First Isolation of Methicillin-Resistant Livestock-Associated Staphylococcus aureus CC398 and CC1 in Intensive Pig Production Farms in Argentina
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sampling and MRSA Identification
2.2. Susceptibility Testing
2.3. Gene Detection by PCR
2.4. Molecular Typing
2.5. Genome Sequencing and Analysis
2.6. Data Availability
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Zarazaga, M.; Gómez, P.; Ceballos, S.; Torres, C. Molecular Epidemiology of Staphylococcus aureus Lineages in the Animal-Human Interface. In Staphylococcus Aureus, 2nd ed.; Fetsch, A., Ed.; Academic Press: London, UK, 2018; pp. 189–214. [Google Scholar] [CrossRef]
- Abdullahi, I.N.; Lozano, C.C.; Saidenberg, A.B.S.; Latorre-Fernández, J.; Zarazaga, M.; Torres, C. Comparative review of the nasal carriage and genetic characteristics of Staphylococcus aureus in healthy livestock: Insight into zoonotic and anthroponotic clones. Infect. Genet. Evol. 2023, 109, 105408. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority—ECDC—European Centre for Disease Prevention and Control. Analysis of the baseline survey on the prevalence of methicillin-resistant Staphylococcus aureus (MRSA) in holdings with breeding pigs, in the EU, 2008−Part A: MRSA prevalence estimates. EFSA J. 2009, 7, 1376. [Google Scholar] [CrossRef]
- Voss, A.; Loeffen, F.; Bakker, J.; Klaassen, C.; Wulf, M. Methicillin-resistant Staphylococcus aureus in Pig Farming. Emerg. Infect. Dis. 2005, 11, 1965–1966. [Google Scholar] [CrossRef] [PubMed]
- Crombé, F.; Argudín, M.A.; Vanderhaeghen, W.; Hermans, K.; Haesebrouck, F.; Butaye, P. Transmission dynamics of methicillin-resistant Staphylococcus aureus in pigs. Front. Microbiol. 2013, 4, 57. [Google Scholar] [CrossRef]
- Aires-de-Sousa, M. Methicillin-resistant Staphylococcus aureus among animals: Current overview. Clin. Microbiol. Infect. 2017, 23, 373–380. [Google Scholar] [CrossRef]
- Khanna, T.; Friendship, R.; Dewey, C.; Weese, J.S. Methicillin resistant Staphylococcus aureus colonization in pigs and pig farmers. Vet. Microbiol. 2008, 128, 298–303. [Google Scholar] [CrossRef]
- Arriola, C.S.; Güere, M.E.; Larsen, J.; Skov, R.L.; Gilman, R.H.; Gonzalez, A.E.; Silbergeld, E.K. Presence of methicillin-resistant Staphylococcus aureus in pigs in Peru. PLoS ONE 2011, 6, e28529. [Google Scholar] [CrossRef]
- Groves, M.D.; O’Sullivan, M.V.; Brouwers, H.J.; Chapman, T.A.; Abraham, S.; Trott, D.J.; Al Jassim, R.; Coombs, G.W.; Skov, R.L.; Jordan, D. Staphylococcus aureus ST398 detected in pigs in Australia. J. Antimicrob. Chemother. 2014, 69, 1426–1428. [Google Scholar] [CrossRef]
- Chuang, Y.Y.; Huang, Y.C. Livestock-associated meticillin-resistant Staphylococcus aureus in Asia: An emerging issue? Int. J. Antimicrob. Agents 2015, 45, 334–340. [Google Scholar] [CrossRef]
- van Cleef, B.A.; Monnet, D.L.; Voss, A.; Krziwanek, K.; Allerberger, F.; Struelens, M.; Zemlickova, H.; Skow, R.L.; Vuopio-Varkila, J.; Cuny, C.; et al. Livestock-associated Methicillin-Resistant Staphylococcus aureus in humans, Europe. Emerg. Infect. Dis. 2011, 17, 502–505. [Google Scholar] [CrossRef]
- World Health Organization. Global Action Plan on Antimicrobial Resistance. 2015. Available online: https://www.who.int/publications-detail-redirect/9789241509763 (accessed on 23 October 2022).
- Giacoboni, G.I.; Nievas, H.D.; Nievas, V.F.; Pérez, E.M.; Armocida, A.D.; Moredo, F.A.; Gagetti, P. Circulación de Staphylococcus aureus resistentes a meticilina y multirresistente, en granjas de cría porcina de Argentina. Analecta Vet. 2022, 42, 063. [Google Scholar] [CrossRef]
- Performance Standards for Antimicrobial Susceptibility Testing, CLSI Supplement M100, 32nd ed.; Clinical and Laboratory Standards Institute: Wayne, PA, USA, 2022.
- Magiorakos, A.P.; Srinivasan, A.; Carey, R.B.; Carmeli, Y.; Falagas, M.E.; Giske, C.G.; Harbarth, S.; Hindler, J.F.; Kahlmeter, G.; Olson-Liljequist, B.; et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 2012, 18, 268–281. [Google Scholar] [CrossRef]
- Vannuffel, P.; Gigi, J.; Ezzedine, H.; Vandercam, B.; Delmee, M.; Wauters, G.; Gala, J.L. Specific detection of methicillin-resistant Staphylococcus species by multiplex PCR. J. Clin. Microbiol. 1995, 33, 2864–2867. [Google Scholar] [CrossRef]
- Stegger, M.; Lindsay, J.A.; Moodley, A.; Skov, R.; Broens, E.M.; Guardabassi, L. Rapid PCR detection of Staphylococcus aureus clonal complex 398 by targeting the restriction-modification system carrying sau1-hsdS1. J. Clin. Microbiol. 2011, 49, 732–734. [Google Scholar] [CrossRef]
- Corso, A.; Santos Sanches, I.; Aires de Sousa, M.; Rossi, A.; de Lencastre, H. Spread of a methicillin-resistant and multiresistant epidemic clone of Staphylococcus aureus in Argentina. Microb. Drug Resist. 1998, 4, 277–288. [Google Scholar] [CrossRef]
- Tenover, F.C.; Arbeit, R.D.; Goering, R.V.; Mickelsen, P.A.; Murray, B.E.; Persing, D.H.; Swaminathan, B. Interpreting chromosomal DNA restriction patterns produced by pulsed-field gel electrophoresis: Criteria for bacterial strain typing. J. Clin. Microbiol. 1995, 33, 2233–2239. [Google Scholar] [CrossRef]
- Larsen, M.; Cosentino, S.; Rasmussen, S.; Rundsten, C.; Hasman, H.; Marvig, R.L.; Jelsbak, L.; Sicheritz-Pontén, T.; Ussery, D.W.; Aarestrup, F.M.; et al. Multilocus Sequence Typing of Total Genome Sequenced Bacteria. J. Clin. Microbiol. 2012, 50, 1355–1361. [Google Scholar] [CrossRef]
- Jolley, K.A.; Bray, J.E.; Maiden, M.C.J. Open-access bacterial population genomics: BIGSdb software, the PubMLST.org website and their applications. Wellcome Open Res. 2018, 24, 124. [Google Scholar] [CrossRef]
- Zankari, E.; Allesøe, R.; Joensen, K.G.; Cavaco, L.M.; Lund, O.; Aarestrup, F.M. PointFinder: A novel web tool for WGS-based detection of antimicrobial resistance associated with chromosomal point mutations in bacterial pathogens. J. Antimicrob. Chemother. 2020, 72, 2764–2768. [Google Scholar] [CrossRef]
- Joensen, K.G.; Scheutz, F.; Lund, O.; Hasman, H.; Kaas, R.S.; Nielsen, E.M.; Aarestrup, F.M. Real-time whole-genome sequencing for routine typing, surveillance, and outbreak detection of verotoxigenic Escherichia coli. J. Clin. Micobiol. 2014, 52, 1501–1510. [Google Scholar] [CrossRef]
- Carattoli, A.; Zankari, E.; Garcia-Fernandez, A.; Voldby Larsen, M.; Lund, O.; Villa, L.; Aarestrup, F.M.; Hasman, H. PlasmidFinder and pMLST: In silico detection and typing of plasmids. Antimicrob. Agents Chemother. 2014, 58, 3895–3903. [Google Scholar] [CrossRef] [PubMed]
- Johansson, M.H.K.; Bortolaia, V.; Tansirichaiya, S.; Aarestrup, F.M.; Roberts, A.P.; Petersen, T.N. Detection of mobile genetic elements associated with antibiotic resistance in Salmonella enterica using a newly developed web tool: MobileElementFinder. J. Antimicrob. Chemother. 2021, 76, 101–109. [Google Scholar] [CrossRef] [PubMed]
- European Food Safety Authority—ECDC—European Centre for Disease Prevention and Control. The European union summary report on antimicrobial resistance in zoonotic and indicator bacteria from humans, animals and food in 2019–2020. EFSA J. 2022, 20, 7209. [Google Scholar] [CrossRef]
- Rodríguez-López, P.; Filipello, V.; Di Ciccio, P.A.; Pitozzi, A.; Ghidini, S.; Scali, F.; Ianieri, A.; Zanardi, E.; Losio, M.N.; Simon, A.C.; et al. Assessment of the Antibiotic Resistance Profile, Genetic Heterogeneity and Biofilm Production of Methicillin-Resistant Staphylococcus aureus (MRSA) Isolated from The Italian Swine Production Chain. Foods 2020, 9, 1141. [Google Scholar] [CrossRef] [PubMed]
- Golob, M.; Pate, M.; Kušar, D.; Zajc, U.; Papić, B.; Ocepek, M.; Zdovc, I.; Avberšek, J. Antimicrobial Resistance and Molecular Characterization of Methicillin-Resistant Staphylococcus aureus from Two Pig Farms: Longitudinal Study of LA-MRSA. Antibiotics 2022, 11, 1532. [Google Scholar] [CrossRef] [PubMed]
- Kadlec, K.; Fessler, A.T.; Hauschild, T.; Schwarz, S. Novel and uncommon antimicrobial resistance genes in livestock-associated methicillin-resistant Staphylococcus aureus. Clin. Microbiol. Infect. 2012, 18, 745–755. [Google Scholar] [CrossRef]
- Leão, C.; Clemente, L.; Cara d’Anjo, M.; Albuquerque, T.; Amaro, A. Emergence of Cfr-Mediated Linezolid Resistance among Livestock-Associated Methicillin-Resistant Staphylococcus aureus (LA-MRSA) from Healthy Pigs in Portugal. Antibiotics. 2022, 11, 1439. [Google Scholar] [CrossRef]
- National Reference Laboratory in Antimicrobial Resistance, Argentina; Servicio Antimicrobianos, Instituto Nacional de Enfermedades Infecciosas ANLIS “Dr. Carlos G. Malbrán”, Buenos Aires, Argentina. Vigilancia de la resistencia a los antimicrobianos Red WHONET Argentina 2010–2021. 2021. Available online: http://antimicrobianos.com.ar/ATB/wp-content/uploads/2022/11/Vigilancia-Nacional-de-la-Resistencia-a-los-Antimicrobianos-Red-WHONET-Argentina-Tendencia-2010-2021.pdf (accessed on 20 December 2022).
- Bens, C.C.; Voss, A.; Klaassen, C.H. Presence of a novel DNA methylation enzyme in methicillin-resistant Staphylococcus aureus isolates associated with pig farming leads to uninterpretable results in standard pulsed-field gel electrophoresis analysis. J. Clin. Microbiol. 2006, 44, 1875–1876. [Google Scholar] [CrossRef]
- Butaye, P.; Argudín, M.A.; Smith, T.C. Livestock-Associated MRSA and Its Current Evolution. Curr. Clin. Micro. Rpt. 2016, 3, 19–31. [Google Scholar] [CrossRef]
- Ruiz-Ripa, L.; Bellés-Bellés, A.; Fernández-Fernández, R.; García, M.; Vilaró, A.; Zarazaga, M.; Torres, C. Linezolid-resistant MRSA-CC398 carrying the cfr gene, and MRSA-CC9 isolates from pigs with signs of infection in Spain. J. Appl. Microbiol. 2021, 131, 615–622. [Google Scholar] [CrossRef]
- Moreno-Flores, A.; Potel-Alvarellos, C.; Francisco-Tomé, M.; Constenla-Caramés, L.; Pérez-Roth, E.; López-Cotón, C.; Comesaña-Da Vila, E.; Eiroa-de la Puente, L.; Álvarez-Fernández, M. Methicillin-resistant Staphylococcus aureus in swine housed indoors in Galicia, Spain. Enferm. Infecc. Microbiol. Clin. 2020, 38, 16–20. [Google Scholar] [CrossRef]
- Reynaga, E.; Navarro, M.; Vilamala, A.; Roure, P.; Quintana, M.; Garcia-Nuñez, M.; Figueras, R.; Torres, C.; Lucchetti, G.; Sabrià, M. Prevalence of colonization by methicillin-resistant Staphylococcus aureus ST398 in pigs and pig farm workers in an area of Catalonia, Spain. BMC Infect. Dis. 2016, 16, 716. [Google Scholar] [CrossRef]
- Lopes, E.; Conceicão, T.; Poirel, L.; de Lencastre, H.; Aires-de-Sousa, M. Epidemiology and antimicrobial resistance of methicillin-resistant Staphylococcus aureus isolates colonizing pigs with different exposure to antibiotics. PLoS ONE 2019, 14, e0225497. [Google Scholar] [CrossRef]
- Price, L.B.; Stegger, M.; Hasman, H.; Aziz, M.; Larsen, J.; Andersen, P.S.; Pearson, T.; Waters, A.E.; Foster, J.T.; Schupp, J.; et al. Staphylococcus aureus CC398: Host adaptation and emergence of methicillin resistance in livestock. mBio 2012, 3, e00305-11. [Google Scholar] [CrossRef]
- Matuszewska, M.; Murray, G.G.R.; Ba, X.; Wood, R.; Holmes, M.A.; Weinert, L.A. Stable antibiotic resistance and rapid human adaptation in livestock-associated MRSA. Elife 2022, 11, e74819. [Google Scholar] [CrossRef]
- Schwendimann, L.; Merda, D.; Berger, T.; Denayer, S.; Feraudet-Tarisse, C.; Kläui, A.J.; Messio, S.; Mistou, M.Y.; Nia, Y.; Hennekinne, J.A.; et al. Staphylococcal Enterotoxin Gene Cluster: Prediction of Enterotoxin (SEG and SEI) Production and of the Source of Food Poisoning on the Basis of vSaβ Typing. Appl. Environ. Microbiol. 2021, 87, e0266220. [Google Scholar] [CrossRef]
- Abreu, R.; Rodríguez-Álvarez, C.; Lecuona, M.; Castro, B.; González, J.C.; Aguirre-Jaime, A.; Arias, Á. Increased Antimicrobial Resistance of MRSA Strains Isolated from Pigs in Spain between 2009 and 2018. Vet. Sci. 2019, 6, 38. [Google Scholar] [CrossRef]
- McDougal, L.K.; Steward, C.D.; Killgore, G.E.; Chaitram, J.M.; McAllister, S.K.; Tenover, F.C. Pulsed-field gel electrophoresis typing of oxacillin-resistant Staphylococcus aureus isolates from the United States: Establishing a national database. J. Clin. Microbiol. 2003, 41, 5113–5120. [Google Scholar] [CrossRef]
- Egea, A.L.; Gagetti, P.; Lamberghini, R.; Faccone, D.; Lucero, C.; Vindel, A.; Tosoroni, D.; Garnero, A.; Saka, H.A.; Galas, M.; et al. New patterns of methicillin-resistant Staphylococcus aureus (MRSA) clones, community-associated MRSA genotypes behave like healthcare-associated MRSA genotypes within hospitals, Argentina. Int. J. Med. Microbiol. 2014, 304, 1086–1099. [Google Scholar] [CrossRef]
- Alba, P.; Feltrin, F.; Cordaro, G.; Porrero, M.C.; Kraushaar, B.; Argudín, M.A.; Nykäsenoja, S.; Monaco, M.; Stegger, M.; Aarestrup, F.M.; et al. Livestock-Associated Methicillin Resistant and Methicillin Susceptible Staphylococcus aureus Sequence Type (CC)1 in European Farmed Animals: High Genetic Relatedness of Isolates from Italian Cattle Herds and Humans. PLoS ONE 2015, 10, e0137143. [Google Scholar] [CrossRef]

| No. of Isolates | Resistance Phenotype | No. of Antimicrobial Classes |
|---|---|---|
| 5 | FOX, PEN, TET, CMP, CIP, ERY, CLI, LIN, GEN, SXT | 7 |
| 2 | FOX, PEN, TET, CMP, CIP, CLI, LIN, SXT | 6 |
| 2 | FOX, PEN, TET, CMP, CIP, ERY, CLI, LIN, GEN | 6 |
| 6 | FOX, PEN, TET, CMP, CIP, ERY, CLI, LIN | 5 |
| 4 | FOX, PEN, TET, CMP, CIP, GEN | 5 |
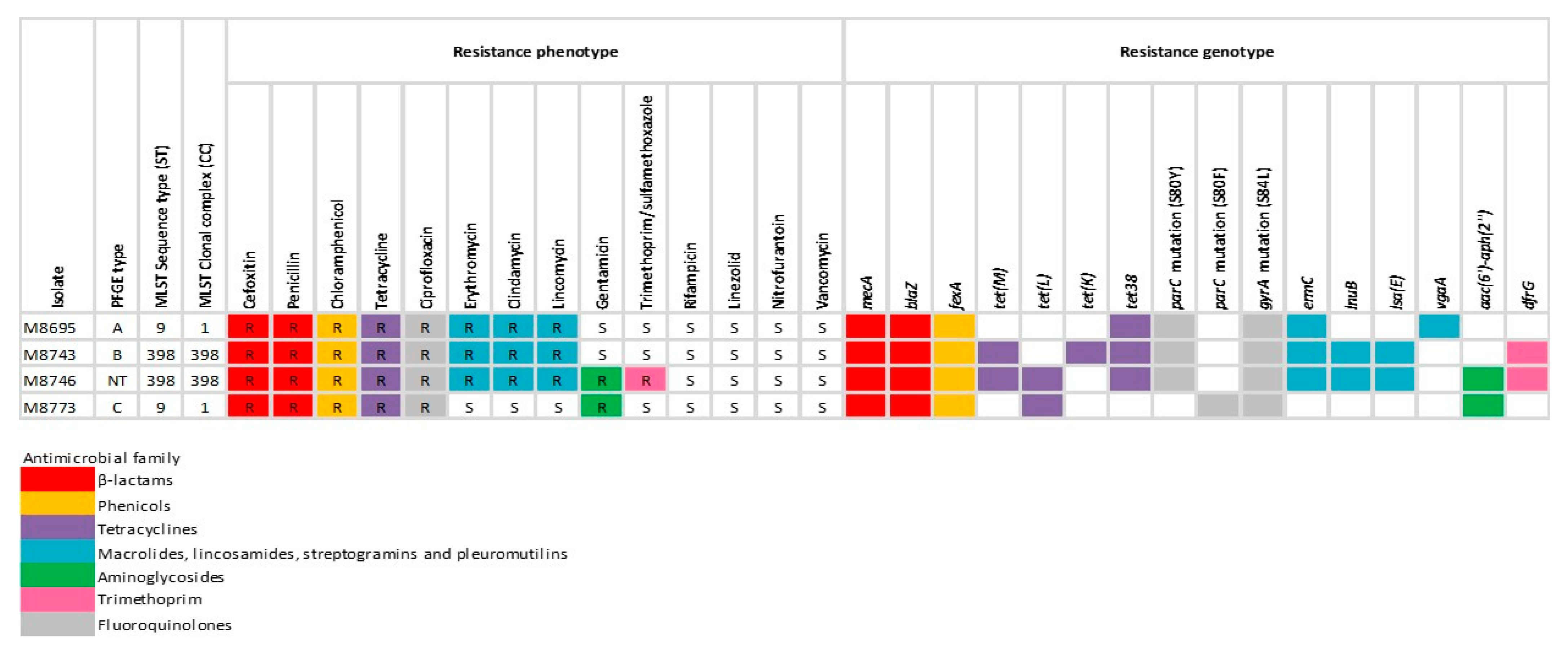
| Features | M8695 | M8743 | M8746 | M8773 |
|---|---|---|---|---|
| Farm region | Buenos Aires | Santa Fe | Buenos Aires | San Luis |
| Genome ID | 128.043.869 | 128.041.875 | 128.043.872 | 128.042.825 |
| Contigs | 74 | 64 | 126 | 54 |
| Genome length (bp) | 2,781,818 | 2,783,192 | 2,794,082 | 2,065,882 |
| GC Content (%) | 32.64 | 32.80 | 32.73 | 33.11 |
| MLST | ST1 (CC9) | ST398 (CC398) | ST398 (CC398) | ST1 (CC9) |
| Resistance genes | mecA; blaZ; fexA; tet38; ermC; vgaA; ant(6)-I; ant(9)-Ia | mecA; blaZ; fexA; tetM; tetK; tet38; ermC; lnuB; lsa(E); aadD; dfrG | mecA; blaZ; fexA; tetM; tetL; tet38; ermC; lnuB; lsa(E); aac(6′)-aph(2″); aadD; dfrG | mecA; blaZ; fexA; tetL; aac(6′)-aph(2″); aadD |
| SCCmec elements | type_V(5C2&5) | type_V(5C2&5) | type_V(5C2&5) | type_V(5C2&5) |
| Virulence genes | aur, hlgA, hlgB, hlgC, seg, sei, sem, sen, seo, seu | aur, hlgA, hlgB, hlgC | aur, hlgA, hlgB, hlgC | aur, hlgA, hlgB, hlgC |
| Plasmid replicons | rep10, rep7b | rep7a, rep10, rep22, repUS43 | rep10, rep13, rep22, repUS43 | rep22, repUS28 |
| MGE * | Tn558, ISSau3 (IS1182) | Tn558, ISSau1 (IS30) | Tn6009, Tn558, IS256, ISSau1 (IS30) | Tn558, ISSau5 (IS30), IS256 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gagetti, P.; Giacoboni, G.I.; Nievas, H.D.; Nievas, V.F.; Moredo, F.A.; Corso, A. First Isolation of Methicillin-Resistant Livestock-Associated Staphylococcus aureus CC398 and CC1 in Intensive Pig Production Farms in Argentina. Animals 2023, 13, 1796. https://doi.org/10.3390/ani13111796
Gagetti P, Giacoboni GI, Nievas HD, Nievas VF, Moredo FA, Corso A. First Isolation of Methicillin-Resistant Livestock-Associated Staphylococcus aureus CC398 and CC1 in Intensive Pig Production Farms in Argentina. Animals. 2023; 13(11):1796. https://doi.org/10.3390/ani13111796
Chicago/Turabian StyleGagetti, Paula, Gabriela Isabel Giacoboni, Hernan Dario Nievas, Victorio Fabio Nievas, Fabiana Alicia Moredo, and Alejandra Corso. 2023. "First Isolation of Methicillin-Resistant Livestock-Associated Staphylococcus aureus CC398 and CC1 in Intensive Pig Production Farms in Argentina" Animals 13, no. 11: 1796. https://doi.org/10.3390/ani13111796
APA StyleGagetti, P., Giacoboni, G. I., Nievas, H. D., Nievas, V. F., Moredo, F. A., & Corso, A. (2023). First Isolation of Methicillin-Resistant Livestock-Associated Staphylococcus aureus CC398 and CC1 in Intensive Pig Production Farms in Argentina. Animals, 13(11), 1796. https://doi.org/10.3390/ani13111796







