2.1. Notations and Definitions
According to the application of this study, cerebrovascular examination data records of patients were collected and denoted as set .
A set of
consists of six features: age
, systolic blood pressure
, body-mass index
, fasting glucose
, and total cholesterol
, which correspond to MRI results as class labels
. Furthermore, there is a collection of distinct items from each feature, denoted
, thus
. Set
can be written as
where
for simplicity. Furthermore, one notes that there is no patient with a
value in both the obese and underweight ranges at the same time in medical case.
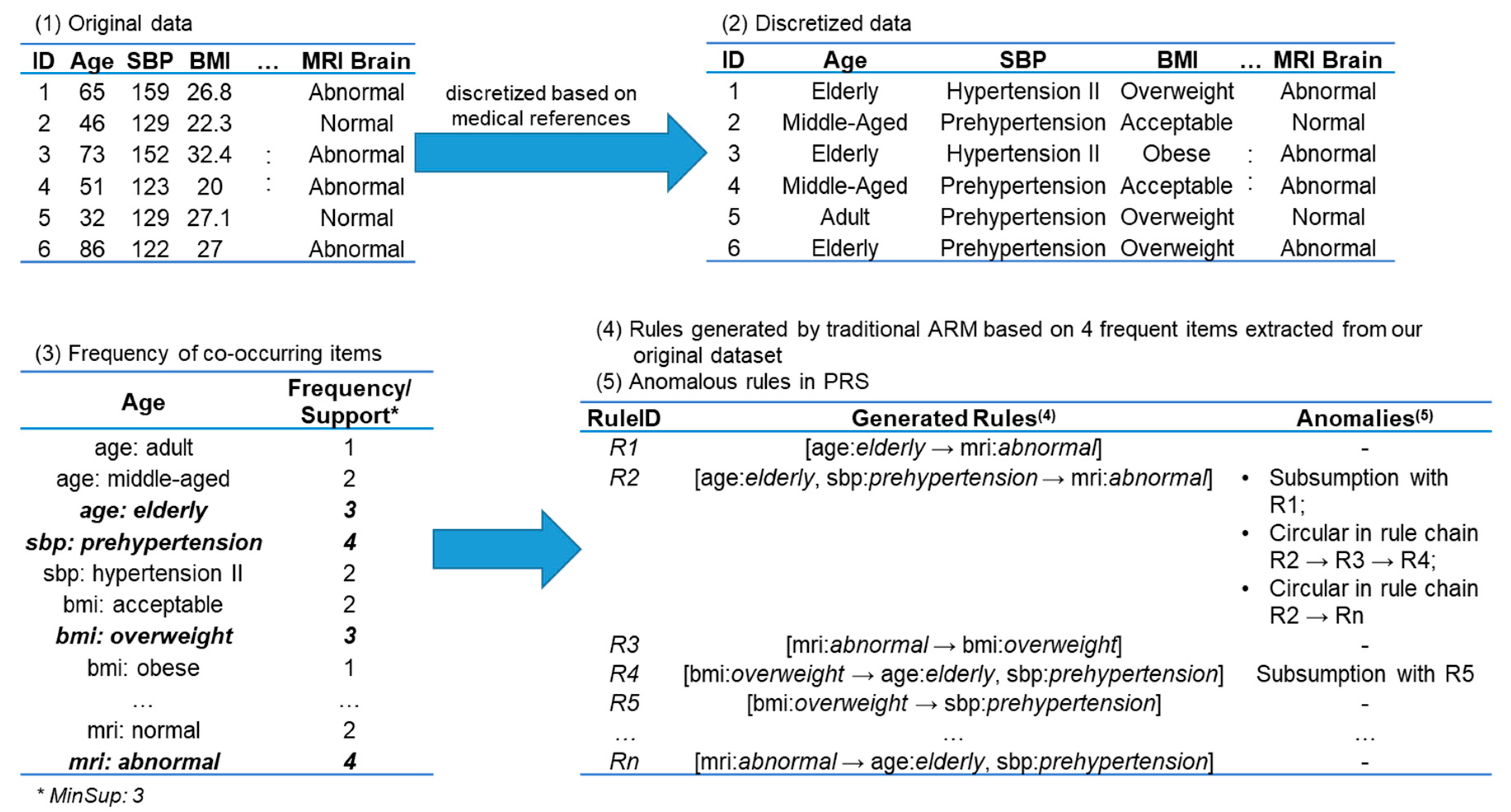
Table 1 gives an illustration of database
.
To discover useful information from the database
, conventional ARM is a popular data-mining technique to assist experts—i.e., medical practitioners, in this case—in decision making. Useful information can be represented as a rule
, where
. Hence,
and
can also be said to be item sets. To determine an association rule
, two measurements need to be defined, e.g., support and confidence values (
and
, respectively), as follows:
Based on [
23],
or
can be considered a frequent itemset if their support values satisfy a predefined minimum support threshold,
, as defined below in Definition 1.
Definition 1. (Frequent itemset)
Given a minimum support threshold , then an itemset is a frequent itemset if .
In conventional ARM, a generate-and-test approach is employed to collect frequent itemsets and subsequently build ARs based on two predefined threshold values. Formally, conventional ARM is defined as follows.
Definition 2. (Conventional ARM)
Given minimum support and confidence thresholds and , and two frequent itemsets and , we extract a set of association rules that fulfill the following conditions:
; and
.
However, conventional ARM is computationally expensive in terms of run-time and memory usage. To overcome this drawback, this study focused on ARM using a hybrid approach [
19]—with a vertical database format (the detailed illustration of a vertical format on
is given in Table 3 in
Section 2.3.1)—and presents NRARs. Furthermore, there are five types of non-redundant frequent item sets discussed in this study: FCI, generator of FCI (GFCI), minimal GFCI (mGFCI), non-subsumption FCI (nSFCI), and non-circular FCI (nCFCI). Henceforth, this study defines all non-redundant frequent item sets and subsequently generates a set of concise rules, so-called non-anomalous association rules (NAARs), based on the aforementioned types of non-redundant frequent item sets. Moreover, this study also takes some properties of PRs into consideration toward NAARs as the main focus.
An FCI is a frequent itemset which has no superset with the same support values. Formally, an FCI is defined as follows:
Definition 3. (Frequent Closed Itemset)
Given a frequent itemset , is an FCI .
One previous study states that a generator item set should be selected according to the minimum description length [
24]. An item set is called a generator if there is no subsequence with the same support values. The formal definition of a generator is presented below.
Definition 4. (Generator Frequent Closed Itemset)
Given an FCI, where is a GFCI .
According to Definition 4, a set of GFCIs may still contain redundant generators, e.g., two GFCIs that share the same information; thus, the longest one can be pruned to enhance the mining process.
Assume we have a set of GFCIs, such that . After that, we have a set of minimal generators , which, by definition, can be described as follows.
Definition 5. (Minimal Generator Frequent Closed Itemset)
Let be a set of GFCIs. can be said to be an mGFCI if , where and .
Regarding Definition 5, we can clearly state that a set of mGFCIs () is not an empty set. Additionally, there is a property of as follows.
Property 1. Ifis a set of mGFCIs, then: (i); and/or (ii)when no proper generatorcan be found.
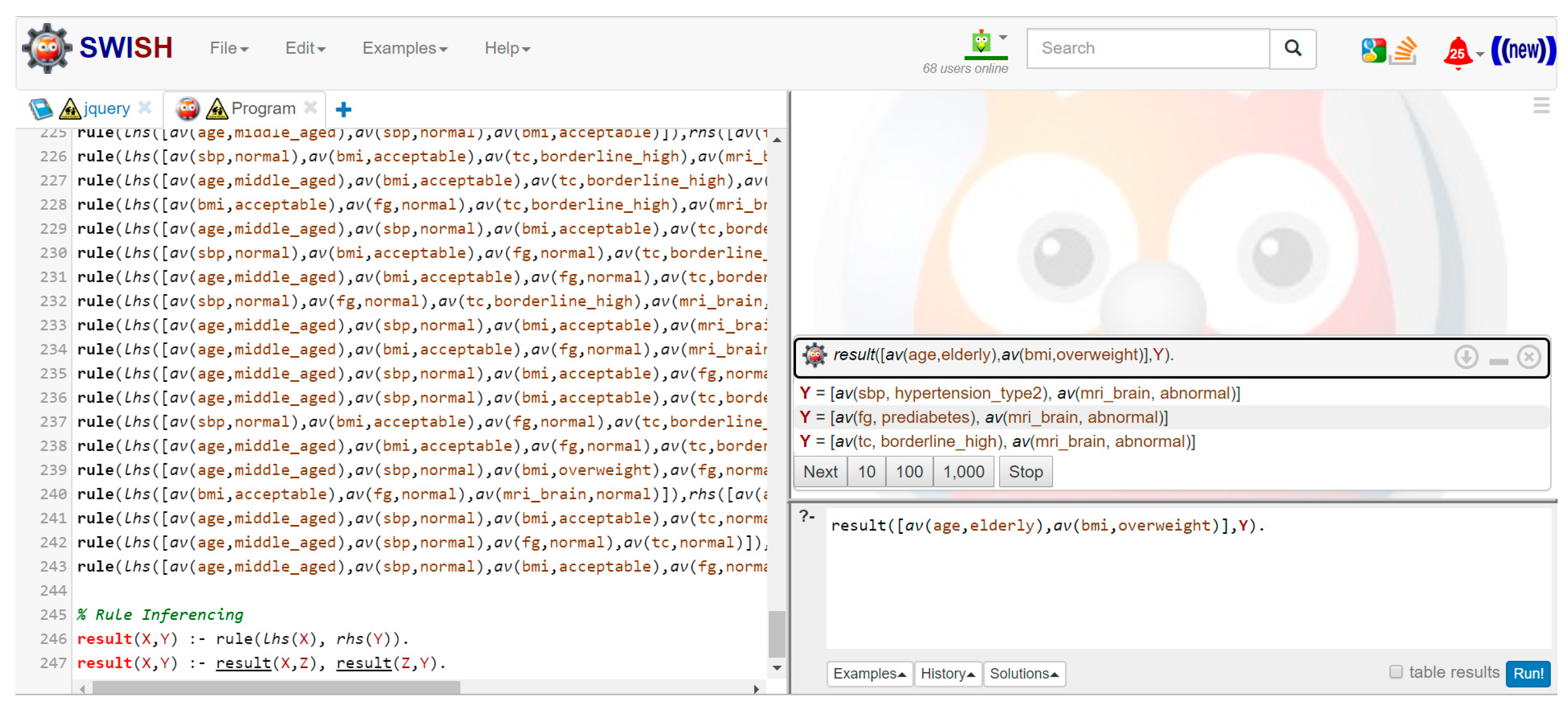
By adopting Definition 2, an AR from an mGFCI set can be generated, which is further called a non-redundant association rule (NRAR). In conventional ARM, ARs can be obtained by randomly putting all possible FCIs on the antecedent or consequent sides. However, because the number of mGFCIs is small, it may difficult to construct a rule only from mGFCIs. Conforming to [
17], a NRAR can be formed by two combinations between FCIs and mGFCIs. To construct a NRAR, mGFCIs are considered a premise, while items in FCI—excluding that of its corresponding mGFCI—are put as the conclusion. The definition of a NRAR is described below.
Definition 6. (A Non-Redundant Association Rule)
Let be a set of mGFCIs and is a set of FCIs. A NRAR can be written as where and .
Regarding Property 1 and Definition 6, we ensure that is a finite set, which means that we do not need to worry about losing any information, e.g., . A construction rule, with , and , can still be applied and considered to be a NRAR, even when . On the other hand, since we are trying to obtain concise rule bases, then PRs are also discussed as one of main focuses of this study.
Definition 7. (Production Rules)
A rule-based mode of knowledge representation is termed a PR. The rule is basically in the form of if <premises> then <conclusions> or similarly if <evidence (E)> then <hypotheses (H)>.
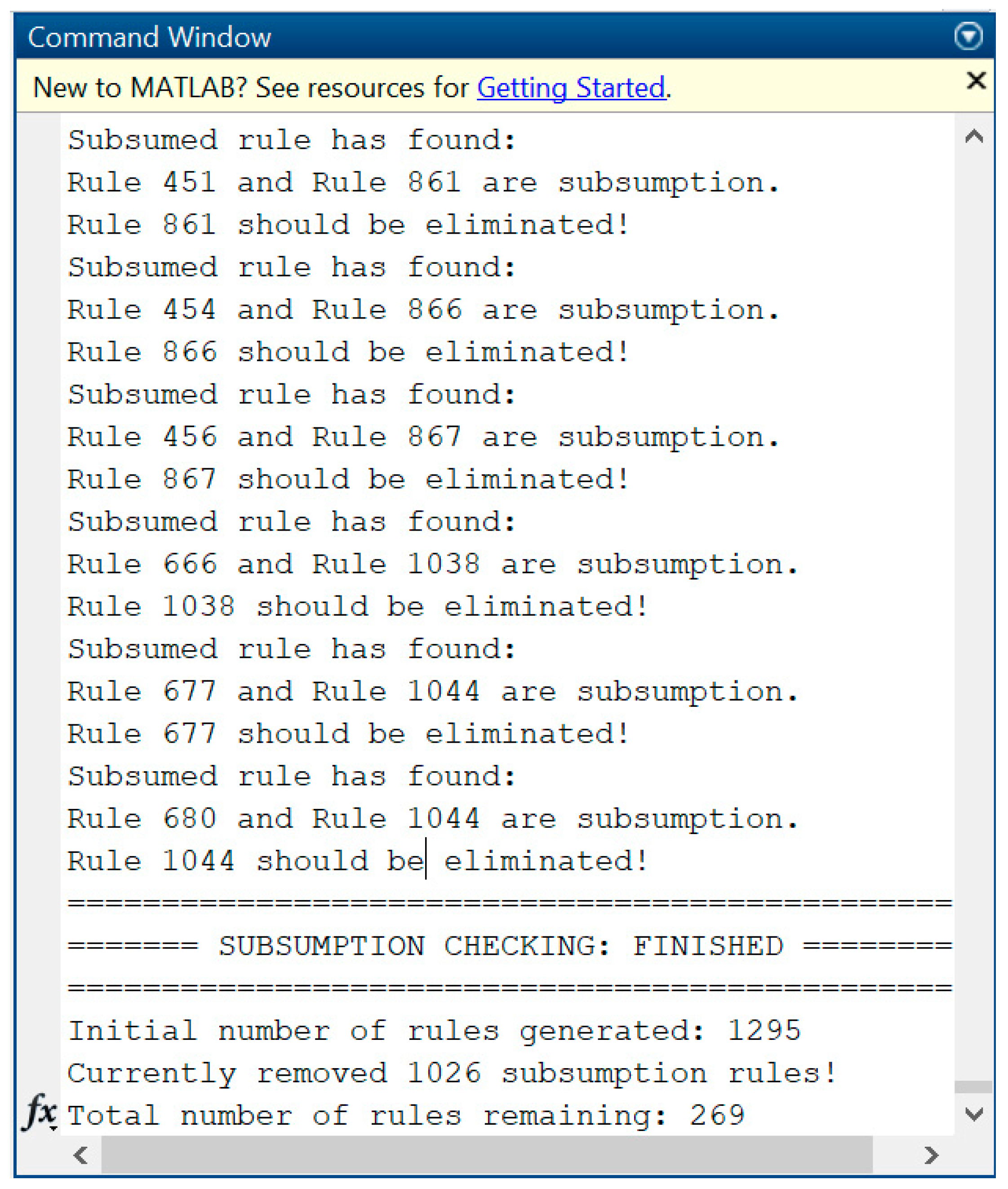
PRs can be applied as rule bases after the collection of rules in PRs is verified through a logical testing procedure. For simplicity, we need to ensure that there is no single rule that can be implied with other rules, called non-subsumption rules (or non-implication rules). For instance, there are two rules:
.
.
From a physician’s viewpoint, both and are equivalent rules, since those share similar symptoms, such that result shows . Hence, a non-subsumption AR can be defined as follows.
Definition 8. (Non-subsumption ARs)
Suppose there is a set of rules . A rule is a non-subsumption rule if .
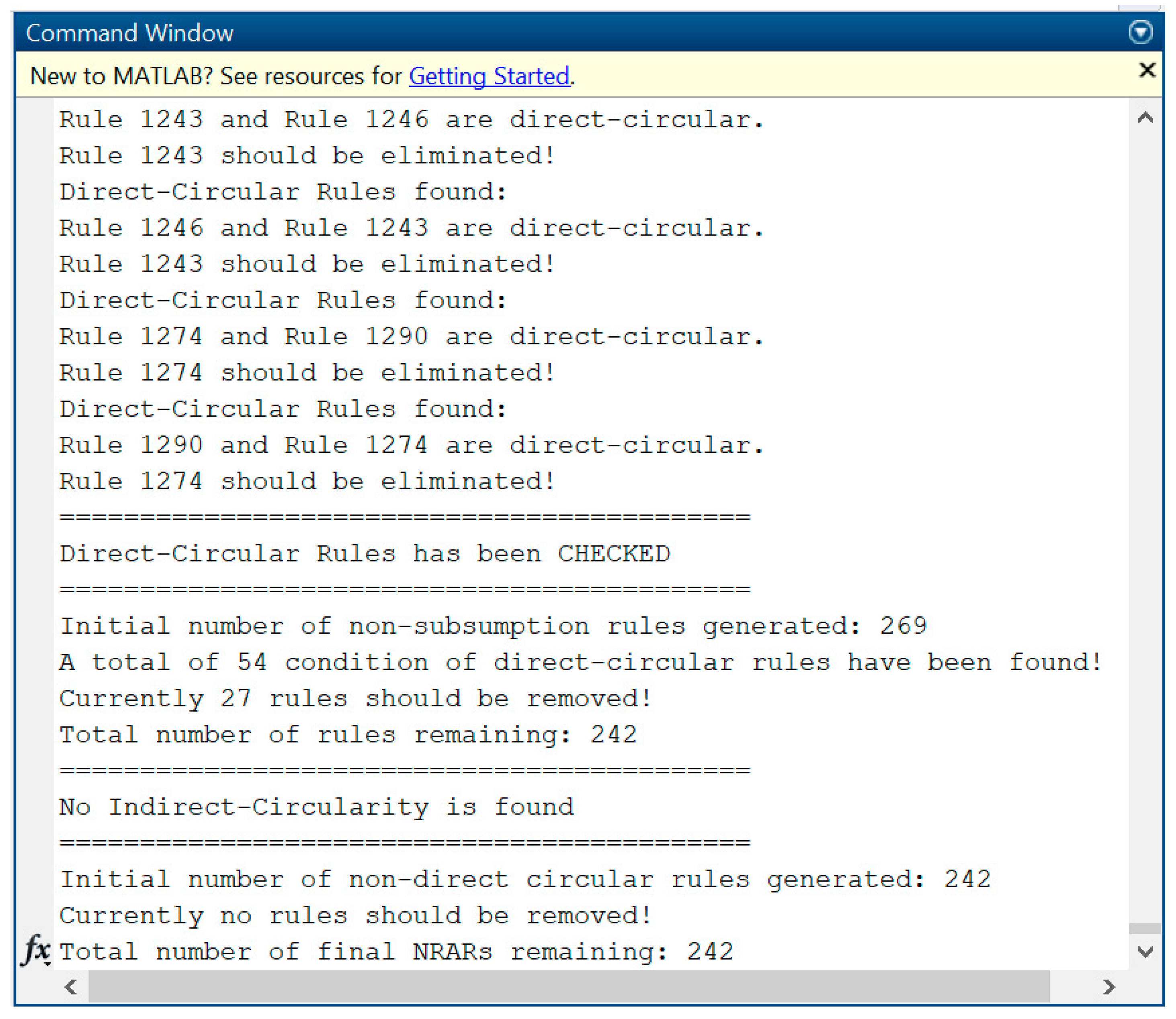
In addition, both ARs and NRARs may still produce rules in which the conclusion of one rule is the same as the premise of another rule or vice versa. Assume there are two rules:
.
.
Regarding [
21], those rules are considered as a circular rule, which tends to be contradictory in meaning or logic. Hence,
can be removed. Consequently, rule
becomes a non-circular rule. The formal definition of non-circular ARs is described below.
Definition 9. (Non-circular ARs)
Given a set of rules
. A rule
is a non-circular rule if
, i.e., the rules create a closed-chain. In this paper, a rule chain might be formed during rule inference and create a closed-chain (i.e., circularity) in any of the following conditions presented in
Table 2.
Afterwards, a set of NRARs is pruned into NAARs that fulfill both non-subsumption and non-circular rule definitions. In this study, some properties of NAAR mining according to CF values are derived to relate between NAARs and PRs in the following section, then the proposed methodology is introduced.
2.2. NAARs as PRs
This study introduces NAARs that satisfy logical testing in KBs; thus, they can be considered PRs. This paper was motivated by project work in the area of medical diagnoses, called MYCIN, which began in 1972 [
20]. It is a rule-based ES for diagnosing infectious blood diseases. MYCIN inferred conclusions from a given condition through certain factor values, which are a calculus of uncertainty for measuring the credibility of a rule, since reasoning in uncertainty is the most important part of MYCIN. Uncertain knowledge representation is also introduced in this PRS by identifying its certainty factor value [
25]. As implied, the generation of an AR differs from PRs as there are two quality measurements (i.e., support and confidence) in ARM. However, we found that quality measurements in ARM and PRs share similar statistical-approach concepts, in which Bayesian probability, i.e., conditional probability, is employed. The certainty factor (
) is denoted by
where
is a measurement of belief and
is a measurement of disbelief which are denoted below.
where,
is a probability of
and
is a conditional probability of
given
.
To relate quality measurements in NRARs with
values in PRs, the gain value of a rule
represents the difference between confidence and support values of an AR, which is denoted as
. Later,
can be utilized to relate to
Based on [
26], we show the derivation of
.
Lemma 1. (Gain Value of an AR)
Suppose that
is an AR. The gain value of
is given by
As in ARM [
23], a set of ARs is extracted by satisfying confidence and/or support threshold values, where
and
. Consequently, the range value of
can be determined as a bounded value of
.
Corollary 1. Given an AR, the bounded value ofis in range.
Proof. Since and , when , and when .
From Equation (6), certainty factors of ARs can be viewed as the value over the value of the consequent part. We denote certainty factor values of ARs as . The derivation of is described as follows.
Theorem 1. (Certainty Factors of ARs)
Let
be an AR. If
, then we have
, such that
And the range interval of is .
Proof. With regard to Corollary 1, it is obvious that as . Hence, there are two main conditions:
According to Theorem 1, we can define NAARs, which represent PRs as rule bases. In detail, NRARs, which are still contained of the subsumption or circular rules, are eliminated by considering the smaller value. Considering all the aforementioned discussion, the definition of NAARs in this study is explained as follows.
Definition 10. (NAARs as PRs)
Let be a set of association rules. An association rule , with are FCI and mGFCI, where , is said to be an NAAR as PR by sequentially satisfying the following conditions:
at this state;
with ; and
with .