Formaldehyde Exposure and Epigenetic Effects: A Systematic Review
Abstract
1. Introduction
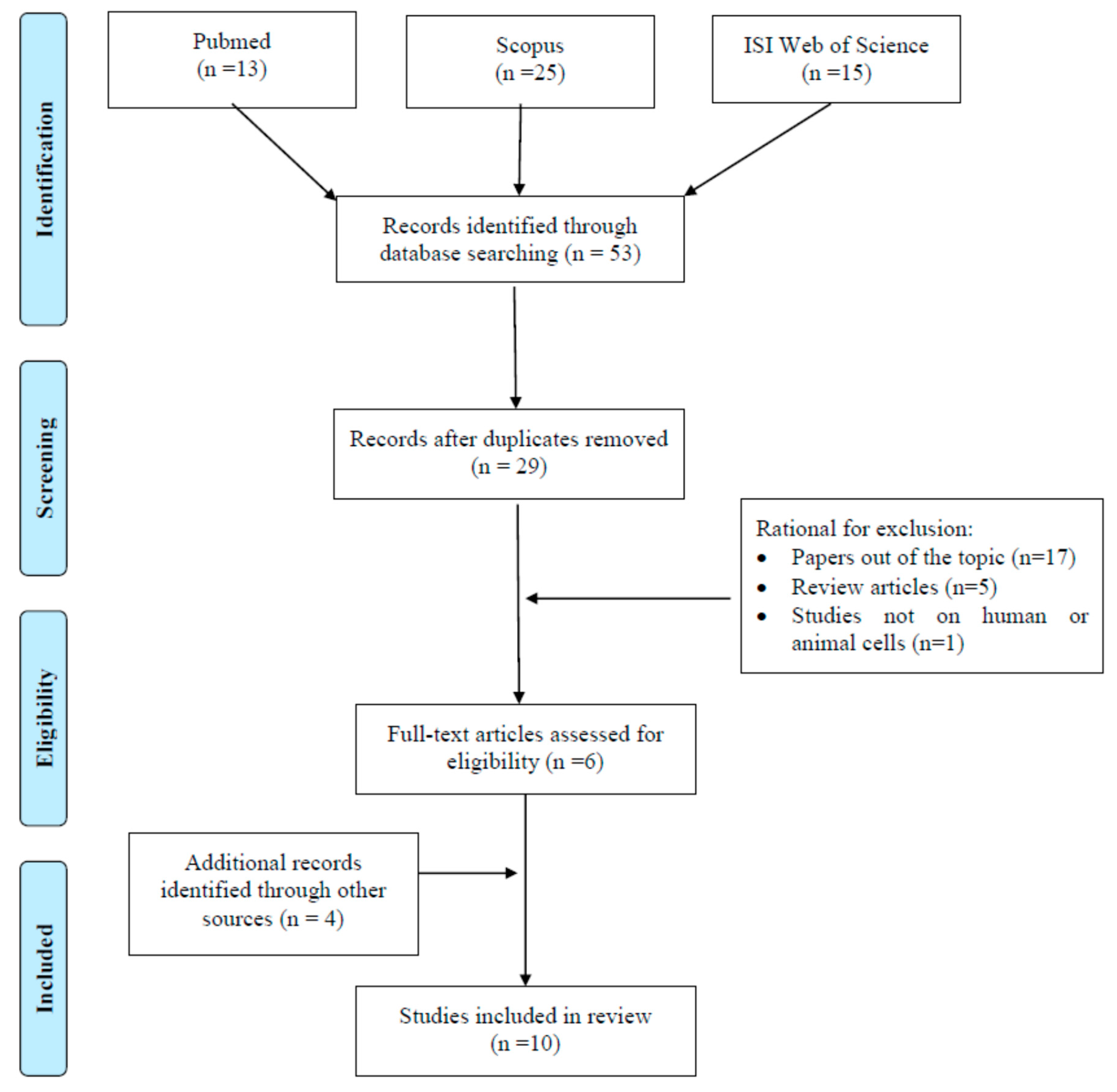
2. Materials and Methods
3. Formaldehyde-Induced Epigenetic Effects
3.1. DNA Methylation
3.2. Histone Modifications
3.3. MicroRNAs
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Salthammer, T.; Mentese, S.; Marutzky, R. Formaldehyde in the indoor environment. Chem. Rev. 2010, 110, 2536–2572. [Google Scholar] [CrossRef]
- World Health Organization (WHO). WHO guidelines for indoor air quality: Selected pollutants. WHO Guide 2010, 9, 454. [Google Scholar]
- IARC, Working Group on the Evaluation of Carcinogenic Risks to Humans Chemical Agents and Related Occupations. Formaldehyde. IARC Monogr. Eval. Carcinog. Risks Hum. 2012, 100, 9–562. [Google Scholar]
- National Toxicology Program (NTP). NTP 12th report on carcinogens. Rep. Carcinog. Carcinog. Profiles 2011, 12, iii-499. [Google Scholar]
- WHO Regional Office for Europe Copenhagen. Formaldehyde. In Air Quality Guidelines for Europe—Second Edition; 2000; chapter 5.8; pp. 87–91. Available online: http://www.euro.who.int/__data/assets/pdf_file/0005/74732/E71922.pdf (accessed on 26 March 2020).
- Godish, T. Formaldehyde exposures from tobacco smoke: A review. Am. J. Public Health 1989, 79, 1044–1045. [Google Scholar] [CrossRef] [PubMed]
- Dahl, A.R.; Hadley, W.M. Formaldehyde production promoted by rat nasal cytochrome P-450-dependent monooxygenases with nasal decongestants, essences, solvents, air pollutants, nicotine, and cocaine as substrates. Toxicol. Appl. Pharmacol. 1983, 67, 200–205. [Google Scholar] [CrossRef]
- Yang, M.; Ospina, M.; Tse, C.; Toth, S.; Caudill, S.P.; Vesper, H.W. Ultraperformance liquid chromatography tandem mass spectrometry method to determine formaldehyde hemoglobin adducts in humans as biomarker for formaldehyde exposure. Chem. Res. Toxicol. 2017, 30, 1592–1598. [Google Scholar] [CrossRef]
- Liteplo, R.G.; Meek, M.E. Inhaled formaldehyde: Exposure estimation, hazard characterization, and exposure-response analysis. J. Toxicol. Environ. Health B Crit. Rev. 2003, 6, 85–114. [Google Scholar] [CrossRef] [PubMed]
- IARC, Working Group on the Evaluation of Carcinogenic Risks to Humans Chemical Agents and Related Occupations. Formaldehyde, 2-butoxyethanol and 1-tertbutoxypropan-2-ol. IARC Monogr. Eval. Carcinog. Risks Hum. 2006, 88, 1–478. [Google Scholar]
- Schmid, O.; Speit, G. Genotoxic effects induced by formaldehyde in human blood and implications for the interpretation of biomonitoring studies. Mutagenesis 2007, 22, 69–74. [Google Scholar] [CrossRef]
- Orsière, T.; Sari-Minodier, I.; Iarmarcovai, G.; Botta, A. Genotoxic risk assessment of pathology and anatomy laboratory workers exposed to formaldehyde by use of personal air sampling and analysis of DNA damage in peripheral lymphocytes. Mutat. Res. 2006, 605, 30–41. [Google Scholar] [CrossRef] [PubMed]
- Ye, X.; Yan, W.; Xie, H.; Zhao, M.; Ying, C. Cytogenetic analysis of nasal mucosa cells and lymphocytes from high level long-term formaldehyde exposed workers and low-level short-term exposed waiters. Mutat. Res. 2005, 588, 22–27. [Google Scholar] [CrossRef] [PubMed]
- Edrissi, B.; Taghizadeh, K.; Dedon, P.C. Quantitative analysis of histone modifications: Formaldehyde is a source of pathological n-formyllysine that is refractory to histone deacetylases. PLoS Genet. 2013, 9, e1003328. [Google Scholar] [CrossRef] [PubMed]
- Edrissi, B.; Taghizadeh, K.; Moeller, B.C.; Kracko, D.; Doyle-Eisele, M.; Swenberg, J.A.; Dedon, P.C. Dosimetry of N-formyllysine adducts following [C H]-formaldehyde exposures in rats. Chem. Res. Toxicol. 2013, 26, 1421–1423. [Google Scholar] [CrossRef] [PubMed]
- Bogdanffy, M.S.; Morgan, P.H.; Starr, T.B.; Morgan, K.T. Binding of formaldehyde to human and rat nasal mucus and bovine serum albumin. Toxicol. Lett. 1987, 38, 145–154. [Google Scholar] [CrossRef]
- Metz, B.; Kersten, G.F.; Baart, G.J.; de Jong, A.; Meiring, H.; ten Hove, J.; van Steenbergen, M.J.; Hennink, W.E.; Crommelin, D.J.; Jiskoot, W. Identification of formaldehyde-induced modifications in proteins: Reactions with insulin. Bioconjugate Chem. 2006, 17, 815–822. [Google Scholar] [CrossRef]
- Guthe, K.F. The formaldehyde−hemoglobin reaction. J. Biol. Chem. 1959, 234, 3169–3173. [Google Scholar]
- Hoberman, H.D.; San George, R.C. Reaction of tobacco smoke aldehydes with human hemoglobin. J. Biochem. Toxicol. 1989, 3, 105–119. [Google Scholar] [CrossRef]
- Swenberg, J.A.; Lu, K.; Moeller, B.C.; Gao, L.; Upton, P.B.; Nakamura, J.; Starr, T.B. Endogenous versus exogenous DNA adducts: Their role in carcinogenesis, epidemiology, and risk assessment. Toxicol. Sci. 2011, 120 (Suppl. 1), S130–S145. [Google Scholar] [CrossRef]
- Tabish, A.M.; Poels, K.; Hoet, P.; Godderis, L. Epigenetic factors in cancer risk: Effect of chemical carcinogens on global DNA methylation pattern in human TK6 cells. PLoS ONE 2012, 7, e34674. [Google Scholar] [CrossRef]
- Swenberg, J.A.; Moeller, B.C.; Lu, K.; Rager, J.E.; Fry, R.C.; Starr, T.B. Formaldehyde carcinogenicity research: 30 years and counting for mode of action, epidemiology, and cancer risk assessment. Toxicol. Pathol. 2013, 41, 181–189. [Google Scholar] [CrossRef] [PubMed]
- Egger, G.; Liang, G.; Aparicio, A.; Jones, P.A. Epigenetics in human disease and prospects for epigenetic therapy. Nature 2004, 429, 457–463. [Google Scholar] [CrossRef] [PubMed]
- Arimondo, P.B.; Barberousse, A.; Pontarotti, G. The Many Faces of Epigenetics Oxford, December 2017. Epigenetics 2019, 14, 623–631. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, A. The key role of epigenetics in human disease. N. Engl. J. Med. 2018, 379, 400–401. [Google Scholar]
- Karpinets, T.V.; Foy, B.D. Tumorigenesis: The adaptation of mammalian cells to sustained stress environment by epigenetic alterations and succeeding matched mutations. Carcinogenesis 2005, 26, 1323–1334. [Google Scholar] [CrossRef]
- Pogribny, I.P.; Muskhelishvili, L.; Tryndyak, V.P.; Beland, F.A. The role of epigenetic events in genotoxic hepatocarcinogenesis induced by 2-acetylaminofluorene. Mutat. Res. 2011, 722, 106–113. [Google Scholar] [CrossRef]
- Barbosa, E.; Dos Santos, A.L.A.; Peteffi, G.P.; Schneider, A.; Müller, D.; Rovaris, D.; Bau, C.H.D.; Linden, R.; Antunes, M.V.; Charão, M.F. Increase of global DNA methylation patterns in beauty salon workers exposed to low levels of formaldehyde. Environ. Sci Pollut. Res. Int. 2019, 26, 1304–1314. [Google Scholar] [CrossRef]
- Liu, Q.; Yang, L.; Gong, C.; Tao, G.; Huang, H.; Liu, J.; Zhang, H.; Wu, D.; Xia, B.; Hu, G.; et al. Effects of long-term low-dose formaldehyde exposure on global genomic hypomethylation in 16HBE cells. Toxicol. Lett. 2011, 205, 235–240. [Google Scholar] [CrossRef]
- Yoshida, I.; Ibuki, Y. Formaldehyde-induced histone H3 phosphorylation via JNK and the expression of proto-oncogenes. Mutat. Res. 2014, 770, 9–18. [Google Scholar] [CrossRef]
- Chen, D.; Fang, L.; Mei, S.; Li, H.; Xu, X.; Des Marais, T.L.; Lu, K.; Liu, X.S.; Jin, C. Regulation of Chromatin Assembly and Cell Transformation by Formaldehyde Exposure in Human Cells. Environ. Health Perspect. 2017, 125, 097019. [Google Scholar] [CrossRef]
- Ibuki, Y.; Toyooka, T.; Zhao, X.; Yoshida, I. Cigarette sidestream smoke induces histone H3 phosphorylation via JNK and PI3K/Akt pathways, leading to the expression of proto-oncogenes. Carcinogenesis 2014, 35, 1228–1237. [Google Scholar] [CrossRef] [PubMed]
- Rager, J.E.; Smeester, L.; Jaspers, I.; Sexton, K.G.; Fry, R.C. Epigenetic changes induced by air toxics: Formaldehyde exposure alters miRNA expression profiles in human lung cells. Environ. Health Perspect. 2011, 119, 494–500. [Google Scholar] [CrossRef] [PubMed]
- Rager, J.E.; Moeller, B.C.; Doyle-Eisele, M.; Kracko, D.; Swenberg, J.A.; Fry, R.C. Formaldehyde and epigenetic alterations: microRna changes in the nasal epithelium of nonhuman primates. Environ. Health Perspect. 2013, 121, 339–344. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Rager, J.E.; Moeller, B.C.; Miller, S.K.; Kracko, D.; Doyle-Eisele, M.; Swenberg, J.A.; Fry, R.C. Formaldehyde-associated changes in microRNAs: Tissue and temporal specificity in the rat nose, white blood cells, and bone marrow. Toxicol. Sci. 2014, 138, 36–46. [Google Scholar] [CrossRef] [PubMed]
- Li, G.; Yang, J.; Ling, S. Formaldehyde exposure alters miRNA expression profiles in the olfactory bulb. Inhal. Toxicol. 2015, 27, 387–393. [Google Scholar] [CrossRef] [PubMed]
- Esteller, M. Relevance of DNA methylation in the management of cancer. Lancet Oncol. 2003, 4, 351–358. [Google Scholar] [CrossRef]
- Pierce, J.S.; Abelmann, A.; Spicer, L.J.; Adams, R.E.; Glynn, M.E.; Neier, K.; Finley, B.L.; Gaffney, S.H. Characterization of formaldehyde exposure resulting from the use of four professional hair straightening products. J. Occup. Environ. Hyg. 2011, 8, 686–699. [Google Scholar] [CrossRef]
- Norma Regulamentadora 15 (NR 15). Atividades e operações insalubres. D.O.U. Portaria MTb 1978, 3, 214. [Google Scholar]
- OSHA, Occupational Safety and Health Administration. Formaldehyde. Available online: https://www.osha.gov/OshDoc/data_General_Facts/formaldehyde-factsheet.pdf (accessed on 25 November 2019).
- ECHA, European Chemicals Agency. Worker Exposure to Formaldehyde and Formaldehyde Releasers. 2019. Available online: https://echa.europa.eu/documents/10162/13641/investigationreport_formaldehyde_workers-exposure_final_en.pdf/ac457a0c-378d-4eae-c602-c7cd59abc4c5 (accessed on 19 March 2020).
- Moore, L.D.; Le, T.; Fan, G. DNA methylation and its basic function. Neuropsychopharmacology 2013, 38, 23–38. [Google Scholar] [CrossRef]
- Chappell, G.; Pogribny, I.P.; Guyton, K.Z.; Rusyn, I. Epigenetic alterations induced by genotoxic occupational and environmental human chemical carcinogens: A systematic literature review. Mutat. Res. Rev. Mutat. Res. 2016, 768, 27–45. [Google Scholar] [CrossRef]
- Weber, D.; Heisig, J.; Kneitz, S.; Wolf, E.; Eilers, M.; Gessler, M. Mechanisms of epigenetic and cell-type specific regulation of Hey target genes in ES cells and cardiomyocytes. J. Mol. Cell. Cardiol. 2015, 79, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Lu, K.; Boysen, G.; Gao, L.; Collins, L.B.; Swenberg, J.A. Formaldehyde-induced histone modifications in vitro. Chem. Res. Toxicol. 2008, 21, 1586–1593. [Google Scholar] [CrossRef] [PubMed]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of post-transcriptional regulation by microRNAs: Are the answers in sight? Nat. Rev. Genet. 2008, 9, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Choi, P.S.; Zakhary, L.; Choi, W.Y.; Caron, S.; Alvarez-Saavedra, E.; Miska, E.A.; McManus, M.; Harfe, B.; Giraldez, A.J.; Horvitz, H.R.; et al. Members of the miRNA-200 family regulate olfactory neurogenesis. Neuron 2008, 57, 41–55. [Google Scholar] [CrossRef] [PubMed]
- Imai, T.; Sakano, H. Axon-axon interactions in neuronal circuit assembly: Lessons from olfactory map formation. Eur. J. Neurosci. 2011, 34, 1647–1654. [Google Scholar] [CrossRef] [PubMed]
- Bartke, N.; Hannun, Y.A. Bioactive sphingolipids: Metabolism and function. J. Lipid Res. 2009, 50, S91–S96. [Google Scholar] [CrossRef]
- Gilcrease, M.Z. Integrin signaling in epithelial cells. Cancer Lett. 2007, 247, 1–25. [Google Scholar] [CrossRef]
- Zhu, X.; Li, D.; Zhang, Z.; Zhu, W.; Li, W.; Zhao, J.; Xing, X.; He, Z.; Wang, S.; Wang, F.; et al. Persistent phosphorylation at specific H3 serine residues involved in chemical carcinogen-induced cell transformation. Mol. Carcinog. 2017, 56, 1449–1460. [Google Scholar] [CrossRef]
- Faucher, D.; Wellinger, R.J. Methylated H3K4, a transcription associated histone modification, is involved in the DNA damage response pathway. PLoS Genet. 2010, 6, e1001082. [Google Scholar] [CrossRef]
- Thompson, L.L.; Guppy, B.J.; Sawchuk, L.; Davie, J.R.; McManus, K.J. Regulation of chromatin structure via histone post-translational modification and the link to carcinogenesis. Cancer Metastasis Rev. 2013, 32, 363–376. [Google Scholar] [CrossRef]
- Li, J.; Gorospe, M.; Barnes, J.; Liu, Y. Tumor promoter arsenite stimulates histone H3 phosphoacetylation of proto-oncogenes c-fos and c-jun chromatin in human diploid fibroblasts. J. Biol. Chem. 2003, 278, 13183–13191. [Google Scholar] [CrossRef] [PubMed]
- Ke, Q.; Li, Q.; Ellen, T.P.; Sun, H.; Costa, M. Nickel compounds induce phosphorylation of histone H3 at serine 10 by activating JNK-MAPK pathway. Carcinogenesis 2008, 29, 1276–1281. [Google Scholar] [CrossRef] [PubMed]
- Keum, Y.S.; Kim, H.G.; Bode, A.M.; Surh, Y.J.; Dong, Z. UVB-induced COX-2 expression requires histone H3 phosphorylation at Ser10 and Ser28. Oncogene 2013, 32, 444–452. [Google Scholar] [CrossRef] [PubMed]
- He, Z.; Cho, Y.Y.; Ma, W.Y.; Choi, H.S.; Bode, A.M.; Dong, Z. Regulation of ultraviolet B-induced phosphorylation of histone H3 at serine 10 by Fyn kinase. J. Biol. Chem. 2005, 280, 2446–2454. [Google Scholar] [CrossRef]
- Banister, C.E.; Koestler, D.C.; Maccani, M.A.; Padbury, J.F.; Houseman, E.A.; Marsit, C.J. Infant growth restriction is associated with distinct patterns of DNA methylation in human placentas. Epigenetics 2011, 6, 920–927. [Google Scholar] [CrossRef]
- Ransom, M.; Dennehey, B.K.; Tyler, J.K. Chaperoning histones during DNA replication and repair. Cell 2010, 140, 183–195. [Google Scholar] [CrossRef]
- Bailey, K.A.; Fry, R.C. Environmental toxicants and perturbation of miRNA signaling. In microRNAs in Toxicology and Medicine; Sahu, S.C., Ed.; John Wiley & Sons, Ltd.: Chichester, UK, 2013. [Google Scholar]
- Cedar, H.; Bergman, Y. Linking DNA methylation and histone modification: Patterns and paradigms. Nat. Rev. Genet. 2009, 10, 295–304. [Google Scholar] [CrossRef]
| EXPERIMENTAL SETTING | EPIGENETIC OUTCOMES | EXPERIMENTAL DESIGN | MAIN RESULTS | REFERENCES |
|---|---|---|---|---|
| HUMAN INVESTIGATION | ||||
| Brazilian hairdressers, hairdresser assistants, receptionists, manicurists, and managers (n. 41F; 8M) Mean age ± SD: 31.04 ± 10.46 years Years working in beauty salons (median: 6 years) No unexposed controls were enrolled. | DNA Methylation | Airborne FA concentrations were measured through personal (samplers fitted in volunteers breathing zone) and area passive samplers (8-h work shift). Global DNA methylation was assessed in whole blood samples were collected after 8-h of work. Biological monitoring of formic acid was performed on urine samples collected at the beginning of the work shift and again 8 h after the first sampling. |
| Barbosa et al. [28] |
| IN VIVO EXPERIMENTS | ||||
| Male Fischer rats | MicroRNAs | Nose-only exposure to 2 ppm formaldehyde for 6 h/day; Four groups of exposure were considered: 7, 28, 28 days plus a 7-day recovery period, unexposed controls. Multiple samples were collected from the nasal epithelium, circulating mononuclear WBCs, and BM cells. Microarray analysis employed to assess 695 rat known miRNAs. |
| Rager et al. [34] |
| ICR male mice | MicroRNAs | Mice treated with 3 ppm FA for 6 h for either 1 or 7 consecutive days via inhalation; controls maintained in the same conditions except for FA. Olfactory bulb homogenized and RNA isolated for microarray analysis. Expression levels of miRNAs tested using quantitative RT-PCR. |
| Li et al. [35] |
| Male cynomolgus macaques (Macaca fascicularis, n. 8) | MicroRNAs | Animals exposed via inhalation to 0, 2, or 6 ppm FA for 6 hr/day for two consecutive days. Small RNAs extracted from nasal samples, collected within 3 h from the last exposure, assessed for genome-wide miRNA expression levels. Transcriptional targets of FA-affected miRNAs analyzed. Expression levels of miRNAs tested using quantitative RT-PCR. |
| Rager et al. [36] |
| IN VITRO EXPERIMENTS | ||||
| Human bronchial epithelial (16HBE) cells | DNA methylation | Cell cultures treated with 10 mM FA for 24 h for 24 weeks. Control cells treated with serum-free MEM. BTC, a malignant transformed 16HBE cell line was used as a positive control. Genomic DNA methylation measured by: anti-5-methyl-C immuno- histochemistry assay, flow cytometric assay and HPCE assay. |
| Liu et al. [29] |
| Human lymphoblastoid cells (TK6) | DNA methylation | Cells were exposed to 3 concentrations of FA (95% cellular viability- high dose; 1/10 of high dose: medium dose; 1/100 of high dose: low dose) for 24 h with or without a mix of S9 metabolic mix required for biotransformation of procarcinogens into active carcinogens. Global DNA methylation was obtained quantifying (5Me)dC and dC using ultrapressure liquid chromatography for separation and tandem mass spectrometry for quantification. |
| Tabish et al. [21] |
| Human lung adenocarcinoma epithelial (A549) cells | Histone Modifications | Cell cultures treated with 0–1 mM FA for 10–600 min Western blotting employed to determine histone modifications. |
| Yoshida et al. [30] |
| Human bronchial epithelial BEAS-2B cells; human osteosarcoma UTA6 cells; human nasal septum quasidiploid tumor RPMI2650 cells | Histone Modifications | Cell cultures treated with 0–0.5 mM FA for 6 h. For chronic FA 0–0.1 mM for 96 h. Protein carbonyl assays employed to determine FA adducts with histone proteins Histone post-transcriptional modifications assessed by Western blot analysis. |
| Chen et al. [31] |
| Human lung adenocarcinoma epithelial (A549) cells; normal human lung fibroblasts, MRC-5 and WI-38 | Histone Modifications | Cells were treated with various concentrations of cigarette side-stream smoke (~6.25%–100%) for ~up to 8 h. Flow cytometric analysis was performed to assess phosphor-H3S10 |
| Ibuki et al. [32] |
| Human lung adenocarcinoma epithelial (A549) cells | MicroRNAs | Cell cultures treated with 1 ppm (1.2 mg/m3) gaseous FA generated by heat in 143 mg paraformaldehyde until powder completely vaporized. Microarray analysis employed to assess >500 known miRNAs. Expression levels of miRNAs also tested using quantitative RT-PCR. |
| Rager et al. [33] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Leso, V.; Macrini, M.C.; Russo, F.; Iavicoli, I. Formaldehyde Exposure and Epigenetic Effects: A Systematic Review. Appl. Sci. 2020, 10, 2319. https://doi.org/10.3390/app10072319
Leso V, Macrini MC, Russo F, Iavicoli I. Formaldehyde Exposure and Epigenetic Effects: A Systematic Review. Applied Sciences. 2020; 10(7):2319. https://doi.org/10.3390/app10072319
Chicago/Turabian StyleLeso, Veruscka, Maria Carmela Macrini, Francesco Russo, and Ivo Iavicoli. 2020. "Formaldehyde Exposure and Epigenetic Effects: A Systematic Review" Applied Sciences 10, no. 7: 2319. https://doi.org/10.3390/app10072319
APA StyleLeso, V., Macrini, M. C., Russo, F., & Iavicoli, I. (2020). Formaldehyde Exposure and Epigenetic Effects: A Systematic Review. Applied Sciences, 10(7), 2319. https://doi.org/10.3390/app10072319







