRef]
- De Serres, G.; Dallaire, F.; Côte, M.; Skowronski, D.M. Bat rabies in the United States and Canada from 1950 through 2007: Human cases with and without bat contact. Clin. Infect. Dis. 2008, 46, 1329–1337. [Google Scholar] [CrossRef]
- Jackson, A.C.; Fenton, M.B. Human rabies and bat bites. Lancet 2001, 357. [Google Scholar] [CrossRef]
- WHO Fact Sheet. “Rabies”. Available online: http://www.who.int/mediacentre/factsheets/fs099/en/ (accessed on 21 May 2013).
- Kuzmin, I.V.; Shi, M.; Orciari, L.A.; Yager, P.A.; Velasco-Villa, A.; Kuzmina, N.A.; Streiker, D.G.; Bergman, D.L.; Rupprecht, C.E. Molecular inferences suggest multiple host shifts of rabies viruses from bats to mesocarnivores in Arizona during 2001–2009. PLoS Pathog. 2012, 8, e1002786. [Google Scholar] [CrossRef]
- Nagorsen, D.W.; Brigham, R.M. Bats of British Columbia. Royal British Columbia Museum Handbook Series; UBC Press: Vancouver, BC, Canada, 1993. [Google Scholar]
- Adams, R.A. Bats of the Rocky Mountain West: Natural History, Ecology, and Conservation. Boulder; University Press of Colorado: Vancouver, BC, Canada, 2003. [Google Scholar]
- National Audubon Society, National Audubon Society Field Guide to North American Mammals; Knopf Doubleday Publishing Group: New York, NY, USA, 1996.
- Environment Canada and Canadian Wildlife Federation. Bats: Hinterland Who’s Who. Available online: http://www.hww.ca/en/species/mammals/bats.html (accessed on 28 February 2013).
- Nadin-Davis, S.A.; Guerrero, E.; Knowles, M.K.; Feng, Y. DNA barcoding facilitates bat species identification for improved surveillance of bat-associated rabies across Canada. Open J. Zool. 2012, 5, 27–37. [Google Scholar] [CrossRef]
- Carnieli, P., Jr.; de Oliveira, F.W.; Castilho, J.G.; Brandão, P.E.; Carrieri, M.L.; Kotait, I. Species determination of Brazilian mammals implicated in the epidemiology of rabies based on the control region of mitochondrial DNA. Braz. J. Infect. Dis. 2008, 12, 462–465. [Google Scholar] [CrossRef] [Green Version]
- Harris, S.L.; Johnson, N.; Brookes, S.M.; Hutson, A.M.; Fooks, A.R.; Jones, G. The application of genetic markers for EBLV surveillance in European bat species. Dev. Biol. 2008, 131, 347–363. [Google Scholar]
- Schena, M.; Heller, R.A.; Theriault, T.P.; Konrad, K.; Lachenmeier, E.; Davis, R.W. Microarrays: Biotechnology’s discovery platform for functional genomics. Trends Biotech. 1998, 16, 301–306. [Google Scholar] [CrossRef]
- Garaizar, J.; Rementeria, A.; Porwollik, S. DNA microarray technology: A new tool for the epidemiological typing of bacterial pathogens? FEMS Immunol. Med. Microbiol. 2006, 47, 178–189. [Google Scholar] [CrossRef]
- Pfunder, M.; Holzgang, O.; Frey, J.E. Development of microarray-based diagnostics of voles and shrews for use in biodiversity monitoring studies, and evaluation of mitochondrial cytochrome oxidase I vs. cytochrome b as genetic markers. Mol. Ecol. 2004, 13, 1277–1286. [Google Scholar] [CrossRef]
- Hajibabaei, M.; Singer, G.A.; Clare, E.L.; Hebert, P.D. Design and applicability of DNA arrays and DNA barcodes in biodiversity monitoring. BMC Biol. 2007, 13. [Google Scholar] [CrossRef] [Green Version]
- Deblauwe, I.; de Witte, J.C.; de Deken, G.; de Deken, R.; Madder, M.; van Erk, S.; Hoza, F.A.; Lathouwers, D.; Geysen, D. A new tool for the molecular identification of Culicoides species of the Obsoletus group: The glass slide microarray approach. Med. Vet. Entomol. 2012, 26, 83–91. [Google Scholar]
- Kochzius, M.; Seidel, C.; Antoniou, A.; Botla, S.K.; Campo, D.; Cariani, A.; Vazquez, E.G.; Hauschild, J.; Hervet, C.; Hjörleifsdottir, S.; et al. Identifying fishes through DNA barcodes and microarrays. PLoS ONE 2010, 5, e12620. [Google Scholar] [CrossRef]
- Chung, I.; Yoo, H.S.; Eah, J.; Yoon, H.; Jung, J.; Hwang, S.Y.; Kim, C. A DNA microarray for identification of selected Korean birds based on mitochondrial Cytochrome c oxidase 1 gene sequences. Mol. Cells 2010, 30, 295–301. [Google Scholar]
- Summerbell, R.C.; Lévesque, C.A.; Seifert, K.A.; Bovers, M.; Fell, J.W.; Diaz, M.R.; Boekhout, T.; de Hoog, G.S.; Stalpers, J.; Crous, P.W. Microcoding: The second step in DNA barcoding. Phil. Trans. Royal Soc. Lond. 2005, 360, 1897–1903. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The ClustalX windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef]
- OligoAnalyzer®. Available online: https://www.idtdna.com/analyzer/Applications/OligoAnalyzer/ (accessed on 28 February 2013).
- Desloire, S.; Moro, C.V.; Chauve, C.; Zenner, L. Comparison of four methods of extracting DNA from D. gallinae (Acari: Dermanyssidae). Vet. Res. 2006, 37, 725–732. [Google Scholar] [CrossRef]
- Lung, O.; Fisher, M.; Beeston, A.; Hughes, K.B.; Clavijo, A.; Goolia, M.; Pasick, J.; Mauro, W.; Deregt, D. Multiplex RT-PCR detection and microarray typing of vesicular disease viruses. J. Virol. Methods 2011, 175, 236–245. [Google Scholar] [CrossRef]
- Meusnier, I.; Singer, G.A.; Landry, J.F.; Hickey, D.A.; Hebert, P.D.; Hajibabaei, M. A universal DNA mini-barcode for biodiversity analysis. BMC Genomics 2008, 12. [Google Scholar] [CrossRef] [Green Version]
- Zinck, J.M.; Duffield, D.A.; Ormsbee, P.C. Primers for identification and polymorphism assessment of Vespertilionid bats in the Pacific Northwest. Mol. Ecol. Notes 2004, 4, 239–242. [Google Scholar] [CrossRef]
- Nadin-Davis, S.A.; Feng, Y.; Mousse, D.; Wandeler, A.I.; Aris-Brosou, S. Spatial and temporal dynamics of rabies virus variants in big brown bat populations across Canada: Footprints of an emerging zoonosis. Mol. Ecol. 2010, 19, 2120–2136. [Google Scholar] [CrossRef]
- Streicker, D.G.; Turmelle, A.S.; Vonhof, M.J.; Kuzmin, I.V.; McCracken, G.F.; Rupprecht, C.E. Host phylogeny constrains cross-species emergence and establishment of rabies virus in bats. Science 2010, 329, 676–679. [Google Scholar] [CrossRef]
- Constantine, D.G. An updated list of rabies-infected bats in North America. J. Wildl. Dis. 1979, 15, 347–349. [Google Scholar]
- Sodré, M.M.; da Gama, A.R.; de Almeida, M.F. Updated list of bat species positive for rabies in Brazil. Rev. Inst. Med. Trop. São Paulo 2010, 52, 75–81. [Google Scholar]
- Spizz, G.; Young, L.; Yasmin, R.; Chen, Z.; Lee, T.; Mahoney, D.; Zhang, X.; Mouchka, G.; Thomas, B.; Honey, W.; et al. Rheonix CARD® technology: An innovative and fully automated molecular diagnostic device. Point Care 2012, 11, 42–51. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lung, O.; Nadin-Davis, S.; Fisher, M.; Erickson, A.; Knowles, M.K.; Furukawa-Stoffer, T.; Ambagala, A. Microarray for Identification of the Chiropteran Host Species of Rabies Virus in Canada. Microarrays 2013, 2, 153-169. https://doi.org/10.3390/microarrays2020153
Lung O, Nadin-Davis S, Fisher M, Erickson A, Knowles MK, Furukawa-Stoffer T, Ambagala A. Microarray for Identification of the Chiropteran Host Species of Rabies Virus in Canada. Microarrays. 2013; 2(2):153-169. https://doi.org/10.3390/microarrays2020153
Chicago/Turabian StyleLung, Oliver, Susan Nadin-Davis, Mathew Fisher, Anthony Erickson, M. Kimberly Knowles, Tara Furukawa-Stoffer, and Aruna Ambagala. 2013. "Microarray for Identification of the Chiropteran Host Species of Rabies Virus in Canada" Microarrays 2, no. 2: 153-169. https://doi.org/10.3390/microarrays2020153







