Effect and Interaction of β-Lactoglobulin, Kappa Casein, and Prolactin Genes on Milk Production and Composition of Awassi Sheep
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Animal and Sample Collection
2.2. Milk Samples and Analysis
2.3. Genomic DNA Extraction and Polymerase Chain Reaction (PCR)
2.4. Restriction Fragment Length Polymorphism (RFLP) Analysis
2.5. Sequencing Analysis
2.6. Statistical Analysis
- -
- Yijklnmop = the studied traits;
- -
- Μ = overall mean of the total milk yield or test-day milk yield;
- -
- BLGi = fixed effect of the ith genotype at β-LG locus (I = AA, AB and BB);
- -
- PRLj = fixed effect of the jth genotype at PRL locus (j = AA, AB and BB);
- -
- CSNk = fixed effect of the kth genotype at CSN3 locus (k = TT and TC);
- -
- Pl = fixed effect of the lth parity or number of lambing (l = 1, 2, 3, 4, 5 and 6);
- -
- Sm= random effect of mth sires (m = 1, 2 to 31);
- -
- SYn = fixed effect of the nth year-season of lambing (n = 2007 to 2011);
- -
- Bo = linear regression coefficient dam weight at lambing;
- -
- DWo = dam weight at lambing as covariate;
- -
- (BLG × PRL)ij = interaction between β-LG genotypes and PRL genotypes (ij = AAAA, AAAB, AABB, ABAA, ABAB, ABBB, BBAA, BBAB, and BBBB);
- -
- (BLG × CSN)ik = interaction between β-LG genotypes and CSN3 genotypes (ik = AATT, AATC, ABTT, ABTC, BBTT, and BBTC);
- -
- (PRL × CSN) jk = interaction between PRL genotypes and CSN3 genotypes (jk = AATT, AATC, ABTT, ABTC, BBTT, BBTC);
- -
- Eijklmnop = random errors with the assumption of N (0, σ2).
- -
- Yijklnmo = the studied traits;
- -
- Μ = overall mean of Fat%; protein%, SNF%, Total solids, lactose%, and density (g/cm2)
- -
- BLGi = fixed effect of the ith genotype at β-LG locus (I = AA, AB and BB);
- -
- PRLj = fixed effect of the jth genotype at PRL locus (j = AA, AB and BB);
- -
- CSNk = fixed effect of the kth genotype at CSN3 locus (k = TT and TC);
- -
- Pl = fixed effect of the lth parity or number of lambing (l = 1, 2, 3, 4, 5 and 6);
- -
- Sm = random effect of mth sires (m = 1, 2, …, 31);
- -
- Bn = linear regression coefficient TDM.
- -
- TDMn = TDM covariant.
- -
- (BLG × PRL)ij = interaction between β-LG genotypes and PRL genotypes (ij = AAAA, AAAB, AABB, ABAA, ABAB, ABBB, BBAA, BBAB, and BBBB);
- -
- (BLG × CSN)ik = interaction between β-LG genotypes and CSN3 genotypes (ik = AATT, AATC, ABTT, ABTC, BBTT, and BBTC);
- -
- (PRL × CSN)jk = interaction between PRL genotypes and CSN3 genotypes (jk = AATT, AATC, ABTT, ABTC, BBTT, and BBTC);
- -
- Eijklmno = random errors with the assumption of N (0, σ2).
3. Results
3.1. Descriptive Statistics
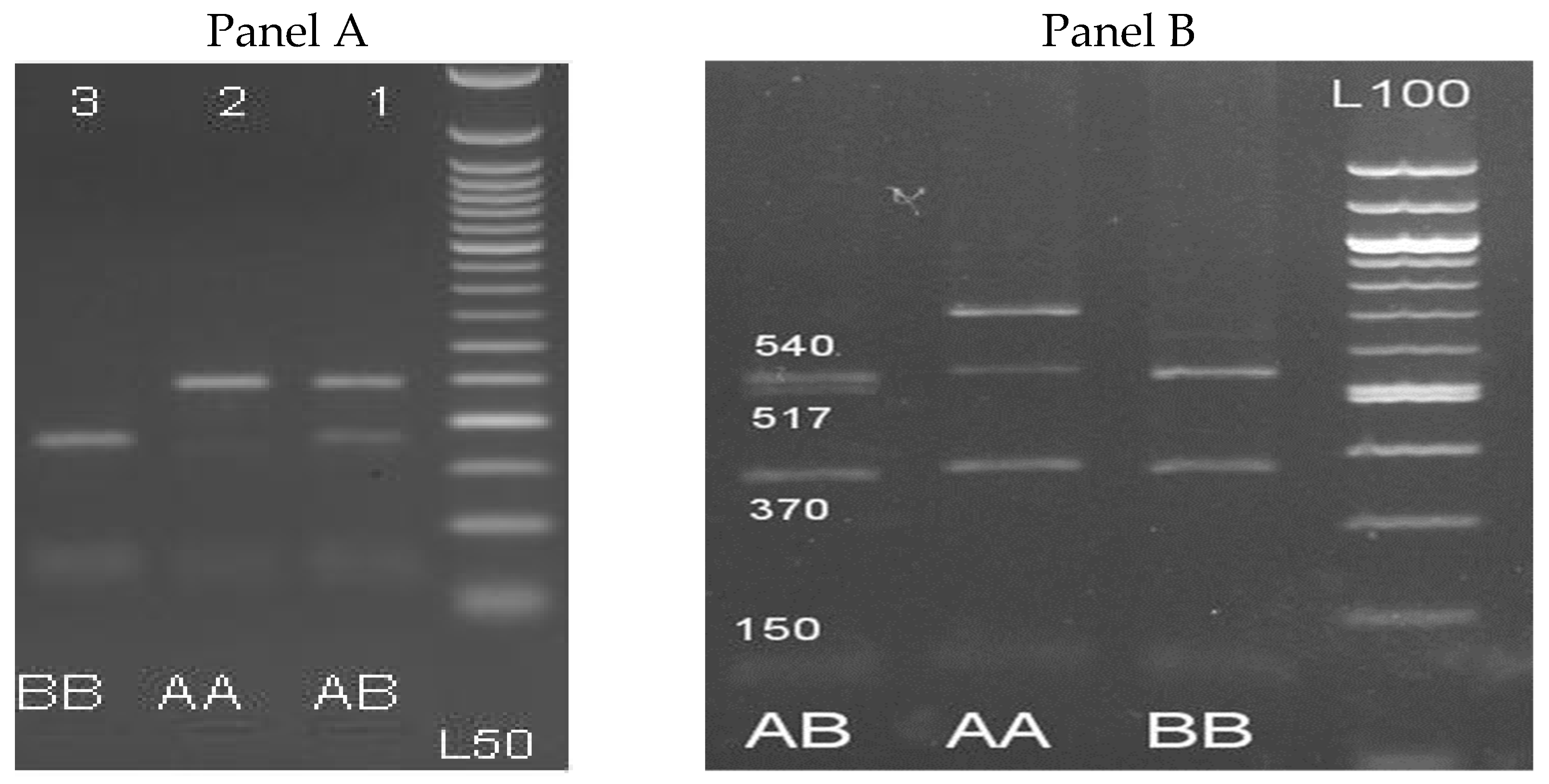
3.2. PCR-RLFP Assay of β-LG and PRL Genes
3.3. Nucleotide Sequence Analysis of the β-LG and CSN3 Gene
3.4. Genotypic and Allelic Frequencies for β-LG, PRL, and CSN3 Genes
3.5. Statistical Analysis of the Effects of β-LG, PRL, and CSN3 Genotypes on Milk Production and Component Traits
3.6. Effects of β-LG, PRL, and CSN3 Genotypes on Milk Production Traits and Their Interactions
3.7. Effects of β-LG, PRL, and CSN3 Genotypes on Milk Composition Traits and Their Interactions
4. Discussion
5. Conclusion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Carta, A.; Casu, S.; Salaris, S. Invited review: Current state of genetic improvement in dairy sheep. J. Dairy Sci. 2009, 92, 5814–5833. [Google Scholar] [CrossRef] [PubMed]
- Selvaggi, M.; Laudadio, V.; Dario, C.; Tufarelli, V. Investigating the genetic polymorphism of sheep milk proteins: A useful tool for dairy production. J. Sci. Food Agric. 2014, 94, 3090–3099. [Google Scholar] [CrossRef] [PubMed]
- Jawasreh, K.I. Quantitative and molecular genetic analysis for some traits in highly selected jordanian awassi sheep for milk production. In Proceedings of the World Congress on Genetics Applied to Livestock Production, Volume Species Breeding: Sheep and Goat Breeding for Milk, Meat and Fibre—Lecture Sessions, Leipzig, Germany, 1–6 August 2010; p. 93. [Google Scholar]
- Ahmed, M.; Abdallah, J. Comparison of milk yield and reproductive performance of sheep breeds in the West Bank, Palestine. An Najah Univ. J. Res. 2013, 27, 11–128. [Google Scholar]
- Al-Samarai, F.R.; Al-Anbari, N.N. Genetic evaluation of rams for total milk yield in Iraqi Awassi sheep. ARPN J. Agri. Biol. Sci. 2009, 4, 54–57. [Google Scholar]
- Dag, B.; Keski, I.; Mikailsoy, F. Application of different models to the lactation curves of unimproved Awassi ewes in Turkey. S. Afr. J. Anim. Sci. 2005, 35, 238–243. [Google Scholar] [CrossRef]
- Epstein, H.E. The Awassi sheep with special reference to the improved dairy type. In FAO Animal Production, Health Paper 57; FAO: Rome, Italy, 1985. [Google Scholar]
- Gootwine, E. Mini review: Breeding Awassi and Assaf sheep for diverse management conditions. Trop. Anim. Health Prod. 2011, 43, 1289–1296. [Google Scholar] [CrossRef] [PubMed]
- Iniguez, L.; Hilali, M.E.D. Evaluation of Awassi genotypes for improved milk production in Syria. Livest. Sci. 2009, 120, 232–239. [Google Scholar] [CrossRef]
- Eggen, A. The development and application of genomic selection as a new breeding paradigm. Anim. Front. 2012, 2, 10–15. [Google Scholar] [CrossRef]
- Barillet, F.; Arranz, J.J.; Carta, A. Mapping quantitative trait loci for milk production and genetic polymorphisms of milk proteins in dairy sheep. Genet. Sel. Evol. 2005, 37, 109–123. [Google Scholar] [CrossRef]
- Gras, M.; Pistol, G.; Pelmus, R.; Lazar, C.; Grosu, H.; Ghita, E. Relationship between gene polymorphism and milk production traits in Teleorman Black Head sheep breed. Rev. MVZ Córdoba 2016, 21, 5124–5136. [Google Scholar] [CrossRef]
- Ozmen, O.; Kul, S. Identification of novel SNPs of ovine PRL gene and their association with milk production traits. Russian J. Genet. 2016, 52, 977–984. [Google Scholar] [CrossRef]
- Padilla, P.; Izquierdo, M.; Martínez-Trancón, M.; Parejo, J.C.; Rabasco, A.; Salazar, J.; Padilla, J.Á. Polymorphisms of α-lactoalbumin, β-lactoglobulin and prolactin genes are highly associated with milk composition traits in Spanish Merino sheep. Livest. Sci. 2018, 217, 26–29. [Google Scholar] [CrossRef]
- Selvaggi, M.; Laudadio, V.; Dario, C.; Tufarelli, V. β-Lactoglobulin gene polymorphisms in sheep and effects on milk production traits: A review. Adv. Anim. Vet. Sci. 2015, 3, 478–484. [Google Scholar] [CrossRef]
- Staiger, E.A.; Thonney, M.L.; Buchanan, J.W.; Rogers, E.R.; Oltenacu, P.A.; Mateescu, R.G. Effect of prolactin, beta-lactoglobulin, and kappa-casein genotype on milk yield in East Friesian sheep. J. Dairy Sci. 2010, 93, 1736–1742. [Google Scholar] [CrossRef]
- Erhardt, G. Evidence for a third allele at the β-lactoglobulin (β-Lg) locus of sheep milk and its occurrence in different breeds. Anim. Genet. 1989, 20, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Moioli, B.; D’Andrea, M.; Pilla, F. Candidate genes affecting sheep and goat milk quality. Small Rumin. Res. 2007, 68, 179–192. [Google Scholar] [CrossRef]
- Triantaphyllopoulos, K.; Koutsouli, P.; Kandris, A.; Papachristou, D.; Markopoulou, K.; Mataragka, A.; Massouras, T.; Bizelis, I. Effect of β-lactoglobulin gene polymorphism, lactation stage and breed on milk traits in Chios and Karagouniko sheep breed. Ann. Anim. Sci. 2017, 17, 371–384. [Google Scholar] [CrossRef]
- Yousefi, S.; Azari, M.A.; Zerehdaran, S.; Samiee, R.; Khataminejhad, R. Effect of β-lactoglobulin and κ-casein genes polymorphism on milk composition in indigenous Zel sheep. Arch. Anim. Breed. 2013, 56, 216–224. [Google Scholar] [CrossRef]
- Orford, M.; Tzamaloukas, O.; Papachristoforou, C.; Miltiadou, D. Technical note: A simplified PCR-based assay for the characterization of two prolactin variants that affect milk traits in sheep breeds. J. Dairy Sci. 2010, 93, 5996–5999. [Google Scholar] [CrossRef]
- Gras, M.A.; Rotar, C.M.; Pelmus, R.S.; Lazar, C.; Ghita, E.; Grosu, H. Prolactin polymorphism effect over production traits types at Transylvanian Merino sheep. Sci. Papers Anim. Sci. Biotechnol. Lucrari Stiintifice Zootehnie si Biotehnologii 2017, 50, 56–60. [Google Scholar]
- Suarez-Vega, A.; Gutierrez-Gil, B.; Klopp, C.; Tosser-Klopp, G.; Arranz, J.J. Variant discovery in the sheep milk transcriptome using RNA sequencing. BMC Genom. 2017, 18, 170. [Google Scholar] [CrossRef] [PubMed]
- Park, Y.W.; Juárez, M.; Ramos, M.; Haenlein, G.F.W. Physico-chemical characteristics of goat and sheep milk. Small Rumin. Res. 2007, 68, 88–113. [Google Scholar] [CrossRef]
- Feligini, M.; Vlaco, S.; Curik, V.C.; Parma, P.; Greppi, G.; Enne, G. A single nucleotide polymorphism in the sheep kappa-casein coding region. J. Dairy Res. 2005, 72, 317–321. [Google Scholar] [CrossRef] [PubMed]
- Othman, O.; El-Fiky, S.A.; Hassan, N.; Mahfouz, E.; Balabel, E. Genetic variations of β- and K-casein genes in Egyptian sheep breeds. J. Appl. Biosci 2013, 64, 4858–4866. [Google Scholar] [CrossRef]
- Nassiri, M.; Heravi-Moussavi, A.; Raoof Alashawkany, A.; Ghovati, S. Leptin gene polymorphism in Iranian native Golpayegani and Taleshi cows. Pakistan J. Biol. Sci. 2007, 10, 3738–3741. [Google Scholar] [CrossRef]
- Vincent, A.L.; Wang, L.; Tuggle, C.K.; Robic, A.; Rothschild, M.F. Prolactin receptor maps to pig chromosome 16. Mamm. Genome 1997, 8, 793–794. [Google Scholar] [CrossRef] [PubMed]
- Hall, T.A. BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucleic Acids Symposium Series 1999, 41, 95–98. [Google Scholar]
- Yeh, F.C.; Yang, R.C.; Boyle, T. POPGENE Version 1.32: Microsoft Window-Based Freeware for Population Genetics Analysis; University of Alberta: Edmonton, AB, Canada, 1999. [Google Scholar] [CrossRef]
- Knight, C. Overview of prolactin’s role in farm animal lactation. Livest. Prod. Sci. 2001, 70, 87–93. [Google Scholar] [CrossRef]
- Georgescu, S.E.; Ene, A.; Dudu, A.; Ghita, E.; Costache, M. Genetic polymorphisms of β-lactoglobulin and α-s1-casein genes in Romanian Racka sheep. Sci. Pap. Anim. Sci. Biotechnol. 2016, 49, 50–53. [Google Scholar]
- Kusza, S.; Sziszkosz, N.; Nagy, K.; Masala, A.; Kukovics, S.; Andras, J. Preliminary result of a genetic polymorphism of beta-lactoglobulin gene and the phylogenetic study of ten balkan and central european indigenous sheep breeds. Acta Biochim. Polon. 2015, 62, 109–112. [Google Scholar] [CrossRef]
- Kawecka, A.; Radko, A. Genetic polymorphism of β-lactoglobulin in sheep raised for milk production. J. Appl. Anim. Res. 2011, 39, 68–71. [Google Scholar] [CrossRef]
- Baranyi, M.; Kerekes, A.; Hiripi, L.; Bősze, Z. Preliminary data on beta-lactoglobulin genetic polymorphisms in Hungarian Awassi and Racka sheep. Anim. Sci. Biotechnol. 2010, 43, 1–4. [Google Scholar]
- Kusza, S.; Ilie, D.; Sauer, M.; Nagy, K.; Atanasiu, T.S.; Dinu, G. Study of LGB gene polymorphisms of small ruminants reared in Eastern Europe. Czech. J. Anim. Sci. 2018, 63, 152–159. [Google Scholar] [CrossRef]
- Jawasreh, K.; Al-Qaisi, A.T.; Awawdeh, F. Association between GDF9, FecB and Prolactin gene polymorphisms and prolificacy of Awassi sheep. In Proceedings of the 10th World Congress of Genetics Applied to Livestock Production, Vancouver, BC, Canada, 18–22 August 2014. [Google Scholar]
- Ramos, A.M.; Matos, C.A.P.; Russo Almeida, P.A.; Bettencourt, C.M.V.; Matos, J.; Martins, Â.; Pinheiro, C.M.S.; Rangel-Figueiredo, T. Candidate genes for milk production traits in Portuguese dairy sheep. Small Rumin. Res. 2009, 82, 117–121. [Google Scholar] [CrossRef]
- Giambra, I.J.; Brandt, H.; Erhardt, G. Milk protein variants are highly associated with milk performance traits in East Friesian Dairy and Lacaune sheep. Small Rumin. Res. 2014, 121, 382–394. [Google Scholar] [CrossRef]
- Çelik, Ş.; Özdemir, S. β-Lactoglobulin variants in awassi and morkaraman sheep and their association with the composition and rennet clotting time of the milk. Turk. J. Vet. Anim. Sci. 2006, 30, 539–544. [Google Scholar] [CrossRef]
- Dario, C.; Carnicella, D.; Dario, M.; Bufano, G. Genetic polymorphism of B-lactoglobulin gene and effect on milk composition in Leccese sheep. Small Rumin. Res. 2008, 74, 270–273. [Google Scholar] [CrossRef]
- Corral, J.M. Associations between milk protein genetic polymorphisms and milk production traits in Merino sheep breed. Livest. Sci. 2010, 129, 73–79. [Google Scholar] [CrossRef]
- Giaccone, P.; Di Stasio, L.; Macciotta, N.; Portolano, B.; Todaro, M.; Cappio-Borlino, A. Effect of β-lactoglobulin polymorphism on milk-related traits of dairy ewes analysed by a repeated measures design. J. Dairy Res. 2000, 67, 443–448. [Google Scholar] [CrossRef]
- Mroczkowski, S.; Korman, K.; Erhardt, G.; Piwczyński, D.; Borys, B. Sheep milk protein polymorphism and its effect on milk performance of Polish Merino. Archiv. Fur Tierzu 2004, 47, 114–121. [Google Scholar]
- Rozbicka-Wieczorek, A.; Radzik-Rant, A.; Rant, W.; Puppel, K. The effect of breed, β lactoglobulin variants and somatic cell count on yield, chemical components and whey protein composition in milk of non-dairy sheep. J. Anim. Plant Sci. 2015, 25, 633–639. [Google Scholar]
- El-Shazly, S.A.; Mahfouz, M.E.; Al-Otaibi, S.A.; Ahmed, M.M. Genetic polymorphism in β-lactoglobulin gene of some sheep breeds in the Kingdom of Saudi Arabia (KSA) and its influence on milk composition. Afr. J. Biotechnol. 2012, 11, 4330–4337. [Google Scholar] [CrossRef]
- Zheng, X.; Ju, Z.; Wang, J.; Li, Q.; Huang, J.; Zhang, A.; Zhong, J.; Wang, C. Single nucleotide polymorphisms, haplotypes and combined genotypes of LAP3 gene in bovine and their association with milk production traits. Mol. Biol. Rep. 2011, 38, 4053–4061. [Google Scholar] [CrossRef] [PubMed]
| Gene | Primers (5′→3′) | TM (°C) | PCR Product (bp) | RE | Reference | |
|---|---|---|---|---|---|---|
| Beta-lactoglobulin (β-LG) | F | CTCTTTGGGTTCAGTGTGAGTCTTG | 58 | 301 | RsaI | [27] |
| R | CACCATTTCTGCAGCAGGATCTC | |||||
| Prolactin (PRL) | F | ACCTCTCCTCGGAAATGTTCA | 56 | 1209 | HaeIII | [28] |
| R | GGGACACTGAAGGACCAGAA | |||||
| Kappa Casein (CSN3) | F | CTGGGTTCACTATTCCCAATG | 57 | 680 | * | Accession # 443394 |
| R | TTGCTCATTTACCTGCGTTG | |||||
| Milk Trait | No. Records | Mean | SE | CV (%) |
|---|---|---|---|---|
| Milk production (Kg) | ||||
| TMY | 391 | 96.8 | 2.60 | 53.0 |
| TDM | 391 | 0.882 | 0.02 | 45.0 |
| Milk composition | ||||
| Fat% | 917 | 5.80 | 0.05 | 25.7 |
| SNF% | 986 | 9.74 | 0.03 | 8.30 |
| Protein% | 986 | 3.90 | 0.02 | 13.0 |
| Lactose% | 986 | 5.10 | 0.02 | 13.9 |
| Density g/cm2 | 986 | 34.3 | 0.10 | 9.4 |
| Gene 1 | Genotype | Observed Number | Expected Number | Genotype Frequency | Allele | Allele Frequency | Value of x2 Test |
|---|---|---|---|---|---|---|---|
| β-LG (n = 159) | AA | 27 | 28.7 | 0.17 | A | 0.42 | 0.29 ns |
| AB | 81 | 77.7 | 0.51 | B | 0.58 | ||
| BB | 51 | 52.7 | 0.32 | ||||
| PRL (n = 158) | AA | 115 | 107 | 0.73 | A | 0.82 | 19.2 ns |
| AB | 30 | 46.1 | 0.19 | B | 0.18 | ||
| BB | 13 | 5 | 0.08 | ||||
| CSN3 (n =156) | TT | 132 | 132.9 | 0.85 | T | 0.92 | 1.1 ns |
| TC | 24 | 22.1 | 0.15 | C | 0.08 |
| Trait | Milk Production 1 | Milk Components 2 | ||||||
|---|---|---|---|---|---|---|---|---|
| Factors | TMY (kg) | TDM (kg) | Fat% | SNF% | Protein% | Lactose% | Density, g/cm2 | |
| β-LG | 0.175 | 0.134 | 0.047 | <0.0001 | 0.046 | 0.03 | 0.002 | |
| PRL | 0.034 | 0.011 | 0.761 | 0.001 | 0.035 | 0.05 | 0.005 | |
| CSN3 | 0.812 | 0.275 | 0.172 | 0.048 | 0.424 | 0.104 | 0.541 | |
| Sire | <0.0001 | <0.0001 | <0.0001 | <0.0001 | 0.019 | 0.035 | 0.0004 | |
| Parity | 0.004 | 0.005 | 0.056 | 0.412 | 0.389 | 0.266 | 0.665 | |
| Year | 0.003 | 0.002 | ||||||
| β-LG × PRL | 0.039 | 0.031 | 0.001 | <0.0001 | 0.008 | 0.035 | 0.05 | |
| β-LG × CSN3 | 0.874 | 0.221 | 0.417 | 0.899 | 0.784 | 0.949 | 0.496 | |
| PRL × CSN3 | 0.177 | 0.104 | 0.002 | 0.02 | 0.228 | 0.115 | 0.767 | |
| Dam weight at lambing | 0.299 | 0.009 | ||||||
| Test-day milk | 0.004 | 0.002 | 0.0004 | 0.481 | <0.0001 | |||
| Gene | Genotype | N | Trait Least Square Means (±SE) | |
|---|---|---|---|---|
| TMY (Kg) | TDM (Kg) | |||
| β-LG | AA | 51 | 100.4 ± 13.4 | 0.718 ± 0.10 |
| AB | 145 | 72.2 ±15.7 | 0.606 ± 0.10 | |
| BB | 96 | 93.2 ± 12.6 | 0.801 ± 0.10 | |
| PRL | AA | 197 | 102.4 ± 9.86 a | 0.814 ± 0.08 a |
| AB | 68 | 71.4 ± 12.6 b | 0.540 ± 0.10 b | |
| BB | 27 | 92.0 ± 14.7 ab | 0.770 ± 0.11 a | |
| CSN3 | TT | 240 | 90.1 ± 11.8 | 0.762 ± 0.09 |
| TC | 52 | 87.1 ± 11.5 | 0.654 ± 0.09 | |
| β-LG × PRL | AAAA | 29 | 96.5 ± 13.0 b | 0.780 ± 0.10 ab |
| AAAB | 12 | 65.0 ± 23.0 bc | 0.359 ± 0.17 c | |
| AABB | 10 | 139.7 ± 25.2 a | 1.02 ± 0.19 a | |
| ABAA | 106 | 99.2 ± 17.3 ab | 0.834 ± 0.13 ab | |
| ABAB | 31 | 72.6 ± 19.3 bc | 0.578 ± 0.15 c | |
| ABBB | 8 | 44.9 ± 18.4 c | 0.406 ± 0.14 c | |
| BBAA | 62 | 111.6 ± 11.1 ab | 0.829 ± 0.08 ab | |
| BBAB | 25 | 76.8 ± 15.4 bc | 0.684 ± 0.12 b | |
| BBBB | 9 | 91.3 ± 27.6 ab | 0.891 ± 0.21 ab | |
| Gene | Genotype | N | Traits Least Square Means (±SE) | ||||
|---|---|---|---|---|---|---|---|
| Fat% | SNF% | Protein% | Lactose% | Density, g/cm2 | |||
| β-LG | AA | 112 | 6.63 ± 0.29 a | 9.50 ± 0.15 b | 3.90 ± 0.10 b | 4.99 ± 0.14 b | 33.0 ± 0.61 b |
| AB | 376 | 5.31 ± 0.43 b | 9.39 ± 0.22 b | 3.66 ± 0.14 b | 4.88 ± 0.20 b | 34.1 ± 0.87 ab | |
| BB | 277 | 6.30 ± 0.35 a | 10.4 ± 0.18 a | 4.13 ± 0.12 a | 5.39 ± 0.16 a | 35.4 ± 0.71 a | |
| PRL | AA | 554 | 6.13 ± 0.17 | 9.86 ± 0.08 a | 4.00 ± 0.06 a | 5.02 ± 0.08 b | 34.2 ± 0.34 a |
| AB | 159 | 5.96 ± 0.19 | 9.44 ± 0.10 b | 3.79 ± 0.06 b | 4.90 ± 0.09 b | 32.9 ± 0.40 b | |
| BB | 52 | 6.15 ± 0.38 | 9.96 ± 0.19 a | 3.91 ± 0.13 ab | 5.35 ± 0.18 a | 35.3 ± 0.77 a | |
| CSN3 | TT | 624 | 6.49 ± 1.45 | 10.1 ± 0.19 a | 3.98 ± 0.12 | 5.32 ± 0.17 | 34.5 ± 0.75 |
| TC | 141 | 5.67 ± 1.05 | 9.45 ± 0.15 b | 3.82 ± 0.10 | 4.86 ± 0.14 | 33.7 ± 0.62 | |
| β-LG × PRL | AAAA | 56 | 6.38 ± 0.34 bc | 9.77 ± 0.17 b | 4.05 ± 0.11 b | 4.87 ± 0.16 c | 33.6 ± 0.70 b |
| AAAB | 29 | 5.56 ± 0.45 c | 8.63 ± 0.21 c | 3.47 ± 0.14 c | 4.46 ± 0.19 d | 30.2 ± 0.84 c | |
| AABB | 27 | 7.95 ± 0.82 a | 10.1 ± 0.42 b | 4.18 ± 0.28 ab | 5.66 ± 0.39 ab | 35.2 ± 1.71 ab | |
| ABAA | 284 | 5.97 ± 0.24 c | 9.87 ± 0.11 b | 3.99 ± 0.08 b | 5.06 ± 0.10 bc | 34.6 ± 0.46 b | |
| ABAB | 78 | 6.67 ± 0.30 b | 9.97 ± 0.15 b | 3.97 ± 0.10 b | 5.24 ± 0.14 b | 34.5 ± 0.61 b | |
| ABBB | 14 | 3.28 ± 1.11 d | 8.33 ± 0.57 c | 3.02 ± 0.38 c | 4.35 ± 0.50 cd | 33.3 ± 2.30 bc | |
| BBAA | 214 | 6.03 ± 0.18 c | 9.92 ± 0.09 b | 3.96 ± 0.06 b | 5.12 ± 0.08 bc | 34.6 ± 0.36 b | |
| BBAB | 52 | 5.63 ± 0.38 c | 9.72 ± 0.15 b | 3.91 ± 0.10 b | 4.99 ± 0.14 bc | 34.1 ± 0.59 b | |
| BBBB | 11 | 7.23 ± 1.03 ab | 11.5 ± 0.52 a | 4.53 ± 0.35 a | 6.05 ± 0.48 a | 37.4 ± 2.10 a | |
| PRL × CSN3 | AATT | 478 | 6.13 ± 0.18 b | 9.89 ± 0.08 b | 3.95 ± 0.05 | 5.11 ± 0.07 | 34.5 ± 0.32 |
| AATC | 75 | 6.12 ± 0.27 b | 9.82 ± 0.13 bc | 4.05 ± 0.09 | 4.93 ± 0.12 | 33.9 ± 0.54 | |
| ABTT | 122 | 5.32 ± 0.22 c | 9.29 ± 0.11 d | 3.68 ± 0.46 | 4.83 ± 0.17 | 32.9 ± 1.02 | |
| ABTC | 37 | 6.59 ± 0.32 b | 9.59 ± 0.16 c | 3.89 ± 0.11 | 4.68 ± 0.15 | 32.9 ± 1.45 | |
| BBTT | 23 | 8.02 ± 1.11 a | 11.0 ± 0.57 a | 4.31 ± 0.38 | 6.03 ± 1.12 | 36.2 ± 2.30 | |
| BBTC | 29 | 4.29 ± 0.75 d | 8.93 ± 0.38 d | 3.51 ± 0.62 | 4.68 ± 0.35 | 34.5 ± 1.55 | |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jawasreh, K.; Amareen, A.A.; Aad, P. Effect and Interaction of β-Lactoglobulin, Kappa Casein, and Prolactin Genes on Milk Production and Composition of Awassi Sheep. Animals 2019, 9, 382. https://doi.org/10.3390/ani9060382
Jawasreh K, Amareen AA, Aad P. Effect and Interaction of β-Lactoglobulin, Kappa Casein, and Prolactin Genes on Milk Production and Composition of Awassi Sheep. Animals. 2019; 9(6):382. https://doi.org/10.3390/ani9060382
Chicago/Turabian StyleJawasreh, Khaleel, Ahmad Al Amareen, and Pauline Aad. 2019. "Effect and Interaction of β-Lactoglobulin, Kappa Casein, and Prolactin Genes on Milk Production and Composition of Awassi Sheep" Animals 9, no. 6: 382. https://doi.org/10.3390/ani9060382
APA StyleJawasreh, K., Amareen, A. A., & Aad, P. (2019). Effect and Interaction of β-Lactoglobulin, Kappa Casein, and Prolactin Genes on Milk Production and Composition of Awassi Sheep. Animals, 9(6), 382. https://doi.org/10.3390/ani9060382





