PICALM rs3851179 Variants Modulate Left Postcentral Cortex Thickness, CSF Amyloid β42, and Phosphorylated Tau in the Elderly
Abstract
:1. Introduction
2. Materials and Methods
2.1. Participants
2.2. MRI Data Acquisition and Processing
2.3. Statistical Analyses
2.3.1. MRI Data Analysis
2.3.2. Clinical Data and Demographic Characteristics
2.3.3. Correlation Analysis
3. Results
3.1. MRI Data Findings
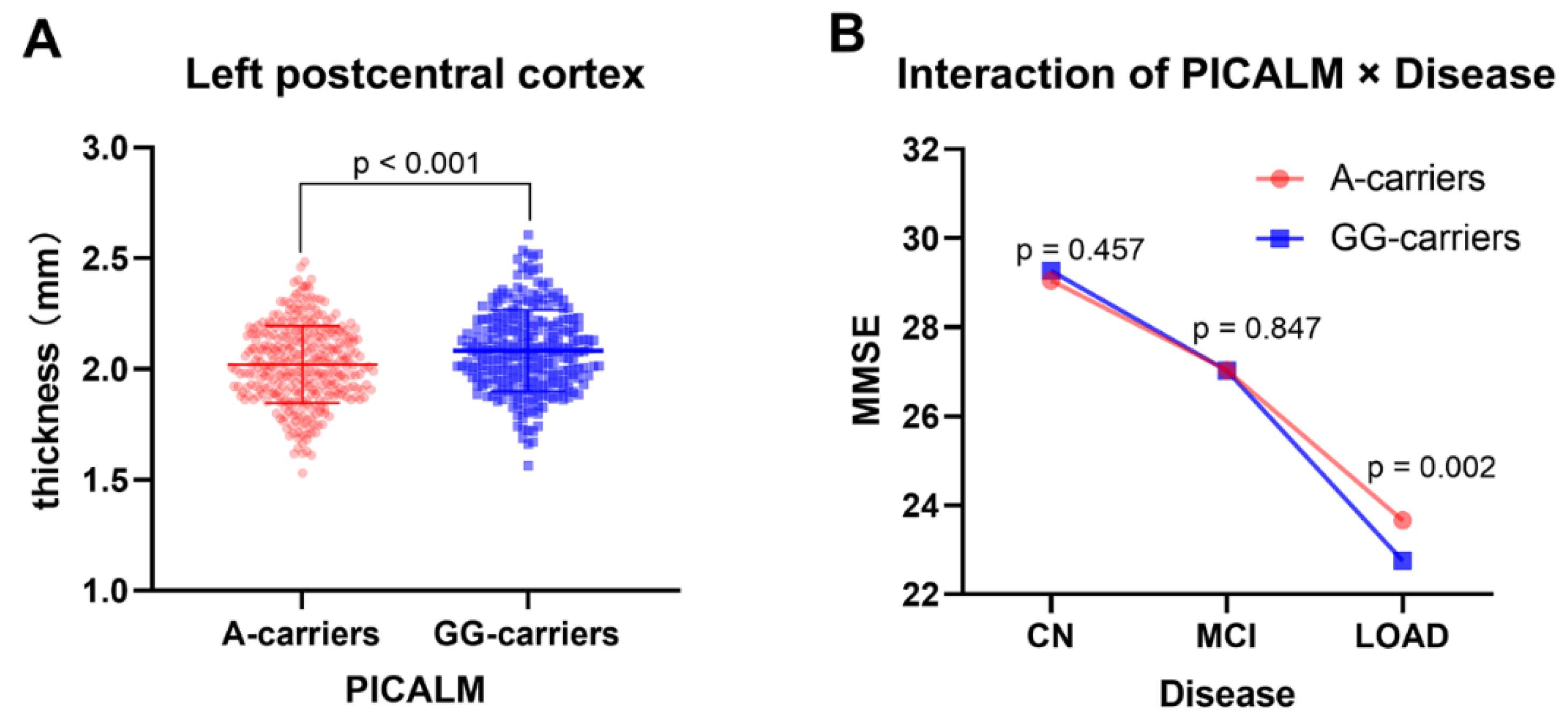
3.1.1. The Interactive Effect of PICALM and Disease on Cortex
3.1.2. The Main Effects of PICALM on Cortex
3.1.3. The Main Effect of Disease on Cortex
3.1.4. The Interactive Effect of PICALM × Disease × AOPE on Cortex
3.2. Demographic and Clinical Data Findings
3.2.1. The Results of Demographic Data
3.2.2. The Interactive Effect of PICALM × Disease on Clinical Data
3.2.3. The Main Effect of Disease on Clinical Data
3.2.4. The Main Effect of PICALM on Clinical Data
3.3. The Results of Correlation Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Mayeda, E.R.; Glymour, M.M.; Quesenberry, C.P.; Johnson, J.K.; Pérez-Stable, E.J.; Whitmer, R.A. Survival after dementia diagnosis in five racial/ethnic groups. Alzheimer’s Dement. 2017, 13, 761–769. [Google Scholar] [CrossRef] [PubMed]
- Rhodius-Meester, H.F.M.; Tijms, B.M.; Lemstra, A.W.; Prins, N.D.; Pijnenburg, Y.A.L.; Bouwman, F.; Scheltens, P.; van der Flier, W.M. Survival in memory clinic cohort is short, even in young-onset dementia. J. Neurol. Neurosurg. Psychiatry 2019, 90, 726–728. [Google Scholar] [CrossRef] [PubMed]
- Dubois, B.; Hampel, H.; Feldman, H.H.; Scheltens, P.; Aisen, P.; Andrieu, S.; Bakardjian, H.; Benali, H.; Bertram, L.; Blennow, K.; et al. Preclinical Alzheimer’s disease: Definition, natural history, and diagnostic criteria. Alzheimer’s Dement. 2016, 12, 292–323. [Google Scholar] [CrossRef] [PubMed]
- Lazarov, O.; Hollands, C. Hippocampal neurogenesis: Learning to remember. Prog. Neurobiol. 2016, 138–140, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Scheltens, P.; De Strooper, B.; Kivipelto, M.; Holstege, H.; Chételat, G.; Teunissen, C.E.; Cummings, J.; van der Flier, W.M. Alzheimer’s disease. Lancet 2021, 397, 1577–1590. [Google Scholar] [CrossRef]
- Jack, C.R., Jr.; Bennett, D.A.; Blennow, K.; Carrillo, M.C.; Dunn, B.; Haeberlein, S.B.; Holtzman, D.M.; Jagust, W.; Jessen, F.; Karlawish, J.; et al. NIA-AA Research Framework: Toward a biological definition of Alzheimer’s disease. Alzheimer’s Dement. 2018, 14, 535–562. [Google Scholar] [CrossRef]
- van der Lee, S.J.; Wolters, F.J.; Ikram, M.K.; Hofman, A.; Ikram, M.A.; Amin, N.; van Duijn, C.M. The effect of APOE and other common genetic variants on the onset of Alzheimer’s disease and dementia: A community-based cohort study. Lancet Neurol. 2018, 17, 434–444. [Google Scholar] [CrossRef]
- Ashford, J.W.; Mortimer, J.A. Non-familial Alzheimer’s disease is mainly due to genetic factors. J. Alzheimer’s Dis. JAD 2002, 4, 169–177. [Google Scholar] [CrossRef] [Green Version]
- DeTure, M.A.; Dickson, D.W. The neuropathological diagnosis of Alzheimer’s disease. Mol. Neurodegener. 2019, 14, 32. [Google Scholar] [CrossRef] [Green Version]
- Jack, C.R., Jr.; Holtzman, D.M.; Sperling, R. Dementia is not synonymous with Alzheimer’s disease. Sci. Transl. Med. 2019, 11, eaav0511. [Google Scholar] [CrossRef]
- Sweeney, M.D.; Sagare, A.P.; Zlokovic, B.V. Blood-brain barrier breakdown in Alzheimer disease and other neurodegenerative disorders. Nat. Rev. Neurol. 2018, 14, 133–150. [Google Scholar] [CrossRef] [PubMed]
- Venegas, C.; Kumar, S.; Franklin, B.S.; Dierkes, T.; Brinkschulte, R.; Tejera, D.; Vieira-Saecker, A.; Schwartz, S.; Santarelli, F.; Kummer, M.P.; et al. Microglia-derived ASC specks cross-seed amyloid-β in Alzheimer’s disease. Nature 2017, 552, 355–361. [Google Scholar] [CrossRef] [PubMed]
- Frisoni, G.B.; Boccardi, M.; Barkhof, F.; Blennow, K.; Cappa, S.; Chiotis, K.; Démonet, J.F.; Garibotto, V.; Giannakopoulos, P.; Gietl, A.; et al. Strategic roadmap for an early diagnosis of Alzheimer’s disease based on biomarkers. Lancet Neurol. 2017, 16, 661–676. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matsuda, H. MRI morphometry in Alzheimer’s disease. Ageing Res. Rev. 2016, 30, 17–24. [Google Scholar] [CrossRef]
- Braak, H.; Braak, E. Staging of Alzheimer’s disease-related neurofibrillary changes. Neurobiol. Aging 1995, 16, 271–278. [Google Scholar] [CrossRef]
- Chen, G.; Shu, H.; Chen, G.; Ward, B.D.; Antuono, P.G.; Zhang, Z.; Li, S.J. Staging Alzheimer’s Disease Risk by Sequencing Brain Function and Structure, Cerebrospinal Fluid, and Cognition Biomarkers. J. Alzheimer’s Dis. JAD 2016, 54, 983–993. [Google Scholar] [CrossRef] [Green Version]
- Grothe, M.J.; Barthel, H.; Sepulcre, J.; Dyrba, M.; Sabri, O.; Teipel, S.J. In vivo staging of regional amyloid deposition. Neurology 2017, 89, 2031–2038. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alexander, G.E.; DeLong, M.R.; Strick, P.L. Parallel organization of functionally segregated circuits linking basal ganglia and cortex. Annu. Rev. Neurosci. 1986, 9, 357–381. [Google Scholar] [CrossRef]
- Draganski, B.; Kherif, F.; Klöppel, S.; Cook, P.A.; Alexander, D.C.; Parker, G.J.; Deichmann, R.; Ashburner, J.; Frackowiak, R.S. Evidence for segregated and integrative connectivity patterns in the human Basal Ganglia. J. Neurosci. 2008, 28, 7143–7152. [Google Scholar] [CrossRef] [Green Version]
- Papma, J.M.; Smits, M.; de Groot, M.; Mattace Raso, F.U.; van der Lugt, A.; Vrooman, H.A.; Niessen, W.J.; Koudstaal, P.J.; van Swieten, J.C.; van der Veen, F.M.; et al. The effect of hippocampal function, volume and connectivity on posterior cingulate cortex functioning during episodic memory fMRI in mild cognitive impairment. Eur. Radiol. 2017, 27, 3716–3724. [Google Scholar] [CrossRef]
- Quan, M.; Zhao, T.; Tang, Y.; Luo, P.; Wang, W.; Qin, Q.; Li, T.; Wang, Q.; Fang, J.; Jia, J. Effects of gene mutation and disease progression on representative neural circuits in familial Alzheimer’s disease. Alzheimers Res. Ther. 2020, 12, 14. [Google Scholar] [CrossRef]
- Jansen, I.E.; Savage, J.E.; Watanabe, K.; Bryois, J.; Williams, D.M.; Steinberg, S.; Sealock, J.; Karlsson, I.K.; Hagg, S.; Athanasiu, L.; et al. Genome-wide meta-analysis identifies new loci and functional pathways influencing Alzheimer’s disease risk. Nat. Genet. 2019, 51, 404–413. [Google Scholar] [CrossRef] [PubMed]
- Harold, D.; Abraham, R.; Hollingworth, P.; Sims, R.; Gerrish, A.; Hamshere, M.L.; Pahwa, J.S.; Moskvina, V.; Dowzell, K.; Williams, A.; et al. Genome-wide association study identifies variants at CLU and PICALM associated with Alzheimer’s disease. Nat. Genet. 2009, 41, 1088–1093. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lambert, J.C.; Ibrahim-Verbaas, C.A.; Harold, D.; Naj, A.C.; Sims, R.; Bellenguez, C.; DeStafano, A.L.; Bis, J.C.; Beecham, G.W.; Grenier-Boley, B.; et al. Meta-analysis of 74,046 individuals identifies 11 new susceptibility loci for Alzheimer’s disease. Nat. Genet. 2013, 45, 1452–1458. [Google Scholar] [CrossRef] [Green Version]
- Baig, S.; Joseph, S.A.; Tayler, H.; Abraham, R.; Owen, M.J.; Williams, J.; Kehoe, P.G.; Love, S. Distribution and expression of picalm in Alzheimer disease. J. Neuropathol. Exp. Neurol. 2010, 69, 1071–1077. [Google Scholar] [CrossRef] [Green Version]
- Parikh, I.; Fardo, D.W.; Estus, S. Genetics of PICALM expression and Alzheimer’s disease. PLoS ONE 2014, 9, e91242. [Google Scholar] [CrossRef]
- Ando, K.; De Decker, R.; Vergara, C.; Yilmaz, Z.; Mansour, S.; Suain, V.; Sleegers, K.; de Fisenne, M.A.; Houben, S.; Potier, M.C.; et al. Picalm reduction exacerbates tau pathology in a murine tauopathy model. Acta Neuropathol. 2020, 139, 773–789. [Google Scholar] [CrossRef] [Green Version]
- Moreau, K.; Fleming, A.; Imarisio, S.; Lopez Ramirez, A.; Mercer, J.L.; Jimenez-Sanchez, M.; Bento, C.F.; Puri, C.; Zavodszky, E.; Siddiqi, F.; et al. PICALM modulates autophagy activity and tau accumulation. Nat. Commun. 2014, 5, 4998. [Google Scholar] [CrossRef] [Green Version]
- Kanatsu, K.; Morohashi, Y.; Suzuki, M.; Kuroda, H.; Watanabe, T.; Tomita, T.; Iwatsubo, T. Decreased CALM expression reduces Aβ42 to total Aβ ratio through clathrin-mediated endocytosis of γ-secretase. Nat. Commun. 2014, 5, 3386. [Google Scholar] [CrossRef] [Green Version]
- Zhao, Z.; Sagare, A.P.; Ma, Q.; Halliday, M.R.; Kong, P.; Kisler, K.; Winkler, E.A.; Ramanathan, A.; Kanekiyo, T.; Bu, G.; et al. Central role for PICALM in amyloid-beta blood-brain barrier transcytosis and clearance. Nat. Neurosci. 2015, 18, 978–987. [Google Scholar] [CrossRef] [PubMed]
- Zeng, F.F.; Liu, J.; He, H.; Gao, X.P.; Liao, M.Q.; Yu, X.X.; Liu, Y.H.; Zhu, S.; Jing, C.X. Association of PICALM Gene Polymorphisms with Alzheimer’s Disease: Evidence from an Updated Meta-Analysis. Curr. Alzheimer Res. 2019, 16, 1196–1205. [Google Scholar] [CrossRef]
- Yang, X.; Li, J.; Liu, B.; Li, Y.; Jiang, T. Impact of PICALM and CLU on hippocampal degeneration. Hum. Brain Mapp. 2016, 37, 2419–2430. [Google Scholar] [CrossRef]
- Morgen, K.; Ramirez, A.; Frolich, L.; Tost, H.; Plichta, M.M.; Kolsch, H.; Rakebrandt, F.; Rienhoff, O.; Jessen, F.; Peters, O.; et al. Genetic interaction of PICALM and APOE is associated with brain atrophy and cognitive impairment in Alzheimer’s disease. Alzheimer’s Dement. 2014, 10, S269–S276. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; Qin, W.; Wang, D.; Liu, B.; Zhang, Y.; Jiang, T.; Yu, C. Impacts of PICALM and CLU variants associated with Alzheimer’s disease on the functional connectivity of the hippocampus in healthy young adults. Brain Struct. Funct. 2015, 220, 1463–1475. [Google Scholar] [CrossRef]
- Petersen, R.C.; Aisen, P.S.; Beckett, L.A.; Donohue, M.C.; Gamst, A.C.; Harvey, D.J.; Jack, C.R., Jr.; Jagust, W.J.; Shaw, L.M.; Toga, A.W.; et al. Alzheimer’s Disease Neuroimaging Initiative (ADNI): Clinical characterization. Neurology 2010, 74, 201–209. [Google Scholar] [CrossRef] [Green Version]
- Ashburner, J. A fast diffeomorphic image registration algorithm. NeuroImage 2007, 38, 95–113. [Google Scholar] [CrossRef] [PubMed]
- Glasser, M.F.; Smith, S.M.; Marcus, D.S.; Andersson, J.L.; Auerbach, E.J.; Behrens, T.E.; Coalson, T.S.; Harms, M.P.; Jenkinson, M.; Moeller, S.; et al. The Human Connectome Project’s neuroimaging approach. Nat. Neurosci. 2016, 19, 1175–1187. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huang, C.C.; Rolls, E.T.; Feng, J.; Lin, C.P. An extended Human Connectome Project multimodal parcellation atlas of the human cortex and subcortical areas. Brain Struct. Funct. 2022, 227, 763–778. [Google Scholar] [CrossRef] [PubMed]
- Penfield, W.; Boldrey, E. Somatic motor and sensory representation in the cerebral cortex of man as studied by electrical stimulation. Brain 1937, 60, 389–443. [Google Scholar] [CrossRef]
- Kaas, J.H.; Nelson, R.J.; Sur, M.; Lin, C.S.; Merzenich, M.M. Multiple representations of the body within the primary somatosensory cortex of primates. Science 1979, 204, 521–523. [Google Scholar] [CrossRef]
- Zhang, Z.; Gadotti, V.M.; Chen, L.; Souza, I.A.; Stemkowski, P.L.; Zamponi, G.W. Role of Prelimbic GABAergic Circuits in Sensory and Emotional Aspects of Neuropathic Pain. Cell Rep. 2015, 12, 752–759. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kropf, E.; Syan, S.K.; Minuzzi, L.; Frey, B.N. From anatomy to function: The role of the somatosensory cortex in emotional regulation. Rev. Bras. De Psiquiatr. 2019, 41, 261–269. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Phillips, M.L.; Drevets, W.C.; Rauch, S.L.; Lane, R. Neurobiology of emotion perception II: Implications for major psychiatric disorders. Biol. Psychiatry 2003, 54, 515–528. [Google Scholar] [CrossRef]
- Damasio, A.R.; Grabowski, T.J.; Bechara, A.; Damasio, H.; Ponto, L.L.; Parvizi, J.; Hichwa, R.D. Subcortical and cortical brain activity during the feeling of self-generated emotions. Nat. Neurosci. 2000, 3, 1049–1056. [Google Scholar] [CrossRef] [PubMed]
- Adolphs, R.; Damasio, H.; Tranel, D.; Cooper, G.; Damasio, A.R. A role for somatosensory cortices in the visual recognition of emotion as revealed by three-dimensional lesion mapping. J. Neurosci. 2000, 20, 2683–2690. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kuhlmann, J.; Andreasson, U.; Pannee, J.; Bjerke, M.; Portelius, E.; Leinenbach, A.; Bittner, T.; Korecka, M.; Jenkins, R.G.; Vanderstichele, H.; et al. CSF Aβ(1-42)—An excellent but complicated Alzheimer’s biomarker—A route to standardisation. Clin. Chim. Acta 2017, 467, 27–33. [Google Scholar] [CrossRef]
- Blennow, K.; Hampel, H.; Weiner, M.; Zetterberg, H. Cerebrospinal fluid and plasma biomarkers in Alzheimer disease. Nat. Rev. Neurol. 2010, 6, 131–144. [Google Scholar] [CrossRef]
- Hansson, O.; Zetterberg, H.; Buchhave, P.; Londos, E.; Blennow, K.; Minthon, L. Association between CSF biomarkers and incipient Alzheimer’s disease in patients with mild cognitive impairment: A follow-up study. Lancet. Neurol. 2006, 5, 228–234. [Google Scholar] [CrossRef] [Green Version]
- Hampel, H.; Frank, R.; Broich, K.; Teipel, S.J.; Katz, R.G.; Hardy, J.; Herholz, K.; Bokde, A.L.; Jessen, F.; Hoessler, Y.C.; et al. Biomarkers for Alzheimer’s disease: Academic, industry and regulatory perspectives. Nat. Rev. Drug Discov. 2010, 9, 560–574. [Google Scholar] [CrossRef]
- Olsson, B.; Lautner, R.; Andreasson, U.; Öhrfelt, A.; Portelius, E.; Bjerke, M.; Hölttä, M.; Rosén, C.; Olsson, C.; Strobel, G.; et al. CSF and blood biomarkers for the diagnosis of Alzheimer’s disease: A systematic review and meta-analysis. Lancet. Neurol. 2016, 15, 673–684. [Google Scholar] [CrossRef]
- Strozyk, D.; Blennow, K.; White, L.R.; Launer, L.J. CSF Abeta 42 levels correlate with amyloid-neuropathology in a population-based autopsy study. Neurology 2003, 60, 652–656. [Google Scholar] [CrossRef] [PubMed]
- Horie, K.; Barthélemy, N.R.; Sato, C.; Bateman, R.J. CSF tau microtubule binding region identifies tau tangle and clinical stages of Alzheimer’s disease. Brain 2021, 144, 515–527. [Google Scholar] [CrossRef] [PubMed]
- Harel, A.; Wu, F.; Mattson, M.P.; Morris, C.M.; Yao, P.J. Evidence for CALM in directing VAMP2 trafficking. Traffic 2008, 9, 417–429. [Google Scholar] [CrossRef] [PubMed]
- Licastro, F.; Carbone, I.; Ianni, M.; Porcellini, E. Gene signature in Alzheimer’s disease and environmental factors: The virus chronicle. J. Alzheimer’s Dis. JAD 2011, 27, 809–817. [Google Scholar] [CrossRef] [PubMed]
- Scotland, P.B.; Heath, J.L.; Conway, A.E.; Porter, N.B.; Armstrong, M.B.; Walker, J.A.; Klebig, M.L.; Lavau, C.P.; Wechsler, D.S. The PICALM protein plays a key role in iron homeostasis and cell proliferation. PLoS ONE 2012, 7, e44252. [Google Scholar] [CrossRef]

| Cluster Peak p-Values | Cluster Peak F-Score | Cluster Size | MNI Coordinates | Overlap of Atlas Region | |||
|---|---|---|---|---|---|---|---|
| x | y | z | DK40 Atlas | HCP MMP Atlas | |||
| 0.00001 | 21.0 | 1336 | −59 | −5 | 17 | postcentral L | area_1 L |
| precentral L | area_3a L | ||||||
| OP4 L | |||||||
| area_3b L | |||||||
| PFop L | |||||||
| area_2 L | |||||||
| CN | MCI | LOAD | |||||||
|---|---|---|---|---|---|---|---|---|---|
| A-Carriers (n = 117) | GG-Carriers (n = 71) | p Value (x2/T Value) | A-Carriers (n = 147) | GG-Carriers (n = 114) | p Value (x2/T Value) | A-Carriers (n = 71) | GG-Carriers (n = 69) | p Value (x2/T value) | |
| APOE ε4 (carriers/noncarriers) | 84/33 | 51/20 | 0.996 (<0.001) | 72/75 | 45/69 | 0.126 (2.346) | 24/47 | 18/51 | 0.319 (0.992) |
| Gender (males/females) | 62/55 | 34/37 | 0.497 (0.461) | 94/53 | 74/40 | 0.872 (0.026) | 38/33 | 35/34 | 0.741 (0.110) |
| Age (years) | 75.92 ± 4.57 | 75.89 ± 4.73 | 0.932 (−0.086) | 76.72 ± 5.25 | 75.56 ± 5.90 | 0.096 (1.672) | 77.47 ± 6.04 | 76.73 ± 5.62 | 0.457 (0.745) |
| Education (years) | 15.73 ± 2.73 | 16.15 ± 2.93 | 0.311 (−1.015) | 15.60 ± 2.93 | 15.81 ± 3.35 | 0.593 (−0.535) | 14.69 ± 3.05 | 14.22 ± 3.42 | 0.389 (0.864) |
| Characteristic | CN | MCI | LOAD | Statistics | p Value | CN vs. MCI p Value | CN vs. LOAD p Value | MCI vs. LOAD p Value |
|---|---|---|---|---|---|---|---|---|
| ADAS13 | 9.59 ± 4.39 | 18.74 ± 5.93 | 28.96 ± 6.82 | F = 357.789 | <0.001 | <0.001 | <0.001 | <0.001 |
| RAVLT immediate | 42.98 ± 9.3 | 30.66 ± 8.93 | 23.12 ± 7.47 | F = 169.504 | <0.001 | <0.001 | <0.001 | <0.001 |
| Amyloid β42 (pg/mL) | 1156.43 ± 555.11 | 797.68 ± 426.07 | 691.86 ± 343.97 | F = 9.797 | <0.001 | <0.001 | <0.001 | 0.629 |
| Amyloid β40 (pg/mL) | 7777.81 ± 2590.03 | 7746.04 ± 2059.81 | 7446.2 ± 2418.3 | F = 0.798 | 0.451 | |||
| Amyloid β42/40 | 0.151 ± 0.054 | 0.105 ± 0.051 | 0.094 ± 0.034 | F = 14.621 | <0.001 | <0.001 | <0.001 | 0.814 |
| Amyloid β38 (pg/mL) | 1843.13 ± 630.32 | 1811.48 ± 525.88 | 1721.26 ± 595.69 | F = 0.907 | 0.405 | |||
| T-tau (pg/mL) | 236.87 ± 86.63 | 313.74 ± 122.17 | 352.35 ± 122.43 | F = 12.410 | <0.001 | <0.001 | <0.001 | 0.207 |
| P-tau (pg/mL) | 22.06 ± 9.17 | 31.09 ± 14.12 | 35.21 ± 13.89 | F = 12.328 | <0.001 | <0.001 | <0.001 | 0.248 |
| P/T-tau | 0.092 ± 0.007 | 0.097 ± 0.009 | 0.099 ± 0.009 | F = 4.141 | 0.017 | 0.010 | 0.015 | 0.754 |
| Characteristic | A-Carriers | GG-Carriers | Statistics | p Values |
|---|---|---|---|---|
| ADAS13 | 17.63 ± 8.68 | 18.9 ± 9.70 | F = 0.026 | 0.871 |
| RAVLT-immediate | 33.1 ± 11.16 | 32.41 ± 12.04 | F = 0.440 | 0.508 |
| Amyloid β42 (pg/mL) | 954.7 ± 531.58 | 807.17 ± 427.31 | F = 4.335 | 0.038 |
| Amyloid β40 (pg/mL) | 7678.27 ± 2322.57 | 7692.8 ± 2331.27 | F = 0.012 | 0.914 |
| Amyloid β42/40 | 0.125 ± 0.055 | 0.108 ± 0.050 | F = 4.733 | 0.039 |
| Amyloid β38 (pg/mL) | 1804.39 ± 588.23 | 1795.11 ± 566.83 | F = 0.020 | 0.889 |
| T-tau (pg/mL) | 283.37 ± 109.25 | 316.03 ± 130.67 | F = 2.110 | 0.147 |
| P-tau (pg/mL) | 27.25 ± 12.14 | 31.45 ± 14.98 | F = 3.164 | 0.076 |
| P/T-tau | 0.094 ± 0.009 | 0.097 ± 0.009 | F = 4.765 | 0.030 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Wu, Z.; Yang, Y.; Song, Z.; Ma, M.; Feng, M.; Liu, Y.; Xing, H.; Chang, Y.; Dai, H. PICALM rs3851179 Variants Modulate Left Postcentral Cortex Thickness, CSF Amyloid β42, and Phosphorylated Tau in the Elderly. Brain Sci. 2022, 12, 1681. https://doi.org/10.3390/brainsci12121681
Wu Z, Yang Y, Song Z, Ma M, Feng M, Liu Y, Xing H, Chang Y, Dai H. PICALM rs3851179 Variants Modulate Left Postcentral Cortex Thickness, CSF Amyloid β42, and Phosphorylated Tau in the Elderly. Brain Sciences. 2022; 12(12):1681. https://doi.org/10.3390/brainsci12121681
Chicago/Turabian StyleWu, Zhiwei, Yiwen Yang, Ziyang Song, Mengya Ma, Mengmeng Feng, Yuanqing Liu, Hanqi Xing, Yue Chang, and Hui Dai. 2022. "PICALM rs3851179 Variants Modulate Left Postcentral Cortex Thickness, CSF Amyloid β42, and Phosphorylated Tau in the Elderly" Brain Sciences 12, no. 12: 1681. https://doi.org/10.3390/brainsci12121681
APA StyleWu, Z., Yang, Y., Song, Z., Ma, M., Feng, M., Liu, Y., Xing, H., Chang, Y., & Dai, H. (2022). PICALM rs3851179 Variants Modulate Left Postcentral Cortex Thickness, CSF Amyloid β42, and Phosphorylated Tau in the Elderly. Brain Sciences, 12(12), 1681. https://doi.org/10.3390/brainsci12121681





