Ecological Niche Models of Four Hard Tick Genera (Ixodidae) in Mexico
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
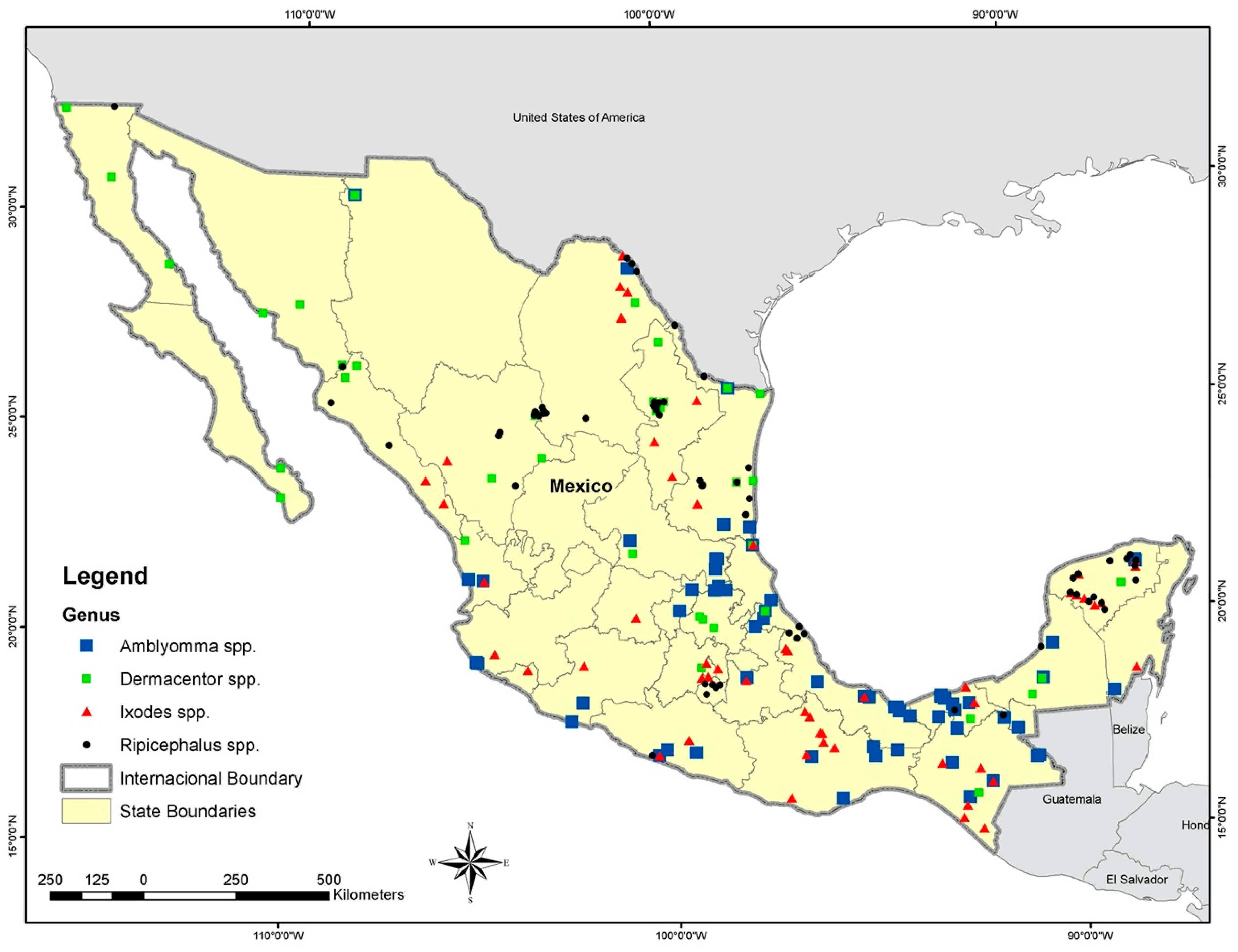
2.1. Tick Occurrence Data
2.2. Pseudoabsence Data
2.3. Environmental Variables
2.4. Ecological Niche Modeling
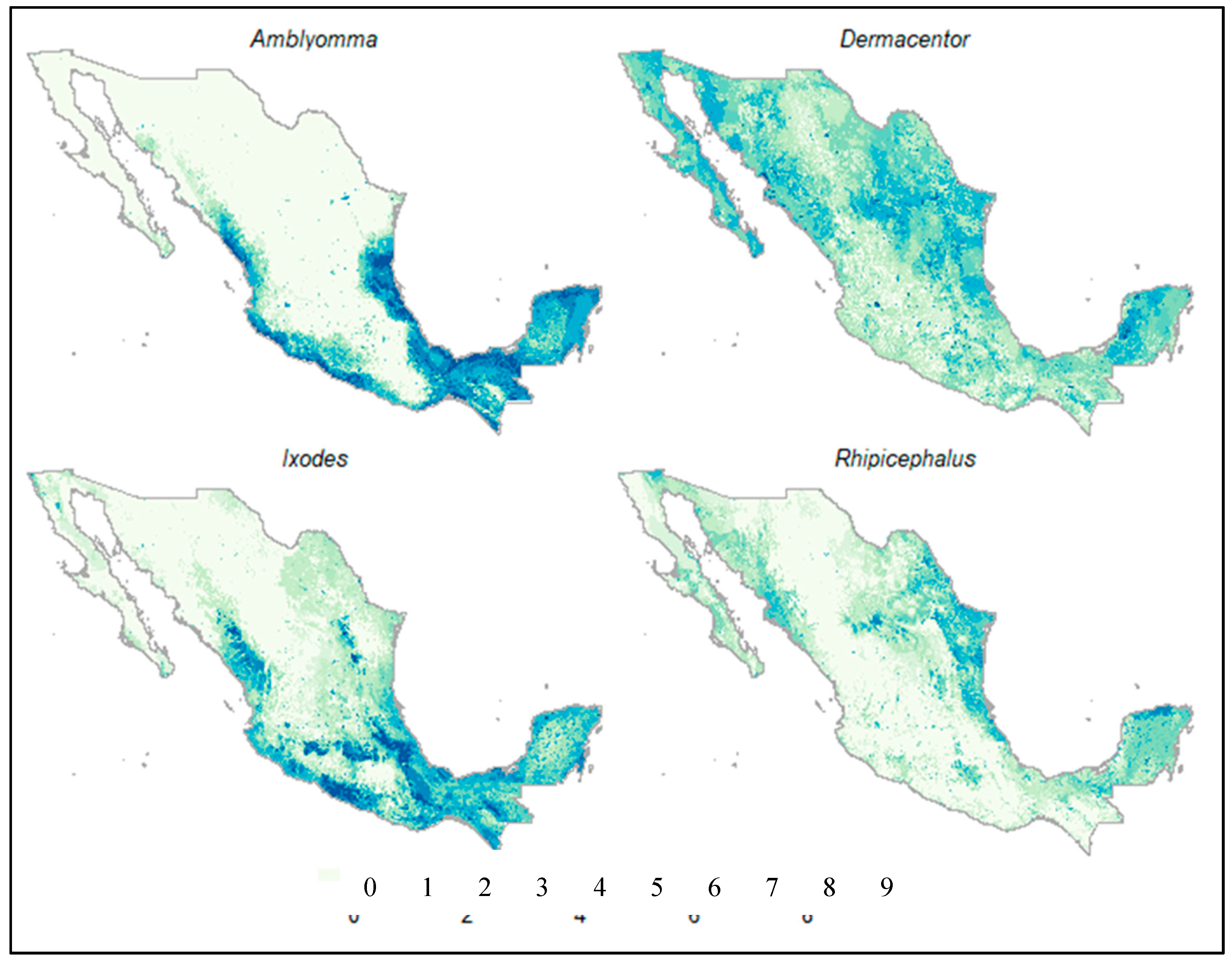
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Tabachnick, W.J. Challenges in predicting climate and environmental effects on vector-borne disease episystems in a changing world. J. Exp. Biol. 2010, 213, 946–954. [Google Scholar] [CrossRef]
- Ramos, N.; Rodrigues, S.; Piovezan, U.; Pablo, M.; Szabó, J. Microhabitat determines uneven distribution of Amblyomma parvum but not of Amblyomma sculptum ticks within forest patches in the Brazilian Pantanal. Exp. Appl. Acarol. 2019, 79, 405–410. [Google Scholar] [CrossRef]
- Peterson, A.T. Ecologic niche modeling and spatial patterns of disease transmission. Emerg. Infect. Dis. 2006, 12, 1822–1826. [Google Scholar] [CrossRef]
- Soberón, J.; Peterson, A.T. Interpretation of models of fundamental ecological niches and species distributional areas. Biodivers. Inf. 2005, 2, 1–10. [Google Scholar] [CrossRef]
- Soberón, J.M. Niche and area of distribution modeling: A population ecology perspective. Ecogr. Cop. 2010, 33, 159–167. [Google Scholar] [CrossRef]
- Rangel, T.F.; Loyola, R.D. Labeling Ecological Niche Models. Nat. Conserv. 2012, 10, 119–126. [Google Scholar] [CrossRef]
- Olden, J.D.; Lawler, J.J.; Poff, N.L. Machine Learning Methods without Tears: A primer for Ecologists. Q. Rev. Biol. 2008, 83, 171–193. [Google Scholar] [CrossRef]
- Parola, P.; Raoult, D. Ticks and tickborne bacterial diseases in humans: An emerging infectious threat. Clin. Infect. Dis. 2001, 32, 897–928. [Google Scholar] [CrossRef] [PubMed]
- Smith, K.F.; Guégan, J.-F. Changing Geographic Distributions of Human Pathogens. Annu. Rev. Ecol. Evol. Syst. 2010, 41, 231–250. [Google Scholar] [CrossRef]
- Jongejan, F.; Uilenberg, G. The global importance of ticks. Parasitology 2004, 129, S3–S14. [Google Scholar] [CrossRef] [PubMed]
- Amsden, J.R.; Warmack, S.; Gubbins, P.O. Tick-borne bacterial, rickettsial, spirochetal, and protozoal infectiouss diseases in the United States: A comprehenssive review. Pharmacotherapy 2005, 25, 191–210. [Google Scholar] [CrossRef]
- Walker, A.R. Ticks and associated diseases: A retrospective review. Med. Vet. Entomol. 2014, 28, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Sprong, H.; Tijsse-Klasen, E.; Langelaar, M.; De Bruin, A.; Fonville, M.; Gassner, F.; Takken, W.; Van Wieren, S.; Nijhof, A.; Jongejan, F.; et al. Prevalence of Coxiella Burnetii in ticks after a large outbreak of Q fever. Zoonoses Public Health 2011, 59, 69–75. [Google Scholar] [CrossRef]
- Chomel, B. Tick-borne infections in dogs-an emerging infectious threat. Vet. Parasitol. 2011, 179, 294–301. [Google Scholar] [CrossRef] [PubMed]
- Dantas-Torres, F. Climate change, biodiversity, ticks and tick-borne diseases: The butterfly effect. Int. J. Parasitol. 2015, 4, 452–461. [Google Scholar] [CrossRef] [PubMed]
- Esteve-Gassent, M.D.; Castro-Arellano, I.; Feria-Arroyo, T.P.; Patino, R.; Li, A.Y.; Medina, R.F.; de León, A.A.P.; Rodríguez-Vivas, R.I. Translating ecology, physiology, biochemistry, and population genetics research to meet the challenge of tick and tick-borne diseases in North America. Arch. Insect Biochem. Physiol. 2016, 92, 38–64. [Google Scholar] [CrossRef]
- Illoldi-rangel, P.; Rivaldi, C.; Sissel, B.; Fryxell, R.T.; Gordillo-Pérez, G.; Rodríguez-Moreno, A.; Williamson, P.; Montiel-Parra, G.; Sánchez-Cordero, V.; Sarkar, S. Species Distribution Models and Ecological Suitability Analysis for Potential Tick Vectors of Lyme Disease in Mexico. J. Trop. Med. 2012, 2012, 959101. [Google Scholar] [CrossRef]
- Feria-Arroyo, T.P.; Castro-Arellano, I.; Gordillo-Perez, G.; Cavazos, A.L.; Vargas-Sandoval, M.; Grover, A.; Torres, J.; Medina, R.F.; de León, A.A.P.; Esteve-Gassent, M.D. Implications of climate change on the distribution of the tick vector Ixodes scapularis and risk for Lyme disease in the Texas-Mexico transboundary region. Parasit. Vectors 2014, 7, 199. [Google Scholar] [CrossRef]
- González, C.; Wang, O.; Strutz, S.E.; González-Salzar, C.; Sánchez-Cordero, V.; Sarkar, S. Climate change and risk of leishmaniasis in North America: Predictions from ecological niche models of vector and reservoir species. PLoS Negl. Trop. Dis. 2010, 4, e585. [Google Scholar] [CrossRef]
- Gurgel-Gonçalves, R.; Galvão, C.; Costa, J.; Peterson, A.T. Geographic distribution of chagas disease vectors in Brazil based on ecological niche modeling. J. Trop. Med. 2012, 2012, 705326. [Google Scholar] [CrossRef]
- Gorla, D.; Noireau, F. Geographic distribution of Triatominae vectors in America. In American Trypanaosomiasis Chagas Disease: One Hundred Years of Research; Telleria, J., Tibayrenc, M., Eds.; Elsevier: Amsterdam, The Netherlands, 2010; pp. 209–231. [Google Scholar]
- Moffett, A.; Strutz, S.; Guda, N.; Gonález, C.; Ferro, M.C.; Sánchez-Cordero, V.; Sarkar, S. A global public database of disease vector and reservoir distributions. PLoS Negl. Trop. Dis. 2009, 3, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Stevens, K.B.; Gilbert, M.; Pfeiffer, D.U. Modeling habitat suitability for occurrence of highly pathogenic avian influenza virus H5N1 in domestic poultry in Asia: A spatial multicriteria decision analysis approach. Spat. Spatio-Tempor. Epidemiol. 2013, 4, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Guzmán-Cornejo, C.; Robbins, R.G. The genus Ixodes (Acari: Ixodidae) in Mexico: Adult identification keys, diagnoses, hosts, and distribution. Rev. Mex. Biodivers. 2010, 81, 289–298. [Google Scholar]
- Guzmán-Cornejo, C.; Robbins, R.G.; Guglielmone, A.A.; Montiel, G.; Pérez, T.M. The Amblyomma (Acari: Ixodida: Ixodidae) of Mexico: Identification keys, distribution and hosts. Zootaxa 2011, 2998, 16–38. [Google Scholar] [CrossRef]
- Guzmán-Cornejo, C.; Robbins, R.G.; Guglielmone, A.A.; Montiel-Parra, G.; Rivas, G.; Pérez, T.M. The Dermacentor (Acari, Ixodida, Ixodidae) of Mexico: Hosts, geographical distribution and new records. Zookeys 2016, 22, 1–22. [Google Scholar] [CrossRef]
- Huberty, C.J. Applied Discriminant Analysis; Wiley-Interscience: New York, NY, USA; University of Michigan: Ann Arbor, MI, USA, 1994; 496p. [Google Scholar]
- Barbet-Massin, M.; Jiguet, F.; Albert, C.H.; Thuiller, W. Selecting pseudo-absences for species distribution models: How, where and how many? Methods Ecol. Evol. 2012, 3, 327–338. [Google Scholar] [CrossRef]
- Di Cola, V.; Broennimann, O.; Petitpierre, B.; Breiner, F.T.; Amen, M.D.; Randin, C.; Engler, R.; Pottier, J.; Pio, D.; Dubuis, A.; et al. ecospat: An R package to support spatial analyses and modeling of species niches and distributions. Ecogr. Cop. 2016, 40, 774–787. [Google Scholar] [CrossRef]
- Jiménez-Valverde, A.; Lobo, J.M.; Hortal, J. The effect of prevalence and its interaction with sample size on the reliability of species distribution models. Community Ecol. 2008, 10, 196–205. [Google Scholar] [CrossRef]
- Stokland, J.N.; Halvorsen, R.; Støa, B. Species distribution modelling—Effect of design and sample size of pseudo-absence observations. Ecol. Model. 2011, 222, 1800–1809. [Google Scholar] [CrossRef]
- Hijmans, R.J.; Cameron, S.E.; Parra, J.L.; Jones, P.G.; Jarvis, A. Very high resolution interpolated climate surfaces for global land areas. Int. J. Clim. 2005, 25, 1965–1978. [Google Scholar] [CrossRef]
- Randolph, S.E. Ecology of non-nidicolous ticks. In Biology of Ticks, 2nd ed.; Sonenshine, D.E., Roe, R.M., Eds.; Oxford University Press: Oxford, UK, 2014; pp. 3–37. [Google Scholar]
- Warren, D.L.; Seifert, S.N. Ecological niche modeling in Maxent: The importance of model complexity and the performance of model. Ecol. Appl. 2011, 21, 335–342. [Google Scholar] [CrossRef] [PubMed]
- Barve, N.; Barve, V.; Jiménez-Valverde, A.; Lira-Noriega, A.; Maher, S.P.; Peterson, A.T.; Soberón, J.; Villalobos, F. The crucial role of the accessible area in ecological niche modeling and species distribution modeling. Ecol. Model. 2011, 222, 1810–1819. [Google Scholar] [CrossRef]
- Naimi, B.; Araújo, M.B. Sdm: A reproducible and extensible R platform for species distribution modelling. Ecogr. Cop. 2016, 39, 368–375. [Google Scholar] [CrossRef]
- Muscarella, R.; Galante, P.J.; Soley-guardia, M.; Boria, R.A.; Kass, J.M.; Anderson, R.P. ENMeval: An R package for conducting spatially independent evaluations and estimating optimal model complexity for MAXENT ecological niche models. Methods Ecol. Evol. 2014, 5, 1198–1205. [Google Scholar] [CrossRef]
- Hastie, T. Generalized Additive Models [Internet]. R Package. 2019. Available online: http://cran.r-project.org/web/packages/gam/gam.pdf (accessed on 8 January 2020).
- Phillips, S.J.; Anderson, R.P.; Schapire, R.E. Maximum entropy modeling of species geographic distributions. Ecol. Model. 2006, 190, 231–259. [Google Scholar] [CrossRef]
- Phillips, S.J.; Ddudík, M. Modeling of species distributions with Maxent: New extensions and a comprehensive evaluation. Echography 2008, 31, 161–175. [Google Scholar] [CrossRef]
- McCullagh, P. Generalized linear models. Eur. J. Oper. Res. 1984, 16, 285–292. [Google Scholar] [CrossRef]
- McCullagh, P.; Nelder, J.A. Generalized Linear Models; Chapman and Hall: Boca Raton, FL, USA, 1989; 532p. [Google Scholar]
- Friedman, H.H. Multivariate adaptive regression splines. Ann. Stat. 1991, 19, 1–141. [Google Scholar] [CrossRef]
- Breiman, L.; Friedman, J.H.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees; Chapman and Hall: Boca Raton, FL, USA, 1984; 386p. [Google Scholar]
- Hastie, T.; Tibshirani, R.; Buja, A. Flexible discriminant analysis by optimal scoring. J. Am. Stat. Assoc. 1994, 89, 1255–1270. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Stat. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]
- Elith, J.; Leathwick, J.R.; Hastie, T. A working guide to boosted regression trees. J. Anim. Ecol. 2008, 77, 802–813. [Google Scholar] [CrossRef] [PubMed]
- Nix, H.A. A biogeographic analysis of Australian Elapid snakes. In Australian Flora and Fauna Series Number 7: Atlas of Elapid Snakes of Australia; Lonngmore, R., Ed.; Australian Goverment Publishing Service: Canberra, Australia, 1986; pp. 4–5. [Google Scholar]
- Booth, T.H.; Nix, H.A.; Busby, J.R.; Hutchinson, M.F. BIOCLIM: The firs species distribution modelling packages, its early applications and relevance to most current MAXENT studies. Divers. Distrib. 2014, 20, 1–9. [Google Scholar] [CrossRef]
- Elith, J.; Graham, C.H.; Anderson, R.P.; Duudík, M.; Ferrier, S.; Guisan, A.; Hijmans, R.J.; Huettmann, F.; Leathwick, J.R.; Lehmann, A.; et al. Novel methods improve prediction of species’ distributions from occurrence data. Ecogr. Cop. 2006, 2, 129–151. [Google Scholar] [CrossRef]
- Guisan, A.; Edwards, T.C.; Hastie, T. Generalized linear and generalized additive models in studies of species distributions: Setting the scene. Ecol. Model. 2002, 157, 89–100. [Google Scholar] [CrossRef]
- Leathwick, J.R.; Elith, J.; Hastie, T. Comparative performance of generalized additive models and multivariate adaptive regression splines for statistical modelling of species distributions. Ecol. Model. 2006, 199, 188–196. [Google Scholar] [CrossRef]
- Hijmans, R.J.; Phillips, S.; Leathwick, J.R.; Elith, J. Package ‘Dismo‘. Available online: https://cran.r-project.org/web/packages/dismo/index.html (accessed on 8 January 2020).
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2013; Available online: http://www.r-project.org/ (accessed on 8 January 2020).
- Swets, J.A. Measuring the accuracy of diagnostic systems. Science 1988, 240, 1285–1293. [Google Scholar] [CrossRef]
- Fiellding, A.H.; Bell, J.F. A review of methods for the assessment of prediction errors in conservation presence/absence models. Environ. Conserv. 1997, 24, 38–49. [Google Scholar] [CrossRef]
- Jiménez-Valverde, A. Insights into the area under the receiver operating characteristic curve (AUC) as a discrimination measure in species distribution modelling. Glob. Biogeogr. 2012, 21, 498–507. [Google Scholar] [CrossRef]
- Guisan, A.; Graham, C.H.; Elith, J.; Huettmann, F. NCEAS Species Distribution Modelling Group. Sensitivity of predictive species distribution models to change in grain size. Divers. Distrib. 2007, 13, 332–340. [Google Scholar] [CrossRef]
- Radosavljevic, A.; Anderson, R.P. Making better Maxent models of species distributions: Complexity, overfitting and evaluation. J. Biogeogr. 2014, 41, 629–643. [Google Scholar] [CrossRef]
- Barker, S.C.; Burger, T.D. Two new genera of hard ticks, Robertsicus n. gen. and Archaeocroton n. gen.; and the solution to the mystery of Hoogstraal’s and Kaufman’s “primitive” tick from the Carpathian Mountains. Zootaxa 2018, 4500, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Estrada-Peña, A.; Guglielmone, A.A.; Mangold, A.J. The distribution and ecological “preferences” of the tick Amblyomma cajennense (Acari: Ixodidae), an ectoparasite of humans and other mammals in the Americas. Ann. Trop. Med. Parasitol. 2004, 98, 283–292. [Google Scholar] [CrossRef]
- Bishopp, F.C.; Hixson, H. Biology and economic importance of the Gulf Coast tick. J. Econ. Entomol. 1936, 29, 1068–1076. [Google Scholar] [CrossRef]
- Goddard, J.; Norment, B.R. Notes on the geographical distribution of the Gulf coast tick, Amblyomma maculatum (Koch) [Acari, Ixodidae]. Entomol. News 1983, 94, 103–104. [Google Scholar]
- Estrada-peña, A.; Tarragona, E.L.; Vesco, U.; Meneghi, D.; De Mastropaolo, M.; Mangold, A.J.; Guglielmone, A.A.; Nava, S. Divergent environmental preferences and areas of sympatry of tick species in the Amblyomma cajennense complex (Ixodidae). Int. J. Parasitol. 2014, 44, 1081–1089. [Google Scholar]
- Pascoe, E.L.; Marcantonio, M.; Caminade, C.; Foley, J.E. Modeling Potential Habitat for Amblyomma Tick Species in California. Insects 2019, 10, 201. [Google Scholar] [CrossRef]
- Strey, O.F.; Teel, P.D.; Longnecker, M.T.; Needham, G.R. Survival and Water-Balance Characteristics of Unfed Adult Amblyomma cajennense (Acari: Ixodidae). J. Med. Entomol. 1996, 33, 63–73. [Google Scholar] [CrossRef]
- Needham, G.R.; Teel, P.D. Off-host physiological ecology of ixodid ticks. Annu. Rev. Entomol. 1991, 36, 659–681. [Google Scholar] [CrossRef]
- Ketchum, H.R.; Teel, P.D.; Coates, C.J.; Strey, O.F.; Longnecker, M.T. Genetic variation in 12S and 16 S mitochondrial r DNA genes of four geographically isolated populations of Gulf Coast ticks (Acari: Ixodidae). J. Med. Entomol. 2009, 46, 482–489. [Google Scholar] [CrossRef]
- Sanders, D.M.; Schuster, A.L.; McCardle, P.W.; Strey, O.F.; Blankenship, T.L.; Teel, P.D. Ixodid ticks associated with feral swine in Texas. J. Vector Ecol. 2013, 38, 361–373. [Google Scholar] [CrossRef] [PubMed]
- Guglielmone, A.A.; Robbins, R.G.; Apanaskevich, D.A.; Petney, T.N.; Estrada-Peña, A.; Hoorak, I.G.; Shao, R.; Barker, S.C. The Argasidae, Ixodidae and Nuttalliellidae (Acari: Ixodida) of the world: A list of valid species names. Zootaxa 2010, 2528, 1–28. [Google Scholar] [CrossRef]
- Keirans, J.E.; Needham, G.R.; Oliver, J.H. The Ixodes (Ixodes) ricinus complex worldwide: Diagnosis of the species in the complex, hosts and distribution. In Acarology IX Symposia; University of Wisconsin-Madison: Madison, WI, USA, 1999; pp. 341–347. [Google Scholar]
- Xu, G.; Fang, Q.Q.; Keirans, J.E.; Durden, L.A. Molecular phylogenetic analyses indicate that the Ixodes ricinus complex is a paraphyletic group. J. Parasitol. 2003, 89, 452–457. [Google Scholar] [CrossRef]
- Beati, L.; Klompen, H. Phylogeography of ticks (Acari: Ixodida). Annu. Rev. Entomol. 2019, 64, 379–397. [Google Scholar] [CrossRef] [PubMed]
- Gray, J.S.; Kahl, O.; Lane, R.S.; Levin, M.L.; Tsao, J.I. Diapause in ticks on the medically important Ixodes ricinus species complex. Ticks Tick-Borne Dis. 2016, 7, 992–1003. [Google Scholar] [CrossRef]
- LoGiudice, K.; Ostfeld, R.S.; Schmidt, K.A.; Keesing, F. The ecology of infectious disease: Effects of host diversity and community composition on Lyme disease risk. Proc. Natl. Acad. Sci. USA 2003, 100, 567–571. [Google Scholar] [CrossRef]
- Eisen, R.J.; Eisen, L.; Lane, R.-S. Habitat related variation in infestation of lizards and rodents with Ixodes ticks in dense woodlands in Mendocino County, California. Exp. Appl. Acarol. 2004, 33, 215–233. [Google Scholar] [CrossRef]
- Salkeld, D.J.; Lane, R.S. Community ecology and disease risk: Lizards, squirrels, and the Lyme disease spirochete in California, USA. Ecology 2010, 91, 293–298. [Google Scholar] [CrossRef]
- Vandyk, J.K.; Bartholomew, D.M.; Rowley, W.A.; Platt, K.B. Survival of Ixodes scapularis (Acari: Ixodidae) exposed to cold. J. Med. Entomol. 1996, 33, 6–10. [Google Scholar] [CrossRef]
- Yoder, J.A.; Chris, B.S.; Croxall, T.J.; Laura, K.S.; Tank, J.L. Moisture requirements for activity/survival of the gulf coast tick Amblyomma maculatum Koch (Acari: Ixodidae), based on a water balance study of all life cycle stages. Int. J. Acarol. 2009, 34, 285–292. [Google Scholar] [CrossRef]
- Wang, F.; Wang, D.; Guo, G.; Hu, Y.; Wei, J.; Liu, J. Species delimitation of the Dermacentor ticks based on phylogenetic clustering and niche modeling. Peer J. 2019, 7, e6911. [Google Scholar] [CrossRef]
- Guglielmone, A.A.; Robbins, R.G.; Apanaskevich, D.A.; Petney, T.N.; Estrada-Peña, A.; Horak, I.G. The Hard Ticks of the World; Springer: New York, NY, USA, 2013; 738p. [Google Scholar]
- Minigan, J.N.; Hager, H.A.; Peregrine, A.S.; Newman, J.A. Current and potential future distribution of the American dog tick (Dermacentor variabilis, Say) in North America. Tick Tick-Borne Dis. 2018, 9, 354–362. [Google Scholar] [CrossRef] [PubMed]
- Süss, J.; Klaus, C.; Gerstengarbe, F.; Werner, P.C. What makes ticks tick? Climate change, ticks, and tick-borne diseases. J. Travel Med. 2008, 15, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Holmes, C.J.; Dobrotka, C.J.; Farrow, D.W.; Rosendale, A.J.; Benoit, J.B.; Pekins, P.J.; Yoder, J.A. Low and high thermal tolerance characteristics for unfed larvae of the winter tick Dermacentor albipictus (Acari: Ixodidae) with special reference to moose. Ticks Tick-Borne Dis. 2018, 9, 25–30. [Google Scholar] [CrossRef] [PubMed]
- Laburna, M.B.; Gerardi, M.; Krawczak, F.S.; Moraes-Filho, J. Comparative biology of the tropical and temperate species of Rhipicephalus sanguineus sensu lato (Acari: Ixodidae) under different laboratory conditions. Ticks Tick-Borne Dis. 2017, 8, 146–156. [Google Scholar] [CrossRef] [PubMed]
- Kessler, W.H.; Ganser, C.; Glass, G.E. Modelling the distribution of medically important tick species in Florida. Insects 2019, 10, 190. [Google Scholar] [CrossRef]
- Guisan, A.; Zimmermann, N. Predictive habitat distribution models in ecology. Ecol. Model. 2000, 135, 147–186. [Google Scholar] [CrossRef]
- Bouchard, C.; Dibernardo, A.; Koffi, J.; Wood, H.; Leighton, P.A.; Lindssay, L.R. Increased risk of tick-borne diseases with climate and environmental changes. Can. Commun. Dis. Rep. 2019, 45, 81–89. [Google Scholar] [CrossRef]
- Marmion, M.; Parviainen, M.; Luoto, M.; Heikkinen, R.K.; Thuiller, W. Evaluation of consensus methods in predictive species distribution modelling. Divers. Distrib. 2009, 15, 59–69. [Google Scholar] [CrossRef]
- Camicas, J.-L.; Hervy, J.-P.; Adam, F.; Morel, P.-C. Les tiques du monde (Acarida, Ixodida). Nomenclature, stades décrits, hotes, répartion; Orstom: Paris, France, 1998; 233p. (In French) [Google Scholar]
- Horak, I.G.; Horak, J.L.; Keiranss, J.E. The Argasidae, Ixodidae, and Nuttalliellidae (Acari: Ixodida): A world list of valid tick names. Exp. Appl. Acarol. 2002, 28, 27–54. [Google Scholar] [CrossRef]
- Barker, S.C.; Murrell, A. Systematics and Evolution of ticks a list of valid genus and species names. In Ticks: Biology Disease and Control; Bowman, A.S., Nuttall, P., Eds.; Cambridge University Press: Cambridge, UK, 2008; pp. 1–39. [Google Scholar]
- Austin, M. Species distribution models and ecological theory: A critical assessment and some possible new approaches. Ecol. Model. 2007, 200, 1–19. [Google Scholar] [CrossRef]
- Mori, E.; Menchetti, M.; Zozzoli, R.; Milaanesi, P. The importance of taxonomy in species distribution models at a global scale: The case of an overlooked alien squirrel facing taxonomic revision. J. Zool. 2018, 307, 43–52. [Google Scholar] [CrossRef]
| Genera | Amblyomma | Dermacentor, Ixodes and Rhipicephalus |
|---|---|---|
| Annual mean temperature | • | |
| Mean diurnal range | • | • |
| Temperature Seasonality | • | |
| Max temperature of warmest month | • | |
| Min temperature of coldest month | • | |
| Mean temperature of wettest quarter | • | |
| Annual precipitation | • | • |
| Precipitation of driest month | • | • |
| Precipitation seasonality | • | • |
| Precipitation of wettest quarter | • | |
| Precipitation of the warmest quarter | • | |
| Type of soil | • | • |
| Type of land use and vegetation | • | • |
| Algorithm | Amblyomma spp. | Dermacentor spp. | Ixodes spp. | Rhipicephalus spp. |
|---|---|---|---|---|
| BIOCLIM | 0.706 | 0.664 | 0.669 | 0.772 |
| BRT | 0.905 | 0.872 | 0.892 | 0.921 |
| CART | 0.913 | 0.856 | 0.883 | 0.965 |
| MDA | 0.883 | 0.797 | 0.789 | 0.869 |
| GAM | 0.93 | 0.871 | 0.962 | 0.98 |
| GLM | 0.888 | 0.804 | 0.792 | 0.878 |
| MARS | 0.92 | 0.941 | 0.947 | 0.962 |
| MAXENT | 0.901 | 0.840 | 0.918 | 0.931 |
| RF | 0.999 | 0.996 | 0.994 | 0.999 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Clarke-Crespo, E.; Moreno-Arzate, C.N.; López-González, C.A. Ecological Niche Models of Four Hard Tick Genera (Ixodidae) in Mexico. Animals 2020, 10, 649. https://doi.org/10.3390/ani10040649
Clarke-Crespo E, Moreno-Arzate CN, López-González CA. Ecological Niche Models of Four Hard Tick Genera (Ixodidae) in Mexico. Animals. 2020; 10(4):649. https://doi.org/10.3390/ani10040649
Chicago/Turabian StyleClarke-Crespo, Emilio, Claudia N. Moreno-Arzate, and Carlos A. López-González. 2020. "Ecological Niche Models of Four Hard Tick Genera (Ixodidae) in Mexico" Animals 10, no. 4: 649. https://doi.org/10.3390/ani10040649
APA StyleClarke-Crespo, E., Moreno-Arzate, C. N., & López-González, C. A. (2020). Ecological Niche Models of Four Hard Tick Genera (Ixodidae) in Mexico. Animals, 10(4), 649. https://doi.org/10.3390/ani10040649






