A Heterozygous Missense Variant in the COL5A2 in Holstein Cattle Resembling the Classical Ehlers–Danlos Syndrome
Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Clinicopathological Investigation
2.3. DNA Samples
2.4. Whole-Genome Sequencing
2.5. Targeted Genotyping
2.6. Protein Predictions
2.7. Sequence Accessions
3. Results
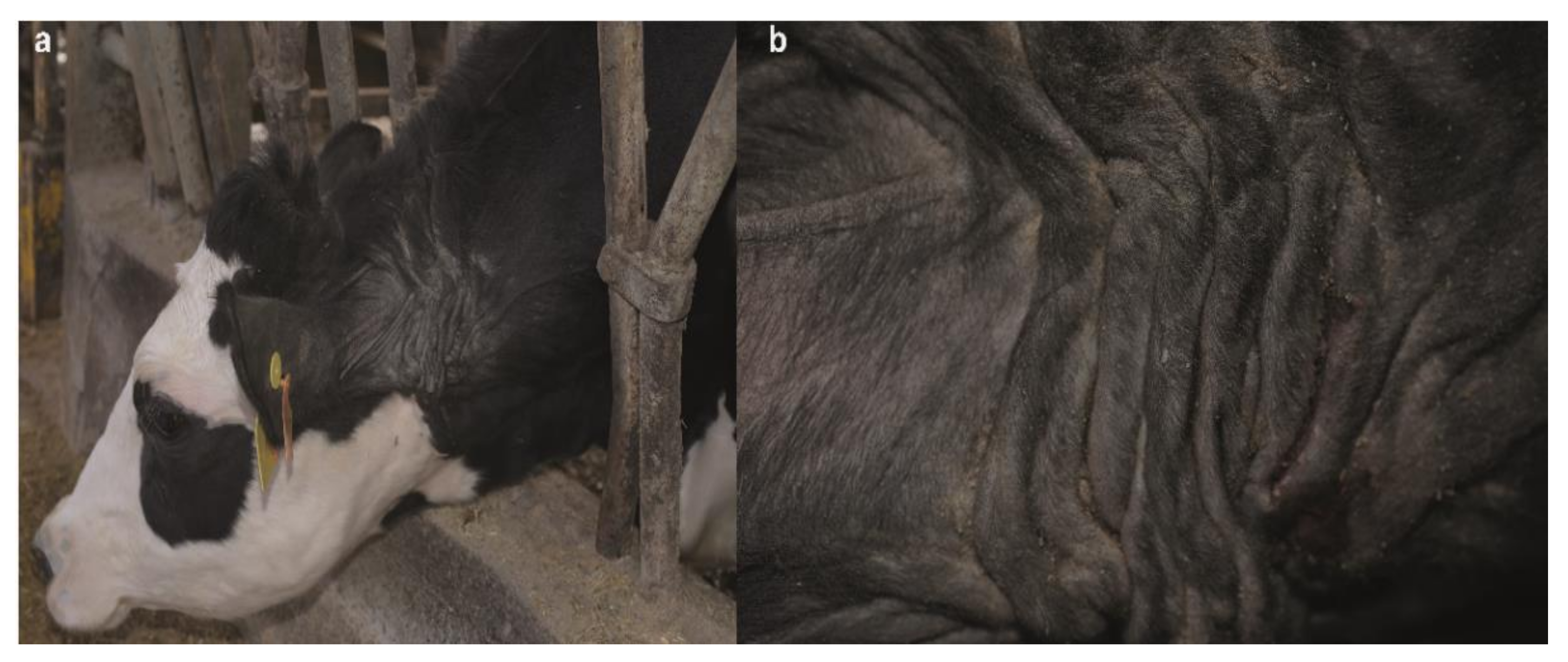
3.1. Clinical Phenotype
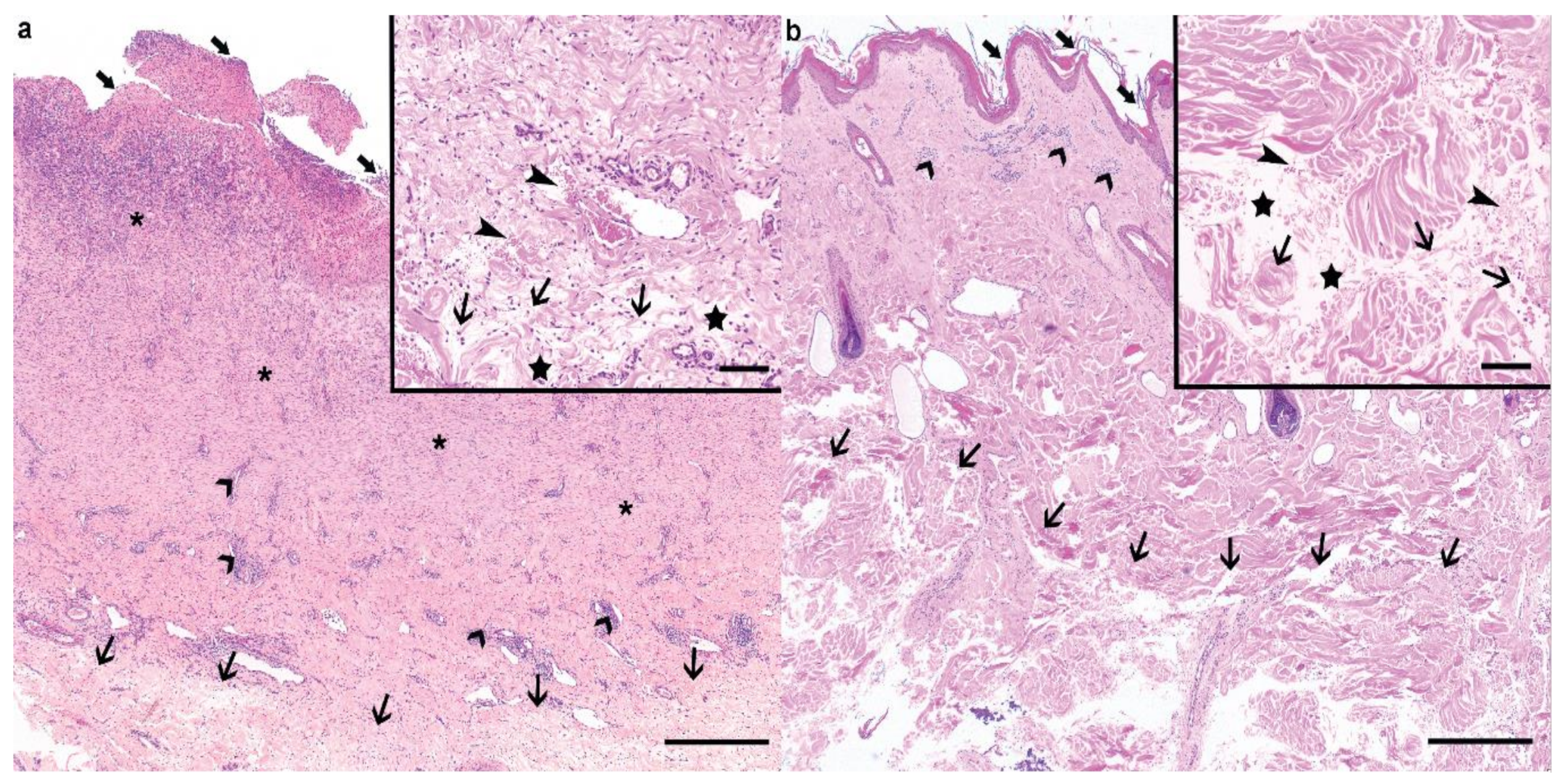
3.2. Histopathological Phenotype
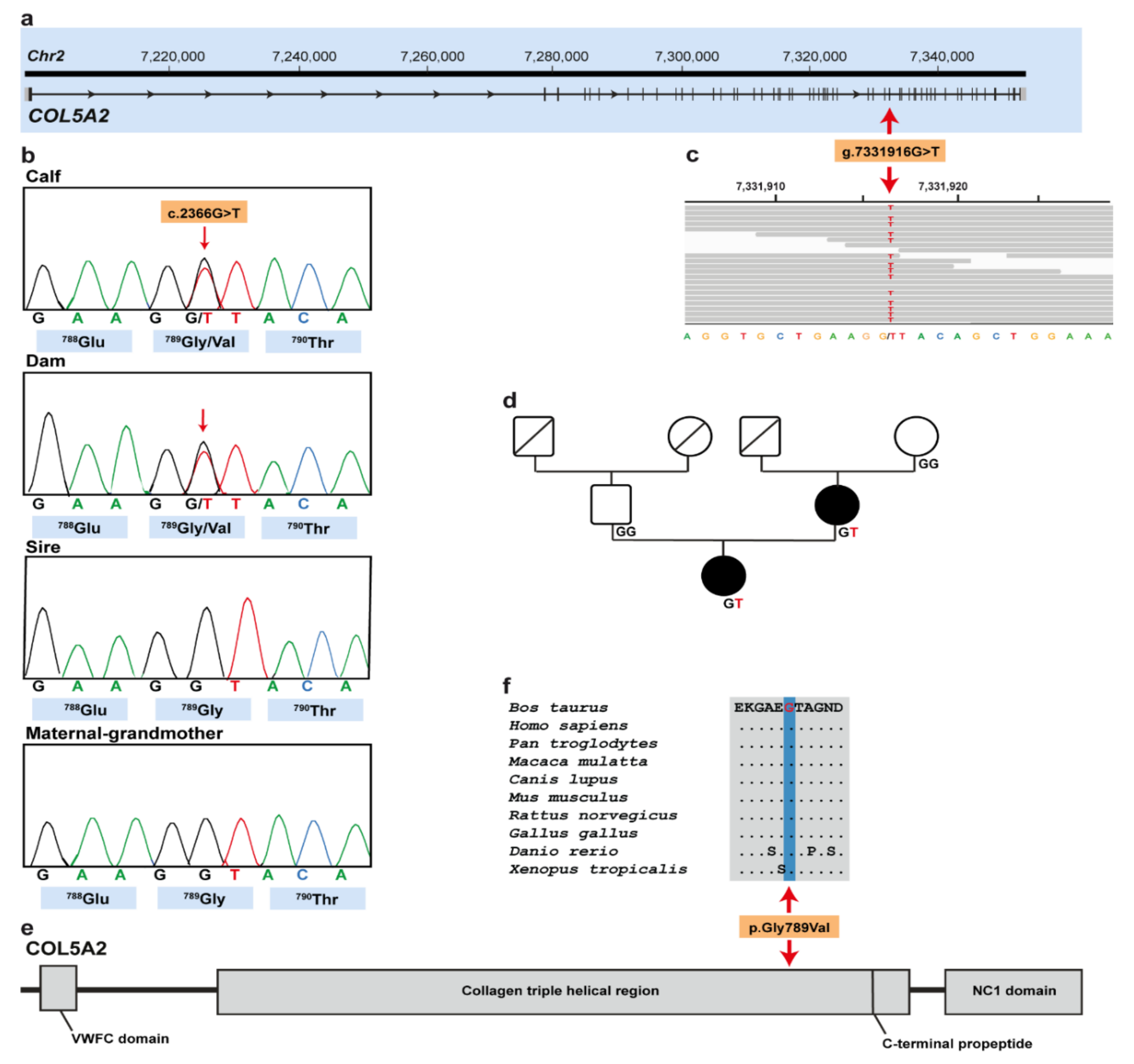
3.3. Genetic Analysis
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Leeb, T.; Müller, E.J.; Roosje, P.; Welle, M. Genetic testing in veterinary dermatology. Vet. Dermatol. 2017, 28, e1. [Google Scholar] [CrossRef] [PubMed]
- Malfait, F.; Francomano, C.; Byers, P.; Belmont, J.; Berglund, B.; Black, J.; Bloom, L.; Bowen, J.M.; Brady, A.F.; Burrows, N.P.; et al. The 2017 international classification of the Ehlers–Danlos syndromes. Am. J. Med. Genet. Part C Semin. Med. Genet. 2017, 175, 8–26. [Google Scholar] [CrossRef] [PubMed]
- Chiarelli, N.; Ritelli, M.; Zoppi, N.; Colombi, M. Cellular and molecular mechanisms in the pathogenesis of classical, vascular, and hypermobile ehlers-danlos syndromes. Genes 2019, 10, 609. [Google Scholar] [CrossRef] [PubMed]
- Ritelli, M.; Dordoni, C.; Venturini, M.; Chiarelli, N.; Quinzani, S.; Traversa, M.; Zoppi, N.; Vascellaro, A.; Wischmeijer, A.; Manfredini, E.; et al. Clinical and molecular characterization of 40 patients with classic Ehlers-Danlos syndrome: Identification of 18 COL5A1 and 2 COL5A2 novel mutations. Orphanet J. Rare Dis. 2013, 8, 1–19. [Google Scholar] [CrossRef] [PubMed]
- Monthoux, C.; de Brot, S.; Jackson, M.; Bleul, U.; Walter, J. Skin malformations in a neonatal foal tested homozygous positive for Warmblood Fragile Foal Syndrome. BMC Vet. Res. 2015, 11, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Tryon, R.C.; White, S.D.; Bannasch, D.L. Homozygosity mapping approach identifies a missense mutation in equine cyclophilin B (PPIB) associated with HERDA in the American Quarter Horse. Genomics 2007, 90, 93–102. [Google Scholar] [CrossRef]
- Jaffey, J.A.; Bullock, G.; Teplin, E.; Guo, J.; Villani, N.A.; Mhlanga-Mutangadura, T.; Schnabel, R.D.; Cohn, L.A.; Johnson, G.S. A homozygous ADAMTS2 nonsense mutation in a Doberman Pinscher dog with Ehlers Danlos syndrome and extreme skin fragility. Anim. Genet. 2019, 50, 543–545. [Google Scholar] [CrossRef]
- Bauer, A.; Bateman, J.F.; Lamandé, S.R.; Hanssen, E.; Kirejczyk, S.G.M.; Yee, M.; Ramiche, A.; Jagannathan, V.; Welle, M.; Leeb, T.; et al. Identification of two independent COL5A1 variants in dogs with ehlers-danlos syndrome. Genes 2019, 10, 731. [Google Scholar] [CrossRef]
- Bauer, A.; de Lucia, M.; Leuthard, F.; Jagannathan, V.; Leeb, T. Compound heterozygosity for TNXB genetic variants in a mixed-breed dog with Ehlers-Danlos syndrome. Anim. Genet. 2019, 50, 546–549. [Google Scholar] [CrossRef]
- Spycher, M.; Bauer, A.; Jagannathan, V.; Frizzi, M.; De Lucia, M.; Leeb, T. A frameshift variant in the COL5A1 gene in a cat with Ehlers-Danlos syndrome. Anim. Genet. 2018, 49, 641–644. [Google Scholar] [CrossRef]
- Hegreberg, G.A.; Padgett, G.A.; Gorham, J.R.; Henson, J.B. A heritable connective tissue disease of dogs and Mink Resembling the Ehlers-Danlos. J. Investig. Dermatol. 1970, 54, 377–380. [Google Scholar] [CrossRef] [PubMed]
- Harvey, R.; Brown, P.; Young, R.; Whitbread, T. A connective tissue defect in two rabbits similar to the Ehlers-Danlos syndrome. Vet. Rec. 1990, 126, 130–132. [Google Scholar] [PubMed]
- Joller, S.; Berenguer Veiga, I.; Drögemüller, C. Dermatosparaxis in White Dorper sheep: Confirmation of a causative nonsense mutation in ADAMTS2. Anim. Genet. 2017, 48, 729–730. [Google Scholar] [CrossRef]
- Zhou, H.; Hickford, J.G.H.; Fang, Q. A premature stop codon in the ADAMTS2 gene is likely to be responsible for dermatosparaxis in Dorper sheep. Anim. Genet. 2012, 43, 471–473. [Google Scholar] [CrossRef] [PubMed]
- Monteagudo, L.V.; Ferrer, L.M.; Catalan-Insa, E.; Savva, D.; Mcguffin, L.J.; Tejedor, M.T. In silico identification and three-dimensional modelling of the missense mutation in ADAMTS2 in a sheep flock with dermatosparaxis. Vet. Dermatol. 2015, 26, 49–52. [Google Scholar] [CrossRef]
- Carty, C.I.; Lee, A.M.; Wienandt, N.A.E.; Stevens, E.L.; Alves, D.A.; Browne, J.A.; Bryan, J.; Ryan, E.G.; Cassidy, J.P. Dermatosparaxis in two Limousin calves. Ir. Vet. J. 2016, 69, 1–5. [Google Scholar] [CrossRef]
- Tajima, M.; Miyake, S.; Takehana, K.; Kobayashi, A.; Yamato, O.; Maede, Y. Gene defect of dermatan sulfate proteoglycan of cattle affected with a variant form of Ehlers-Danlos syndrome. J. Vet. Intern. Med. 1999, 13, 202–205. [Google Scholar] [CrossRef]
- Colige, A.; Sieron, A.L.; Li, S.W.; Schwarze, U.; Petty, E.; Wertelecki, W.; Wilcox, W.; Krakow, D.; Cohn, D.H.; Reardon, W.; et al. Human ehlers-danlos syndrome type VII C and bovine dermatosparaxis are caused by mutations in the procollagen I N-proteinase gene. Am. J. Hum. Genet. 1999, 65, 308–317. [Google Scholar] [CrossRef]
- Rosen, B.D.; Bickhart, D.M.; Schnabel, R.D.; Koren, S.; Elsik, C.G.; Tseng, E.; Rowan, T.N.; Low, W.Y.; Zimin, A.; Couldrey, C.; et al. De novo assembly of the cattle reference genome with single-molecule sequencing. Gigascience 2020, 9, 1–9. [Google Scholar] [CrossRef]
- Hayes, B.J.; Daetwyler, H.D. 1000 Bull Genomes Project to Map Simple and Complex Genetic Traits in Cattle: Applications and Outcomes. Annu. Rev. Anim. Biosci. 2019, 7, 89–102. [Google Scholar] [CrossRef]
- Chen, S.; Zhou, Y.; Chen, Y.; Gu, J. Fastp: An ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 2018, 34, i884–i890. [Google Scholar] [CrossRef] [PubMed]
- Häfliger, I.M.; Wiedemar, N.; Švara, T.; Starič, J.; Cociancich, V.; Šest, K.; Gombač, M.; Paller, T.; Agerholm, J.S.; Drögemüller, C. Identification of small and large genomic candidate variants in bovine pulmonary hypoplasia and anasarca syndrome. Anim. Genet. 2020, 51, 382–390. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.T.; Thorvaldsdóttir, H.; Wenger, A.M.; Zehir, A.; Mesirov, J.P. Variant review with the integrative genomics viewer. Cancer Res. 2017, 77, e31–e34. [Google Scholar] [CrossRef]
- Choi, Y.; Chan, A.P. PROVEAN web server: A tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015, 31, 2745–2747. [Google Scholar] [CrossRef]
- Pejaver, V.; Urresti, J.; Lugo-Martinez, J.; Pagel, K.; Lin, G.N.; Nam, H.-J.; Mort, M.; Cooper, D.; Sebat, J.; Iakoucheva, L.; et al. MutPred2: Inferring the molecular and phenotypic impact of amino acid variants. bioRxiv 2017, 134981. [Google Scholar] [CrossRef]
- Bendl, J.; Stourac, J.; Salanda, O.; Pavelka, A.; Wieben, E.D.; Zendulka, J.; Brezovsky, J.; Damborsky, J. PredictSNP: Robust and Accurate Consensus Classifier for Prediction of Disease-Related Mutations. PLoS Comput. Biol. 2014, 10, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Dalgleish, R. The Human Collagen Mutation Database 1998. Nucleic Acids Res. 1998, 26, 253–255. [Google Scholar] [CrossRef] [PubMed]
- Dalgleish, R.; Teh, W.K. Ehlers Danlos Syndrome Variant Database. Available online: https://eds.gene.le.ac.uk/home.php?select_db=COL5A1 (accessed on 29 September 2020).
- Symoens, S.; Syx, D.; Malfait, F.; Callewaert, B.; de Backer, J.; Vanakker, O.; Coucke, P.; de Paepe, A. Comprehensive molecular analysis demonstrates type V collagen mutations in over 90% of patients with classic EDS and allows to refine diagnostic criteria. Hum. Mutat. 2012, 33, 1485–1493. [Google Scholar] [CrossRef]
- Zoppi, N.; Chiarelli, N.; Cinquina, V.; Ritelli, M.; Colombi, M. GLUT10 deficiency leads to oxidative stress and non-canonical αvβ3 integrin-mediated TGFβ signalling associated with extracellular matrix disarray in arterial tortuosity syndrome skin fibroblasts. Hum. Mol. Genet. 2015, 24, 6769–6787. [Google Scholar] [CrossRef]
- Richards, A.J.; Martin, S.; Nicholls, A.C.; Harrison, J.B.; Pope, F.M.; Burrows, N.P. A single base mutation in COL5A2 causes Ehlers-Danlos syndrome type II. J. Med. Genet. 1998, 35, 846–848. [Google Scholar] [CrossRef]
- Zaffino Heyerhoff, J.C.; LeBlanc, S.J.; DeVries, T.J.; Nash, C.G.R.; Gibbons, J.; Orsel, K.; Barkema, H.W.; Solano, L.; Rushen, J.; de Passillé, A.M.; et al. Prevalence of and factors associated with hock, knee, and neck injuries on dairy cows in freestall housing in Canada. J. Dairy Sci. 2014, 97, 173–184. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, A.L.; Schwarze, U.; Jennings, J.F.; Byers, P.H. Molecular Mechanisms of Classical Ehlers-Danlos Syndrome. Hum. Mutat. 2009, 30, 995–1002. [Google Scholar] [CrossRef]
- Lamandé, S.R.; Bateman, J.F. Genetic Disorders of the Extracellular Matrix. Anat. Rec. 2020, 303, 1527–1542. [Google Scholar] [CrossRef] [PubMed]
- DE, B. Type V collagen: Heterotypic type I/V collagen interactions in the regulation of fibril assembly. Micron 2001, 32, 223–237. [Google Scholar]

| Inheritance | Autosomal dominant |
| Molecular Basis | COL5A1; COL5A2; COL1A1; COL3A1 |
| Major Criteria | 1. Skin hyperextensibility and atrophic scarring |
| 2. Generalized joint hypermobility | |
| Minor Criteria | 1. Easy bruising |
| 2. Soft, doughy skin | |
| 3. Skin fragility (or traumatic splitting) | |
| 4. Molluscoid pseudotumors | |
| 5. Subcutaneous spheroids | |
| 6. Hernia | |
| 7. Epicanthal folds | |
| 8. Complications of joint hypermobility | |
| 9. Family history of first-degree relative who meets clinical criteria |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jacinto, J.G.P.; Häfliger, I.M.; Veiga, I.M.B.; Letko, A.; Benazzi, C.; Bolcato, M.; Drögemüller, C. A Heterozygous Missense Variant in the COL5A2 in Holstein Cattle Resembling the Classical Ehlers–Danlos Syndrome. Animals 2020, 10, 2002. https://doi.org/10.3390/ani10112002
Jacinto JGP, Häfliger IM, Veiga IMB, Letko A, Benazzi C, Bolcato M, Drögemüller C. A Heterozygous Missense Variant in the COL5A2 in Holstein Cattle Resembling the Classical Ehlers–Danlos Syndrome. Animals. 2020; 10(11):2002. https://doi.org/10.3390/ani10112002
Chicago/Turabian StyleJacinto, Joana G. P., Irene M. Häfliger, Inês M. B. Veiga, Anna Letko, Cinzia Benazzi, Marilena Bolcato, and Cord Drögemüller. 2020. "A Heterozygous Missense Variant in the COL5A2 in Holstein Cattle Resembling the Classical Ehlers–Danlos Syndrome" Animals 10, no. 11: 2002. https://doi.org/10.3390/ani10112002
APA StyleJacinto, J. G. P., Häfliger, I. M., Veiga, I. M. B., Letko, A., Benazzi, C., Bolcato, M., & Drögemüller, C. (2020). A Heterozygous Missense Variant in the COL5A2 in Holstein Cattle Resembling the Classical Ehlers–Danlos Syndrome. Animals, 10(11), 2002. https://doi.org/10.3390/ani10112002






