TickPhone App: A Smartphone Application for Rapid Tick Identification Using Deep Learning
Abstract
:1. Introduction
2. Methods
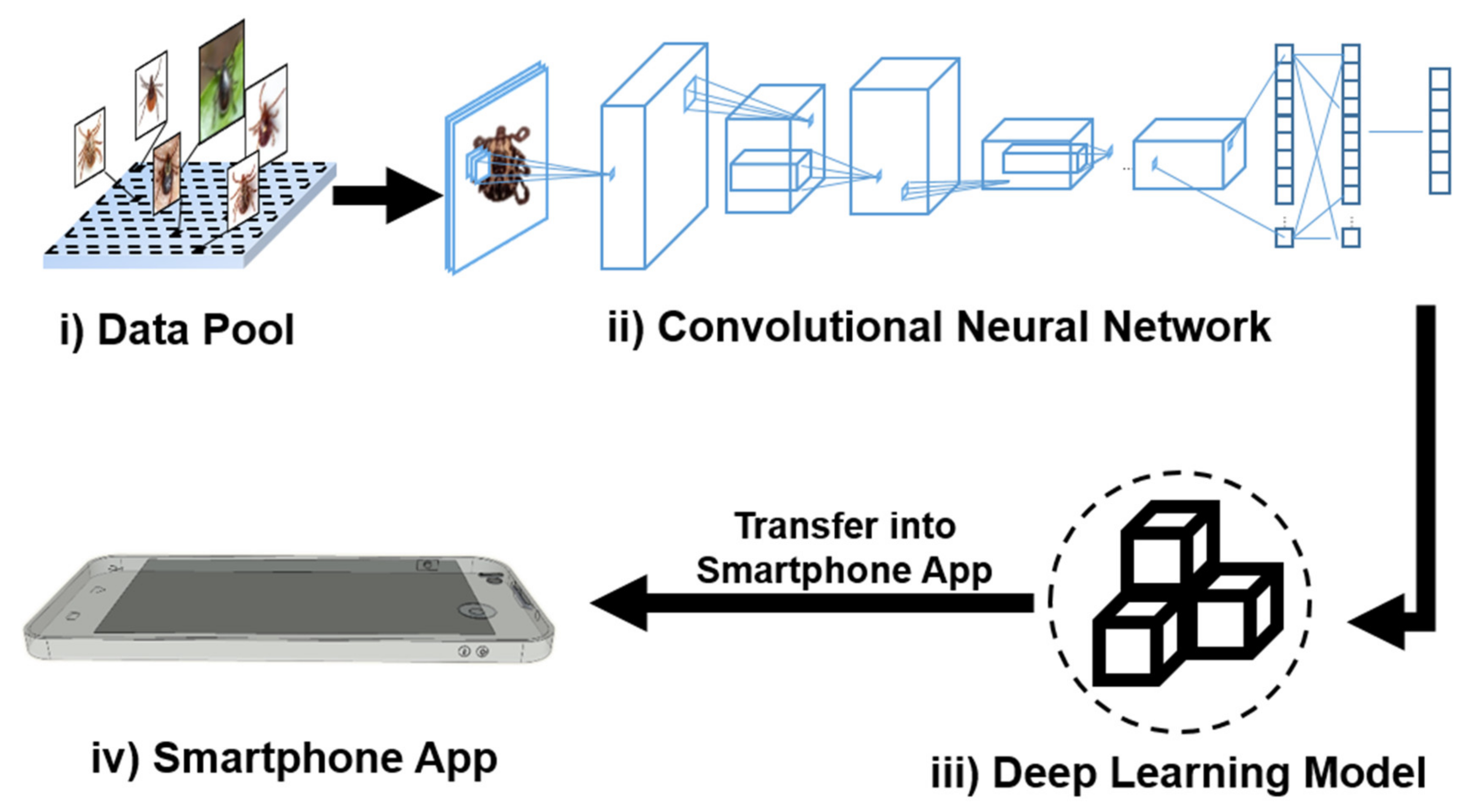
2.1. Development Process of TickPhone App
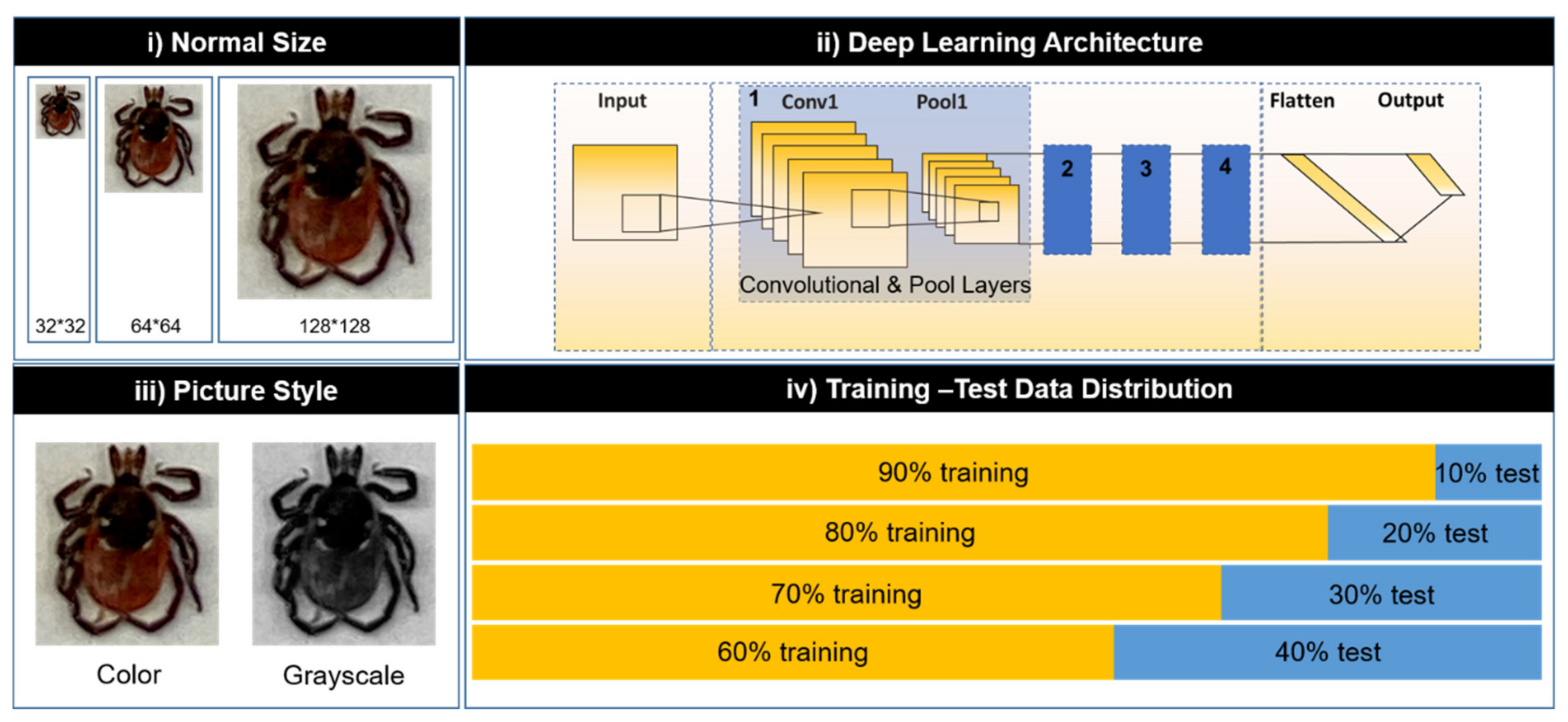
2.2. Dataset Description
2.3. Deep Learning Model Optimization
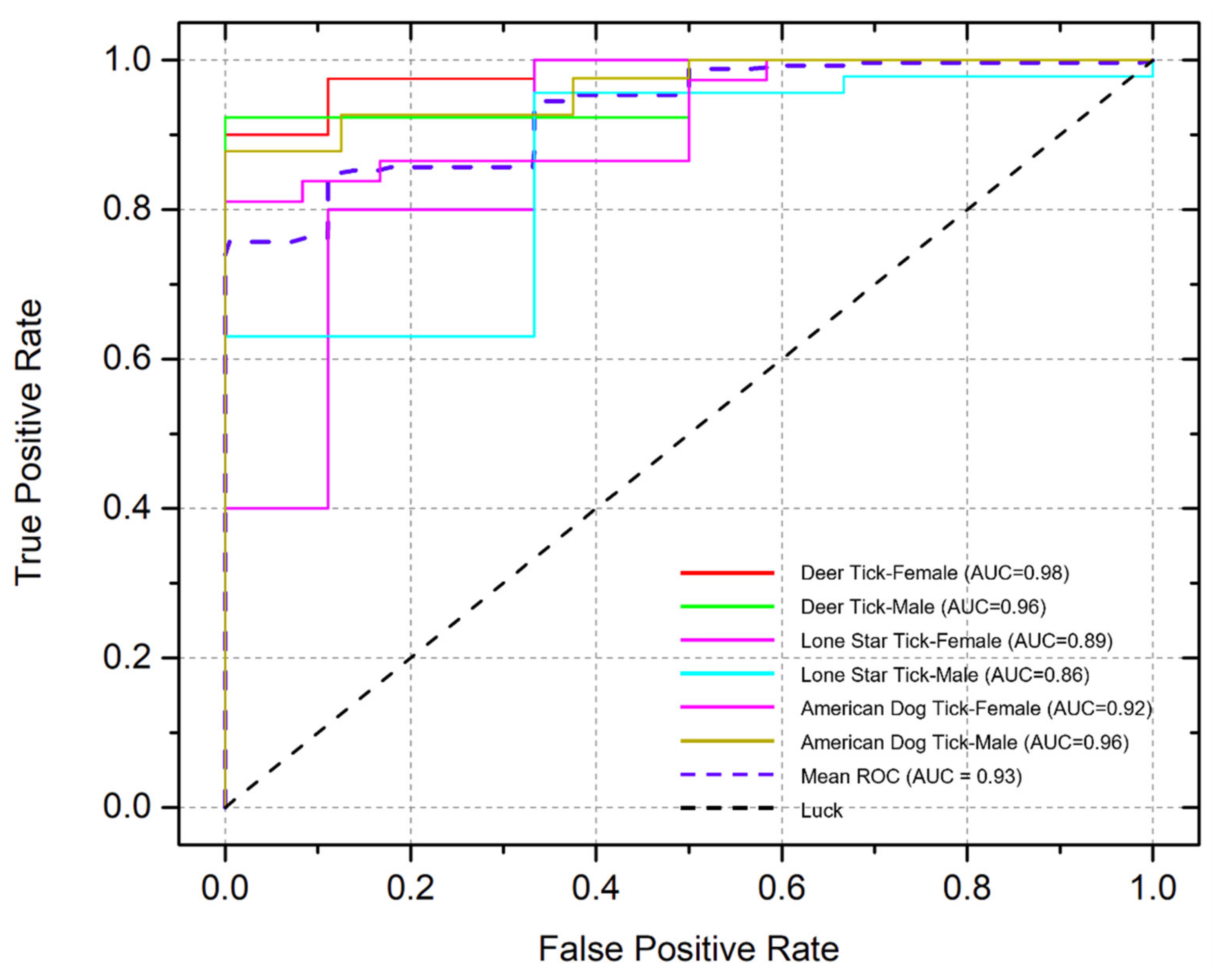
2.4. Deep Learning Model Evaluation
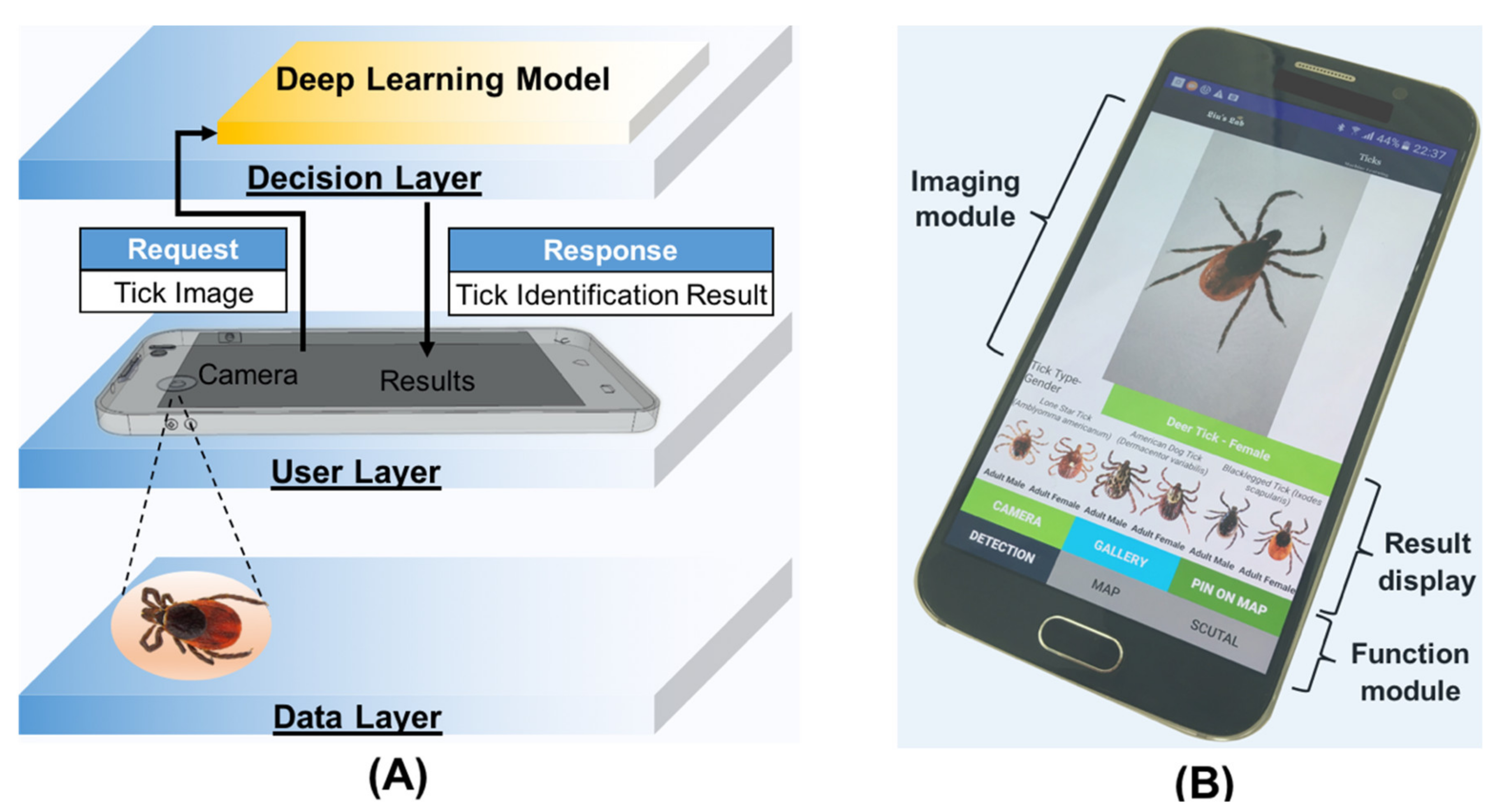
2.5. TickPhone App Design
3. Results and Discussion
3.1. Deep Learning Network
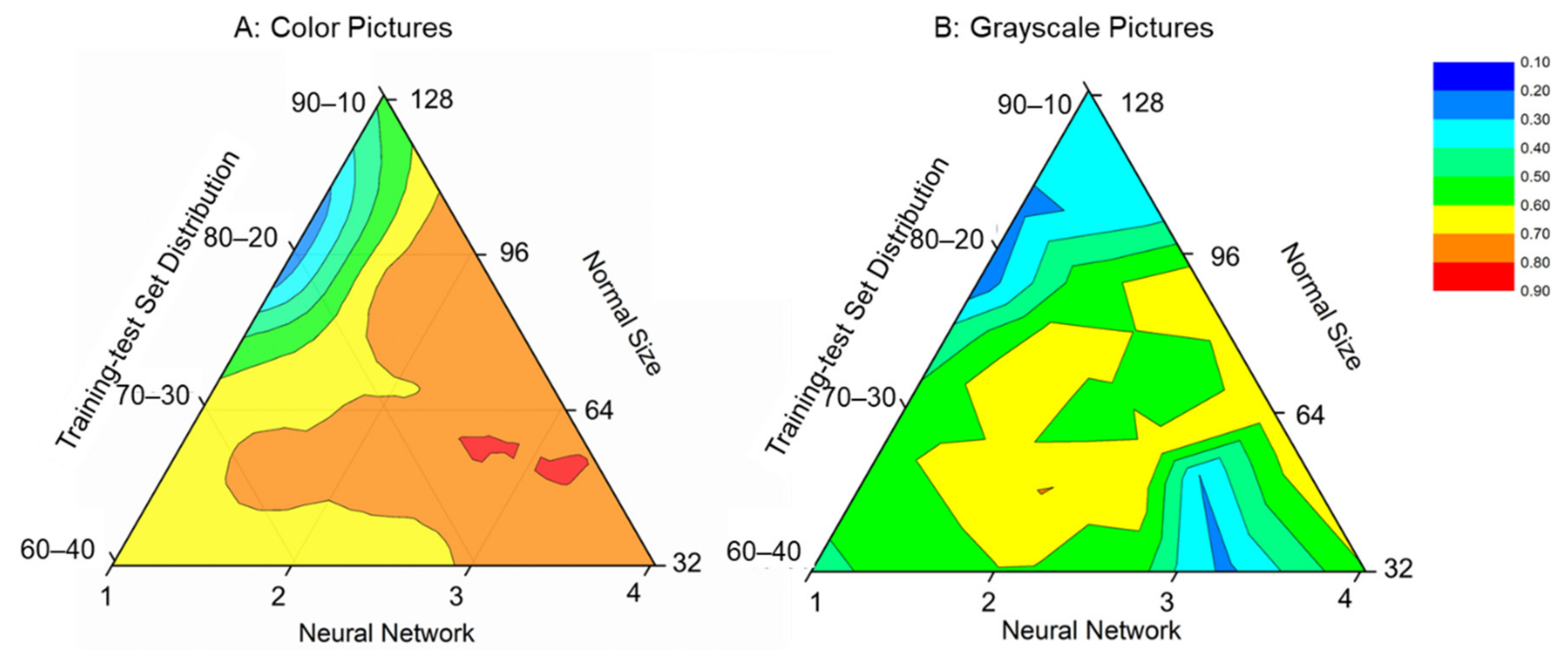
3.2. Parameter Optimization of Deep Learning Model
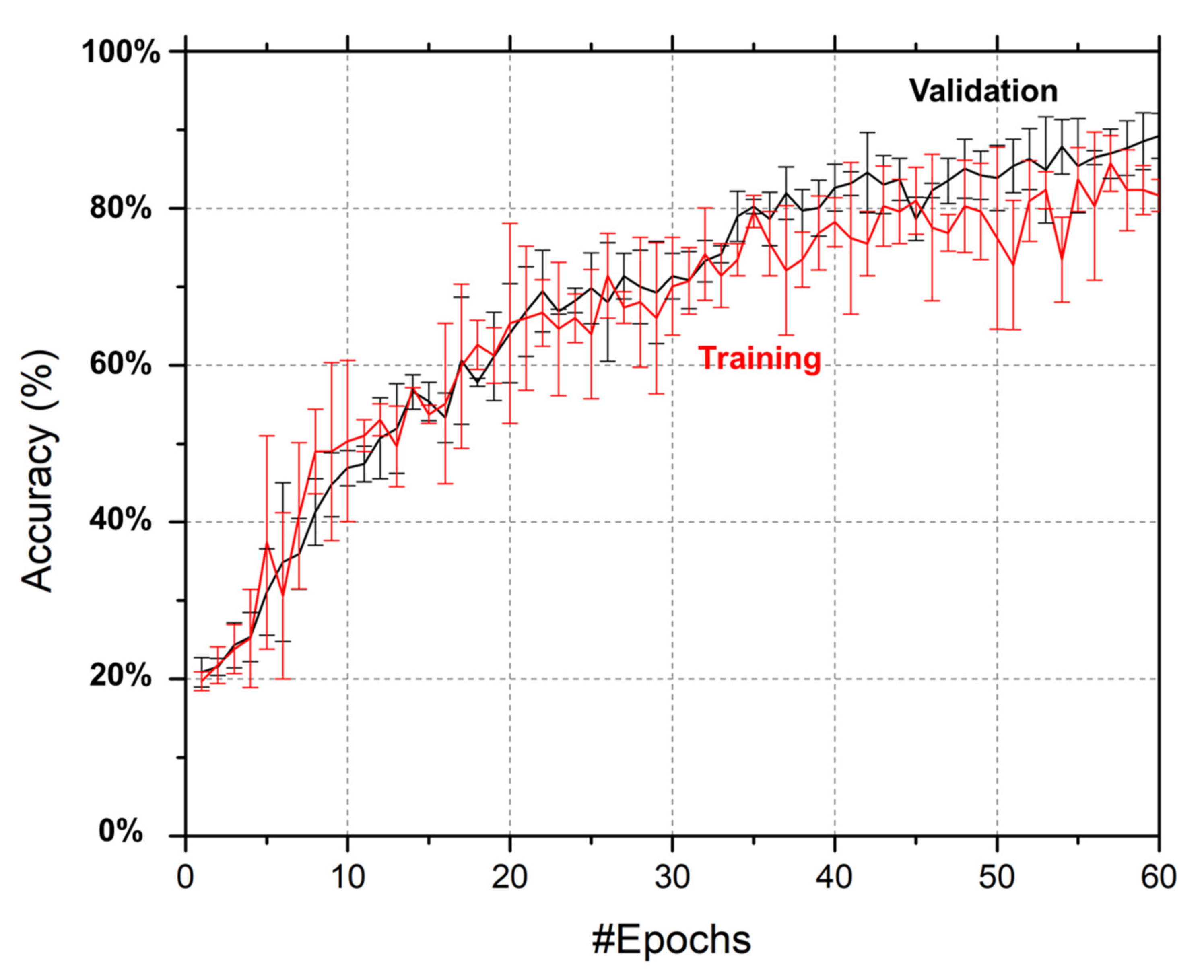
3.3. Training and Validation Accuracy
3.4. Tick Identification with TickPhone App
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Sonenshine, D.E. Range Expansion of Tick Disease Vectors in North America: Implications for Spread of Tick-Borne Disease. Int. J. Environ. Res. Public Health 2018, 15, 478. [Google Scholar] [CrossRef] [Green Version]
- Eisen, R.J.; Kugeler, K.; Eisen, L.; Beard, C.B.; Paddock, C.D. Tick-Borne Zoonoses in the United States: Persistent and Emerging Threats to Human Health. ILAR J. 2017, 58, 319–335. [Google Scholar] [CrossRef] [Green Version]
- Hook, S.A.; Nelson, C.A.; Mead, P.S. U.S. public’s experience with ticks and tick-borne diseases: Results from national HealthStyles surveys. Ticks Tick-Borne Dis. 2015, 6, 483–488. [Google Scholar] [CrossRef]
- Michelet, L.; Joncour, G.; Devillers, E.; Torina, A.; Vayssier-Taussat, M.; Bonnet, S.I.; Moutailler, S. Tick species, tick-borne pathogens and symbionts in an insular environment off the coast of Western France. Ticks Tick-Borne Dis. 2016, 7, 1109–1115. [Google Scholar] [CrossRef] [PubMed]
- Davies, S.; Abdullah, S.; Helps, C.; Tasker, S.; Newbury, H.; Wall, R. Prevalence of ticks and tick-borne pathogens: Babesia and Borrelia species in ticks infesting cats of Great Britain. Vet. Parasitol. 2017, 244, 129–135. [Google Scholar] [CrossRef] [Green Version]
- Estrada-Peña, A.; Roura, X.; Sainz, A.; Miró, G.; Solano-Gallego, L. Species of ticks and carried pathogens in owned dogs in Spain: Results of a one-year national survey. Ticks Tick-Borne Dis. 2017, 8, 443–452. [Google Scholar] [CrossRef] [PubMed]
- Centers for Disease Control and Prevention. Lyme Disease Data Tables: Historical Data Reported Cases of Lyme Disease by State or Locality, 2007–2017. Available online: https://www.cdc.gov/lyme/stats/tables.html?CDC_AA_refVal=https%3A%2F%2Fwww.cdc.gov%2Flyme%2Fstats%2Fchartstables%2Freportedcases_statelocality.html (accessed on 8 August 2020).
- Adrion, E.R.; Aucott, J.; Lemke, K.W.; Weiner, J.P. Health Care Costs, Utilization and Patterns of Care following Lyme Disease. PLoS ONE 2015, 10, e0116767. [Google Scholar] [CrossRef]
- Nadelman, R.B.; Nowakowski, J.; Fish, D.; Falco, R.C.; Freeman, K.; McKenna, D.; Welch, P.; Marcus, R.; Agüero-Rosenfeld, M.E.; Dennis, D.T.; et al. Prophylaxis with Single-Dose Doxycycline for the Prevention of Lyme Disease after anIxodes scapularisTick Bite. N. Engl. J. Med. 2001, 345, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Delaney, M. Treating Lyme disease: When will science catch up? Pharm. J. 2016, 14, 34. [Google Scholar] [CrossRef]
- Marques, A.R. Laboratory Diagnosis of Lyme Disease: Advances and Challenges. Infect. Dis. Clin. N. Am. 2015, 29, 295–307. [Google Scholar] [CrossRef] [Green Version]
- Nicholson, M.C.; Mather, T.N. Methods for Evaluating Lyme Disease Risks Using Geographic Information Systems and Geospatial Analysis. J. Med. Entomol. 1996, 33, 711–720. [Google Scholar] [CrossRef]
- Ostfeld, R.S.; Levi, T.; Keesing, F.; Oggenfuss, K.; Canham, C.D. Tick-borne disease risk in a forest food web. Ecology 2018, 99, 1562–1573. [Google Scholar] [CrossRef]
- University of Rhode Island TickEncounter Resource Center. TickSpotters. Available online: https://tickencounter.org/tickspotters/submit_form (accessed on 8 August 2020).
- University of Rhode Island TickEncounter Resource Center. Tick Identification Chart. Available online: https://tickencounter.org/tick_identification (accessed on 8 August 2020).
- Butler, A.D.; Carlson, M.L.; Nelson, C.A. Use of a tick-borne disease manual increases accuracy of tick identification among primary care providers in Lyme disease endemic areas. Ticks Tick-Borne Dis. 2017, 8, 262–265. [Google Scholar] [CrossRef]
- Falco, R.C.; Fish, D.; D’Amico, V. Accuracy of tick identification in a Lyme disease endemic area. J. Am. Med. Assoc. 1998, 280, 602–603. [Google Scholar] [CrossRef] [PubMed]
- Koffi, J.K.; Savage, J.; Thivierge, K.; Lindsay, L.R.; Bouchard, C.; Pelcat, Y.; Ogden, N.H. Evaluating the submission of digital images as a method of surveillance for Ixodes scapularis ticks. Parasitology 2017, 144, 877–883. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Erickson, D.; O’Dell, D.; Jiang, L.; Oncescu, V.; Gumus, A.; Lee, S.; Mancuso, M.; Mehta, S. Smartphone technology can be transformative to the deployment of lab-on-chip diagnostics. Lab Chip 2014, 14, 3159–3164. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ding, X.; Mauk, M.G.; Yin, K.; Kadimisetty, K.; Liu, C. Interfacing Pathogen Detection with Smartphones for Point-of-Care Applications. Anal. Chem. 2018, 91, 655–672. [Google Scholar] [CrossRef]
- Sun, Y.; Liang, D.; Wang, X.; Tang, X. DeepID3: Face Recognition with Very Deep Neural Networks. arXiv 2015, arXiv:1502.00873. [Google Scholar]
- Zhong, Z.; Jin, L.; Xie, Z. High performance offline handwritten Chinese character recognition using GoogLeNet and directional feature maps. In Proceedings of the 2015 13th International Conference on Document Analysis and Recognition (ICDAR), Tunis, Tunisia, 23–26 August 2015; pp. 846–850. [Google Scholar] [CrossRef] [Green Version]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2017 IEEE 2nd Information Technology, Networking, Electronic and Automation Control Conference (ITNEC), Chengdu, China, 15–17 December 2017; pp. 770–778. [Google Scholar] [CrossRef] [Green Version]
- Dourado, C.M.J.M.; Da Silva, S.P.P.; Da Nobrega, R.V.M.; Filho, P.P.R.; Muhammad, K.; De Albuquerque, V.H.C. An Open IoHT-Based Deep Learning Framework for Online Medical Image Recognition. IEEE J. Sel. Areas Commun. 2020, 39, 541–548. [Google Scholar] [CrossRef]
- Quiroz, I.; Alférez, G.H. Image recognition of Legacy blueberries in a Chilean smart farm through deep learning. Comput. Electron. Agric. 2020, 168, 105044. [Google Scholar] [CrossRef]
- Grinblat, G.L.; Uzal, L.C.; Larese, M.G.; Granitto, P. Deep learning for plant identification using vein morphological patterns. Comput. Electron. Agric. 2016, 127, 418–424. [Google Scholar] [CrossRef] [Green Version]
- Atila, Ü.; Uçar, M.; Akyol, K.; Uçar, E. Plant leaf disease classification using EfficientNet deep learning model. Ecol. Inform. 2021, 61, 101182. [Google Scholar] [CrossRef]
- Khare, S.; Gupta, N.; Srivastava, V. Optimization technique, curve fitting and machine learning used to detect Brain Tumor in MRI. In Proceedings of the IEEE International Conference on Computer Communication and Systems ICCCS14, Chennai, India, 20–21 February 2014; pp. 254–259. [Google Scholar] [CrossRef]
- Dai, X.; Spasic, I.; Meyer, B.; Chapman, S.; Andres, F. Machine Learning on Mobile: An On-device Inference App for Skin Cancer Detection. In Proceedings of the 2019 Fourth International Conference on Fog and Mobile Edge Computing (FMEC), Rome, Italy, 10–13 June 2019; pp. 301–305. [Google Scholar] [CrossRef]
- Liu, Q.; Zhang, N.; Yang, W.; Wang, S.; Cui, Z.; Chen, X.; Chen, L. A review of image recognition with deep convolutional neural network. In International Conference on Intelligent Computing; Springer: Cham, Switzerland, 2017; pp. 69–80. [Google Scholar] [CrossRef]
- Holzmann, H. Diagnosis of tick-borne encephalitis. Vaccine 2003, 21, S36–S40. [Google Scholar] [CrossRef]
- Berlin, J.; Motro, A. Database Schema Matching Using Machine Learning with Feature Selection. In International Conference on Advanced Information Systems Engineering; Springer: Berlin/Heidelberg, Germany, 2013; pp. 452–466. [Google Scholar] [CrossRef]
- Socher, R.; Huval, B.; Bhat, B.; Manning, C.D.; Ng, A.Y. Convolutional-recursive deep learning for 3D object classification. Adv. Neural Inf. Process. Syst. 2012, 25, 656–664. [Google Scholar]
- El-Sawy, A.; El-Bakry, H.; Loey, M. CNN for Handwritten Arabic Digits Recognition Based on LeNet-5. In Advances in Intelligent Systems and Computing; Springer: Cham, Switzerland, 2016; pp. 566–575. [Google Scholar] [CrossRef] [Green Version]
- Iandola, F.N.; Han, S.; Moskewicz, M.W.; Ashraf, K.; Dally, W.J.; Keutzer, K. SqueezeNet: AlexNet-level accuracy with 50× fewer parameters and <0.5MB model size. arXiv 2016, arXiv:1602.07360. [Google Scholar]
- Al-Jawfi, R. Handwriting Arabic character recognition lenet using neural network. Int. Arab. J. Inf. Technol. 2009, 6, 304–309. [Google Scholar]
- Hand, D.J.; Till, R.J. A Simple Generalisation of the Area under the ROC Curve for Multiple Class Classification Problems. Mach. Learn. 2001, 45, 171–186. [Google Scholar] [CrossRef]
- Horng, M.-H. Multi-class support vector machine for classification of the ultrasonic images of supraspinatus. Expert Syst. Appl. 2009, 36, 8124–8133. [Google Scholar] [CrossRef]
- Li, Y.; Huang, H.; Xie, Q.; Yao, L.; Chen, Q. Research on a Surface Defect Detection Algorithm Based on MobileNet-SSD. Appl. Sci. 2018, 8, 1678. [Google Scholar] [CrossRef] [Green Version]
- Xiong, D.; Huang, K.; Jiang, H.; Li, B.; Chen, S.; Jiang, X. IdleSR: Efficient Super-Resolution Network with Multi-scale IdleBlocks. In European Conference on Computer Vision; Springer: Cham, Switzerland, 2020; pp. 136–151. [Google Scholar] [CrossRef]
- Ucar, F.; Korkmaz, D. COVIDiagnosis-Net: Deep Bayes-SqueezeNet based diagnosis of the coronavirus disease 2019 (COVID-19) from X-ray images. Med. Hypotheses 2020, 140, 109761. [Google Scholar] [CrossRef]
- Kwon, H.; Yoon, H.; Park, K.-W. Multi-Targeted Backdoor: Indentifying Backdoor Attack for Multiple Deep Neural Networks. IEICE Trans. Inf. Syst. 2020, 103, 883–887. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, Z.; Ding, X.; Yin, K.; Li, Z.; Smyth, J.A.; Sims, M.B.; McGinnis, H.A.; Liu, C. TickPhone App: A Smartphone Application for Rapid Tick Identification Using Deep Learning. Appl. Sci. 2021, 11, 7355. https://doi.org/10.3390/app11167355
Xu Z, Ding X, Yin K, Li Z, Smyth JA, Sims MB, McGinnis HA, Liu C. TickPhone App: A Smartphone Application for Rapid Tick Identification Using Deep Learning. Applied Sciences. 2021; 11(16):7355. https://doi.org/10.3390/app11167355
Chicago/Turabian StyleXu, Zhiheng, Xiong Ding, Kun Yin, Ziyue Li, Joan A. Smyth, Maureen B. Sims, Holly A. McGinnis, and Changchun Liu. 2021. "TickPhone App: A Smartphone Application for Rapid Tick Identification Using Deep Learning" Applied Sciences 11, no. 16: 7355. https://doi.org/10.3390/app11167355
APA StyleXu, Z., Ding, X., Yin, K., Li, Z., Smyth, J. A., Sims, M. B., McGinnis, H. A., & Liu, C. (2021). TickPhone App: A Smartphone Application for Rapid Tick Identification Using Deep Learning. Applied Sciences, 11(16), 7355. https://doi.org/10.3390/app11167355





