GraphMS: Drug Target Prediction Using Graph Representation Learning with Substructures
Abstract
Featured Application
Abstract
1. Introduction
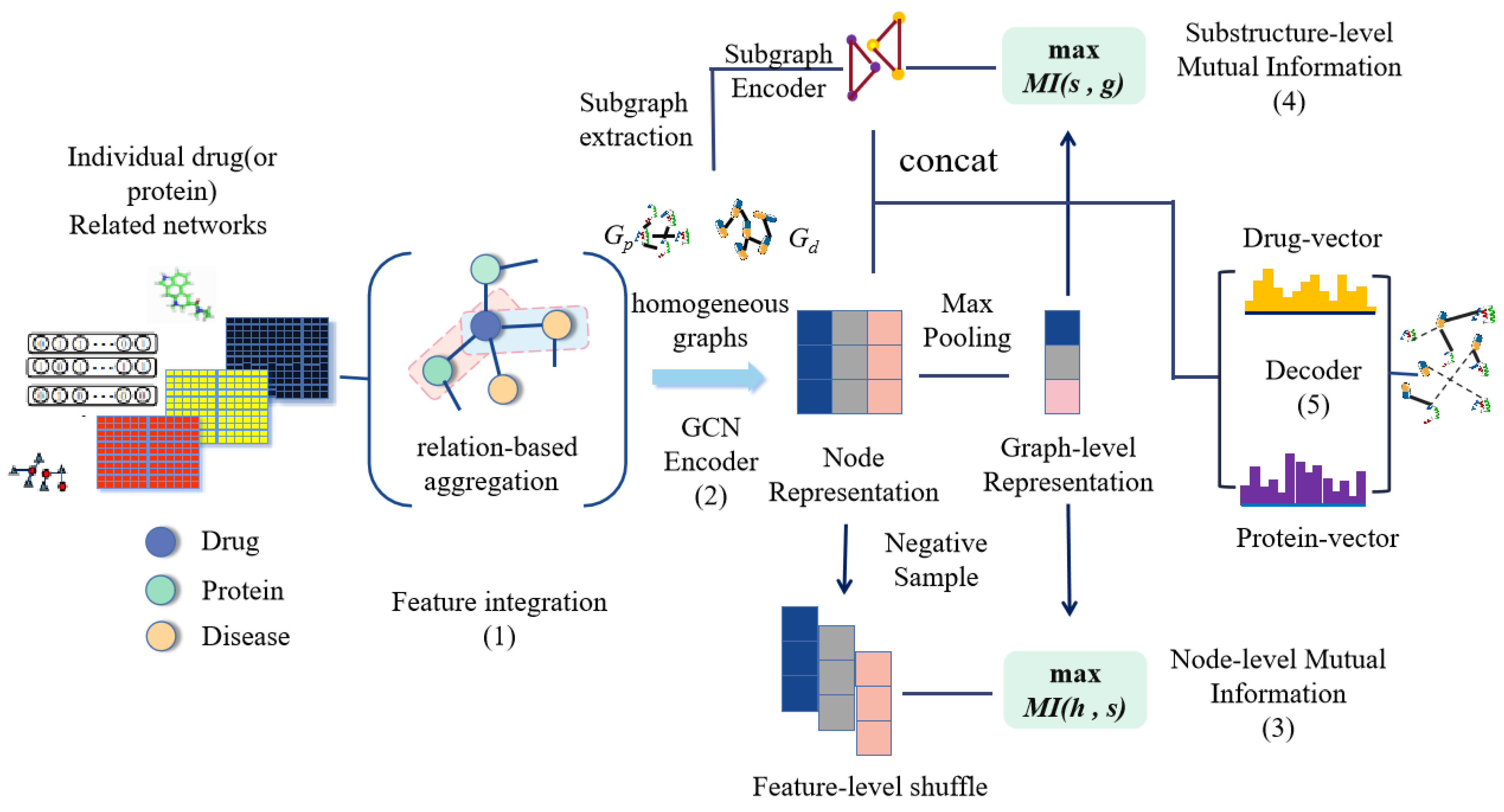
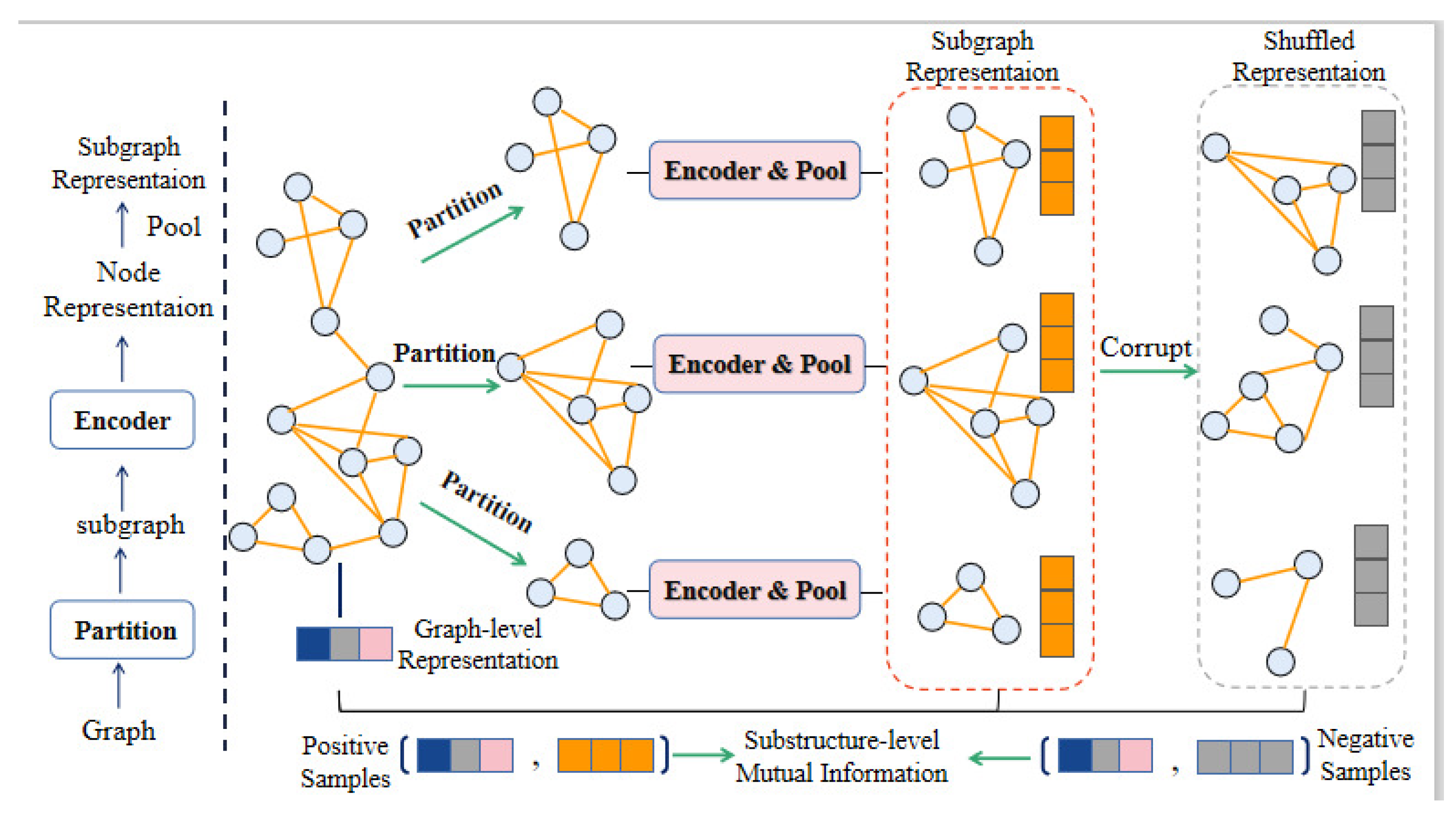
- We apply the substructure embedding to DTI prediction, and remove certain noise in the graph network. The subgraph comparison strengthens the correlation between graph-level representation and subgraph representation to capture substructure information;
- We maximize the mutual information of node representation and graph–level representation. This allows the graph–level representation to contain more information about the node itself, and it will be more concentrated on the representative nodes in the embedded representation;
- Case study and comparison method experiments also show that our model is effective.
2. Related Work
2.1. DTI Prediction
2.2. Graph Representation Learning
3. Our Approach
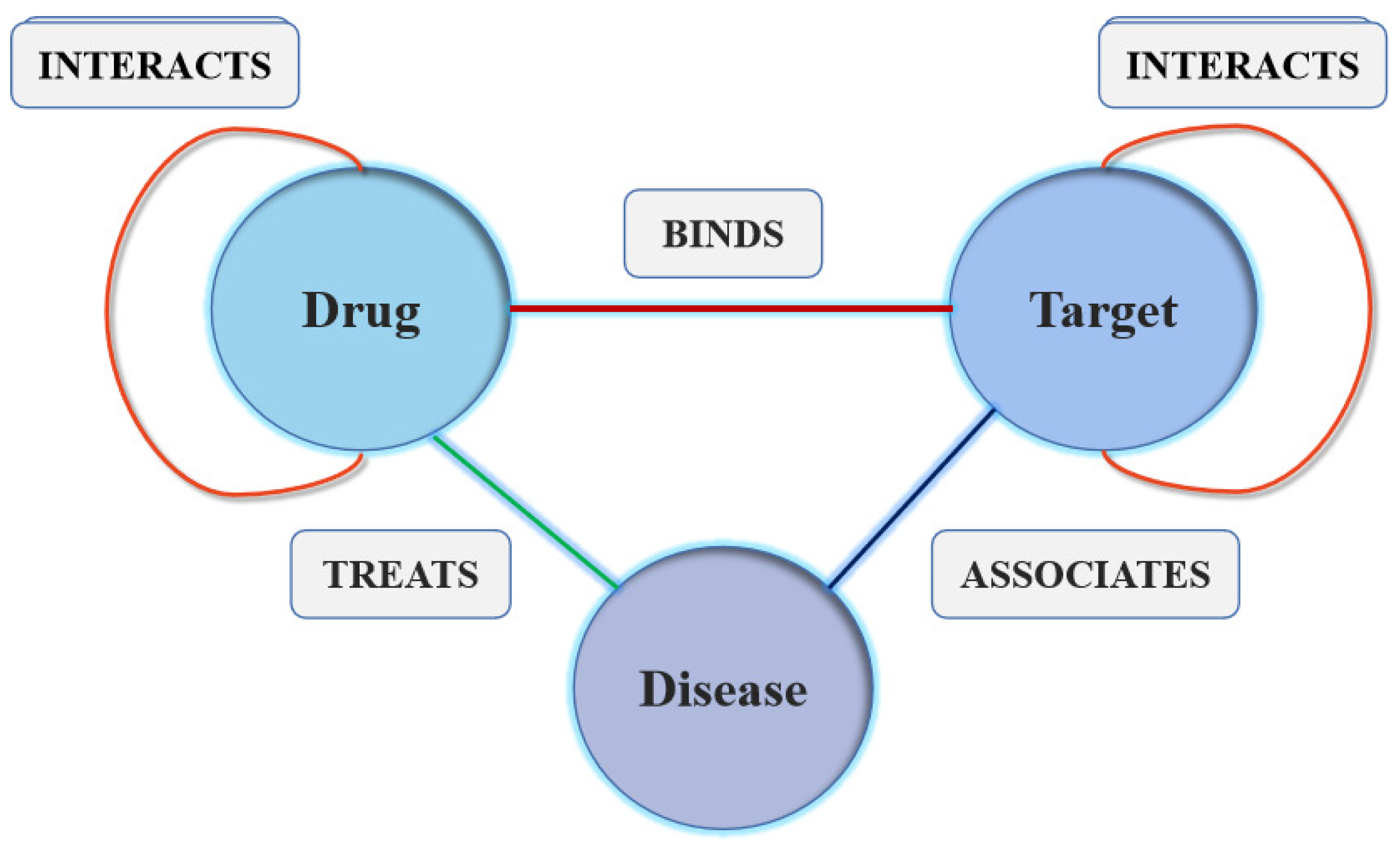
3.1. Problem Formulation
| Algorithm 1 GraphMS |
| Input: A Heterogeneous Graph |
| Output: Drug–Protein Reconstruction Matrix M |
| 1: Perform feature integration on nodes whose node type according to Equation (1); |
| 2: Select the relationship type and convert to homogeneous graphs ,; |
| 3: while not Convergence do |
| 4: Shuffle X on row dimension to obtain ; |
| 5: for node type do |
| 6: ; //Obtain Node-level Representation (Positive) |
| 7: ; //Obtain Node-level Representation (Negative) |
| 8: ; //Obtain Graph-level Representation |
| 9: Partition , nodes into k subgraphs , , ⋯, by METIS separately; |
| 10: for all each subgraph do |
| 11: Form the subgraph with nodes and edges into ; |
| 12: Shuffle other nodes except nodes of the current subgraph and select k nodes randomly to obtain the corresponding negative subgraph ; |
| 13: ; //Obtain Substructure-level Representation (Positive) |
| 14: //Obtain Substructure-level Representation (Negative); |
| 15: end for |
| 16: ; |
| 17: end for |
| 18: ; |
| 19: Compute the final loss and update parameters according to Equation (14); |
| 20: ; |
| 21: end while |
| 22: return M |
3.2. Information Fusion on Heterogeneous Graph
3.3. GCN Encoder
3.4. Mutual Information between Node–Level and Graph–Level Representation
3.5. Mutual Information Between Graph–Level Representation and Substructure Representation
3.6. Automatic Decoder for Prediction
4. Experiments and Results
4.1. Datasets
4.2. Experimental Settings
4.3. Baselines
- NeoDTI [27] integrates the neighborhood information constructed by different data sources through a large number of information transmission and aggregation operations.
- DTINet [29] aggregates information on heterogeneous data sources, and can tolerate large amounts of noise and incompleteness by learning low–dimensional vector representations of drugs and proteins.
- LightGCN [30] simplified the design of GCN to make it more concise. This model only contains the most important part of GCN–neighborhood aggregation for collaborative filtering.
- GAT [26] proposes to use the attention mechanism to weight and sum the features of neighboring nodes. The weight of features of neighboring nodes depends entirely on the features of the nodes and is independent of the graph structure. GAT uses the attention mechanism to replace the fixed standardized operations in GCN. In essence, GAT just replaces the original GCN standardization function with a neighbor node feature aggregation function using attention weights.
4.4. Comparative Experiment
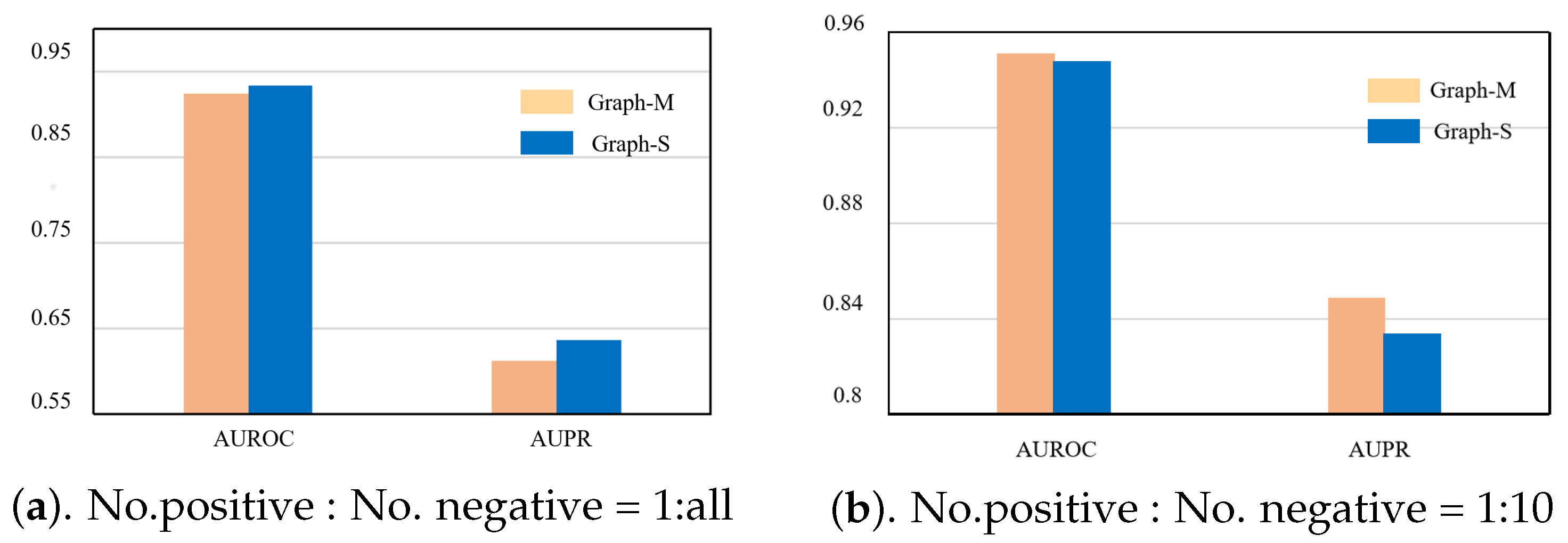
4.5. Ablation Experiment
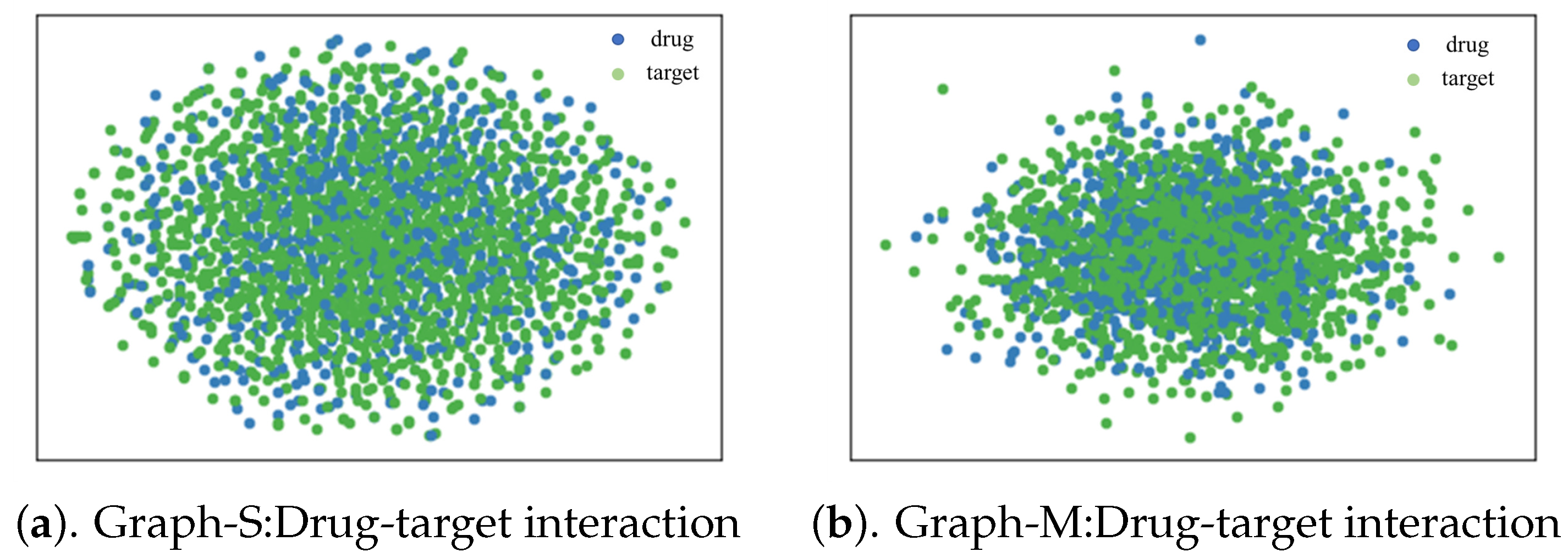
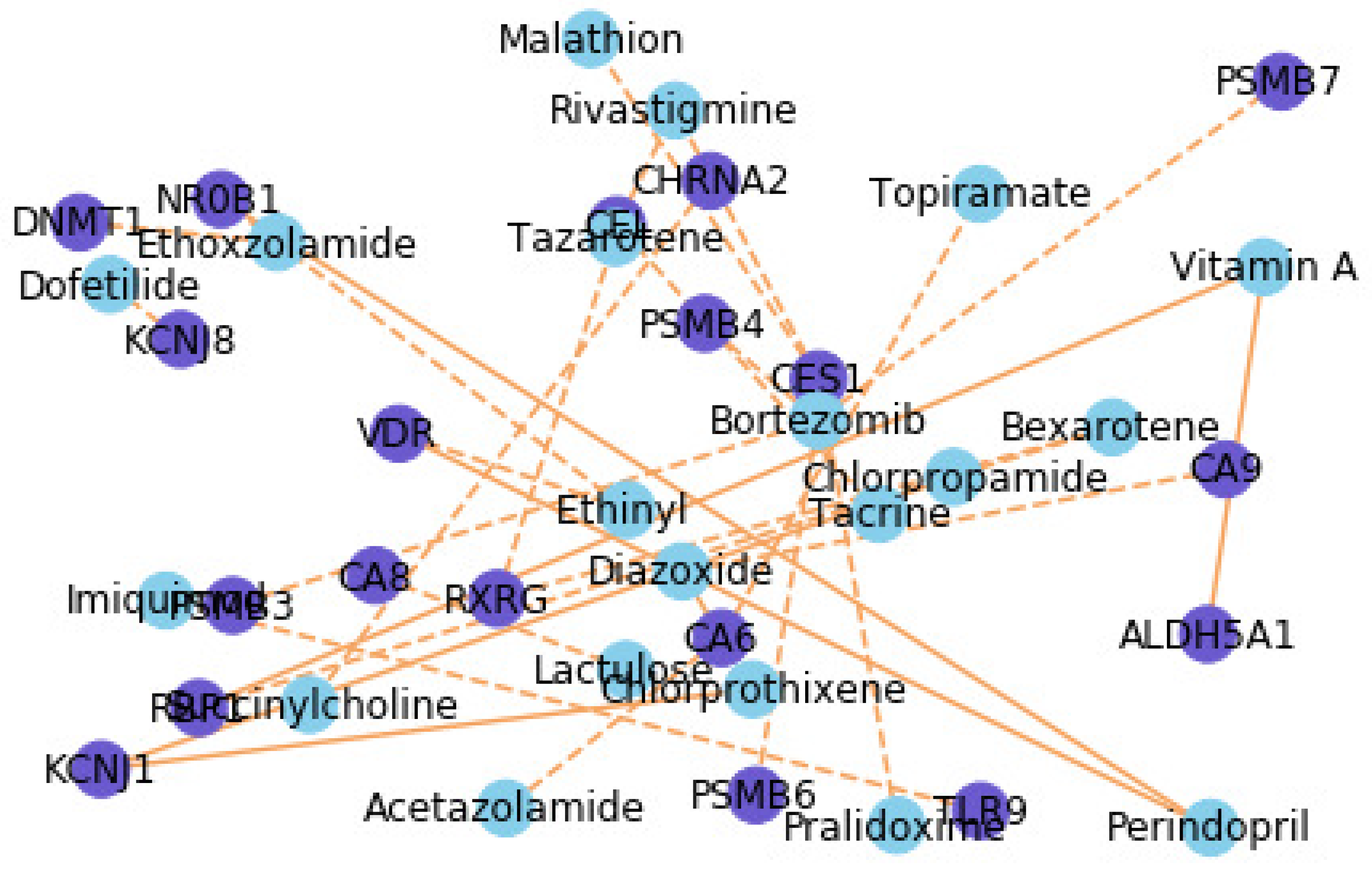
4.6. Case Study for Interpretability
5. Discussion
6. Patents
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Altae-Tran, H.; Ramsundar, B.; Pappu, A.S.; Pande, V. Low data drug discovery with one-shot learning. ACS Cent. Sci. 2017, 3, 283–293. [Google Scholar] [CrossRef]
- Bleakley, K.; Yamanishi, Y. Supervised prediction of drug–target interactions using bipartite local models. Bioinformatics 2009, 25, 2397–2403. [Google Scholar] [CrossRef]
- Ali, F.; El-Sappagh, S.; Islam, S.; Ali, A.; Attique, M.; Imran, M.; Kwak, K. An intelligent healthcare monitoring framework using wearable sensors and social networking data. Future Gener. Comput. Syst. 2021, 114, 23–43. [Google Scholar] [CrossRef]
- Xie, Y.; Yao, C.; Gong, M.; Chen, C.; Qin, A. Graph convolutional networks with multi-level coarsening for graph classification. Knowl. Based Syst. 2020, 194, 105578. [Google Scholar] [CrossRef]
- Ali, F.; El-Sappagh, S.; Islam, S.; Kwak, D.; Ali, A.; Imran, M.; Kwak, K. A smart healthcare monitoring system for heart disease prediction based on ensemble deep learning and feature fusion. Inf. Fusion 2020, 63, 208–222. [Google Scholar] [CrossRef]
- Zhao, T.; Hu, Y.; Valsdottir, L.R.; Zang, T.; Peng, J. Identifying drug-target interactions based on graph convolutional network and deep neural network. Briefings Bioinform. 2020, 22, 2141–2150. [Google Scholar] [CrossRef] [PubMed]
- Ashburner, M.; Ball, C.A.; Blake, J.A.; Botstein, D.; Butler, H.; Cherry, J.M.; Davis, A.P.; Dolinski, K.; Dwight, S.S.; Eppig, J.T.; et al. Gene ontology: Tool for the unification of biology. Nat. Genet. 2000, 25, 25–29. [Google Scholar] [CrossRef]
- Kipf, T.N.; Welling, M. Semi-supervised classification with graph convolutional networks. arXiv 2016, arXiv:1609.02907. [Google Scholar]
- Cao, S.; Lu, W.; Xu, Q. Grarep: Learning graph representations with global structural information. In Proceedings of the 24th ACM International on Conference on Information and Knowledge Management, Melbourne, VIC, Australia, 19–23 October 2015; pp. 891–900. [Google Scholar]
- Chiang, W.L.; Liu, X.; Si, S.; Li, Y.; Bengio, S.; Hsieh, C.J. Cluster-gcn: An efficient algorithm for training deep and large graph convolutional networks. In Proceedings of the 25th ACM SIGKDD International Conference on Knowledge Discovery & Data Mining, Anchorage, AK, USA, 4–8 August 2019; pp. 257–266. [Google Scholar]
- Jiao, Y.; Xiong, Y.; Zhang, J.; Zhang, Y.; Zhang, T.; Zhu, Y. Sub-graph Contrast for Scalable Self-Supervised Graph Representation Learning. arXiv 2020, arXiv:2009.10273. [Google Scholar]
- Velickovic, P.; Fedus, W.; Hamilton, W.L.; Liò, P.; Bengio, Y.; Hjelm, R.D. Deep Graph Infomax. arXiv 2019, arXiv:1809.10341. [Google Scholar]
- Park, C.; Han, J.; Yu, H. Deep multiplex graph infomax: Attentive multiplex network embedding using global information. Knowl. Based Syst. 2020, 197, 105861. [Google Scholar] [CrossRef]
- Zhu, Y.; Che, C.; Jin, B.; Zhang, N.; Su, C.; Wang, F. Knowledge-driven drug repurposing using a comprehensive drug knowledge graph. Health Inform. J. 2020, 26, 2737–2750. [Google Scholar] [CrossRef] [PubMed]
- Wan, F.; Zeng, J.M. Deep learning with feature embedding for compound-protein interaction prediction. bioRxiv 2016, 086033. [Google Scholar] [CrossRef]
- Faulon, J.L.; Misra, M.; Martin, S.; Sale, K.; Sapra, R. Genome scale enzyme–metabolite and drug–target interaction predictions using the signature molecular descriptor. Bioinformatics 2008, 24, 225–233. [Google Scholar] [CrossRef]
- Mei, J.P.; Kwoh, C.K.; Yang, P.; Li, X.L.; Zheng, J. Drug–target interaction prediction by learning from local information and neighbors. Bioinformatics 2013, 29, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Wen, M.; Zhang, Z.; Niu, S.; Sha, H.; Yang, R.; Yun, Y.; Lu, H. Deep-learning-based drug–target interaction prediction. J. Proteome Res. 2017, 16, 1401–1409. [Google Scholar] [CrossRef]
- Hu, P.W.; Chan, K.C.; You, Z.H. Large-scale prediction of drug-target interactions from deep representations. In Proceedings of the 2016 International Joint Conference on Neural Networks (IJCNN), Vancouver, BC, Canada, 24–29 July 2016; pp. 1236–1243. [Google Scholar]
- Gao, K.Y.; Fokoue, A.; Luo, H.; Iyengar, A.; Dey, S.; Zhang, P. Interpretable Drug Target Prediction Using Deep Neural Representation. IJCAI 2018, 2018, 3371–3377. [Google Scholar]
- Duvenaud, D.; Maclaurin, D.; Aguilera-Iparraguirre, J.; Gómez-Bombarelli, R.; Hirzel, T.; Aspuru-Guzik, A.; Adams, R.P. Convolutional networks on graphs for learning molecular fingerprints. arXiv 2015, arXiv:1509.09292. [Google Scholar]
- Che, M.; Yao, K.; Che, C.; Cao, Z.; Kong, F. Knowledge-Graph-Based Drug Repositioning against COVID-19 by Graph Convolutional Network with Attention Mechanism. Future Internet 2021, 13, 13. [Google Scholar] [CrossRef]
- Perozzi, B.; Al-Rfou, R.; Skiena, S. Deepwalk: Online learning of social representations. In Proceedings of the 20th ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, New York, NY, USA, 24–27 August 2014; pp. 701–710. [Google Scholar]
- Grover, A.; Leskovec, J. node2vec: Scalable feature learning for networks. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 855–864. [Google Scholar]
- Tang, J.; Qu, M.; Wang, M.; Zhang, M.; Yan, J.; Mei, Q. Line: Large-scale information network embedding. In Proceedings of the 24th International Conference on World Wide Web, Florence, Italy, 18–22 May 2015; pp. 1067–1077. [Google Scholar]
- Veličković, P.; Cucurull, G.; Casanova, A.; Romero, A.; Lio, P.; Bengio, Y. Graph attention networks. arXiv 2017, arXiv:1710.10903. [Google Scholar]
- Wan, F.; Hong, L.; Xiao, A.; Jiang, T.; Zeng, J. NeoDTI: Neural integration of neighbor information from a heterogeneous network for discovering new drug–target interactions. Bioinformatics 2019, 35, 104–111. [Google Scholar] [CrossRef] [PubMed]
- Karypis, G.; Kumar, V. A Fast and High Quality Multilevel Scheme for Partitioning Irregular Graphs. SIAM J. Sci. Comput. 1998, 20, 359–392. [Google Scholar] [CrossRef]
- Luo, Y.; Zhao, X.; Zhou, J.; Yang, J.; Zhang, Y.; Kuang, W.; Peng, J.; Chen, L.; Zeng, J. A network integration approach for drug-target interaction prediction and computational drug repositioning from heterogeneous information. Nat. Commun. 2017, 8, 1–13. [Google Scholar] [CrossRef] [PubMed]
- He, X.; Deng, K.; Wang, X.; Li, Y.; Zhang, Y.; Wang, M. Lightgcn: Simplifying and powering graph convolution network for recommendation. In Proceedings of the 43rd International ACM SIGIR Conference on Research and Development in Information Retrieval, Xi’an, China, 25–30 July 2020; pp. 639–648. [Google Scholar]
- Takeshima, H.; Niwa, T.; Yamashita, S.; Takamura-Enya, T.; Iida, N.; Wakabayashi, M.; Nanjo, S.; Abe, M.; Sugiyama, T.; Kim, Y.J.; et al. TET repression and increased DNMT activity synergistically induce aberrant DNA methylation. J. Clin. Investig. 2020, 130, 10. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Tikhomirova, A.; Modak, J.K.; Hutton, M.L.; Supuran, C.T.; Roujeinikova, A. Antibacterial activity of ethoxzolamide against Helicobacter pylori strains SS1 and 26695. Gut Pathog. 2020, 12, 1–7. [Google Scholar] [CrossRef]
Short Biography of Authors
 | Bo Jin Professor, PhD supervisor, innovative talents in colleges and universities in Liaoning Province, special consultant of Baidu Research Institute, outstanding member of CCF of China Computer Society, senior member of American ACM and IEEE. He graduated from Dalian University of Technology with a Ph.D., and visited Rutgers, the State University of New Jersey in the United States under the tutelage of Professor Xiong Hui, an authoritative scholar in the field of international big data. Served as Chair at ICDM, a top conference in the field of data mining for two consecutive years, and only two domestic experts hold relevant positions each year. The main research direction is deep learning, big data mining, artificial intelligence, and is committed to the analysis and mining methods of multi-source heterogeneous networked and serialized data. |

| Node Type | Drug | Protein | Disease |
|---|---|---|---|
| Number | 708 | 1512 | 5603 |
| Edge Type | Number |
|---|---|
| Drug-Protein | 1923 |
| Drug-Disease | 199,214 |
| Protein-Disease | 1,596,745 |
| Drug-Drug | 10,036 |
| Protein-Protein | 7363 |
| Model | Multi-View | Single-View | ||
|---|---|---|---|---|
| 1:10 | 1:all | 1:10 | 1:all | |
| GraphMS | 0.959 ± 0.002 | 0.943 ± 0.001 | 0.933 ± 0.003 | 0.914 ± 0.002 |
| LightGCN | 0.940 ± 0.002 | 0.929 ± 0.001 | 0.922 ± 0.001 | 0.895 ± 0.002 |
| GAT | 0.937 ± 0.001 | 0.927 ± 0.001 | 0.920 ± 0.001 | 0.893 ± 0.001 |
| NeoDTI | 0.929 ± 0.003 | 0.919 ± 0.002 | 0.908 ± 0.001 | 0.880 ± 0.001 |
| DTINet | 0.896 ± 0.001 | 0.862 ± 0.002 | 0.872 ± 0.001 | 0.867 ± 0.001 |
| Model | Multi-View | Single-View | ||
|---|---|---|---|---|
| 1:10 | 1:all | 1:10 | 1:all | |
| GraphMS | 0.847 ± 0.002 | 0.622 ± 0.001 | 0.760 ± 0.002 | 0.594 ± 0.002 |
| LightGCN | 0.834 ± 0.001 | 0.608 ± 0.001 | 0.734 ± 0.001 | 0.582 ± 0.001 |
| GAT | 0.832 ± 0.002 | 0.608 ± 0.001 | 0.731 ± 0.001 | 0.581 ± 0.001 |
| NeoDTI | 0.815 ± 0.003 | 0.587 ± 0.001 | 0.714 ± 0.001 | 0.559 ± 0.002 |
| DTINet | 0.743 ± 0.001 | 0.452 ± 0.001 | 0.693 ± 0.002 | 0.313 ± 0.001 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, S.; Zhang, L.; Jin, B.; Zhang, Q.; Lu, X.; You, M.; Tian, X. GraphMS: Drug Target Prediction Using Graph Representation Learning with Substructures. Appl. Sci. 2021, 11, 3239. https://doi.org/10.3390/app11073239
Cheng S, Zhang L, Jin B, Zhang Q, Lu X, You M, Tian X. GraphMS: Drug Target Prediction Using Graph Representation Learning with Substructures. Applied Sciences. 2021; 11(7):3239. https://doi.org/10.3390/app11073239
Chicago/Turabian StyleCheng, Shicheng, Liang Zhang, Bo Jin, Qiang Zhang, Xinjiang Lu, Mao You, and Xueqing Tian. 2021. "GraphMS: Drug Target Prediction Using Graph Representation Learning with Substructures" Applied Sciences 11, no. 7: 3239. https://doi.org/10.3390/app11073239
APA StyleCheng, S., Zhang, L., Jin, B., Zhang, Q., Lu, X., You, M., & Tian, X. (2021). GraphMS: Drug Target Prediction Using Graph Representation Learning with Substructures. Applied Sciences, 11(7), 3239. https://doi.org/10.3390/app11073239








