Coexistence of Three Divergent mtDNA Lineages in Northeast Asia Provides New Insights into Phylogeography of Goldfish (Carssius auratus)
Abstract
:Simple Summary
Abstract
1. Introduction
2. Materials and Methods
2.1. Sample Collection
2.2. Molecular Methods
2.3. Population Genetic Analyses
2.4. Phylogenetic Analyses
3. Results
3.1. Genetic Diversity
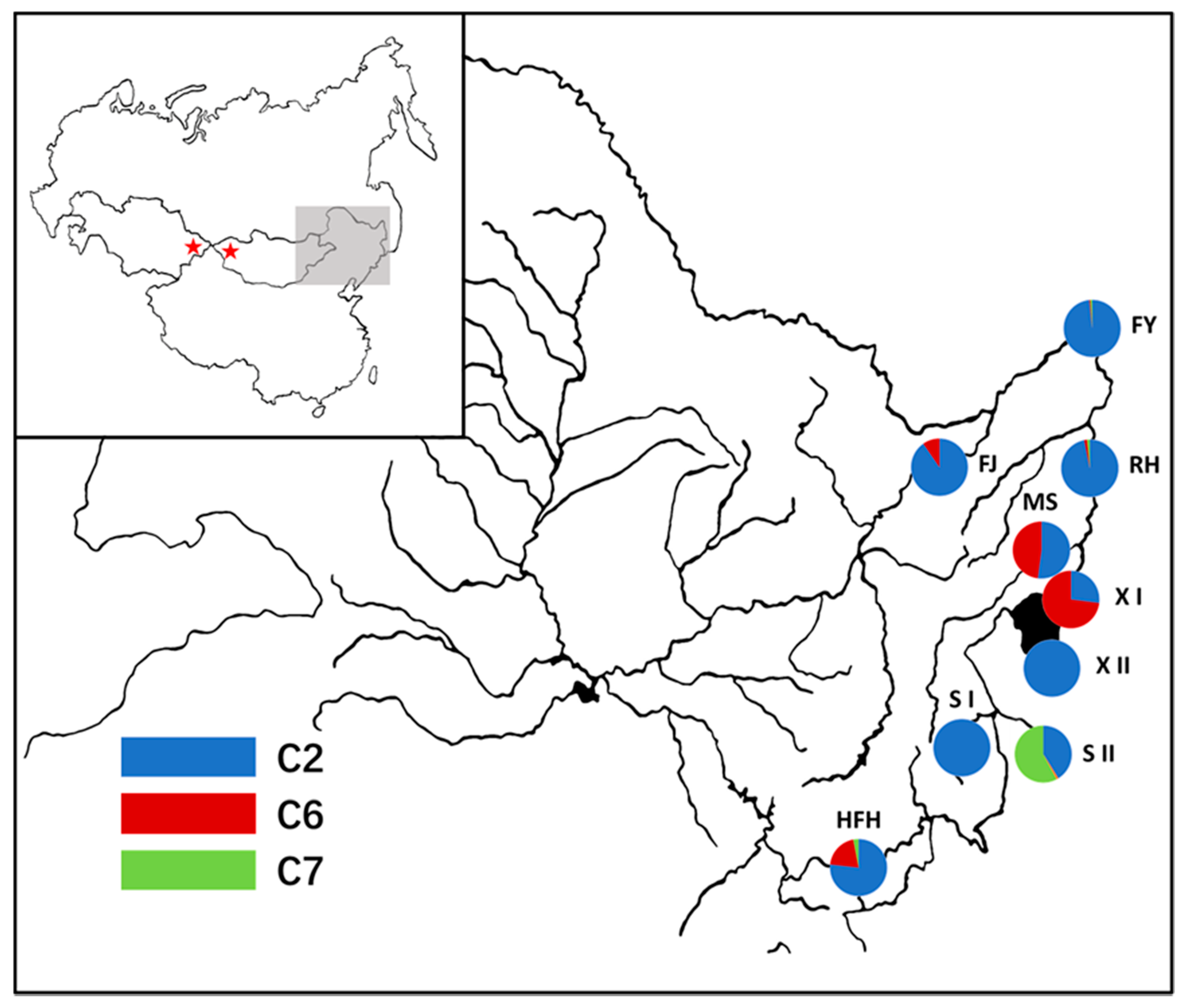
3.2. Genetic Structure
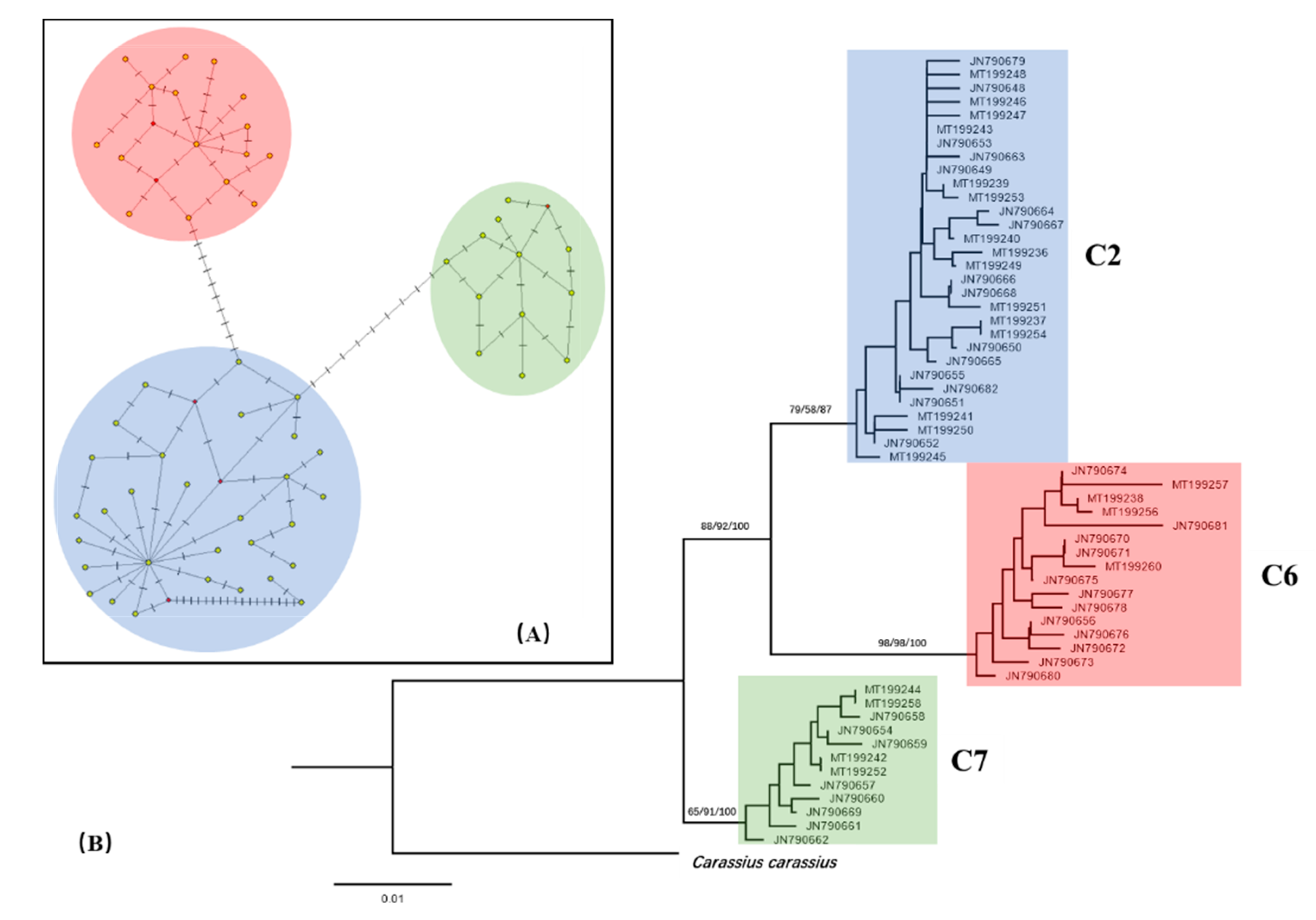
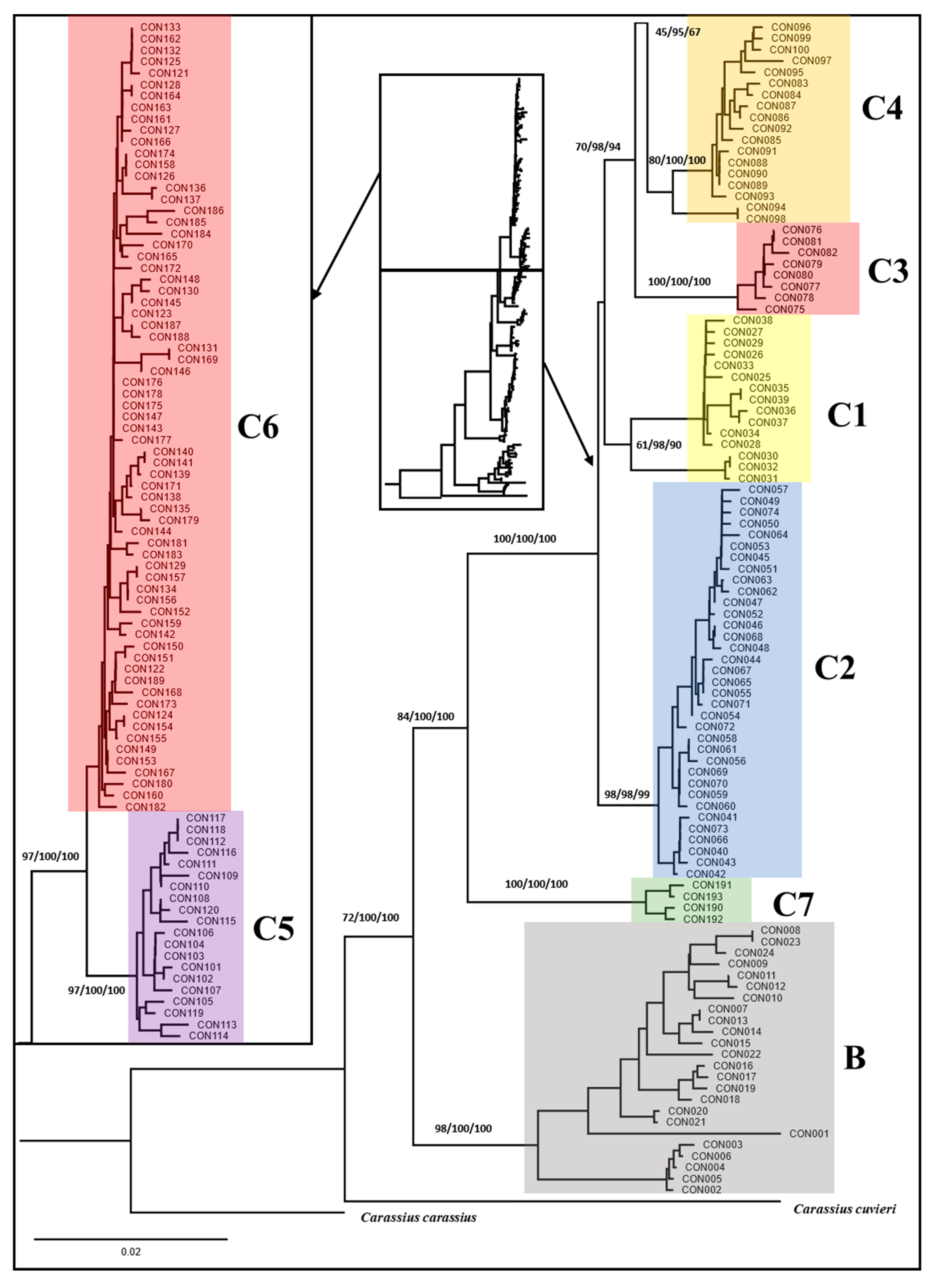
3.3. Phylogeny of Carassius Auratus Complex
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Luo, Y.L.; Yue, P.Q. Cyprininae in Fauna Sinica, Osteichthyes, Cypriniformes; Yue, P.Q., Ed.; Science Press: Beijing, China, 2000; pp. 427–434. (In Chinese) [Google Scholar]
- Hosoya, K. Cyprinidae in Fishes of Japan with Pictorial Keys to the Species; Nakabo, T., Ed.; Tokai University Press: Tokyo, Japan, 2002; pp. 253–254. (In Japanese) [Google Scholar]
- Kottelat, M.; Freyhof, J. Handbook of European Freshwater Fishes; Kottelat, Cornol and Freyhof: Berlin, German, 2007; pp. 142–147. [Google Scholar]
- Cheng, L.; Chang, Y.M.; Lu, C.Y.; Cao, D.C.; Sun, X.W. DNA barcoding and species and subspecies classification within genus Carassius. Zool. Res. 2012, 33, 463–472. (In Chinese) [Google Scholar] [CrossRef] [Green Version]
- Golovinskaya, K.A.; Romashov, D.D.; Cherfas, N.B. Unisexual and Bisexual Forms of Silver Crucian Carp (Carassius auratus gibelio Bloch). Vopr. Ikhtiol. 1965, 5, 614–629. [Google Scholar]
- Kobayasi, H.; Kawashima, Y.; Takeuchi, N. Comparative chromosome studies in the genus Carassius, especially with a finding of polyploidy in the ginbuna (C. auratus langsdorfii). Jpn. J. Ichthyol. 1970, 17, 153–160. [Google Scholar]
- Xiao, J.; Zou, T.M.; Chen, Y.B.; Chen, L.; Liu, S.J.; Tao, M.; Zhang, C.; Zhao, R.R.; Zhou, Y.; Long, Y.; et al. Coexistence of diploid, triploid and tetraploid crucian carp (Carassius auratus) in natural waters. BMC Genet. 2011, 12, e20. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Nakamura, M. Keys to the Freshwater Fishes of Japan Fully Illustrated in Colors; Hokuryukan: Tokyo, Japan, 1982; pp. 140–142. (In Japanese) [Google Scholar]
- Murakami, M.; Matsuba, C.; Fujitani, H. The maternal origins of the triploid ginbuna (Carassius auratus langsdorfi): Phylogenetic relationships within the C. auratus taxa by partial mitochondrial D-loop sequencing. Gen. Genet. Syst. 2001, 76, 25–32. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Iguchi, K.; Yamamoto, G.; Matsubara, N.; Nishida, M. Morphological and genetic analysis of fish of a Carassius complex (Cyprinidae) in Lake Kasumigaura with reference to the taxonomic status of two all-female triploid morphs. Biol. J. Linn. Soc. 2003, 79, 351–357. [Google Scholar] [CrossRef] [Green Version]
- Yamamoto, G.; Takada, M.; Iguchi, K.; Nishida, M. Genetic constitution and phylogenetic relationships among Japanese crucian carps (Carassius). Ichthyol. Res. 2010, 57, 215–222. [Google Scholar] [CrossRef]
- Galtier, N.; Nabholz, B.; Glémin, S.; Hurst, G.D. Mitochondrial DNA as a marker of molecular diversity: A reappraisal. Mol. Ecol. 2009, 18, 4541–4550. [Google Scholar] [CrossRef]
- Borzée, A.; Fong, J.J.; Nguyen, H.Q.; Jang, Y. Large-Scale Hybridisation as an Extinction Threat to the Suweon Treefrog (Hylidae: Dryophytes suweonensis). Animals 2020, 10, 764. [Google Scholar] [CrossRef]
- Takada, M.; Tachihara, K.; Kon, T.; Yamamoto, G.; Iguchi, K.; Miya, M.; Nishida, M. Biogeography and evolution of the Carassius auratus-complex in East Asia. BMC Evol. Biol. 2010, 10, e7. [Google Scholar] [CrossRef] [Green Version]
- Gao, Y.; Wang, S.Y.; Luo, J.; Murphy, R.W.; Du, R.; Wu, S.F.; Zhu, C.L.; Li, Y.; Poyarkov, A.D.; Nguyen, S.N.; et al. Quaternary palaeoenvironmental oscillations drove the evolution of the Eurasian Carassius auratus complex (Cypriniformes, Cyprinidae). J. Biogeogr. 2012, 39, 2264–2278. [Google Scholar] [CrossRef]
- Sakai, H.; Iguchi, K.; Yamazaki, Y.; Sideleva, V.G.; Goto, A. Morphological and mtDNA sequence studies on three crucian carps (Carassius: Cyprinidae) including a new stock from the Ob River system, Kazakhstan. J. Fish Biol. 2009, 74, 1756–1773. [Google Scholar] [CrossRef] [PubMed]
- Kalous, L.; Bohlen, J.; Rylková, K.; Petrtýl, M. Hidden diversity within the Prussian carp and designation of a neotype for Carassius gibelio (Teleostei: Cyprinidae). Ichthyol. Explor. Fres. 2012, 23, 11–18. [Google Scholar]
- Jiang, F.F.; Wang, Z.W.; Zhou, L.; Jiang, L.; Zhang, X.J.; Apalikova, O.V.; Brykov, V.A.; Gui, J.F. High male incidence and evolutionary implications of triploid form in northeast Asia Carassius auratus complex. Mol. Phylogenet. Evol. 2013, 66, 350–359. [Google Scholar] [CrossRef] [PubMed]
- Iguchi, K.; Tanimura, Y.; Nishida, M. Sequence divergence in the mtDNA control region of amphidromous and landlocked forms of ayu. Fish Sci. 1997, 63, 901–905. [Google Scholar] [CrossRef] [Green Version]
- Nishida, M.; Ohkawa, T.; Iwata, H. Methods of Analysis of genetic population structure with mitochondrial DNA markers. Fish Genet. Breed Sci. 1998, 26, 81–100. [Google Scholar]
- Xiao, W.; Zhang, Y.; Liu, H. Molecular systematics of Xenocyprinae (Teleostei: Cyprinidae): Taxonomy, biogeography, and coevolution of a special group restricted in East Asia. Mol. Phylogenet. Evol. 2001, 18, 163–173. [Google Scholar] [CrossRef] [Green Version]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [Green Version]
- Julio, R.; Albert, F.M.; Juan, C.S.; Sara, G.R.; Pablo, L.; Sebastián, E.R.; Alejandro, S. DnaSP v6: DNA Sequence Polymorphism Analysis of Large Datasets. Mol. Biol. Evol. 2017, 34, 3299–3302. [Google Scholar]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin ver. 3.0: An integrated software package for population genetics data analysis. Evol. Bioinform. 2005, 1, 47–50. [Google Scholar] [CrossRef] [Green Version]
- Kumar, S.; Stecher, G.; Li, M.; Knyaz, C.; Tamura, K. MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol. Biol. Evol. 2018, 35, 1547–1549. [Google Scholar] [CrossRef]
- Nguyen, L.T.; Schmidt, H.A.; von Haeseler, A.; Minh, B.Q. IQ-TREE: A fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 2015, 32, 268–274. [Google Scholar] [CrossRef]
- Ronquist, F.; Teslenko, M.; van der Mark, P.; Ayres, L.D.; Darling, A.; Hohna, S.; Larget, B.; Liu, L.; Suchard, A.M.; Huelsenbeck, P.J. MrBayes 3.2: Efficient Bayesian phylogenetic inference and model choice across a large model space. Syst. Biol. 2012, 61, 539–542. [Google Scholar] [CrossRef] [Green Version]
- Nylander, J.A.A. MrModeltest v2. Program Distributed by the Author; Evolutionary Biology Centre, Uppsala University: Uppsala, Sweden, 2004. [Google Scholar]
- Liu, X.L.; Li, X.Y.; Jiang, F.F.; Wang, Z.W.; Li, Z.; Zhang, X.J.; Zhou, L.; Gui, J.F. Numerous mitochondrial DNA haplotypes reveal multiple independent polyploidy origins of hexaploids in Carassius species complex. Ecol. Evol. 2017, 7, 10604–10615. [Google Scholar] [CrossRef] [PubMed]
- Rylkova, K.; Kalous, L.; Bohlen, J.; Lamatsch, D.K.; Petrtýl, M. Phylogeny and biogeographic history of the cyprinid fish genus Carassius (Teleostei: Cyprinidae) with focus on natural and anthropogenic arrivals in Europe. Aquaculture 2013, 380–383, 13–20. [Google Scholar] [CrossRef]
- Ribeiro, F.; Rylkova, K.; Morenovalcarcel, R.; Carrapato, C.; Kalous, L. Prussian carp Carassius gibelio: A silent invader arriving to the Iberian Peninsula. Aquat. Ecol. 2015, 49, 99–104. [Google Scholar] [CrossRef]
- Wang, S.Y.; Luo, J.; Murphy, R.W.; Wu, S.F.; Zhu, C.L.; Gao, Y.; Zhang, Y.P. Origin of Chinese goldfish and sequential loss of genetic diversity accompanies new breeds. PLoS ONE 2013, 8, e59571. [Google Scholar] [CrossRef] [PubMed]
- Kottelat, M. Fishes of Mongolia. A Check-List of the Fishes Known to Occur in Mongolia with Comments on Systematics and Nomenclature; World Bank: Washington, DC, USA, 2006; pp. 27–28. [Google Scholar]
| Target | Primer | Sequence (5′-3′) | References |
|---|---|---|---|
| control region | L15923 | TTAAAGCATCGGTCTTGTAA | [19] |
| Hl6500 | GCCCTGAAATAGGAACCAGA | [20] | |
| 16500F | AGCGCCCAGAAAAGAGAGAT | This study | |
| cytochrome b gene | L14724 | GACTTGAAAAACCACCGTTG | [21] |
| H15915 | CTCCGATCTCCGGATTACAAGAC | [21] | |
| 15372F | GACCTACCCACACCATCCAA | This study |
| Population | Code | n | h | S | K | Hd | π (%) |
|---|---|---|---|---|---|---|---|
| Fujin, China | FJ | 135 | 15 | 38 | 4.191 ± 2.095 | 0.673 ± 0.042 | 1.174 ± 0.650 |
| Fuyuan, China | FY | 188 | 15 | 31 | 1.430 ± 0.876 | 0.466 ± 0.044 | 0.399 ± 0.271 |
| Mishan, China | MS | 119 | 8 | 22 | 7.263 ± 3.426 | 0.591 ± 0.024 | 2.035 ± 1.063 |
| Raohe, China | RH | 127 | 16 | 43 | 4.065 ± 2.041 | 0.602 ± 0.050 | 1.139 ± 0.633 |
| Huifahe, China | HFH | 99 | 14 | 46 | 6.908 ± 3.277 | 0.689 ± 0.047 | 1.935 ± 1.017 |
| Suifenhe, China | SI | 100 | 7 | 21 | 1.942 ± 1.112 | 0.208 ± 0.054 | 0.544 ± 0.345 |
| Suifenhe, Russia | SII | 90 | 11 | 24 | 7.512 ± 3.541 | 0.780 ± 0.023 | 2.104 ± 1.099 |
| Xingkaihu, China | XI | 100 | 15 | 24 | 7.842 ± 3.680 | 0.878 ± 0.013 | 2.197 ± 1.142 |
| Xingkaihu, Russia | XII | 96 | 13 | 28 | 1.418 ± 0.875 | 0.563 ± 0.057 | 0.396 ± 0.271 |
| Lineage | C2 | 822 | 30 | 38 | 1.564 ± 0.934 | 0.479 ± 0.022 | 0.439 ± 0.290 |
| C6 | 173 | 16 | 18 | 2.327 ± 1.279 | 0.756 ± 0.029 | 0.657 ± 0.400 | |
| C7 | 59 | 12 | 8 | 1.193 ± 0.775 | 0.679 ± 0.056 | 0.336 ± 0.242 | |
| Total | 1054 | 58 | 57 | 6.417 ± 3.042 | 0.676 ± 0.016 | 1.798 ± 0.942 |
| Lineage | GenBank NO. | FJ | FY | MS | RH | HFH | SI | SII | XI | XII |
|---|---|---|---|---|---|---|---|---|---|---|
| C2 | JN790648 | 5 | 5 | 7 | 13 | |||||
| JN790649 | 74 | 136 | 56 | 79 | 53 | 89 | 22 | 18 | 62 | |
| JN790650 | 5 | 18 | 1 | 4 | 2 | 4 | ||||
| JN790651 | 1 | |||||||||
| JN790652 | 6 | 9 | 3 | 6 | 7 | 1 | 1 | |||
| JN790653 | 2 | 1 | 1 | |||||||
| JN790655 | 4 | 7 | 15 | 1 | 1 | |||||
| JN790663 | 1 | |||||||||
| JN790664 | 6 | |||||||||
| JN790665 | 6 | 2 | ||||||||
| JN790666 | 20 | 4 | 3 | 1 | 2 | 2 | ||||
| JN790667 | 1 | |||||||||
| JN790668 | 2 | 1 | ||||||||
| JN790679 | 5 | |||||||||
| JN790682 | 1 | 2 | ||||||||
| MT199236 | 2 | 1 | ||||||||
| MT199237 | 5 | |||||||||
| MT199239 | 1 | 9 | 2 | |||||||
| MT199240 | 2 | |||||||||
| MT199241 | 2 | |||||||||
| MT199243 | 2 | 2 | ||||||||
| MT199245 | 1 | |||||||||
| MT199246 | 1 | |||||||||
| MT199247 | 1 | |||||||||
| MT199248 | 1 | |||||||||
| MT199249 | 1 | |||||||||
| MT199250 | 1 | |||||||||
| MT199251 | 2 | |||||||||
| MT199253 | 1 | |||||||||
| MT199254 | 7 | |||||||||
| C6 | JN790656 | 6 | 52 | 2 | 12 | 3 | 1 | 3 | ||
| JN790670 | 2 | 16 | ||||||||
| JN790671 | 1 | 17 | ||||||||
| JN790672 | 17 | |||||||||
| JN790673 | 8 | |||||||||
| JN790674 | 3 | 1 | 4 | 2 | 1 | |||||
| JN790675 | 6 | |||||||||
| JN790676 | 1 | 1 | 1 | |||||||
| JN790677 | 1 | |||||||||
| JN790678 | 2 | 3 | ||||||||
| JN790680 | 1 | |||||||||
| JN790681 | 2 | |||||||||
| MT199238 | 1 | |||||||||
| MT199256 | 1 | |||||||||
| MT199257 | 1 | |||||||||
| MT199260 | 1 | |||||||||
| C7 | JN790654 | 31 | ||||||||
| JN790657 | 3 | |||||||||
| JN790658 | 13 | |||||||||
| JN790659 | 1 | |||||||||
| JN790660 | 1 | |||||||||
| JN790661 | 1 | |||||||||
| JN790662 | 1 | |||||||||
| JN790669 | 1 | |||||||||
| MT199242 | 1 | |||||||||
| MT199244 | 1 | 1 | ||||||||
| MT199252 | 2 | |||||||||
| MT199258 | 2 | |||||||||
| Total | 135 | 188 | 119 | 127 | 99 | 100 | 90 | 100 | 96 |
| Population Code | FJ | FY | MS | RH | HFH | SI | SII | XI | XII |
|---|---|---|---|---|---|---|---|---|---|
| FJ | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.001 ±0.001 | 0.004 ± 0.002 | 0.014 ± 0.003 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | |
| FY | 0.060 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.005 ± 0.003 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.002 ± 0.001 | |
| MS | 0.245 | 0.442 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | |
| RH | 0.034 | 0.032 | 0.335 | 0.000 ± 0.000 | 0.002 ± 0.001 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.007 ± 0.003 | |
| HFH | 0.028 | 0.150 | 0.110 | 0.092 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.000 ± 0.000 | |
| SI | 0.026 | 0.027 | 0.338 | 0.029 | 0.089 | 0.000 ± 0.000 | 0.000 ± 0.000 | 0.003 ± 0.002 | |
| SII | 0.405 | 0.558 | 0.346 | 0.434 | 0.296 | 0.484 | 0.000 ± 0.000 | 0.000 ± 0.000 | |
| XI | 0.475 | 0.656 | 0.139 | 0.540 | 0.326 | 0.563 | 0.436 | 0.000 ± 0.000 | |
| XII | 0.059 | 0.024 | 0.398 | 0.029 | 0.138 | 0.029 | 0.510 | 0.605 |
| Partitions | Source of Variation | d.f. | Sum of Squares | Variance Components | Percentage of Variation | Fixation Indices | p-Value |
|---|---|---|---|---|---|---|---|
| I | |||||||
| Among groups | 1 | 92.32 | −0.083 | −2.51 | ΦCT = −0.025 | 0.423 ± 0.015 | |
| Among populations within groups | 7 | 962.80 | 1.159 | 35.13 | ΦSC = 0.343 | 0.000 ± 0.000 | |
| Within populations | 1045 | 2323.58 | 2.224 | 67.38 | ΦST = 0.326 | 0.000 ± 0.000 | |
| Total | 1053 | 3378.70 | 3.300 | ||||
| II | |||||||
| Among lineages | 2 | 2502.10 | 6.559 | 88.72 | ΦST = 0.887 | 0.000 ± 0.000 | |
| Within lineages | 1051 | 876.60 | 0.834 | 11.28 | |||
| Total | 1053 | 3378.70 | 7.393 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, L.; Lu, C.; Wang, L.; Li, C.; Yu, X. Coexistence of Three Divergent mtDNA Lineages in Northeast Asia Provides New Insights into Phylogeography of Goldfish (Carssius auratus). Animals 2020, 10, 1785. https://doi.org/10.3390/ani10101785
Cheng L, Lu C, Wang L, Li C, Yu X. Coexistence of Three Divergent mtDNA Lineages in Northeast Asia Provides New Insights into Phylogeography of Goldfish (Carssius auratus). Animals. 2020; 10(10):1785. https://doi.org/10.3390/ani10101785
Chicago/Turabian StyleCheng, Lei, Cuiyun Lu, Le Wang, Chao Li, and Xiaoli Yu. 2020. "Coexistence of Three Divergent mtDNA Lineages in Northeast Asia Provides New Insights into Phylogeography of Goldfish (Carssius auratus)" Animals 10, no. 10: 1785. https://doi.org/10.3390/ani10101785




