Microsatellite Diversity and Phylogenetic Relationships among East Eurasian Bos taurus Breeds with an Emphasis on Rare and Ancient Local Cattle
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Information and Microsatellite Data
2.2. DNA Extraction and Fragment Analysis
2.3. Statistical Analysis
3. Results
3.1. Genetic Variability
3.2. Pairwise Fst and Ast Values
3.3. Wright’s F Statistics for Each Locus
3.4. Bayesian Clustering Analysis
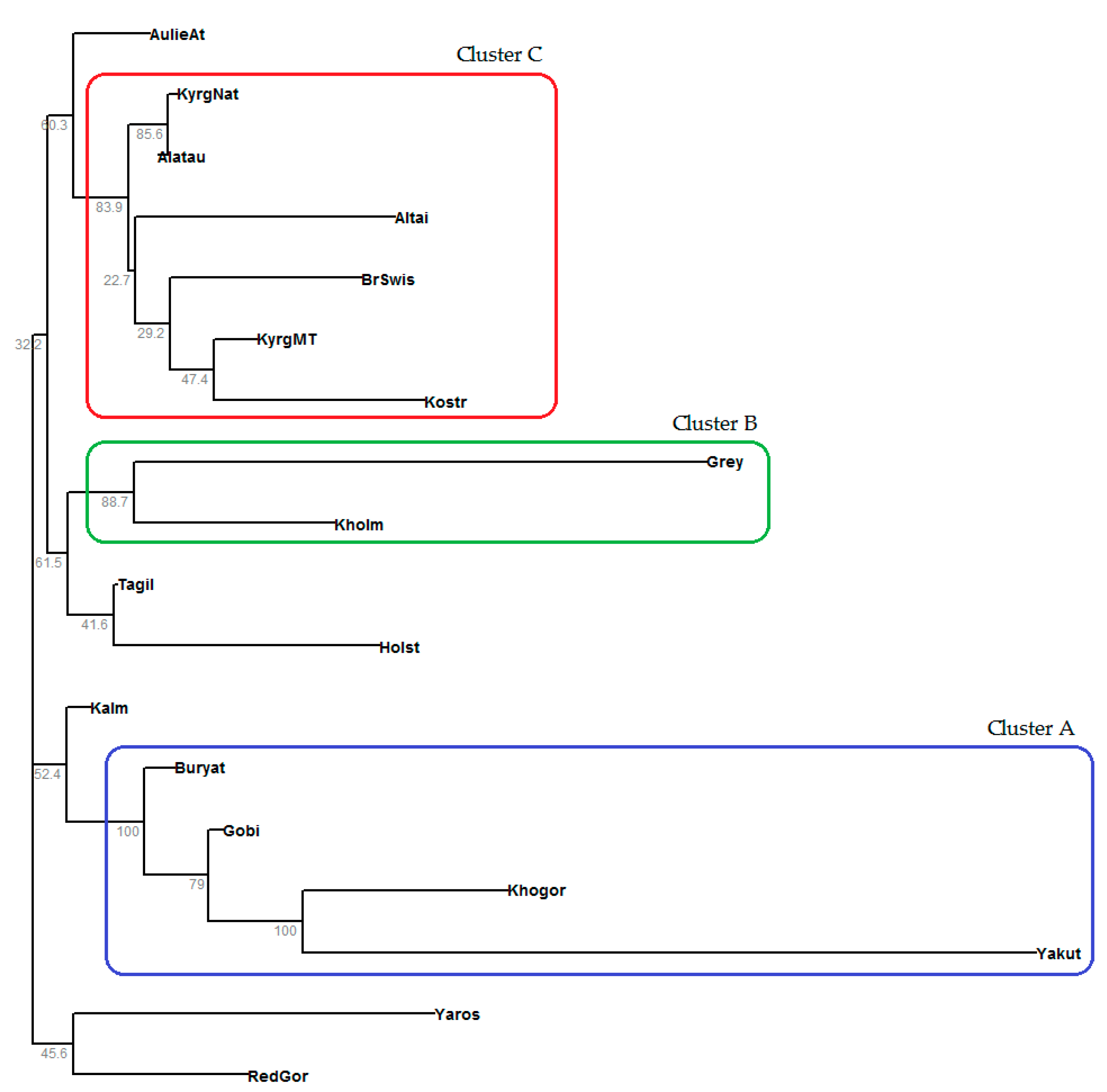
3.5. Phylogenetic Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Ethical Statement
Appendix A
| Locus and Source Reference | Position on Chromosome | Repeating Sequences | Sequences Forward (F) and Inverse (R) Primers | Length of Amplicons (bp) |
|---|---|---|---|---|
| BM1824 [68] | D1S34 | (GT)n | F: GAGCAAGGTGTTTTTCCAATC R: CATTCTCCAACTGCTTCCTTG | 176–188 |
| BM2113 [69] | D2S26 | (CA)n | F: GCTGCCTTCTACCAAATACCC R: CTTCCTGAGAGAAGCAACACC | 124–146 |
| CSRM60 [70] | D10S5 | (AC)n | F: AAGATGTGATCCAAGAGAGAGGCA R: AGGACCAGATCGTGAAAGGCATAG | 91–117 |
| CSSM66 [68] | D14S31 | (AC)n | F: AATTTAATGCACTGAGGAGCTTGG R: ACACAAATCCTTTCTGCCAGCTGA | 177–203 |
| ETH3 [71] | D19S2 | (GT)nAC(GT)6 | F: GAACCTGCCTCTCCTGCATTGG R: ACTCTGCCTGTGGCCAAGTAGG | 100–128 |
| ETH10 [71] | D5S3 | (AC)n | F: GTTCAGGACTGGCCCTGCTAACA R: CCTCCAGCCCACTTTCTCTTCTC | 206–222 |
| ETH225 [72] | D9S2 | (TG)4CG(TG)(CA)n | F: GATCACCTTGCCACTATTTCCT R: ACATGACAGCCAGCTGCTACT | 139–157 |
| ILSTS006 [73] | D7S8 | (GT)n | F: TGTCTGTATTTCTGCTGTGG R: ACACGGAAGCGATCTAAACG | 279–297 |
| INRA023 [74] | D3S10 | (AC)n | F: GAGTAGAGCTACAAGATAAACTTC R: TAACTACAGGGTGTTAGATGAACTC | 201–225 |
| SPS115 [75] | D15 | (CA)nTA(CA)6 | F: AAAGTGACACAACAGCTTCACCAG R: AACCGAGTGTCCTAGTTTGGCTGTG | 247–261 |
| TGLA53 [76] | D16S3 | (TG)6CG(TG)4(TA)n | F: GCTTTCAGAAATAGTTTGCATTCA R: ATCTTCACATGATATTACAGCAGA | 151–187 |
| TGLA122 [76] | D21S6 | (AC)n(AT)n | F: AATCACATGGCAAATAAGTACATAC R: CCCTCCTCCAGGTAAATCAGC | 136–182 |
| TGLA126 [76] | D20S1 | (TG)n | F: CTAATTTAGAATGAGAGAGGCTTCT R: TTGGTCCTCTATTCTCTGAATATTCC | 111–127 |
| TGLA227 [76] | D18S1 | (TG)n | F: GGAATTCCAAATCTGTTAATTTGCT R: ACAGACAGAAACTCAATGAAAGCA | 76–104 |
| Breed | ETH3 | CSSM66 | INRA023 | ILSTS006 | TGLA227 | TGLA126 | TGLA122 | SPS115 | ETH225 | TGLA53 | CSRM60 | BM2113 | BM1824 | ETH10 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Holst | 0.142 | 0.743 | 0.546 | 0.617 | 0.721 | 0.429 | 0.479 | 0.433 | 0.394 | 0.134 | 0.986 | 0.77 | 0.225 | 0.849 |
| Tagil | 0.354 | 0.411 | 0.969 | 0.411 | 0.493 | 0.559 | 0.595 | 0.293 | 0.133 | 0.28 | 0.725 | 0.465 | 0.342 | 0.298 |
| Kholm | 0.325 | 0.348 | 0.141 | 0.844 | 0.058 | 0.818 | 0.181 | 0.277 | 0.106 | 0.114 | 0.794 | 0.646 | 0.673 | 0.212 |
| RedGor | 0.6 | 0.267 | 0.241 | 0.668 | 0.5 | 0.005 | 0.282 | 0.614 | 0.767 | 0.776 | 0.502 | 0.832 | 0.282 | 0.45 |
| BrSwis | 0.102 | 0 | 0.314 | 0.004 | 0 | 0.025 | 0.001 | 0.623 | 0.001 | 0 | 0.006 | 0 | 0.07 | 0.163 |
| Grey | 0.321 | 0.331 | 0.764 | 0.341 | 0.737 | 0.889 | 0.844 | 0.184 | 0.295 | 0.917 | 0.457 | 0.666 | 0.14 | 0.57 |
| Yaros | 0.212 | 0.224 | 0.986 | 0.135 | 0.322 | 0.865 | 0.867 | 0.352 | 0.186 | 0.182 | 0.155 | 0.855 | 0.559 | 0.294 |
| Kostr | 0.497 | 0.562 | 0.025 | 0.625 | 0.092 | 1 | 0.397 | 0.701 | 0.169 | 0.267 | 0.233 | 0.823 | 0.1 | 0.364 |
| Altai | 0.008 | 0.301 | 0.012 | 0.257 | 0.409 | 0.694 | 0.914 | 0.029 | 0.538 | 0.282 | 0.911 | 0.84 | 0.953 | 0.494 |
| Kalm | 0.464 | 0.084 | 0.048 | 0.491 | 0.473 | 0.239 | 0.555 | 0.103 | 0.184 | 0.122 | 0.565 | 0.165 | 0.098 | 0.861 |
| AulieAt | 0.799 | 0.09 | 0.182 | 0.071 | 0.405 | 0.391 | 0.74 | 0.333 | 0.321 | 0.844 | 0.808 | 0.054 | 0.694 | 0.68 |
| Alatau | 0.844 | 0.207 | 0.334 | 0.975 | 0.647 | 0.652 | 0.015 | 0.094 | 0.247 | 0.015 | 0.626 | 0.5 | 0.594 | 0.527 |
| KyrgBT | 0.089 | 0.909 | 0.319 | 0.837 | 0.504 | 0.007 | 0.121 | 0.027 | 0.337 | 0.483 | 0.22 | 0.712 | 0.888 | 0.32 |
| KyrgNat | 0.094 | 0.465 | 0.898 | 0.218 | 0.53 | 0.541 | 0.927 | 0.337 | 0.418 | 0.372 | 0.304 | 0.731 | 0.833 | 0.927 |
| Yakut | 0.085 | 0.785 | 0.359 | 0.918 | 0.018 | 0.981 | 0.005 | 1 | 0.619 | 0.755 | 0.619 | 0.9 | 0.559 | 0.91 |
| Khogor | 0.345 | 0.912 | 0.269 | 0.005 | 0.095 | 0.115 | 0.037 | 0.155 | 0.605 | 0.465 | 0.186 | 0.697 | 0.456 | 0.348 |
| Gobi | 0.301 | 0.392 | 0.25 | 0.067 | 0.156 | 0.579 | 0.457 | 0.146 | 0.248 | 0.595 | 0.84 | 0.388 | 0.14 | 0.315 |
| Buryat | 0.761 | 0.133 | 0.552 | 0 | 0.898 | 0.321 | 0.017 | 0.819 | 0.034 | 0.048 | 0.322 | 0.029 | 0.969 | 0.578 |
| Breed | ETH3 | CSSM66 | INRA023 | ILSTS006 | TGLA227 | TGLA126 | TGLA122 | SPS115 | ETH225 | TGLA53 | CSRM60 | BM2113 | BM1824 | ETH10 |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Holst | 0.628 | 0.731 | 0.713 | 0.534 | 0.789 | 0.542 | 0.826 | 0.545 | 0.644 | 0.805 | 0.646 | 0.662 | 0.633 | 0.660 |
| Tagil | 0.646 | 0.803 | 0.668 | 0.726 | 0.850 | 0.564 | 0.856 | 0.603 | 0.723 | 0.851 | 0.632 | 0.783 | 0.695 | 0.721 |
| Kholm | 0.710 | 0.736 | 0.788 | 0.727 | 0.774 | 0.705 | 0.533 | 0.700 | 0.550 | 0.869 | 0.633 | 0.728 | 0.637 | 0.592 |
| RedGor | 0.704 | 0.788 | 0.725 | 0.737 | 0.787 | 0.558 | 0.757 | 0.399 | 0.771 | 0.804 | 0.637 | 0.759 | 0.745 | 0.601 |
| BrSwis | 0.590 | 0.771 | 0.492 | 0.723 | 0.830 | 0.462 | 0.552 | 0.471 | 0.785 | 0.862 | 0.638 | 0.682 | 0.636 | 0.676 |
| Grey | 0.621 | 0.627 | 0.776 | 0.672 | 0.519 | 0.544 | 0.626 | 0.481 | 0.450 | 0.581 | 0.696 | 0.769 | 0.571 | 0.505 |
| Yaros | 0.671 | 0.784 | 0.693 | 0.683 | 0.737 | 0.579 | 0.746 | 0.260 | 0.733 | 0.661 | 0.763 | 0.705 | 0.608 | 0.617 |
| Kostr | 0.445 | 0.822 | 0.629 | 0.666 | 0.804 | 0.386 | 0.655 | 0.541 | 0.649 | 0.784 | 0.571 | 0.745 | 0.559 | 0.571 |
| Altai | 0.462 | 0.642 | 0.725 | 0.528 | 0.784 | 0.467 | 0.666 | 0.637 | 0.680 | 0.881 | 0.726 | 0.723 | 0.677 | 0.684 |
| Kalm | 0.750 | 0.806 | 0.832 | 0.736 | 0.806 | 0.692 | 0.746 | 0.630 | 0.780 | 0.871 | 0.769 | 0.843 | 0.659 | 0.614 |
| AulieAt | 0.710 | 0.851 | 0.705 | 0.734 | 0.852 | 0.550 | 0.635 | 0.645 | 0.719 | 0.833 | 0.734 | 0.789 | 0.687 | 0.771 |
| Alatau | 0.720 | 0.820 | 0.705 | 0.734 | 0.862 | 0.589 | 0.803 | 0.555 | 0.738 | 0.874 | 0.638 | 0.806 | 0.704 | 0.718 |
| KyrgBT | 0.575 | 0.838 | 0.755 | 0.722 | 0.844 | 0.586 | 0.728 | 0.572 | 0.683 | 0.806 | 0.734 | 0.774 | 0.676 | 0.604 |
| KyrgNat | 0.662 | 0.800 | 0.690 | 0.726 | 0.852 | 0.568 | 0.827 | 0.653 | 0.743 | 0.882 | 0.635 | 0.822 | 0.700 | 0.653 |
| Yakut | 0.563 | 0.629 | 0.371 | 0.675 | 0.688 | 0.667 | 0.455 | 0.303 | 0.651 | 0.538 | 0.482 | 0.627 | 0.240 | 0.540 |
| Khogor | 0.597 | 0.782 | 0.632 | 0.559 | 0.791 | 0.536 | 0.812 | 0.581 | 0.544 | 0.794 | 0.812 | 0.668 | 0.411 | 0.550 |
| Gobi | 0.677 | 0.698 | 0.717 | 0.729 | 0.858 | 0.547 | 0.687 | 0.619 | 0.716 | 0.838 | 0.808 | 0.762 | 0.669 | 0.600 |
| Buryat | 0.728 | 0.768 | 0.810 | 0.709 | 0.856 | 0.632 | 0.804 | 0.691 | 0.706 | 0.911 | 0.727 | 0.765 | 0.660 | 0.658 |
| Overall | 0.740 | 0.837 | 0.790 | 0.763 | 0.870 | 0.643 | 0.822 | 0.647 | 0.759 | 0.903 | 0.756 | 0.817 | 0.706 | 0.692 |
| Breed | Holst | Tagil | Kholm | RedGor | BrSwis | Grey | Yaros | Kostr | Altai | Kalm | AulieAt | Alatau | KyrgBT | KyrgNat | Yakut | Khogor | Gobi |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tagil | 0.0163 | ||||||||||||||||
| Kholm | 0.0333 | 0.0325 | |||||||||||||||
| RedGor | 0.033 | 0.0268 | 0.0417 | ||||||||||||||
| BrSwis | 0.0329 | 0.0321 | 0.0558 | 0.0431 | |||||||||||||
| Grey | 0.047 | 0.0628 | 0.0665 | 0.0779 | 0.0876 | ||||||||||||
| Yaros | 0.0498 | 0.0407 | 0.0583 | 0.0468 | 0.0587 | 0.0922 | |||||||||||
| Kostr | 0.0229 | 0.0367 | 0.0567 | 0.0416 | 0.0319 | 0.087 | 0.0654 | ||||||||||
| Altai | 0.0268 | 0.0359 | 0.0522 | 0.0372 | 0.0426 | 0.0836 | 0.0588 | 0.0489 | |||||||||
| Kalm | 0.0268 | 0.0155 | 0.0322 | 0.0291 | 0.0306 | 0.0652 | 0.0341 | 0.0351 | 0.0323 | ||||||||
| AulieAt | 0.0208 | 0.0182 | 0.0312 | 0.0308 | 0.0311 | 0.0687 | 0.0476 | 0.0275 | 0.0348 | 0.0204 | |||||||
| Alatau | 0.0235 | 0.0193 | 0.0398 | 0.0283 | 0.0276 | 0.071 | 0.0508 | 0.0244 | 0.025 | 0.0163 | 0.0198 | ||||||
| KyrgBT | 0.0301 | 0.0281 | 0.0472 | 0.0311 | 0.0304 | 0.0658 | 0.0587 | 0.0231 | 0.0298 | 0.028 | 0.0272 | 0.0136 *** | |||||
| KyrgNat | 0.0267 | 0.0198 | 0.0402 | 0.031 | 0.0291 | 0.0696 | 0.0535 | 0.0271 | 0.0291 | 0.0156 | 0.0204 | 0.0049* | 0.0147 | ||||
| Yakut | 0.0634 | 0.0796 | 0.0953 | 0.1 | 0.1161 | 0.1459 | 0.1127 | 0.1527 | 0.1275 | 0.0761 | 0.0913 | 0.0912 | 0.1102 | 0.0803 | |||
| Khogor | 0.0512 | 0.0409 | 0.0645 | 0.059 | 0.0762 | 0.1074 | 0.0667 | 0.0815 | 0.0642 | 0.0352 | 0.0523 | 0.0503 | 0.0695 | 0.0487 | 0.0763 | ||
| Gobi | 0.0338 | 0.0238 | 0.05 | 0.0342 | 0.0525 | 0.066 | 0.0531 | 0.0494 | 0.0369 | 0.0209 | 0.0328 | 0.0266 | 0.0391 | 0.0251 | 0.0675 | 0.0299 | |
| Buryat | 0.0385 | 0.0087 * | 0.0173 | 0.0188 | 0.0218 | 0.0317 | 0.0242 | 0.0127 ** | 0.0118 ** | 0.0073 * | 0.0108 ** | 0.0097 * | 0.0165 | 0.0094 * | 0.0231 | 0.0136 ** | 0.0089 * |
| Holst | Tagil | Kholm | RedGor | BrSwis | Grey | Yaros | Kostr | Altai | Kalm | AulieAt | Alatau | KyrgBT | KyrgNat | Yakut | Khogor | Gobi | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tagil | 18.5 | ||||||||||||||||
| Kholm | 19.5 | 20 | |||||||||||||||
| RedGor | 18.5 | 21 | 22 | ||||||||||||||
| BrSwis | 19.5 | 25 | 21 | 21 | |||||||||||||
| Grey | 29.5 * | 30 * | 28 * | 31 * | 25 | ||||||||||||
| Yaros | 24.5 | 25 | 23 | 22 | 23 | 31 ** | |||||||||||
| Kostr | 25 | 24.5 | 18.5 | 23.5 | 18.5 | 26.5 * | 22.5 | ||||||||||
| Altai | 24.5 | 29 * | 22 | 23 | 23 | 27 * | 23 | 21.5 | |||||||||
| Kalm | 29.5 * | 24 | 24 | 21 | 31 ** | 35 ** | 29 * | 28.5 * | 27 * | ||||||||
| AulieAt | 23.5 | 24 | 29 * | 22 | 32 ** | 33 ** | 29 * | 31.5 ** | 30 ** | 25 * | |||||||
| Alatau | 26 | 25.5 | 27.5 * | 23.5 | 29.5* | 36.5** | 27.5* | 27 * | 28.5 ** | 16.5 | 21.5 | ||||||
| KyrgBT | 24.5 | 19 | 17 | 22 | 26 * | 33 ** | 25 * | 23.5 | 26 * | 23 | 23 | 19.5 | |||||
| KyrgNat | 28 | 23.5 | 25.5 | 19.5 | 27.5 * | 36.5 ** | 29.5 * | 26 * | 27.5 * | 19.5 | 23.5 | 14 | 19.5 | ||||
| Yakut | 30 * | 29.5 * | 23.5 | 30.5 * | 25.5 | 30.5 ** | 24.5 | 27 ** | 26.5 * | 37.5 ** | 39.5 ** | 39 ** | 32.5 ** | 39 ** | |||
| Khogor | 28 | 26.5 | 27.5 * | 26.5 | 25.5 | 31.5 * | 26.5 * | 28 * | 28.5 ** | 30.5 ** | 27.5 ** | 28 ** | 26.5 ** | 28 ** | 29 ** | ||
| Gobi | 26.5 | 24 | 22 | 20 | 22 | 30 * | 25 | 20.5 | 20 | 23 | 28 ** | 22.5 | 23 | 22.5 | 26.5 * | 20.5 | |
| Buryat | 34.5 * | 29 | 35 * | 32 * | 41 ** | 45 ** | 38 ** | 38.5 ** | 37 ** | 25 | 27 * | 24.5 | 30 ** | 26.5 * | 48.5 ** | 36.5 ** | 33 ** |
| Holst | Tagil | Kholm | RedGor | BrSwis | Grey | Yaros | Kostr | Altai | Kalm | AulieAt | Alatau | KyrgBT | KyrgNat | Yakut | Khogor | Gobi | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Tagil | 0.2259 | ||||||||||||||||
| Kholm | 0.1895 | 0.2357 | |||||||||||||||
| RedGor | 0.2431 | 0.2536 | 0.2524 | ||||||||||||||
| BrSwis | 0.2043 | 0.235 | 0.218 | 0.2369 | |||||||||||||
| Grey | 0.2067 | 0.2144 | 0.2264 | 0.238 | 0.2058 | ||||||||||||
| Yaros | 0.2984 | 0.2935 | 0.2908 | 0.2892 | 0.2976 | 0.2856 | |||||||||||
| Kostr | 0.262 | 0.2639 | 0.2349 | 0.2523 | 0.2555 | 0.2679 | 0.315 | ||||||||||
| Altai | 0.1809 | 0.2156 | 0.2069 | 0.2467 | 0.1926 | 0.205 | 0.2992 | 0.27 | |||||||||
| Kalm | 0.2438 | 0.2828 | 0.2514 | 0.2823 | 0.2452 | 0.2296 | 0.3215 | 0.2706 | 0.248 | ||||||||
| AulieAt | 0.234 | 0.2288 | 0.2329 | 0.2441 | 0.228 | 0.244 | 0.2853 | 0.239 | 0.2264 | 0.2806 | |||||||
| Alatau | 0.2162 | 0.23 | 0.2392 | 0.2202 | 0.2379 | 0.2228 | 0.29 | 0.2848 | 0.2192 | 0.2882 | 0.2198 | ||||||
| KyrgBT | 0.1765 | 0.2322 | 0.2001 | 0.2311 | 0.2048 | 0.2171 | 0.284 | 0.2501 | 0.1978 | 0.2512 | 0.1984 | 0.1989 | |||||
| KyrgNat | 0.1602 | 0.2276 | 0.2023 | 0.2363 | 0.2017 | 0.2029 | 0.3028 | 0.2717 | 0.1854 | 0.2465 | 0.2269 | 0.2129 | 0.1799 | ||||
| Yakut | 0.2096 | 0.2333 | 0.2048 | 0.2288 | 0.2278 | 0.2166 | 0.2956 | 0.2276 | 0.2075 | 0.2601 | 0.2198 | 0.2367 | 0.2053 | 0.2066 | |||
| Khogor | 0.2155 | 0.2486 | 0.1971 | 0.2333 | 0.2017 | 0.2199 | 0.2847 | 0.2098 | 0.2058 | 0.2447 | 0.1971 | 0.2294 | 0.1913 | 0.2091 | 0.195 | ||
| Gobi | 0.3141 | 0.3047 | 0.3102 | 0.3118 | 0.2991 | 0.2775 | 0.3447 | 0.3185 | 0.3081 | 0.3132 | 0.2747 | 0.3092 | 0.2982 | 0.3095 | 0.297 | 0.2768 | |
| Buryat | 0.2417 | 0.2396 | 0.2509 | 0.2592 | 0.2494 | 0.245 | 0.3089 | 0.2684 | 0.238 | 0.2763 | 0.2459 | 0.2413 | 0.2271 | 0.2584 | 0.233 | 0.2408 | 0.2895 |
References
- Kashi, Y.; Tikochinsky, Y.; Genislav, E.; Lraqi, F.; Nave, A.; Beckmann, J.S.; Gruenbaum, Y.; Soller, M. Large restriction fragments containing poly-TG are highly polymorphic in a variety of vertebrates. Nucleic Acids Res. 1990, 18, 1129–1132. [Google Scholar] [CrossRef] [PubMed]
- Tautz, D.; Trick, M.; Dover, G.A. Cryptic simplicity in DNA is a major source of genetic variation. Nature 1986, 322, 652–656. [Google Scholar] [CrossRef] [PubMed]
- Litt, M.; Luty, J.A. A Hypervariable Microsatellite Revealed by Invitro Amplification of a Dinucleotide Repeat within the Cardiac-Muscle Actin Gene. Am. J. Hum. Genet. 1989, 44, 397–401. [Google Scholar] [PubMed]
- Tautz, D. Hypervariability of simple sequences as a general source for polymorphic DNA markers. Nucleic Acids Res. 1989, 17, 6463–6471. [Google Scholar] [CrossRef]
- Vargas, J.; Landi, V.; Martínez, A.; Gómez, M.; Camacho, M.E.; Álvarez, L.Á.; Aguirre, L.; Delgado, J.V. Molecular Study of the Amazonian Macabea Cattle History. PLoS ONE 2016, 11, e0165398. [Google Scholar] [CrossRef]
- Goldstein, D.B.; Linares, A.R.; Cavalli-Sforza, L.L.; Feldman, M.W. Genetic absolute dating based on microsatellites and the origin of modern humans. Proc. Natl. Acad. Sci. USA 1995, 92, 6723–6727. [Google Scholar] [CrossRef]
- Hillel, J.; Groenen, M.A.M.; Tixier-Bochard, M.; Korol, A.B.; David, L.; Kirzhner, V.M.; Burke, T.; Barre-Dirie, A.; Crooijmans, R.P.M.A.; Elo, K.; et al. Biodiversity of 52 chicken populations assessed by microsatellite typing of DNA pools. Genet. Sel. Evol. 2003, 35, 533–557. [Google Scholar] [CrossRef]
- Kantanen, J.; Brooks, M. Genetic diversity and population structure of 20 north European cattle breeds. J. Hered. 2000, 91, 446–457. [Google Scholar] [CrossRef]
- SanCristobal, M.; Chevalet, C.; Haley, C.S.; Joosten, R.; Rattink, A.P.; Harlizius, B.; Groenen, M.A.M.; Amigues, Y.; Boscher, M.-Y.; Russell, G.; et al. Genetic diversity within and between European pig breeds using microsatellite markers. Anim. Genet. 2006, 37, 189–198. [Google Scholar] [CrossRef]
- Tapio, I.; Värv, S.; Bennewitz, J.; Maleviciute, J.; Fimland, E.; Grislis, Z.; Meuwissen, T.; Miceikiene, I.; Olsaker, I.; Viinalass, H.; et al. Prioritization for Conservation of Northern European Cattle Breeds Based on Analysis of Microsatellite Data. Conserv. Biol. 2006, 20, 1768–1779. [Google Scholar] [CrossRef]
- Cortés, A.V.; Dünner, S.; Gama, L.T.; Martínez, A.; Delgado, J.; Ginja, C.; Jiménez, L.; Jordana, J.; Luís, C.; Oom, M.D.M.; et al. The legacy of Columbus in American horse populations assessed by microsatellite markers. J. Anim. Breed. Genet. 2017, 134, 340–350. [Google Scholar] [CrossRef] [PubMed]
- Bulut, Z.; Kurar, E.; Ozsensoy, Y.; Altunok, V.; Nizamlioglu, M. Genetic Diversity of Eight Domestic Goat Populations Raised in Turkey. BioMed Res. Int. 2016, 2016, 1–6. [Google Scholar] [CrossRef]
- Montenegro, M.; Llambí, S.; Castro, G.; Barlocco, N.; Vadell, A.; Landi, V.; Delgado, J.; Martinez, A. Genetic characterization of Uruguayan Pampa Rocha pigs with microsatellite markers. Genet. Mol. Biol. 2014, 38, 48–54. [Google Scholar] [CrossRef] [PubMed]
- Barker, S.J.; Bradley, D.G.; Fries, R. An Integrated Global Program to Establish the Genetic Relationships among the Breeds of Each Domestic Animals Species. In Animal Health and Production Division; FAO: Rome, Italy, 1993. [Google Scholar]
- Cymbron, T.; Freeman, A.R.; Malheiro, M.I.; Vigne, J.-D.; Bradley, D.G. Microsatellite diversity suggests different histories for Mediterranean and Northern European cattle populations. Proc. R. Soc. B Biol. Sci. 2005, 272, 1837–1843. [Google Scholar] [CrossRef] [PubMed]
- Hanotte, O. African Pastoralism: Genetic Imprints of Origins and Migrations. Science 2002, 296, 336–339. [Google Scholar] [CrossRef] [PubMed]
- Metta, M.; Kanginakudru, S.; Gudiseva, N.; Nagaraju, J. Genetic characterization of the Indian cattle breeds, Ongole and Deoni (Bos indicus), using microsatellite markers—A preliminary study. BMC Genet. 2004, 5, 16. [Google Scholar] [CrossRef]
- Nagarajan, M.; Kumar, N.; Nishanth, G.; Haribaskar, R.; Paranthaman, K.; Gupta, J.; Mishra, M.; Vaidhegi, R.; Kumar, S.; Ranjan, A.; et al. Microsatellite markers of water buffalo, Bubalus bubalis—Development, characterisation and linkage disequilibrium studies. BMC Genet. 2009, 10, 68. [Google Scholar] [CrossRef]
- Nishimaki, T.; Ibi, T.; Tanabe, Y.; Miyazaki, Y.; Kobayashi, N.; Matsuhashi, T.; Akiyama, T.; Yoshida, E.; Imai, K.; Matsui, M.; et al. The assessment of genetic diversity within and among the eight subpopulations of Japanese Black cattle using 52 microsatellite markers. Anim. Sci. J. 2013, 84, 585–591. [Google Scholar] [CrossRef]
- Sanarana, Y.; Visser, C.; Bosman, L.; Nephawe, K.; Maiwashe, A.; Van Marle-Köster, E. Genetic diversity in South African Nguni cattle ecotypes based on microsatellite markers. Trop. Anim. Health Prod. 2015, 48, 379–385. [Google Scholar] [CrossRef]
- Sharma, R.; Kishore, A.; Mukesh, M.; Ahlawat, S.; Maitra, A.; Pandey, A.K.; Tantia, M.S. Genetic diversity and relationship of Indian cattle inferred from microsatellite and mitochondrial DNA markers. BMC Genet. 2015, 16, 73. [Google Scholar] [CrossRef]
- Suh, S.; Kim, Y.-S.; Cho, C.-Y.; Byun, M.-J.; Choi, S.-B.; Ko, Y.-G.; Lee, C.W.; Jung, K.-S.; Bae, K.H.; Kim, J.-H. Assessment of Genetic Diversity, Relationships and Structure among Korean Native Cattle Breeds Using Microsatellite Markers. Asian Australas. J. Anim. Sci. 2014, 27, 1548–1553. [Google Scholar] [CrossRef] [PubMed]
- Zhao, J.; Zhu, C.; Xu, Z.; Jiang, X.; Yang, S.-M.; Chen, A. Microsatellite markers for animal identification and meat traceability of six beef cattle breeds in the Chinese market. Food Control. 2017, 78, 469–475. [Google Scholar] [CrossRef]
- Qi, X.B.; Han, J.-L.; Wang, G.; Rege, J.E.O.; Hanotte, O. Assessment of cattle genetic introgression into domestic yak populations using mitochondrial and microsatellite DNA markers. Anim. Genet. 2010, 41, 242–252. [Google Scholar] [CrossRef] [PubMed]
- Fan, B.; Du, Z.-Q.; Gorbach, D.M.; Rothschild, M.F. Development and Application of High-density SNP Arrays in Genomic Studies of Domestic Animals. Asian Australas. J. Anim. Sci. 2010, 23, 833–847. [Google Scholar] [CrossRef]
- Edea, Z.; Dadi, H.; Dessie, T.; Uzzaman, M.R.; Rothschild, M.F.; Kim, E.-S.; Sonstegard, T.S.; Kim, K.-S. Genome-wide scan reveals divergent selection among taurine and zebu cattle populations from different regions. Anim. Genet. 2018, 49, 550–563. [Google Scholar] [CrossRef] [PubMed]
- Yurchenko, A.A.; Yudin, N.; Aitnazarov, R.; Plyusnina, A.; Brukhin, V.; Soloshenko, V.; Lhasaranov, B.; Popov, R.; Paronyan, I.A.; Plemyashov, K.V.; et al. Genome-wide genotyping uncovers genetic profiles and history of the Russian cattle breeds. Hereditary 2017, 120, 125–137. [Google Scholar] [CrossRef]
- Baruch, E.; Weller, J.I. Estimation of the number of SNP genetic markers required for parentage verification. Anim. Genet. 2008, 39, 474–479. [Google Scholar] [CrossRef]
- Honda, T.; Katsuta, T.; Mukai, F. Simulation Study on Parentage Analysis with SNPs in the Japanese Black Cattle Population. Asian Australas. J. Anim. Sci. 2009, 22, 1351–1358. [Google Scholar] [CrossRef]
- Agrafioti, I.; Stumpf, M.P. SNPSTR: A database of compound microsatellite-SNP markers. Nucleic Acids Res. 2007, 35, D71–D75. [Google Scholar] [CrossRef]
- Yu, G.; Tang, Q.; Long, K.; Che, T.; Li, M.; Shuai, S. Effectiveness of microsatellite and single nucleotide polymorphism markers for parentage analysis in European domestic pigs. Genet. Mol. Res. 2015, 14, 1362–1370. [Google Scholar] [CrossRef]
- Chistiakov, D.A.; Hellemans, B.; Volckaert, F.A. Microsatellites and their genomic distribution, evolution, function and applications: A review with special reference to fish genetics. Aquaculture 2006, 255, 1–29. [Google Scholar] [CrossRef]
- Ellegren, H. Microsatellites: Simple sequences with complex evolution. Nat. Rev. Genet. 2004, 5, 435–445. [Google Scholar] [CrossRef] [PubMed]
- Selkoe, K.A.; Toonen, R.J. Microsatellites for ecologists: A practical guide to using and evaluating microsatellite markers. Ecol. Lett. 2006, 9, 615–629. [Google Scholar] [CrossRef] [PubMed]
- Kushnir, A.V.; Глaзкo, T. Gray Ukrainian cattle and their closely related forms. Contemp. Probl. Ecol. 2009, 2, 288–295. [Google Scholar] [CrossRef]
- Kantanen, J.; Edwards, C.J.; Bradley, D.G.; Viinalass, H.; Thessler, S.; Ivanova, Z.; Kiselyova, T.; Ćinkulov, M.; Popov, R.; Stojanović, S.; et al. Maternal and paternal genealogy of Eurasian taurine cattle (Bos taurus). Hereditary 2009, 103, 404–415. [Google Scholar] [CrossRef]
- Granberg, L.; Soini, K.; Osva, A.; Kantanen, J. A New Millennium for the Yakutian Cattle. Sakha Ynaga: Cattle of the Yakuts; University of Helsinki: Helsinki, Finland, 2009; Volume 355, pp. 189–197. [Google Scholar]
- Soini, K.; Ovaska, U.; Kantanen, J. Spaces of Conservation of Local Breeds: The Case of Yakutian Cattle. Sociol. Rural. 2012, 52, 170–191. [Google Scholar] [CrossRef]
- Li, M.-H.; Kantanen, J. Genetic structure of Eurasian cattle (Bos taurus) based on microsatellites: Clarification for their breed classification. Anim. Genet. 2010, 41, 150–158. [Google Scholar] [CrossRef]
- Ruzina, M.N.; Shtyfurko, T.A.; Mohammadabadi, M.; Gendzhieva, O.B.; Tsedev, T.; Sulimova, G.E. Polymorphism of the BoLA-DRB3 gene in the Mongolian, Kalmyk, and Yakut cattle breeds. Russ. J. Genet. 2010, 46, 456–463. [Google Scholar] [CrossRef]
- Porter, V.; Anderson, L.; Hall, S.J.; Sponenberg, D.P. Mason’s World Encyclopedia of Livestock Breeds and Breeding; CABI: Wallingford, UK, 2016; Volume 1. [Google Scholar]
- Ernst, L.; Dmitriev, N. 1. CATTLE (excluding zebus). Anim. Gen. Res. USSR 1989, 65, 1. [Google Scholar]
- Van De Goor, L.H.P.; Panneman, H.; Van Haeringen, W.A. A proposal for standardization in forensic bovine DNA typing: Allele nomenclature of 16 cattle-specific short tandem repeat loci. Anim. Genet. 2009, 40, 630–636. [Google Scholar] [CrossRef]
- Jombart, T. Adegenet: A R package for the multivariate analysis of genetic markers. Bioinformatics 2008, 24, 1403–1405. [Google Scholar] [CrossRef] [PubMed]
- Keenan, K.; McGinnity, P.; Cross, T.F.; Crozier, W.W.; Prodöhl, P. diveRsity: AnRpackage for the estimation and exploration of population genetics parameters and their associated errors. Methods Ecol. Evol. 2013, 4, 782–788. [Google Scholar] [CrossRef]
- Gruber, B.; Adamack, A. Landgenreport: A new r function to simplify landscape genetic analysis using resistance surface layers. Mol. Ecol. Resour. 2015, 15, 1172–1178. [Google Scholar] [CrossRef] [PubMed]
- Adamack, T.A.; Gruber, B. PopGenReport: Simplifying basic population genetic analyses in R. Methods Ecol. Evol. 2014, 5, 384–387. [Google Scholar] [CrossRef]
- Mousadik, E.A.; Petit, R.J. High level of genetic differentiation for allelic richness among populations of the argan tree [Argania spinosa (L.) Skeels] endemic to Morocco. Theor. Appl. Genet. 1996, 92, 832–839. [Google Scholar] [CrossRef]
- Botstein, D.; White, R.L.; Skolnick, M.; Davis, R.W. Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am. J. Hum. Genet. 1980, 32, 314–331. [Google Scholar]
- Clark, L.V.; Jasieniuk, M. polysat: An R package for polyploid microsatellite analysis. Mol. Ecol. Resour. 2011, 11, 562–566. [Google Scholar] [CrossRef]
- Goudet, J. hierfstat, a package for r to compute and test hierarchical F-statistics. Mol. Ecol. Notes 2005, 5, 184–186. [Google Scholar] [CrossRef]
- Caballero, A.; Rodríguez-Ramilo, S.T.; Ávila, V.; Fernández, J. Management of genetic diversity of subdivided populations in conservation programmes. Conserv. Genet. 2009, 11, 409–419. [Google Scholar] [CrossRef]
- Caballero, A.; Rodríguez-Ramilo, S.T. A new method for the partition of allelic diversity within and between subpopulations. Conserv. Genet. 2010, 11, 2219–2229. [Google Scholar] [CrossRef]
- López-Cortegano, E.; Pérez-Figueroa, A.; Caballero, A. metapop 2: Re-implementation of software for the analysis and management of subdivided populations using gene and allelic diversity. Mol. Ecol. Resour. 2019, 19, 1095–1100. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, J.K.; Stephens, M.; Donnelly, P. Inference of population structure using multilocus genotype data. Genetics 2000, 155, 945–959. [Google Scholar] [PubMed]
- Evanno, G.; Regnaut, S.; Goudet, J. Detecting the number of clusters of individuals using the software structure: A simulation study. Mol. Ecol. 2005, 14, 2611–2620. [Google Scholar] [CrossRef] [PubMed]
- Earl, D.A.; Vonholdt, B.M. Structure Harvester: A website and program for visualizing Structure output and implementing the Evanno method. Conserv. Genet. Resour. 2011, 4, 359–361. [Google Scholar] [CrossRef]
- Nei, M. Analysis of Gene Diversity in Subdivided Populations. Proc. Natl. Acad. Sci. USA 1973, 70, 3321–3323. [Google Scholar] [CrossRef]
- Kamvar, Z.N.; Tabima, J.F.; Grünwald, N.J. Poppr: An R package for genetic analysis of populations with clonal, partially clonal, and/or sexual reproduction. PeerJ 2014, 2, 281. [Google Scholar] [CrossRef]
- Comps, B.; Gömöry, D.; Letouzey, J.; Thiébaut, B.; Petit, R.J. Diverging trends between heterozygosity and allelic richness during postglacial colonization in the European beech. Genetics 2001, 157, 389–397. [Google Scholar]
- Nei, M.; Maruyama, T.; Chakraborty, R. The bottleneck effect and genetic variability in populations. Evolution 1975, 29, 1–10. [Google Scholar] [CrossRef]
- Weitzman, M.L. The Noah’s ark problem. Econometrica 1998, 66, 1279–1298. [Google Scholar] [CrossRef]
- Foulley, J.-L.; Ollivier, L. Estimating allelic richness and its diversity. Livest. Sci. 2006, 101, 150–158. [Google Scholar] [CrossRef]
- Weir, S.B.; Cockerham, C.C. Estimating F-Statistics for the Analysis of Population Structure. Evolution 1984, 38, 1358–1370. [Google Scholar]
- Felius, M. Cattle Breeds: An Encyclopedia, 1st ed.; Trafalgar Square Books: North Pomfret, VT, USA, 1995. [Google Scholar]
- Upadhyay, M.; Eriksson, S.; Mikk, S.; Strandberg, E.; Stålhammar, H.; Groenen, M.A.M.; Crooijmans, R.P.M.A.; Andersson, L.; Johansson, A.M. Genomic relatedness and diversity of Swedish native cattle breeds. Genet. Sel. Evol. 2019, 51, 56. [Google Scholar] [CrossRef]
- Mwai, O.; Hanotte, O.; Kwon, Y.-J.; Cho, S. Invited Review—African Indigenous Cattle: Unique Genetic Resources in a Rapidly Changing World. Asian Australas. J. Anim. Sci. 2015, 28, 911–921. [Google Scholar] [CrossRef]
- Barendse, W.; Armitage, S.; Kossarek, L.; Shalom, A.; Kirkpatrick, B.; Ryan, A.; Clayton, D.; Li, L.; Neibergs, H.; Zhang, N.; et al. A genetic linkage map of the bovine genome. Nat. Genet. 1994, 6, 227–235. [Google Scholar] [CrossRef]
- Sunden, S.L.F.; Stone, R.T.; Bishop, M.D.; Kappes, S.M.; Keele, J.W.; Beattie, C.W. A highly polymorphic bovine microsatellite locus: BM2113. Anim. Genet. 2009, 24, 69. [Google Scholar] [CrossRef]
- Moore, S.; Byrne, K.; Berger, K.T.; Barendse, W.; McCarthy, F.; Womack, J.E.; Hetzel, D.J.S. Characterization of 65 bovine microsatellites. Mamm. Genome 1994, 5, 84–90. [Google Scholar] [CrossRef]
- Toldo, S.S.; Fries, R.; Steffen, P.; Neiberg, H.L.; Barendse, W.; Womack, J.E.; Hetzel, D.J.S.; Stranzinger, G. Physically mapped, cosmid-derived microsatellite markers as anchor loci on bovine chromosomes. Mamm. Genome 1993, 4, 720–727. [Google Scholar] [CrossRef]
- Steffen, P.; Eggen, A.; Dietz, A.B.; E Womack, J.; Stranzinger, G.; Fries, R. Isolation and mapping of polymorphic microsatellites in cattle. Anim. Genet. 1993, 24, 121–124. [Google Scholar] [CrossRef]
- Brezinsky, L.; Kemp, S.J.; Teale, A.J. Ilsts005—A Polymorphic Bovine Microsatellite. Anim. Genet. 1993, 24, 73. [Google Scholar] [CrossRef]
- Vaiman, D.; Mercier, D.; Moazami-Goudarzi, K.; Eggen, A.; Ciampolini, R.; Lépingle, A.; Velmala, R.; Kaukinen, J.; Varvio, S.L.; Martin, P. A set of 99 cattle microsatellites: Characterization, synteny mapping, and polymorphism. Mamm. Genome 1994, 5, 288–297. [Google Scholar] [CrossRef]
- Moore, S.S. Dinucleotide polymorphism at the bovine calmodulin independent adenylcyclase locus. Anim. Genet. 2009, 24, 150. [Google Scholar] [CrossRef] [PubMed]
- Georges, M.; Massey, J.M. Polymorphic DNA Markers in Bovidae; W.I.P. Organization Editor: Geneva, Switzerland, 1992. [Google Scholar]
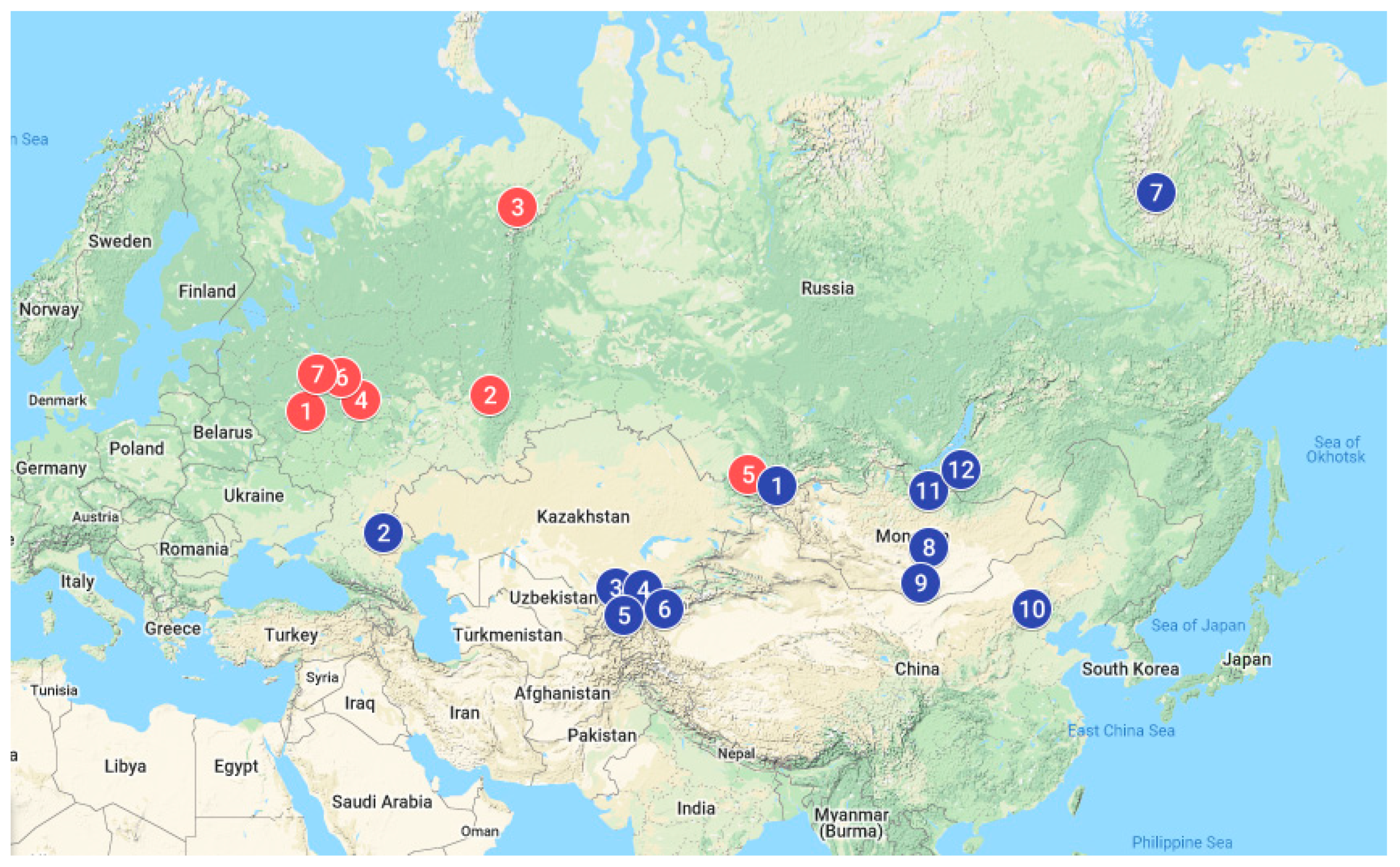

| Breed (Code) | Breeding Purpose | Category | n | Location of Sample | Latitude, Longitude |
|---|---|---|---|---|---|
| European origin | |||||
| Brown Swiss (BrSwis) | Dual | IT * | 50 | Kostroma region, Kostroma district (Russia) | 57.77, 40.93 |
| Holstein (Holst) | Milk | IT | 176 | Moscow region (Russia) | 55.4, 37.27 |
| Kostroma (Kostr) | Dual | RT ** | 20 | Kostroma Region, Kostroma district, (Russia) | 57.77, 40.93 |
| Kholmogory (Kholm) | Milk | RT | 50 | Komi republic, Inta (Russia) | 66.03, 60.17 |
| Yaroslavl (Yaros) | Milk | RT | 50 | Yaroslavl region, Yaroslavl district (Russia) | 57.73, 39.83 |
| Tagil (Tagil) | Milk | Native | 49 | Perm region, Oktyabrsky District (Russia) | 56.51, 57.2 |
| Red Gorbatov (RedGor) | Milk | Native | 50 | Nizhny Novgorod region, Pavlovsky district (Russia) | 56.03, 43.16 |
| Ukrainian Grey (Grey) | Working | Native | 44 | Altai republic, Shebalinsky district (Russia) | 51.34, 85.41 |
| Asian origin | |||||
| Aulie-Ata (AulieAt) | Milk | RT | 42 | Talas region, Talas District, (Kyrgyzstan) | 42.76, 71.41 |
| Alatau (Alatau) | Dual | RT | 49 | Chui region, Zhayilsky district (Kyrgyzstan) | 42.81, 71.41 |
| Kyrgyz Beef-type (KyrgBT) | Meat | Native | 48 | Chui region, Panfilovsky district (Kyrgyzstan) | 42.82, 73.67 |
| Kyrgyz native (KyrgNat) | Dual | Native | 49 | Naryn region, At-Bashinsky District (Kyrgyzstan) | 41.24, 76.13 |
| Yakut (Yakut) | Dual | Native | 30 | Yakutia republic (Russia) | 67.63, 130.86 |
| Altai (Altai) | Dual | Native | 21 | Altai, Ulagan district, Yazula, (Russia) | 50.63, 88.77 |
| Kalmyk (Kalm) | Meat | Native | 54 | Kalmykia republic, Yustinsky district (Russia) | 47.11, 45.97 |
| Khogorogo (Khogor) | Dual | Native | 50 | Khuvsgul aimag (Mongolia) | 46.00, 105.00 |
| Gobi (Gobi) | Milk | Native | 50 | South Gobi aimag (Mongolia) | 43.34, 104.25 |
| Buryat (Buryat) | Dual | Native | 24 | Khuvsgul aimag (Mongolia) | 46.00, 105.00 |
| 10 | Inner Mongolia (China) | 41.13, 116.38 | |||
| 252 | Buryatia Republic, Dzhidinsky District (Russia) | 50.65, 105.22 | |||
| Breed | N | A | % | Ar | Ho | He | Fis | HWE |
|---|---|---|---|---|---|---|---|---|
| European origin | ||||||||
| Brown Swiss | 49.43 | 79 | 42.27 | 5.29 | 0.77 | 0.7 | −1.00 × 10−1 | 0 × 100 |
| Holstein | 176 | 98 | 52.57 | 5.53 | 0.72 | 0.71 | −1.41 × 10−2 | 8.07 × 10−1 |
| Kostroma | 20 | 86 | 45.55 | 5.43 | 0.71 | 0.68 | −4.41 × 10−2 | 6.16 × 10−1 |
| Kholmogory | 49.86 | 95 | 50.9 | 5.79 | 0.73 | 0.73 | 0 × 100 | 9.80 × 10−2 |
| Yaroslavl | 50 | 89 | 48.36 | 5.56 | 0.72 | 0.7 | −2.86 × 10−2 | 6.59 × 10−1 |
| Tagil | 48.21 | 113 | 60.19 | 6.69 | 0.77 | 0.76 | −1.32 × 10−2 | 7.04 × 10−2 |
| Red Gorbatov | 50 | 109 | 58.02 | 6.21 | 0.78 | 0.73 | −6.85 × 10−2 | 9.99 × 10−1 |
| Asian origin | ||||||||
| Aulie−Ata | 40.71 | 129 | 69.11 | 7.21 | 0.75 | 0.76 | 1.32 × 10−2 | 5.70 × 10−3 |
| Alatau | 49 | 132 | 70.13 | 7.52 | 0.76 | 0.76 | 0 × 100 | 4.58 × 10−1 |
| Kyrgyz Beef−type | 48 | 115 | 61.96 | 6.77 | 0.75 | 0.74 | −1.35 × 10−2 | 1.40 × 10−1 |
| Kyrgyz native | 48.93 | 128 | 66.93 | 7.23 | 0.77 | 0.76 | −1.32 × 10−2 | 9.75 × 10−1 |
| Yakut | 30 | 66 | 35.9 | 4.29 | 0.61 | 0.58 | −5.17 × 10−2 | 9.53 × 10−1 |
| Altai | 21 | 87 | 46.71 | 5.57 | 0.71 | 0.7 | −1.43 × 10−2 | 6.60 × 10−1 |
| Kalmyk | 54 | 131 | 69.8 | 7.51 | 0.78 | 0.78 | 0 × 100 | 2.00 × 10−4 |
| Khogorogo | 50 | 98 | 52.38 | 5.8 | 0.69 | 0.69 | 0 × 100 | 4.51 × 10−2 |
| Gobi | 50 | 99 | 52.16 | 6.12 | 0.75 | 0.75 | 0 × 100 | 6.50 × 10−1 |
| Buryat | 285.64 | 155 | 81.51 | 7.36 | 0.77 | 0.78 | 1.28 × 10−2 | 0 × 100 |
| Breed | % * | Locus | Allele | AF |
|---|---|---|---|---|
| Grey (1) ** | 11.1 | BM1824 | 176 | 0.011 |
| Buryat (9) | CSSM66 | 177 | 0.002 | |
| ILSTS006 | 276 | 0.019 | ||
| TGLA227 | 69 | 0.009 | ||
| TGLA227 | 71 | 0.005 | ||
| 15.5 | TGLA122 | 137 | 0.002 | |
| SPS115 | 238 | 0.002 | ||
| ETH225 | 160 | 0.031 | ||
| TGLA53 | 190 | 0.007 | ||
| BM1824 | 192 | 0.011 | ||
| Gobi (1) | 9.7 | TGLA227 | 85 | 0.020 |
| Kalm (2) | 18 | TGLA126 | 129 | 0.028 |
| CSRM60 | 86 | 0.009 | ||
| Khogor (1) | 9.7 | CSRM60 | 88 | 0.040 |
| KyrgNat (2) | 19.9 | CSSM66 | 207 | 0.010 |
| INRA023 | 194 | 0.010 |
| Locus | Fst(se) | Fis(se) | Fit(se) |
|---|---|---|---|
| ILSTS006 | 0.0955(0.0132) | −0.0022 (0.0312) | 0.0935 (0.0306) |
| SPS115 | 0.0896(0.0140) | −0.0019 (0.0336) | 0.0879 (0.0333) |
| BM1824 | 0.0837(0.0111) | 0.0041 (0.0346) | 0.0875 (0.0332) |
| TGLA122 | 0.0821(0.0107) | −0.0137 (0.0257) | 0.0695 (0.0255) |
| ETH3 | 0.0793(0.0108) | −0.0040 (0.0326) | 0.0756 (0.0318) |
| ETH225 | 0.0781(0.0099) | −0.0169 (0.0312) | 0.0625 (0.0297) |
| BM2113 | 0.0762(0.0099) | −0.0050 (0.0288) | 0.0716 (0.0278) |
| TGLA126 | 0.0759 (0.0123) | −0.0086 (0.0374) | 0.0680 (0.0364) |
| CSSM66 | 0.0740 (0.0096) | −0.0185 (0.0258) | 0.0568 (0.0252) |
| TGLA53 | 0.0715 (0.0079) | 0.0036 (0.0222) | 0.0749 (0.0220) |
| INRA023 | 0.0694 (0.0105) | −0.0047 (0.0285) | 0.0651 (0.0282) |
| CSRM60 | 0.0659 (0.0094) | −0.0349(0.0289) | 0.0333 (0.0287) |
| ETH10 | 0.0615 (0.0109) | −0.0215(0.0320) | 0.0413 (0.0317) |
| TGLA227 | 0.0608 (0.0077) | 0.0059(0.0236) | 0.0663 (0.0234) |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Svishcheva, G.; Babayan, O.; Lkhasaranov, B.; Tsendsuren, A.; Abdurasulov, A.; Stolpovsky, Y. Microsatellite Diversity and Phylogenetic Relationships among East Eurasian Bos taurus Breeds with an Emphasis on Rare and Ancient Local Cattle. Animals 2020, 10, 1493. https://doi.org/10.3390/ani10091493
Svishcheva G, Babayan O, Lkhasaranov B, Tsendsuren A, Abdurasulov A, Stolpovsky Y. Microsatellite Diversity and Phylogenetic Relationships among East Eurasian Bos taurus Breeds with an Emphasis on Rare and Ancient Local Cattle. Animals. 2020; 10(9):1493. https://doi.org/10.3390/ani10091493
Chicago/Turabian StyleSvishcheva, Gulnara, Olga Babayan, Bulat Lkhasaranov, Ariuntuul Tsendsuren, Abdugani Abdurasulov, and Yurii Stolpovsky. 2020. "Microsatellite Diversity and Phylogenetic Relationships among East Eurasian Bos taurus Breeds with an Emphasis on Rare and Ancient Local Cattle" Animals 10, no. 9: 1493. https://doi.org/10.3390/ani10091493
APA StyleSvishcheva, G., Babayan, O., Lkhasaranov, B., Tsendsuren, A., Abdurasulov, A., & Stolpovsky, Y. (2020). Microsatellite Diversity and Phylogenetic Relationships among East Eurasian Bos taurus Breeds with an Emphasis on Rare and Ancient Local Cattle. Animals, 10(9), 1493. https://doi.org/10.3390/ani10091493




