EpicPCR 2.0: Technical and Methodological Improvement of a Cutting-Edge Single-Cell Genomic Approach
Abstract
:1. Introduction
2. Materials and Methods
2.1. Bacterial Strain
2.2. Sample Collection and Microbial Cells Extraction
2.3. EpicPCR
2.3.1. Cell Enumeration and Bead Formation
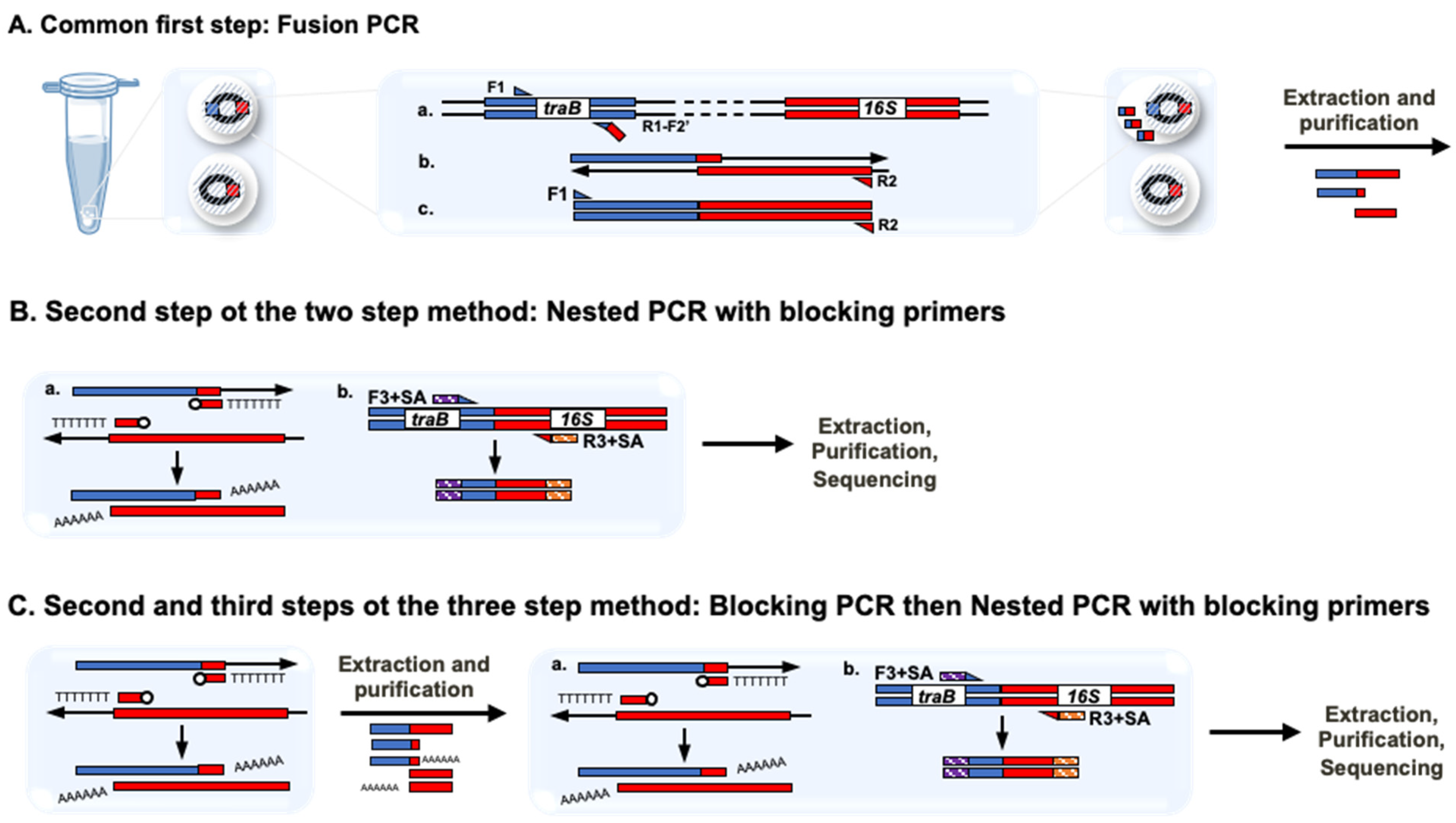
2.3.2. Fusion, Nested and Blocking PCRs and Sequencing
2.4. 16S rRNA Gene Amplification and Sequencing
2.5. Sequencing Data Analysis
3. Results and Discussion
3.1. Elaboration of Polyacrylamide Beads Carrying Isolated Environmental Cells
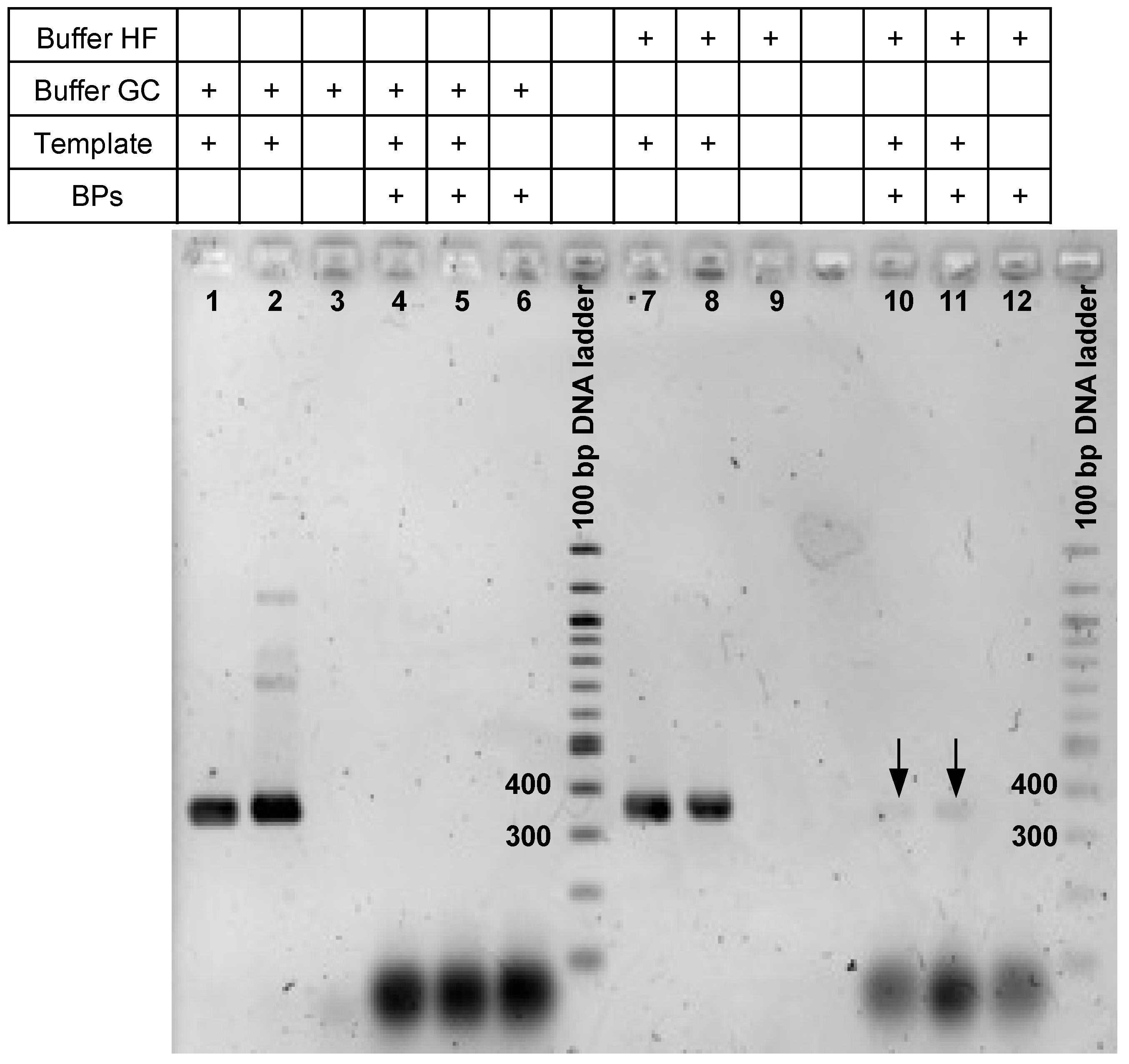
3.2. Checking epicPCR Design on Pure Culture
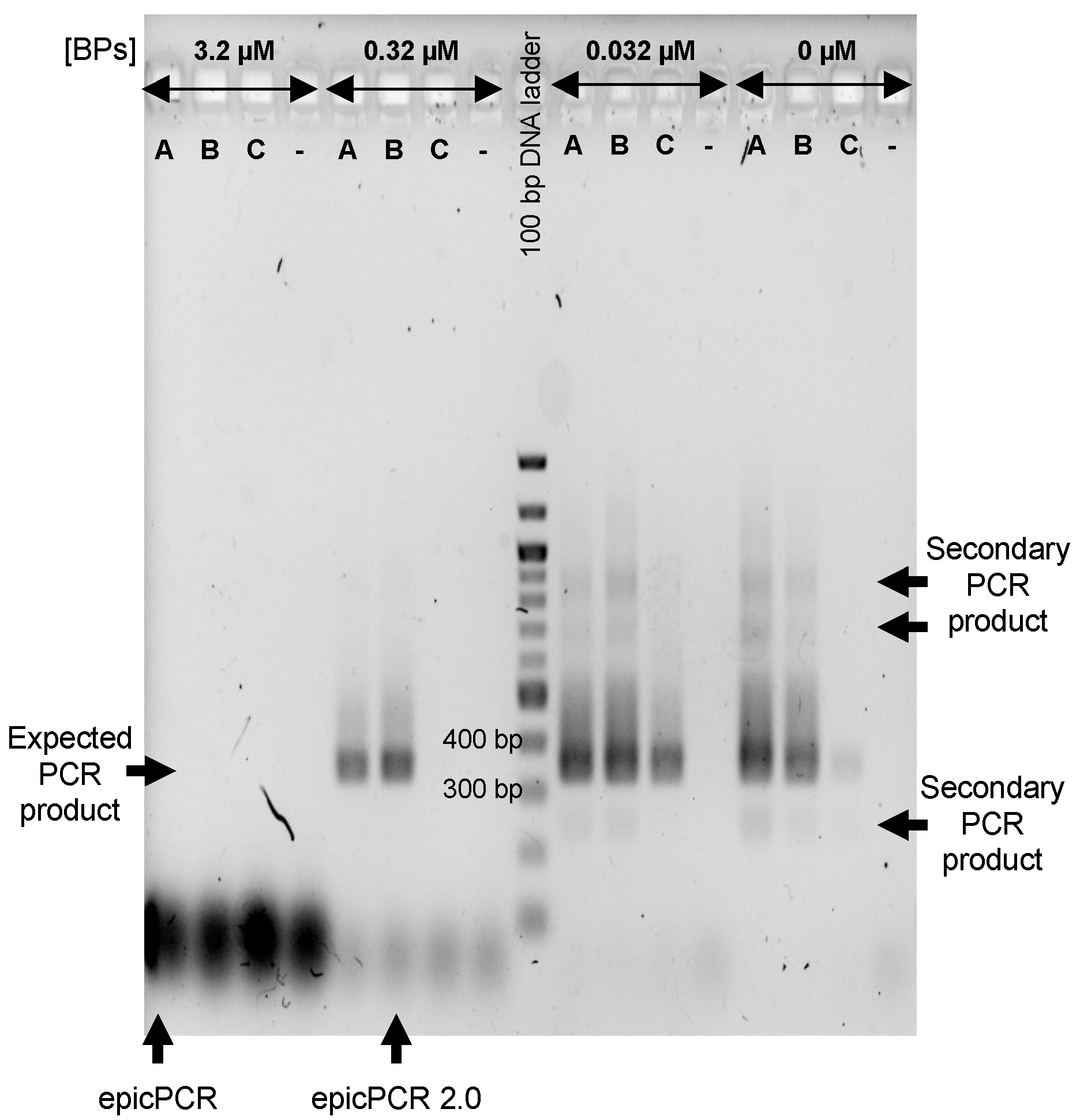
3.3. Tuning the Use of BP to Maximize epicPCR Efficiency on Environmental Cells
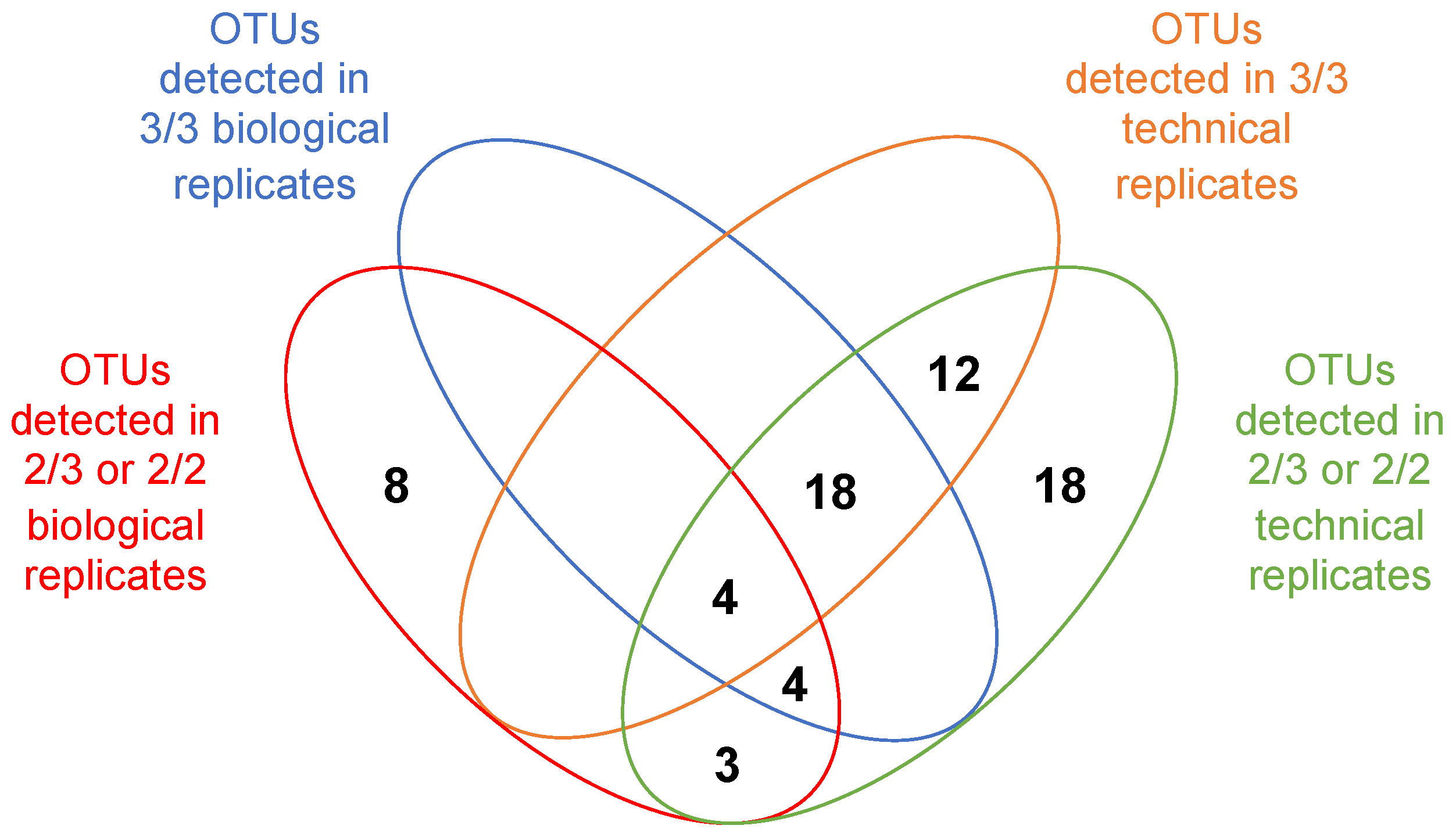
3.4. Exploring the Way to Validate OTUs in epicPCR Experiments
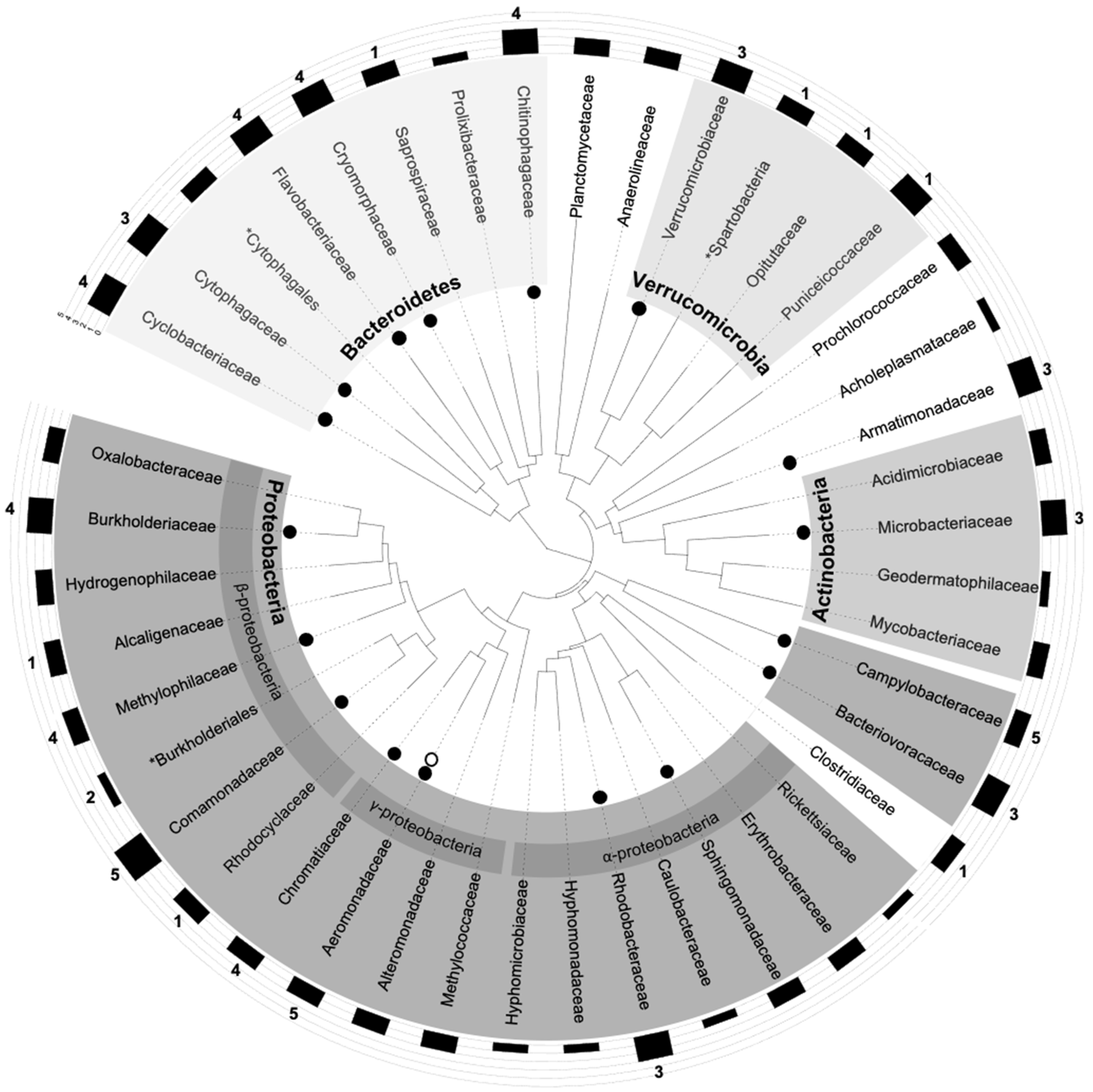
3.5. Plotting epicPCR Results on a Gold-Standard 16S-rDNA Tree
3.6. Synthetic Comparison of epicPCR and epicPCR 2.0 Methodologies
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Collignon, P.; Beggs, J.J.; Walsh, T.R.; Gandra, S.; Laxminarayan, R. Anthropological and socioeconomic factors contributing to global antimicrobial resistance: A univariate and multivariable analysis. Lancet Planet. Health 2018, 2, e398–e405. [Google Scholar] [CrossRef]
- Partridge, S.R.; Kwong, S.M.; Firth, N.; Jensen, S.O. Mobile genetic elements associated with antimicrobial resistance. Clin. Microbiol. Rev. 2018, 31, e00088-17. [Google Scholar] [CrossRef] [Green Version]
- Bengtsson-Palme, J.; Larsson, D.J.; Kristiansson, E. Using metagenomics to investigate human and environmental resistomes. J. Antimicrob. Chemother. 2017, 72, 2690–2703. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Li, J.; Chen, H.; Bond, P.L.; Yuan, Z. Metagenomic analysis reveals wastewater treatment plants as hotspots of antibiotic resistance genes and mobile genetic elements. Water Res. 2017, 123, 468–478. [Google Scholar] [CrossRef]
- Larsson, D.J.; Andremont, A.; Bengtsson-Palme, J.; Brandt, K.K.; de Roda Husman, A.M.; Fagerstedt, P.; Kvint, K.; Laxminarayan, R.; Manaia, C.M.; Nielsenn, K.M.; et al. Critical knowledge gaps and research needs related to the environmental dimensions of antibiotic resistance. Environ. Int. 2018, 117, 132–138. [Google Scholar] [CrossRef]
- Stalder, T.; Press, M.O.; Sullivan, S.; Liachko, I.; Top, E.M. Linking the resistome and plasmidome to the microbiome. ISME J. 2019, 13, 2437–2446. [Google Scholar] [CrossRef] [Green Version]
- Spencer, S.J.; Tamminen, M.V.; Preheim, S.P.; Guo, M.T.; Briggs, A.W.; Brito, I.L.; Weitz, D.A.; Pitkänen, L.K.; Vigneault, F.; Virta, M.P.J. Massively parallel sequencing of single cells by epicPCR links functional genes with phylogenetic markers. ISME J. 2016, 10, 427–436. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hultman, J.; Tamminen, M.; Pärnänen, K.; Cairns, J.; Karkman, A.; Virta, M.P.J. Host range of antibiotic resistance genes in wastewater treatment plant influent and effluent. FEMS Microbiol. Ecol. 2018, 94, fiy038. [Google Scholar] [CrossRef]
- Roman, V.L.; Merlin, C.; Baron, S.; Larvor, E.; Ledevendec, L.; Virta, M.P.J.; Bellanger, X. Abundance and environmental host range of the SXT/R391 ICE family of in aquatic microbial communities. Environ. Pollut. 2021, 117673. [Google Scholar] [CrossRef] [PubMed]
- Bioteau, A.; Durand, R.; Burrus, V. Redefinition and unification of the SXT/R391 family of integrative and conjugative elements. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [Green Version]
- Espéli, O.; Moulin, L.; Boccard, F. Transcription attenuation associated with bacterial repetitive extragenic BIME elements. J. Mol. Biol. 2001, 314, 375–386. [Google Scholar] [CrossRef]
- Hochhut, B.; Waldor, M.K. Site-specific integration of the conjugal Vibrio cholerae SXT element into prfC. Mol. Microbiol. 1999, 32, 99–110. [Google Scholar] [CrossRef]
- Federation, W.E.; APH Association. Standard Methods for the Examination of Water and Wastewater; American Public Health Association (APHA): Washington, DC, USA, 2005. [Google Scholar]
- Kirschner, A.K.T.; Pleininger, S.; Jakwerth, S.; Rehak, S.; Farnleitner, A.H.; Huhulescu, S.; Indra, A. Application of three different methods to determine the prevalence, the abundance and the environmental drivers of culturable Vibrio cholerae in fresh and brackish bathing waters. J. Appl. Microbiol. 2018, 125, 1186–1198. [Google Scholar] [CrossRef] [PubMed]
- Boulos, L.; Prevost, M.; Barbeau, B.; Coallier, J.; Desjardins, R. LIVE/DEAD® BacLight™: Application of a new rapid staining method for direct enumeration of viable and total bacteria in drinking water. J. Microbiol. Methods 1999, 37, 77–86. [Google Scholar] [CrossRef]
- Zhang, J.; Kobert, K.; Flouri, T.; Stamatakis, A. PEAR: A fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics 2014, 30, 614–620. [Google Scholar] [CrossRef] [Green Version]
- Andrews, S. FastQC: A Quality Control Tool for High Throughput Sequence Data. 2010. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 21 October 2020).
- Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011, 17, 10–12. [Google Scholar] [CrossRef]
- Edgar, R.C. Search and clustering orders of magnitude faster than BLAST. Bioinformatics 2010, 26, 2460–2461. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2017; Available online: https://www.R-project.org/ (accessed on 23 June 2020).
- Madeira, F.; Park, Y.M.; Lee, J.; Buso, N.; Gur, T.; Madhusoodanan, N.; Basutkar, P.; Tivey, A.R.N.; Potter, C.S.; Finn, R.D. The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res. 2019, 47, W636–W641. [Google Scholar] [CrossRef] [Green Version]
- Price, M.N.; Dehal, P.S.; Arkin, A.P. FastTree: Computing large minimum evolution trees with profiles instead of a distance matrix. Mol. Biol. Evol. 2009, 26, 1641–1650. [Google Scholar] [CrossRef] [PubMed]
- Letunic, I.; Bork, P. Interactive Tree of Life (iTOL) v4: Recent updates and new developments. Nucleic Acids Res. 2019, 47, W256–W259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Duhamel, S.; Jacquet, S. Flow cytometric analysis of bacteria- and virus-like particles in lake sediments. J. Microbiol. Methods 2006, 64, 316–332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Patel, A.; Noble, R.T.; Steele, J.A.; Schwalbach, M.S.; Hewson, I.; Fuhrman, J.A. Virus and prokaryote enumeration from planktonic aquatic environments by epifluorescence microscopy with SYBR Green I. Nat. Protoc. 2007, 2, 269. [Google Scholar] [CrossRef] [PubMed] [Green Version]

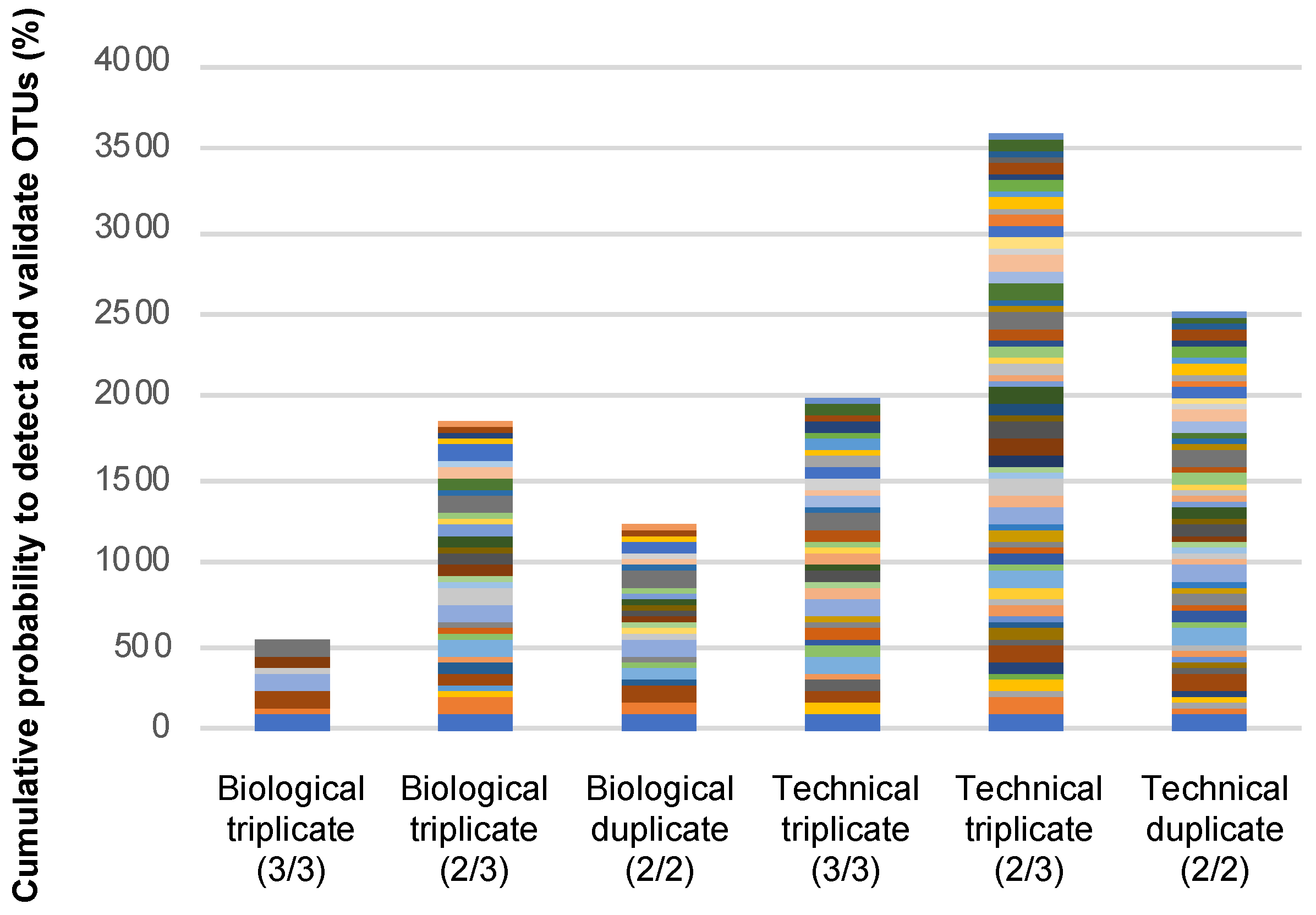
| Characteristics | epicPCR | epicPCR 2.0 |
|---|---|---|
| Polyacrylamide beads formation | Done once per sample | Done three times in parallel per samples |
| Beads formation quality check (by microscopy after cell staining) | More than 90% of beads are empty | More than 90% of beads are empty and 85% of non-empty beads only carry one cell |
| Replicate strategy for performing the fusion-PCR | Triplicated amplification using a single beads batch per sample as template (=technical triplicates) | Single non-replicated amplification performed on each of the 3 beads batches (=biological triplicates) |
| Fusion-PCR 1,2 | Performed with primers F1, R1-F2′ and R2 used at 1, 0.01 and 1 µM, respectively | |
| Step between the fusion- and the nested-PCRs | None | Additional “blocking PCR” step using the blocking primers at 3.2 µM alone |
| Nested-PCR2 | Performed with the primers F3 + SA and R3 + SA at 0.3 µM and blocking primers at 3.2 µM | Performed with the primers F3 + SA and R3 + SA at 0.3 µM and blocking primers at 0.32 µM |
| Minimum of reads for creating an OTU in each replicate | 100 | 10 |
| Preferential OTU validation method | OTU detected in each of the 3 technical replicates | OTU detected in each of the 3 biological replicates |
| Confidence levels for characterizing OTU validation | None | Yes; from 5, the best one, to 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Roman, V.L.; Merlin, C.; Virta, M.P.J.; Bellanger, X. EpicPCR 2.0: Technical and Methodological Improvement of a Cutting-Edge Single-Cell Genomic Approach. Microorganisms 2021, 9, 1649. https://doi.org/10.3390/microorganisms9081649
Roman VL, Merlin C, Virta MPJ, Bellanger X. EpicPCR 2.0: Technical and Methodological Improvement of a Cutting-Edge Single-Cell Genomic Approach. Microorganisms. 2021; 9(8):1649. https://doi.org/10.3390/microorganisms9081649
Chicago/Turabian StyleRoman, Véronica L., Christophe Merlin, Marko P. J. Virta, and Xavier Bellanger. 2021. "EpicPCR 2.0: Technical and Methodological Improvement of a Cutting-Edge Single-Cell Genomic Approach" Microorganisms 9, no. 8: 1649. https://doi.org/10.3390/microorganisms9081649
APA StyleRoman, V. L., Merlin, C., Virta, M. P. J., & Bellanger, X. (2021). EpicPCR 2.0: Technical and Methodological Improvement of a Cutting-Edge Single-Cell Genomic Approach. Microorganisms, 9(8), 1649. https://doi.org/10.3390/microorganisms9081649






