Federating Medical Deep Learning Models from Private Jupyter Notebooks to Distributed Institutions
Abstract
1. Introduction
- Reduce the complexity of implementing a federated learning framework by making use of a Jupyter notebook code template for artificial intelligence experts;
- Improve the deployment efficiency of cross-silo federated learning systems by automatically containerizing the Jupyter components to provide to distributed users;
- Seamlessly bridge the gap between Jupyter and FL systems by providing a generic solution that is both easy-to-use and reproducible;
- Give an example tutorial of the proposed solution, the Notebook Federator, by applying it to a real-world cross-countries computational pathology classification task.
2. Related Work
3. Notebook Federator
3.1. System Definition and Requirements
3.2. Challenges and Assumptions
- All the participating institutions are already registered as part of the federation;
- Local data cannot be shipped outside the institution;
- Local data are stored on each client machine following the same—previously defined—nomenclature for labels and metadata across institutions. As some institutions may have their own tools or methods to process raw data, we assume the data have passed the quality check;
- Each medical collaborator either has local machines (e.g., local computers) to run the code or an agreement with a research group that acts as a local infrastructure (e.g., high-performance servers, or computer clusters) and is thus able to train models;
- Each participating entity has a Docker cluster or is able to build Docker images to run the corresponding code.
3.3. Architecture
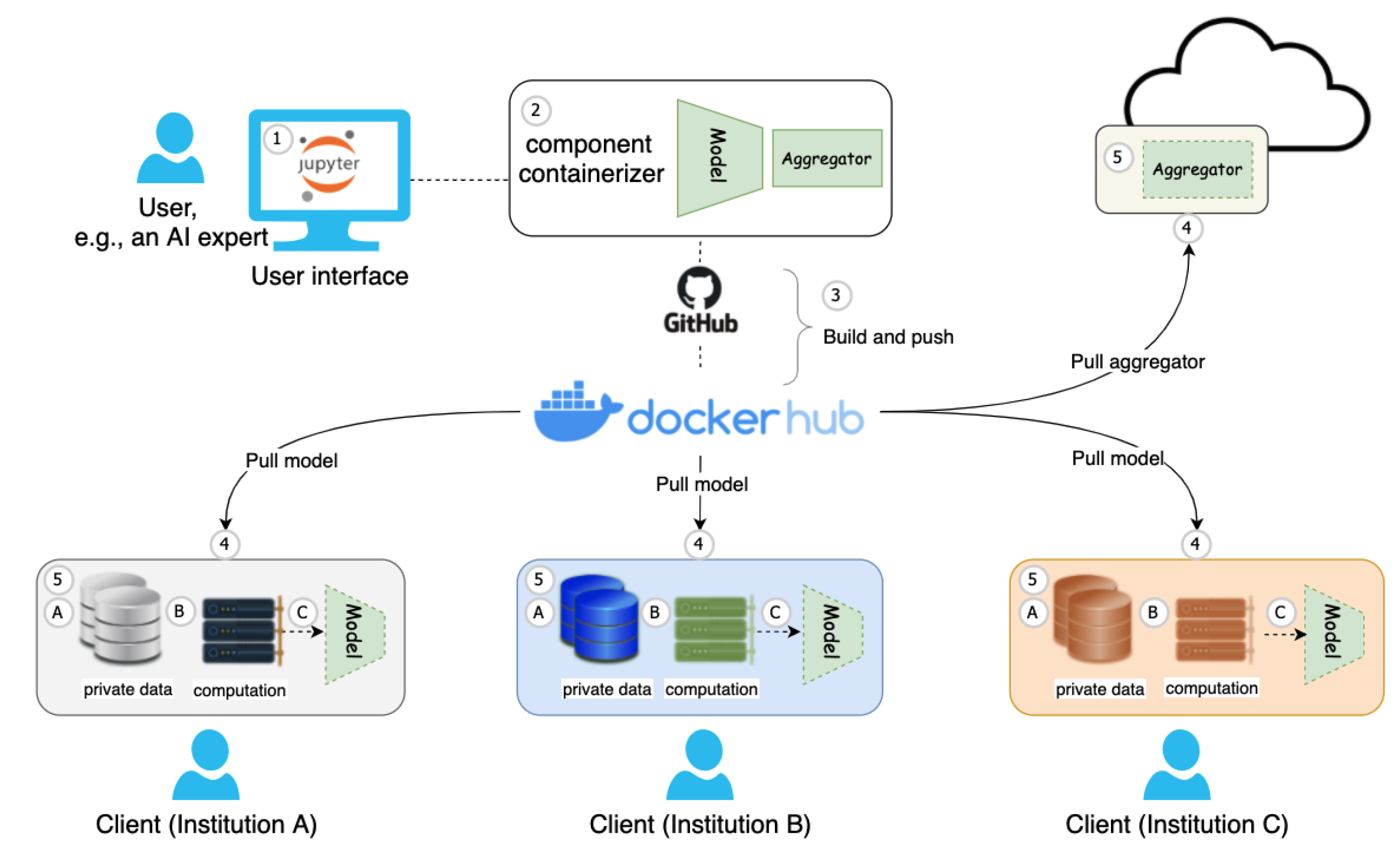
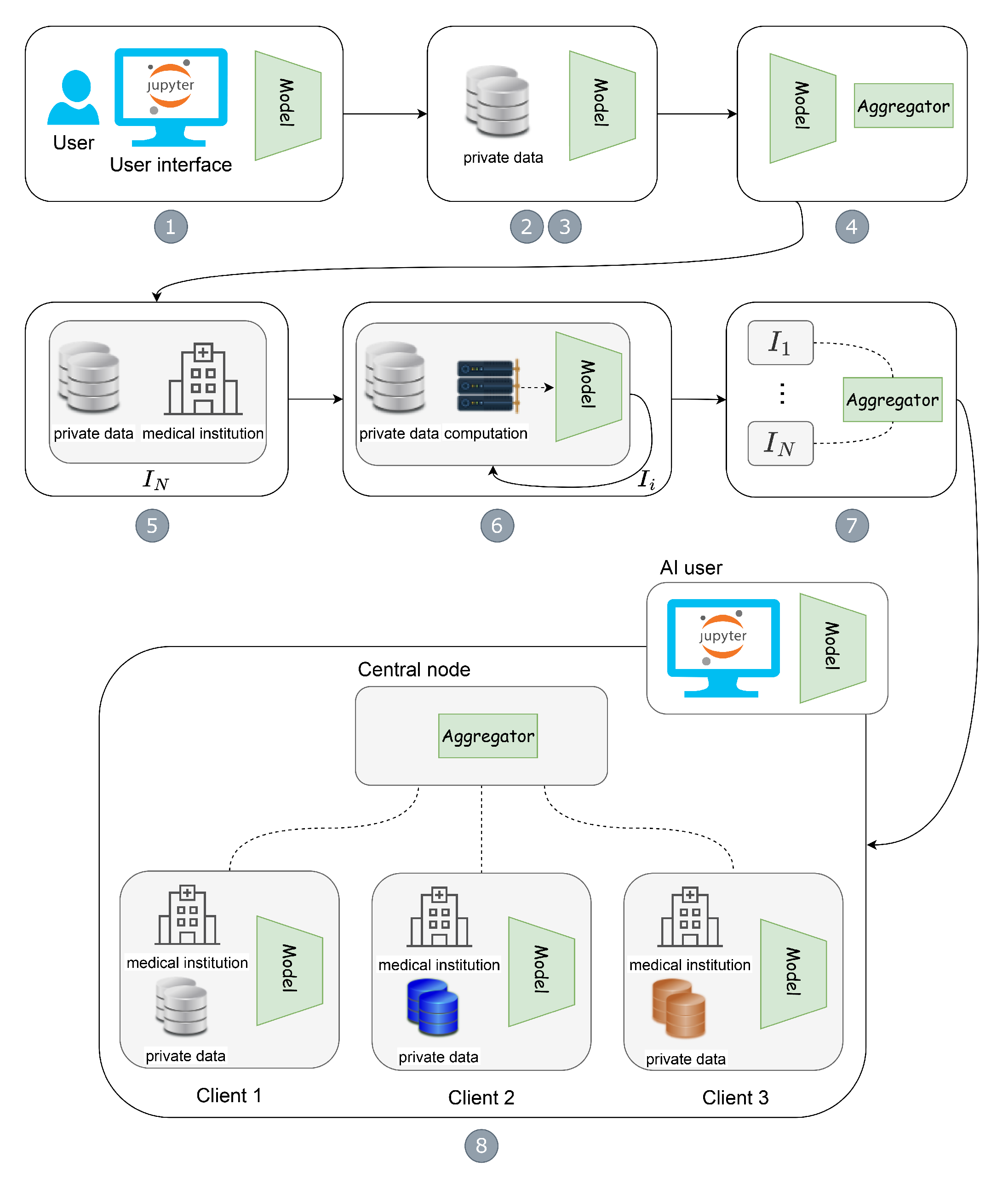
- Model and aggregator definition: Suppose a Jupyter Notebook user, e.g., an AI expert, develops a cutting-edge model architecture, aggregation function, and other machine learning-related code fragments for the FL training process.
- Create FL pipeline building blocks: For flexibility purposes when it comes to handling collaborating institutions and updates, as well as reducing the complexity of FL deployment and execution, we propose to adopt one Jupyter Notebook extension named component containerizer on the local experimental environment, e.g., NaaVRE [8], to encapsulate FL pipeline building blocks as reusable services, such as model and aggregator.
- Build and push job automation: Once the model and aggregator are ready, with the workflow of the GitHub project and docker registry, such components can be automatically built and pushed to the Docker Hub.
- Deliver FL building blocks to distributed resources: The model can be delivered to distributed client users worldwide, i.e., geographically distributed clients in institutions A, B, and C, by pulling the model to local sites. At the same time, the aggregator can be easily delivered to the cloud infrastructure by pulling the aggregator to the cloud virtual machine.
- Federation setup: Once the aggregator is assigned to the cloud infrastructure, e.g., a cloud virtual machine, it is easy to start the Docker container with specific IP and port number. For geographically distributed users, i.e., clients across wide-area institutions, the local AI model training process mainly contains (A) feeding with local data, (B) assigning suitable computation, and (C) starting the model training with specific data and computation (e.g., GPU resources) on site. By this time, the FL starts.
3.4. Technology Considerations
- Jupyter environment—We build the FL pipeline with Jupyter Notebook. By default, we suppose that AI experts use a Jupyter environment such as Notebook to design the architecture of the model and aggregation for FL experiments.
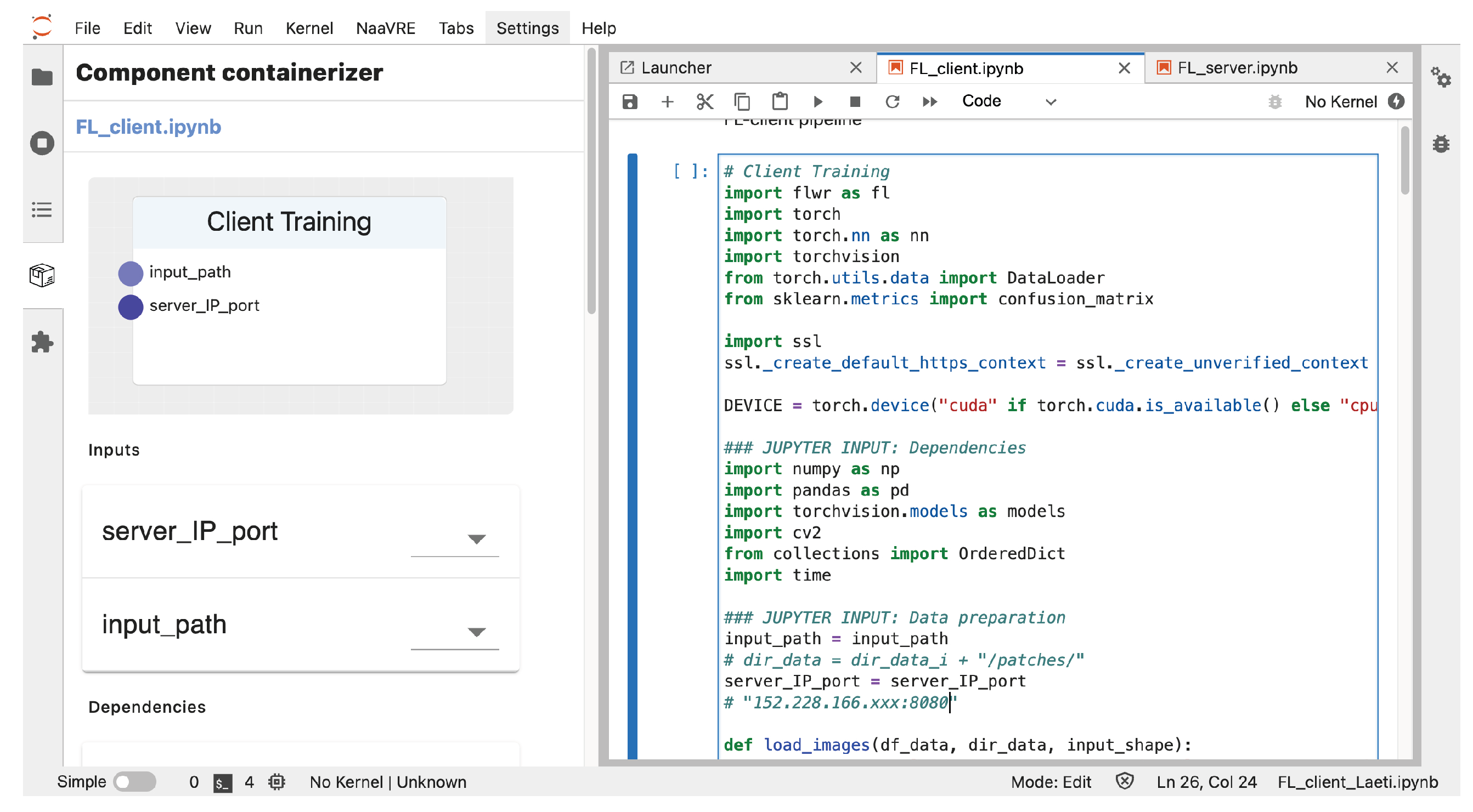
- Component containerizer—We encapsulate the model and aggregator as reusable FL building blocks based on a Jupyter Notebook. The component containerizer module is one of the Jupyter extensions in NaaVRE.
- Docker Hub—We share model and aggregator with distributed users around the world. Docker Hub, i.e., a central repository of containers, is the easiest way to deliver reusable container applications anywhere.
- Docker Engine—We enhance the automation of the FL deployment and execution. In this work, we consider the Docker Engine as the critical technology for FL pipeline automation because of its popularity in the community and flourishing software tools such as Docker container, docker-nvidia, docker-compose, or even Docker Swarm. In this paper, we mainly utilize docker-nvidia for local client training with CUDA GPU resources for automating FL pipelines, as we suppose that client users have their demands for local autonomy, e.g., controlling their own data and computation for AI model training, although using Docker Swarm can also achieve large-scale automated deployment for client users.
4. Case Study: Histological Image Analysis
4.1. Use Case Scenarios
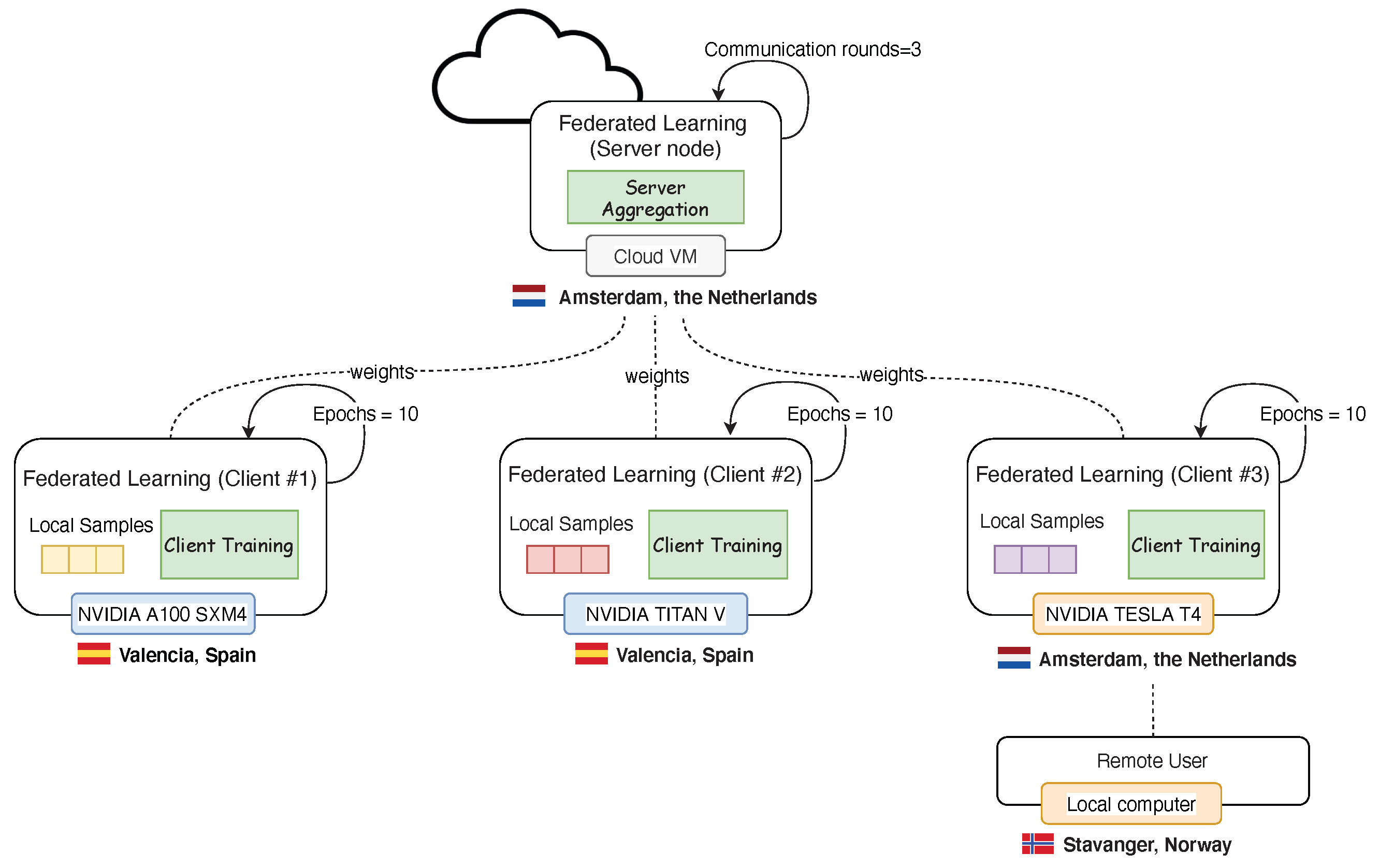
4.2. Federation
4.3. Federated Implementation
4.4. Experimental Results
4.4.1. FL System Setup
- Automated build–push job for the server-aggregation image container (approx. 1.9 GB): 4 m 57 s;
- Building and pushing client-training image container to the Docker hub (approx. 3.16 GB): 10 m 48 s.
4.4.2. AI Training
5. Discussion
5.1. Achievements
5.2. Weaknesses and Future Work
6. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| ACC | Accuracy |
| AI | Artificial Intelligence |
| DOAJ | Directory of Open Access Journals |
| F1S | F1-score |
| FL | Federated Learning |
| MDPI | Multidisciplinary Digital Publishing Institute |
| NaaVRE | Notebook-as-a-VRE |
| NPV | Negative Predictive Value |
| PPV | Positive Predictive Value |
| ROI | Region of Interest |
| SN | Sensitivity |
| SPC | Specificity |
| VRE | Virtual Research Environment |
| WSI | Whole Slide Image |
References
- Chen, R.J.; Lu, M.Y.; Chen, T.Y.; Williamson, D.F.; Mahmood, F. Synthetic data in machine learning for medicine and healthcare. Nat. Biomed. Eng. 2021, 5, 493–497. [Google Scholar] [CrossRef] [PubMed]
- Oza, P.; Sharma, P.; Patel, S.; Adedoyin, F.; Bruno, A. Image Augmentation Techniques for Mammogram Analysis. J. Imaging 2022, 8, 141. [Google Scholar] [CrossRef] [PubMed]
- Rocher, L.; Hendrickx, J.M.; de Montjoye, Y.A. Estimating the success of re-identifications in incomplete datasets using generative models. Nat. Commun. 2019, 10, 1–9. [Google Scholar] [CrossRef]
- Konecný, J.; McMahan, H.B.; Ramage, D.; Richtárik, P. Federated Optimization: Distributed Machine Learning for On-Device Intelligence. arXiv 2016, arXiv:abs/1610.02527. [Google Scholar]
- Li, T.; Sahu, A.K.; Talwalkar, A.; Smith, V. Federated Learning: Challenges, Methods, and Future Directions. IEEE Signal Process. Mag. 2020, 37, 50–60. [Google Scholar] [CrossRef]
- Li, W.; Milletarì, F.; Xu, D.; Rieke, N.; Hancox, J.; Zhu, W.; Baust, M.; Cheng, Y.; Ourselin, S.; Cardoso, M.J.; et al. Privacy-preserving federated brain tumour segmentation. In Proceedings of the International Workshop on Machine Learning in Medical Imaging; Springer International Publishing: Cham, Switzerland, 2019; pp. 133–141. [Google Scholar]
- Sheller, M.J.; Edwards, B.; Reina, G.A.; Martin, J.; Pati, S.; Kotrotsou, A.; Milchenko, M.; Xu, W.; Marcus, D.; Colen, R.R.; et al. Federated learning in medicine: Facilitating multi-institutional collaborations without sharing patient data. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Z.; Koulouzis, S.; Bianchi, R.; Farshidi, S.; Shi, Z.; Xin, R.; Wang, Y.; Li, N.; Shi, Y.; Timmermans, J.; et al. Notebook-as-a-VRE (NaaVRE): From private notebooks to a collaborative cloud virtual research environment. Softw. Pract. Exp. 2022, 52, 1947–1966. [Google Scholar] [CrossRef]
- Kairouz, P.; McMahan, H.B.; Avent, B.; Bellet, A.; Bennis, M.; Bhagoji, A.N.; Bonawitz, K.; Charles, Z.; Cormode, G.; Cummings, R.; et al. Advances and open problems in federated learning. Found. Trends® Mach. Learn. 2021, 14, 1–210. [Google Scholar] [CrossRef]
- Rieke, N.; Hancox, J.; Li, W.; Milletari, F.; Roth, H.R.; Albarqouni, S.; Bakas, S.; Galtier, M.N.; Landman, B.A.; Maier-Hein, K.; et al. The future of digital health with federated learning. NPJ Digit. Med. 2020, 3, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Roy, A.G.; Siddiqui, S.; Pölsterl, S.; Navab, N.; Wachinger, C. BrainTorrent: A Peer-to-Peer Environment for Decentralized Federated Learning. arXiv 2019, arXiv:abs/1905.06731. [Google Scholar]
- Li, X.; Gu, Y.; Dvornek, N.; Staib, L.H.; Ventola, P.; Duncan, J.S. Multi-site fMRI analysis using privacy-preserving federated learning and domain adaptation: ABIDE results. Med. Image Anal. 2020, 65, 101765. [Google Scholar] [CrossRef] [PubMed]
- Ju, C.; Gao, D.; Mane, R.; Tan, B.; Liu, Y.; Guan, C. Federated Transfer Learning for EEG Signal Classification. In Proceedings of the Annual International Conference of the IEEE Engineering in Medicine and Biology Society, Montreal, QC, Canada, 20–24 July 2020. [Google Scholar] [CrossRef]
- Andreux, M.; du Terrail, J.O.; Beguier, C.; Tramel, E.W. Siloed Federated Learning for Multi-centric Histopathology Datasets. In Proceedings of the Lecture Notes in Computer Science (Including subseries Lecture Notes in Artificial Intelligence and Lecture Notes in Bioinformatics); Springer International Publishing: Cham, Switzerland, 2020. [Google Scholar] [CrossRef]
- Lu, M.Y.; Chen, R.J.; Kong, D.; Lipkova, J.; Singh, R.; Williamson, D.F.; Chen, T.Y.; Mahmood, F. Federated learning for computational pathology on gigapixel whole slide images. Med. Image Anal. 2022, 76, 102298. [Google Scholar] [CrossRef] [PubMed]
- Ziller, A.; Trask, A.; Lopardo, A.; Szymkow, B.; Wagner, B.; Bluemke, E.; Nounahon, J.M.; Passerat-Palmbach, J.; Prakash, K.; Rose, N.; et al. PySyft: A Library for Easy Federated Learning. In Federated Learning Systems: Towards Next-Generation AI; ur Rehman, M.H., Gaber, M.M., Eds.; Springer International Publishing: Cham, Switzerland, 2021; pp. 111–139. [Google Scholar] [CrossRef]
- Reina, G.A.; Gruzdev, A.; Foley, P.; Perepelkina, O.; Sharma, M.; Davidyuk, I.; Trushkin, I.; Radionov, M.; Mokrov, A.; Agapov, D.; et al. OpenFL: An open-source framework for Federated Learning. arXiv 2021, arXiv:2105.06413. [Google Scholar] [CrossRef]
- Beutel, D.J.; Topal, T.; Mathur, A.; Qiu, X.; Parcollet, T.; Lane, N.D. Flower: A Friendly Federated Learning Research Framework. arXiv 2020, arXiv:2007.14390. [Google Scholar]
- Xie, Y.; Wang, Z.; Chen, D.; Gao, D.; Yao, L.; Kuang, W.; Li, Y.; Ding, B.; Zhou, J. FederatedScope: A Flexible Federated Learning Platform for Heterogeneity. arXiv 2022, arXiv:2204.05011. [Google Scholar]
- Lee, H.; Chai, Y.J.; Joo, H.; Lee, K.; Hwang, J.Y.; Kim, S.M.; Kim, K.; Nam, I.C.; Choi, J.Y.; Yu, H.W.; et al. Federated learning for thyroid ultrasound image analysis to protect personal information: Validation study in a real health care environment. JMIR Med. Inform. 2021, 9, e25869. [Google Scholar] [CrossRef] [PubMed]
- Florescu, L.M.; Streba, C.T.; Şerbănescu, M.S.; Mămuleanu, M.; Florescu, D.N.; Teică, R.V.; Nica, R.E.; Gheonea, I.A. Federated Learning Approach with Pre-Trained Deep Learning Models for COVID-19 Detection from Unsegmented CT images. Life 2022, 12, 958. [Google Scholar] [CrossRef] [PubMed]
- Lo, J.; Timothy, T.Y.; Ma, D.; Zang, P.; Owen, J.P.; Zhang, Q.; Wang, R.K.; Beg, M.F.; Lee, A.Y.; Jia, Y.; et al. Federated learning for microvasculature segmentation and diabetic retinopathy classification of OCT data. Ophthalmol. Sci. 2021, 1, 100069. [Google Scholar] [CrossRef] [PubMed]
- Lodha, S.; Saggar, S.; Celebi, J.T.; Silvers, D.N. Discordance in the histopathologic diagnosis of difficult melanocytic neoplasms in the clinical setting. J. Cutan. Pathol. 2008, 35, 349–352. [Google Scholar] [CrossRef] [PubMed]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. In Proceedings of the 3rd International Conference on Learning Representations, ICLR 2015—Conference Track Proceedings, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Brendan McMahan, H.; Moore, E.; Ramage, D.; Hampson, S.; Agüera y Arcas, B. Communication-efficient learning of deep networks from decentralized data. In Proceedings of the 20th International Conference on Artificial Intelligence and Statistics (AISTATS 2017), Fort Lauderdale, FL, USA, 20–22 April 2017. [Google Scholar]
- Kanagavelu, R.; Li, Z.; Samsudin, J.; Yang, Y.; Yang, F.; Goh, R.S.; Cheah, M.; Wiwatphonthana, P.; Akkarajitsakul, K.; Wang, S. Two-Phase Multi-Party Computation Enabled Privacy-Preserving Federated Learning. In Proceedings of the 20th IEEE/ACM International Symposium on Cluster, Cloud and Internet Computing (CCGRID 2020), Melbourne, VIC, Australia, 11–14 May 2020; pp. 410–419. [Google Scholar] [CrossRef]

| # WSIs | 10 | 6 | 8 |
| # tumorous patches | 1554 | 2627 | 1005 |
| # non-tumorous patches | 5609 | 4694 | 3979 |
| Federation | ||||
|---|---|---|---|---|
| Sensitivity | ||||
| Specificity | ||||
| Positive predictive value | ||||
| Negative predictive value | ||||
| F1-Score | ||||
| Accuracy |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Launet, L.; Wang, Y.; Colomer, A.; Igual, J.; Pulgarín-Ospina, C.; Koulouzis, S.; Bianchi, R.; Mosquera-Zamudio, A.; Monteagudo, C.; Naranjo, V.; et al. Federating Medical Deep Learning Models from Private Jupyter Notebooks to Distributed Institutions. Appl. Sci. 2023, 13, 919. https://doi.org/10.3390/app13020919
Launet L, Wang Y, Colomer A, Igual J, Pulgarín-Ospina C, Koulouzis S, Bianchi R, Mosquera-Zamudio A, Monteagudo C, Naranjo V, et al. Federating Medical Deep Learning Models from Private Jupyter Notebooks to Distributed Institutions. Applied Sciences. 2023; 13(2):919. https://doi.org/10.3390/app13020919
Chicago/Turabian StyleLaunet, Laëtitia, Yuandou Wang, Adrián Colomer, Jorge Igual, Cristian Pulgarín-Ospina, Spiros Koulouzis, Riccardo Bianchi, Andrés Mosquera-Zamudio, Carlos Monteagudo, Valery Naranjo, and et al. 2023. "Federating Medical Deep Learning Models from Private Jupyter Notebooks to Distributed Institutions" Applied Sciences 13, no. 2: 919. https://doi.org/10.3390/app13020919
APA StyleLaunet, L., Wang, Y., Colomer, A., Igual, J., Pulgarín-Ospina, C., Koulouzis, S., Bianchi, R., Mosquera-Zamudio, A., Monteagudo, C., Naranjo, V., & Zhao, Z. (2023). Federating Medical Deep Learning Models from Private Jupyter Notebooks to Distributed Institutions. Applied Sciences, 13(2), 919. https://doi.org/10.3390/app13020919







