Mapping Cancer Registry Data to the Episode Domain of the Observational Medical Outcomes Partnership Model (OMOP)
Abstract
:1. Introduction
2. Materials and Methods
2.1. Source System
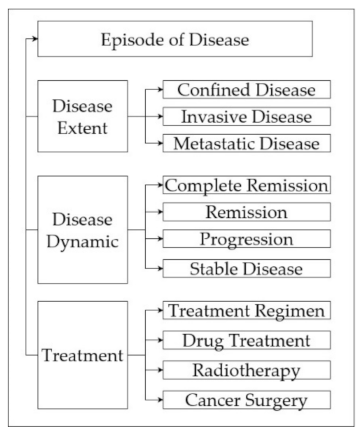
2.2. Episodic Modelling
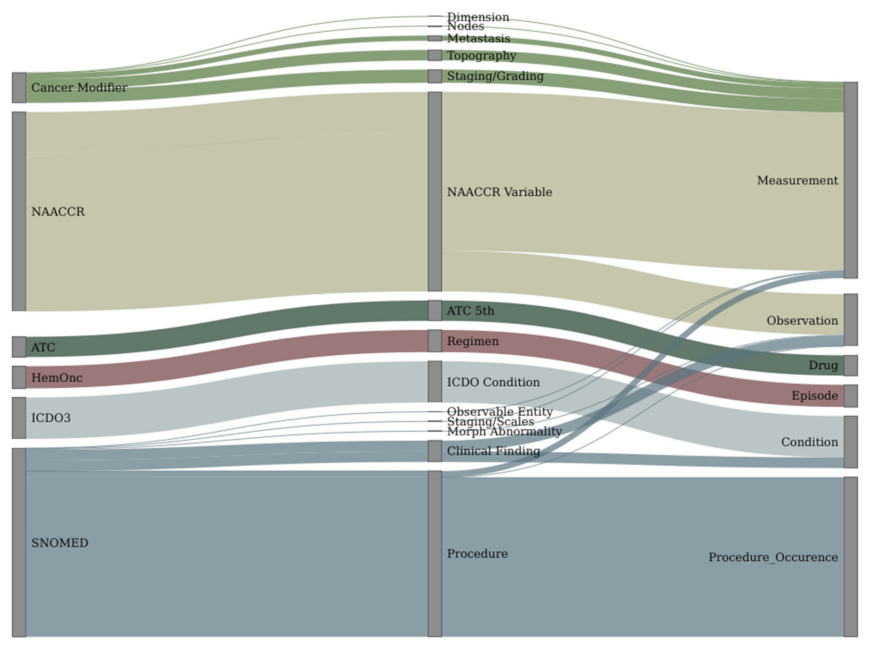
2.3. Vocabulary Mapping
2.4. Linkage between Episodes and Underlying Clinical Events
2.5. CDM Application and Comparison
3. Results
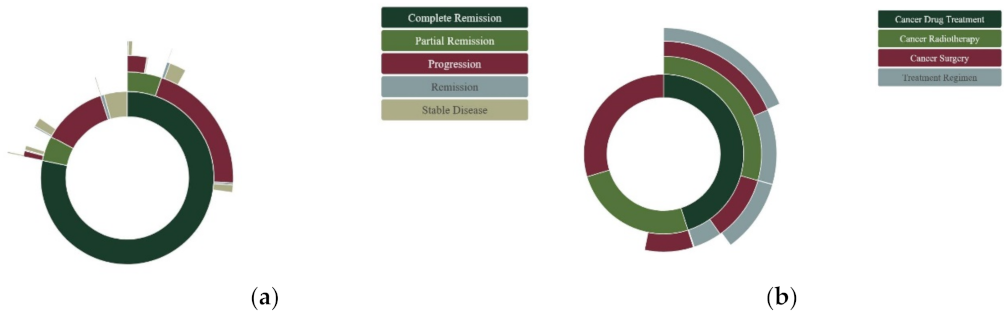
3.1. Episodic Modeling
3.2. Vocabulary Mapping
3.3. Linkage between Episodes and Underlying Clinical Events
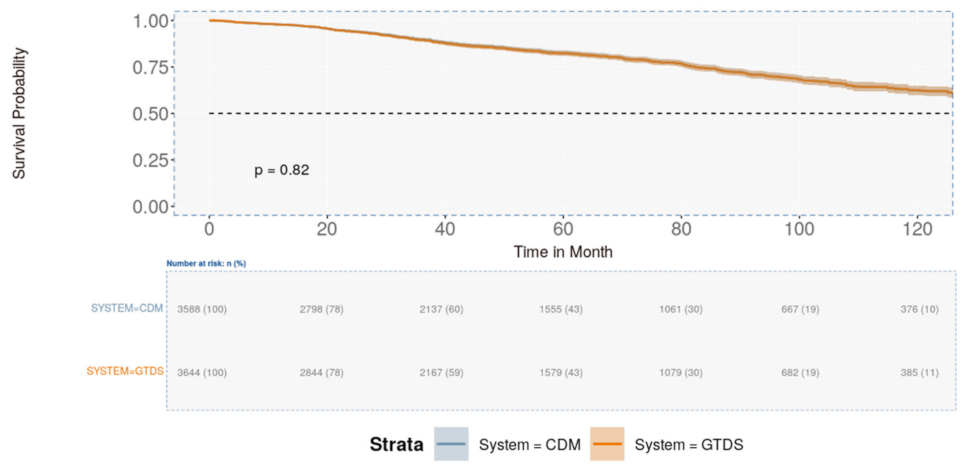
3.4. CDM Application and Comparison
4. Discussion
Limitations
5. Conclusions
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Malone, E.R.; Oliva, M.; Sabatini, P.J.B.; Stockley, T.L.; Siu, L.L. Molecular profiling for precision cancer therapies. Genome Med. 2020, 12, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Liu, S.; Kurzrock, R. Toxicity of targeted therapy: Implications for response and impact of genetic polymorphisms. Cancer Treat. Rev. 2014, 40, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Gao, K.; Liu, B.; Pan, C.; Liang, K.; Yan, L.; Ma, J.; He, F.; Zhang, S.; Pan, S.; et al. Advances in Deep Learning-Based Medical Image Analysis. Health Data Sci. 2021, 2021, 1–14. [Google Scholar] [CrossRef]
- Hawkins, S.; Wang, H.; Liu, Y.; Garcia, A.; Stringfield, O.; Krewer, H.; Li, Q.; Cherezov, D.; Gatenby, R.A.; Balagurunathan, Y.; et al. Predicting Malignant Nodules from Screening CT Scans. J. Thorac. Oncol. 2016, 11, 2120–2128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nagpal, K.; Foote, D.; Liu, Y.; Chen, P.-H.C.; Wulczyn, E.; Tan, F.; Olson, N.; Smith, J.L.; Mohtashamian, A.; Wren, J.H.; et al. Development and validation of a deep learning algorithm for improving Gleason scoring of prostate cancer. NPJ Digit. Med. 2019, 2, 48. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shimizu, H.; Nakayama, K.I. Artificial intelligence in oncology. Cancer Sci. 2020, 111, 1452–1460. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Eggermont, A.M.M.; Apolone, G.; Baumann, M.; Caldas, C.; Celis, J.E.; de Lorenzo, F.; Ernberg, I.; Ringborg, U.; Rowell, J.; Tabernero, J.; et al. Cancer Core Europe: A translational research infrastructure for a European mission on cancer. Mol. Oncol. 2019, 13, 521–527. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lablans, M.; Schmidt, E.E.; Ückert, F. An Architecture for Translational Cancer Research As Exemplified by the German Cancer Consortium. JCO Clin. Cancer Inform. 2018, 2, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Fleurence, R.L.; Curtis, L.H.; Califf, R.M.; Platt, R.; Selby, J.V.; Brown, J.S. Launching PCORnet, a national patient-centered clinical research network. J. Am. Med. Inform. Assoc. 2014, 21, 578–582. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Garza, M.; Del Fiol, G.; Tenenbaum, J.; Walden, A.; Zozus, M.N. Evaluating common data models for use with a longitudinal community registry. J. Biomed. Inform. 2016, 64, 333–341. [Google Scholar] [CrossRef] [PubMed]
- OHDSI. The Book of OHDSI; Observational Health Data Sciences and Informatics. 2021. Available online: https://ohdsi.github.io/TheBookOfOhdsi/ (accessed on 15 January 2022).
- Reps, J.M.; Schuemie, M.J.; Suchard, M.A.; Ryan, P.B.; Rijnbeek, P.R. Design and implementation of a standardized framework to generate and evaluate patient-level prediction models using observational healthcare data. J. Am. Med. Inform. Assoc. 2018, 25, 969–975. [Google Scholar] [CrossRef] [PubMed]
- Belenkaya, R.; Gurley, M.J.; Golozar, A.; Dymshyts, D.; Miller, R.T.; Williams, A.E.; Ratwani, S.; Siapos, A.; Korsik, V.; Warner, J.; et al. Extending the OMOP Common Data Model and Standardized Vocabularies to Support Observational Cancer Research. JCO Clin. Cancer Inform. 2021, 5, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Therasse, P.; Arbuck, S.G.; Eisenhauer, E.A.; Wanders, J.; Kaplan, R.S.; Rubinstein, L.; Verweij, J.; Van Glabbeke, M.; van Oosterom, A.T.; Christian, M.C. New guidelines to evaluate the response to treatment in solid tumors. J. Natl. Cancer Inst. 2000, 92, 205–216. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Altmann, U.; Katz, F.R.; Tafazzoli, A.G.; Haeberlin, V.; Dudeck, J. GTDS--a tool for tumor registries to support shared patient care. In Proceedings of the Proc AMIA Annu Fall Symp, Washington, DC, USA, 30 October 1996; pp. 512–516. [Google Scholar]
- OncoRegimenFinder. Available online: https://github.com/OHDSI/OncologyWG/tree/master/OncoRegimenFinder (accessed on 15 January 2022).
- HemOnc. Available online: https://hemonc.org/wiki/Main_Page (accessed on 15 January 2022).
- Fritz, A.; Percy, C.; Jack, A.; Shanmugaratnam, K.; Sobin, L.; Parkin, D.M.; Whelan, S. International Classification of Diseases for Oncology; World Health Organization: Geneva, Switzerland, 2000. [Google Scholar]
- NAACCR. Standards for Cancer Registries Volume II. Data Standards and Data Dictionary. Available online: http://datadictionary.naaccr.org/default.aspx?c=1&Version=21 (accessed on 24 February 2022).
- NIH. National Library of Medicine. In Mission. Available online: https://www.nih.gov/about-nih/what-we-do/nih-almanac/national-library-medicine-nlm (accessed on 24 February 2022).
- Stearns, M.; Price, C.; Spackman, K.; Wang, A.Y. SNOMED clinical terms: Overview of the development process and project status. In Proceedings of the AMIA Annual Symposium, Washington, DC, USA, 3–7 November 2001; pp. 662–666. [Google Scholar]
- Warner, J.L.; Cowan, A.J.; Hall, A.C.; Yang, P.C. HemOnc.org: A Collaborative Online Knowledge Platform for Oncology Professionals. J. Oncol. Pract. 2015, 11, e336–e350. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Warner, J.L.; Dymshyts, D.; Reich, C.G.; Gurley, M.J.; Hochheiser, H.; Moldwin, Z.H.; Belenkaya, R.; Williams, A.E.; Yang, P.C. HemOnc: A new standard vocabulary for chemotherapy regimen representation in the OMOP common data model. J. Biomed. Inform. 2019, 96, 103239. [Google Scholar] [CrossRef] [PubMed]
| Domain | Vocabulary | Version | Items |
|---|---|---|---|
| Underlying Observation Events | SNOMED-CT | 31 July 2020 SNOMED CT International Edition; 1 August 2020 SNOMED CT US Edition; 28 October 2020 SNOMED CT UK Edition | ECOG, Histology |
| NAACCR | NAACCR v18 | Primary Site, histology, behavior | |
| Underlying Measurement Events | Cancer Modifier | Cancer Modifier 20201014 | topography, metastasis–topography, grading, lymph nodes, other classification (Gleason score, Fuhrman, WHO-ISUP, Durie and Salmon, Clark level, Masaoka staging) |
| NAACCR | NAACCR v18 | Tumor board, regional nodes, metastasis, pathological grade, c/p TNM, c/p stage group, residual classification, Her2 | |
| SNOMED-CT | 31 July 2020 SNOMED CT International Edition; 1 August 2020 SNOMED CT US Edition; 28 October 2020 SNOMED CT UK Edition | morphology, Ann Arbor Classification, estregone/progesterone Receptor, tumor size (mm) | |
| Underlying Diagnosis Events | ICDO-3 | ICDO3 SEER Site/Histology Released 06/2020 | Diagnosis |
| Abstracted Episodes | HemOnc | 26 January 2021 HemOnc | Treatment Regimen |
| OPS | OPS Version 2020 | Cancer Surgeries | |
| ATC | 4 May 2020 ATC | Drugs | |
| SNOMED-CT | 31 July 2020 SNOMED CT International Edition; 1 August 2020 SNOMED CT US Edition; 28 October 2020 SNOMED CT UK Edition | Radiotherapy |
| Relationship_ID | n | Percent (≈%) |
|---|---|---|
| Maps to | 463,880 | 17 |
| Mapped from | 422,273 | 15 |
| Is a | 214,233 | 8 |
| Variable to Schema | 212,210 | 8 |
| Has Answer | 160,424 | 6 |
| Has parent item | 134,531 | 5 |
| Has start date | 134,531 | 5 |
| Subsumes | 105,450 | 4 |
| Has method | 87,313 | 3 |
| Concept same_as from | 58,737 | 2 |
| Domain | Vocabulary | Concept Class | Distinct Relations | n (With Linked Concepts) | n (Without Linked Concepts) | n (Source System: GTDS) | Mapping (%) | Mapping (%)/Domain |
|---|---|---|---|---|---|---|---|---|
| Condition | ICDO3 | ICDO Condition | Maps to, Mapped from, Is a, ICDO to Schema, ICDO to Proc Schema, Has variant, Has Topography ICDO, Has Histology ICDO, Has finding Site, Has asso morph, Concept replaces, Concept replaced by | 210,354 | 28,322 | 28,541 | 99.2 | 84.2 |
| SNOMED | Clinical Finding | Is a, Mapped from, Maps to | 13,074 | 5419 | 7828 | 69.2 | ||
| Measurement | NAACR | NAACCR Variable | Has Answer, Has parent item, Has start date, Variable to Schema, Maps to, Mapped from | 807,186 | 147,145 | 219,660 | 67.0 | 71.1 |
| Cancer Modifier | Dimension | Maps to, Mapped from | 1694 | 865 | 1247 | 69.4 | ||
| Metastasis | Maps to, Mapped from | 26,522 | 13,861 | 14,273 | 97.1 | |||
| Nodes | Maps to, Mapped from | 3958 | 2033 | 7208 | 28.2 | |||
| Staging/Grading | Maps to, Mapped from | 67,572 | 35,414 | 56,679 | 62.5 | |||
| Topography | Maps to, Mapped from | 53,744 | 27,595 | 29,856 | 92.4 | |||
| SNOMED-CT | Staging/Scales | Is a, Mapped from, Subsumes, Maps to | 3828 | 991 | 991 | 100 | ||
| Procedure | Maps to, Mapped from, Has component, Value mapped from, Has method, Is a | 31,154 | 5941 | 6140 | 96.8 | |||
| Observable Entity | Is a, Subsumes, Maps to, Mapped from | 334 | 1247 | 1235 | 26.8 | |||
| Observation | NAACCR | NAACCR Variable | Variable to Schema, Mapped from, Maps to, Parent item of, Has Answer | 207,144 | 85,614 | 86,442 | 99.0 | 63.0 |
| SNOMED-CT | Morph Abnormality | Asso morph of, Maps to, Mapped from, Subsumes, Concept same_as from, Concept replaces, Is a, Concept poss_eq from | 154,358 | 30,444 | 32,022 | 95.1 | ||
| Clinical Finding | Maps to, Has interprets, Mapped from, Has interpretation, Is a | 54,523 | 24,748 | 80,320 | 30.8 | |||
| Drug | ATC | ATC 5th | Drug class of drug, Is a, Maps to, ATC—RxNorm pr lat, ATC—SNOMED eq, ATC—RxNorm | 140,182 | 26,972 | 29,205 | 92.4 | 92.4 |
| Procedure | SNOMED | Procedure | Concept replaces, Due to of, Occurs before, Has indir proc site, Maps to, Follows, Mapped frim, Value mapped from, Has surgical appr, Has access, Has temp finding, Interprets of, Has method, Has revision status, Has dir device, Has proc morph, Concept poss_eq from, Asso proc of, Focus of, Has dir porph, Has dir subst, Has proc site, Asso with finding, Using device, Has indir morph, Has complication, Has proc device, Using subst, Has intent, Has priority, Concept was_a from, Has focus, Using acc device, Subsumes, Is a, Has dir proc site, Using energy, Ha route of admin, Specimen proc of, Comoncept same_as from, Has property | 851,813 | 116 458 | 118,039 | 98.7 | 98.7 |
| Episode | HemOnc | Regimen | Is a, Mapped from, Has antineopl Rx, Has modality, Maps to, Has accepted use, Has antineoplastic, Has context, Is historical in, Has supportive med, Is current in, Concept replaces, Has support med Rx, Has local therapy, Has immunosuppr Rx, Has local therap Rx, Has immunosuppressor | 137,512 | 11,474 | 25,714 | 44.6 | 44.6 |
| Total | 2,765,942 | 545,784 | 739,204 | 73.8 | 73.8 | |||
| Domain | Events per Episode | Total Events per Episode |
|---|---|---|
| Measurement | 8.23 | 820,013 |
| Procedure | 3.29 | 477,966 |
| Observation | 4.79 | 399,128 |
| Drug | 2.94 | 238,664 |
| Condition | 1.16 | 121,044 |
| System | N | Events | Median | Standard Error | 0.95 Lower CL | 0.95 Upper CL |
|---|---|---|---|---|---|---|
| CDM | 3588 | 784 | 164.2 | 0.02 | 155.0 | 175.9 |
| GTDS | 3644 | 806 | 162.6 | 0.02 | 155.0 | 172.1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Carus, J.; Nürnberg, S.; Ückert, F.; Schlüter, C.; Bartels, S. Mapping Cancer Registry Data to the Episode Domain of the Observational Medical Outcomes Partnership Model (OMOP). Appl. Sci. 2022, 12, 4010. https://doi.org/10.3390/app12084010
Carus J, Nürnberg S, Ückert F, Schlüter C, Bartels S. Mapping Cancer Registry Data to the Episode Domain of the Observational Medical Outcomes Partnership Model (OMOP). Applied Sciences. 2022; 12(8):4010. https://doi.org/10.3390/app12084010
Chicago/Turabian StyleCarus, Jasmin, Sylvia Nürnberg, Frank Ückert, Catarina Schlüter, and Stefan Bartels. 2022. "Mapping Cancer Registry Data to the Episode Domain of the Observational Medical Outcomes Partnership Model (OMOP)" Applied Sciences 12, no. 8: 4010. https://doi.org/10.3390/app12084010
APA StyleCarus, J., Nürnberg, S., Ückert, F., Schlüter, C., & Bartels, S. (2022). Mapping Cancer Registry Data to the Episode Domain of the Observational Medical Outcomes Partnership Model (OMOP). Applied Sciences, 12(8), 4010. https://doi.org/10.3390/app12084010






