Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota
Abstract
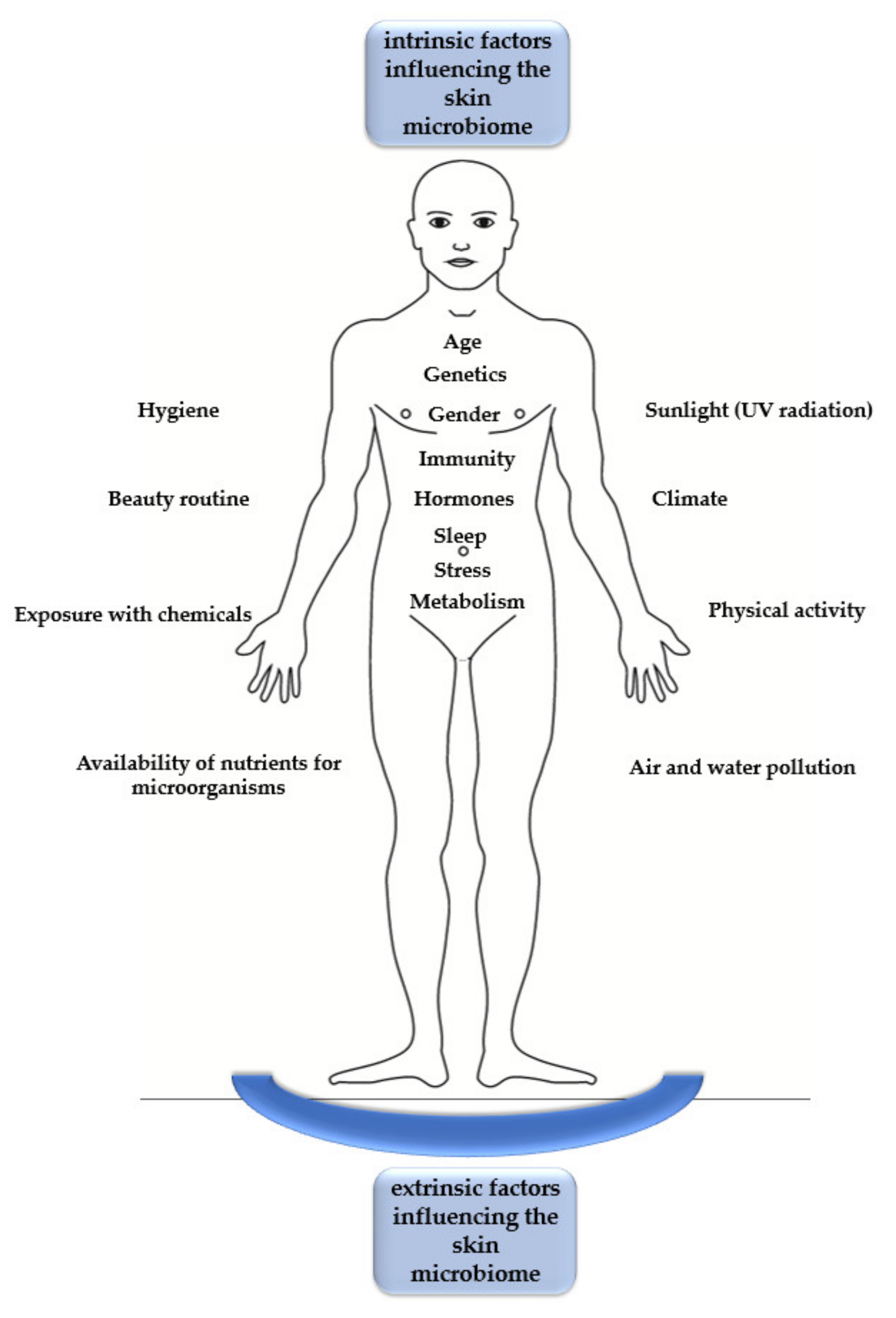
1. Introduction
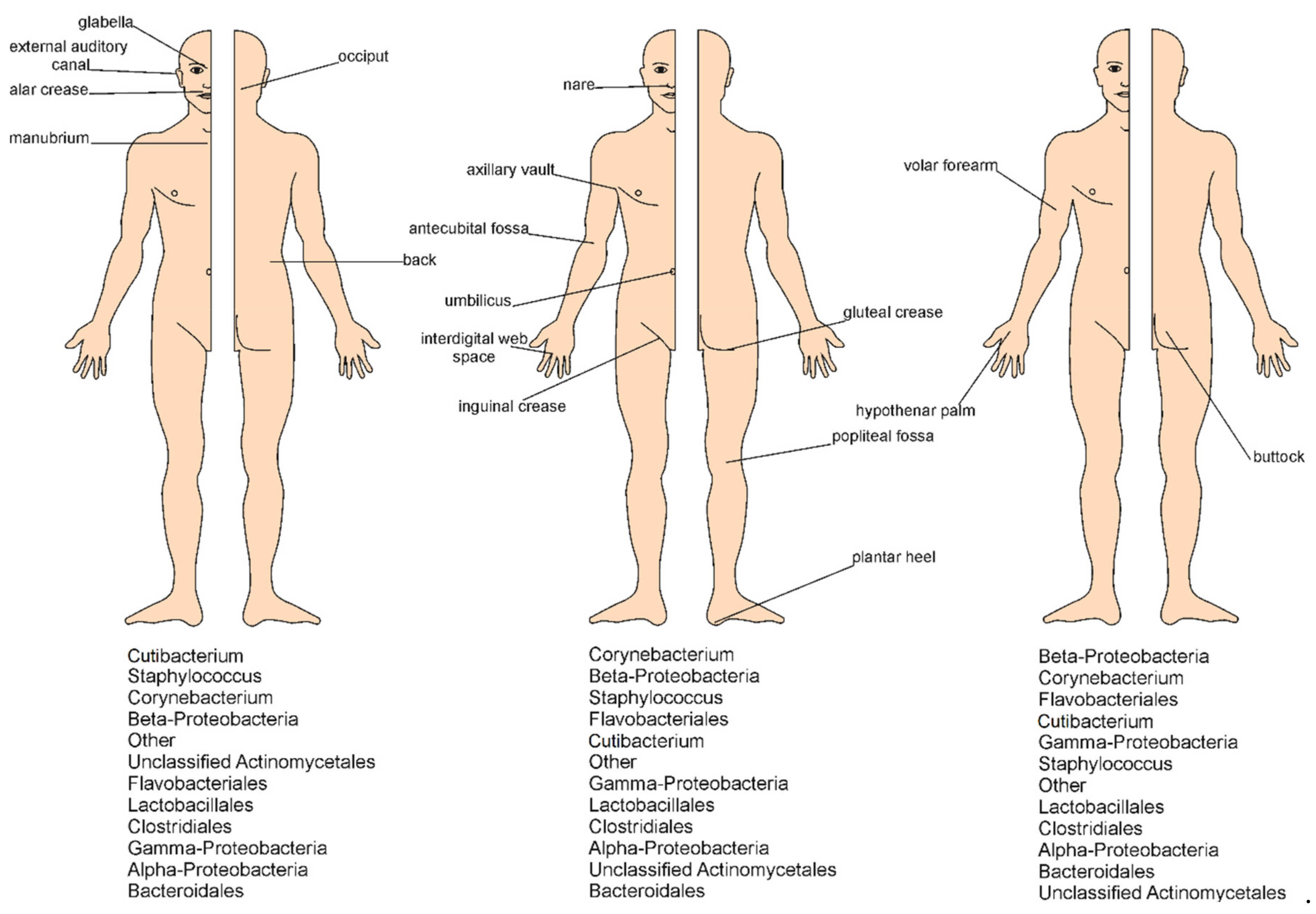
2. Microorganisms Inhabiting the Skin and Methods of Their Identification
3. Interactions between Skin Microorganisms
4. Skin as a Living Environment for Microorganisms
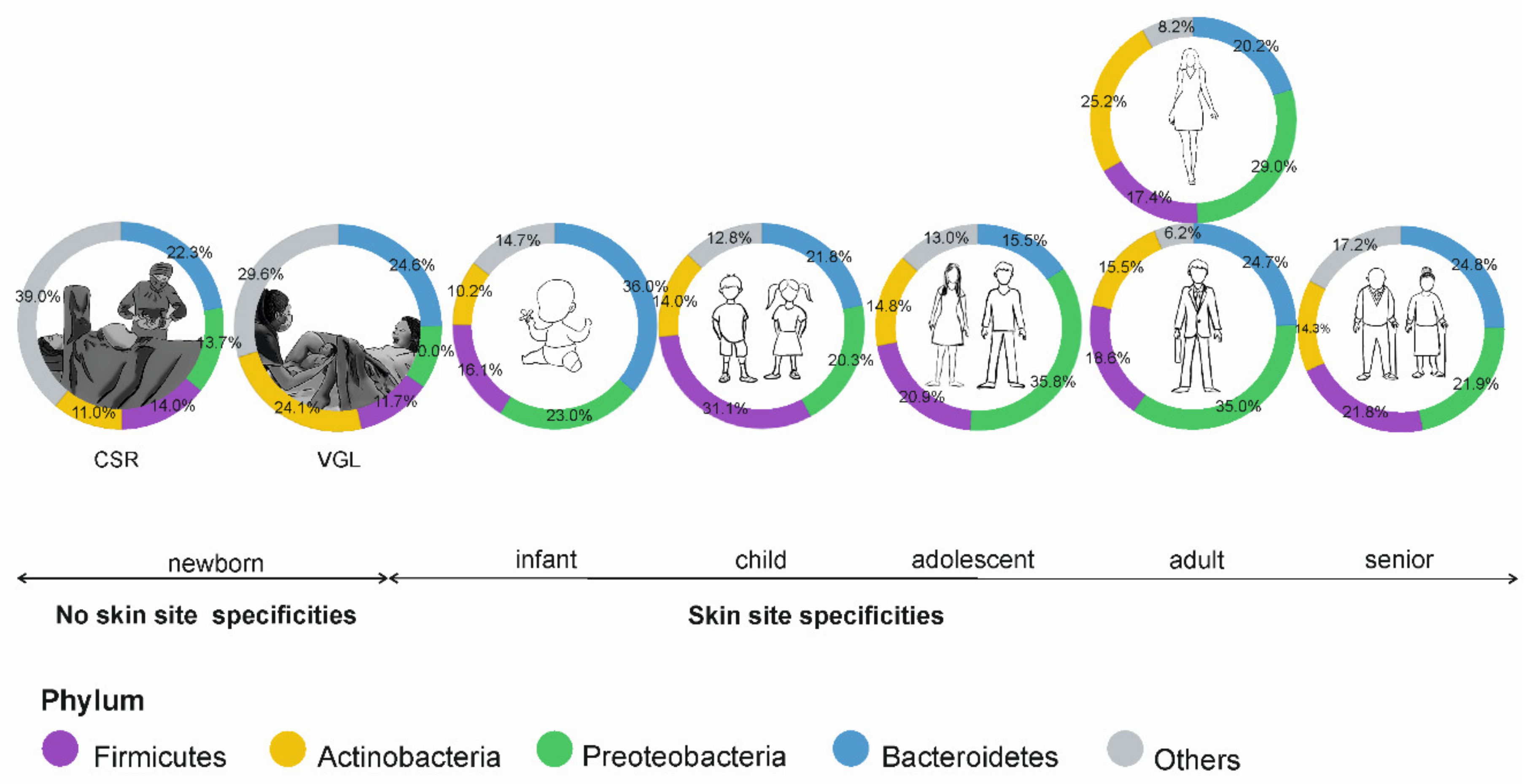
5. The Skin Microbiome in Different Stages of Human Development
5.1. Prenatal Stage and Childhood
5.2. Puberty and Adulthood
6. Gender and the Skin Microbiome
7. Ethnicity and the Skin Microbiome
8. Living and Working Environment
9. Antibiotics
10. Microbiome Transplantation—An Alternative to Antibiotics in the Treatment of Skin Diseases
11. Cosmetics and Clothing Textiles
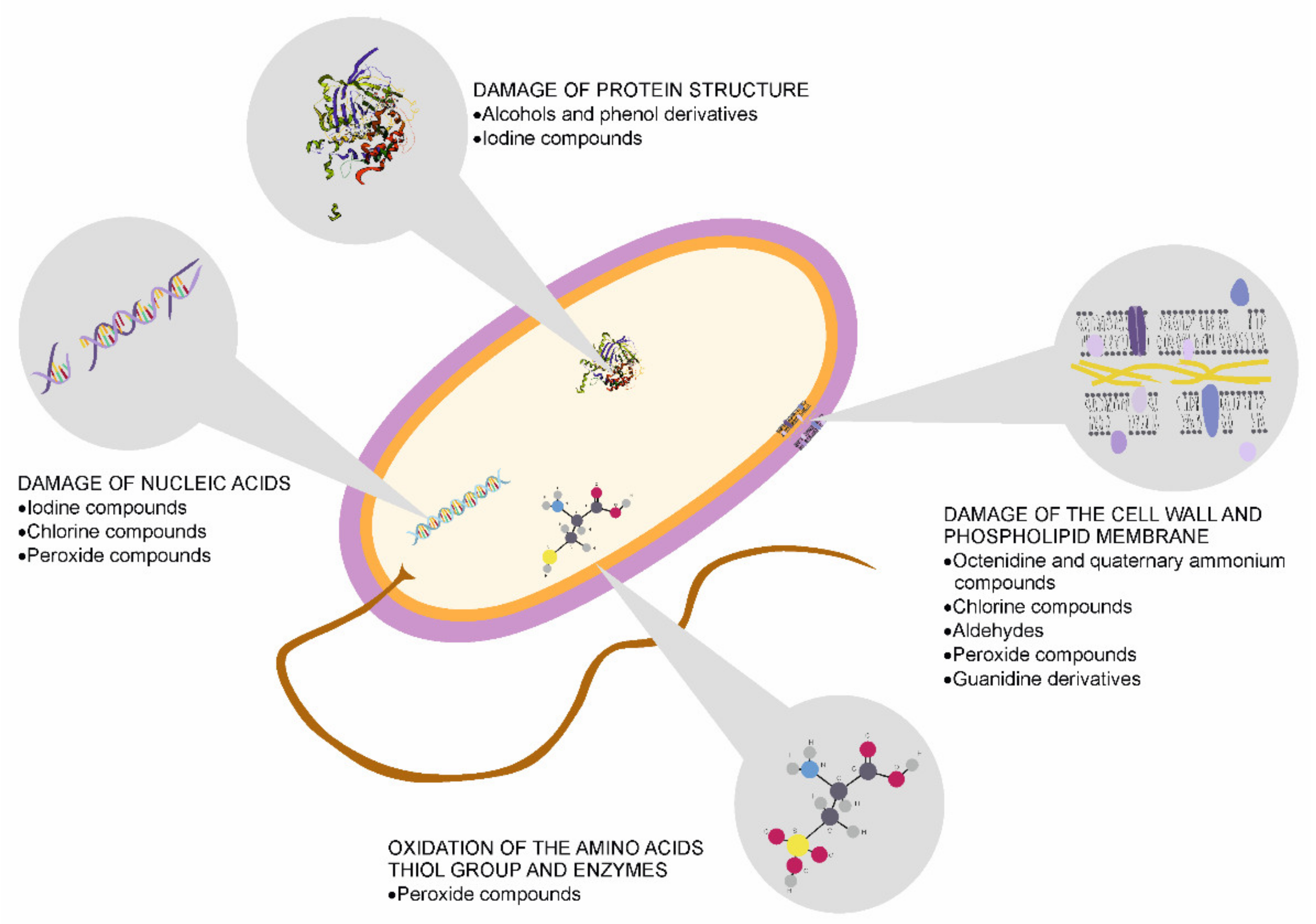
12. Skin Disinfection and Its Influence on the Microbiome Condition
13. Gut–Skin Axis
14. Summary
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Gallo, R.L. Human Skin Is the Largest Epithelial Surface for Interaction with Microbes. J. Investig. Dermatol. 2017, 137, 1213–1214. [Google Scholar] [CrossRef]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Cundell, A.M. Microbial Ecology of the Human Skin. Microb. Ecol. 2018, 76, 113–120. [Google Scholar] [CrossRef]
- Malinowska, M.; Tokarz-Deptuła, B.; Deptuła, W. Mikrobiom człowieka. Post. Mikrobiol. 2017, 56, 33–42. [Google Scholar]
- Byrd, A.L.; Belkaid, Y.; Segre, J.A. The human skin microbiome. Nat. Rev. Microbiol. 2018, 16, 143–155. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Hata, T. Dysbiosis of the Skin Microbiome in Atopic Dermatitis. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020. [Google Scholar]
- Probst, A.J.; Auerbach, A.K.; Moissl-Eichinger, C. Archaea on human skin. PLoS ONE 2013, 8, e65388. [Google Scholar] [CrossRef] [PubMed]
- Patra, V.; Byrne, S.N.; Wolf, P. The skin microbiome: Is it affected by UV-induced immune suppression? Front. Microbiol. 2016, 7, 1235. [Google Scholar] [CrossRef] [PubMed]
- Manrique, P.; Bolduc, B.; Walk, S.T.; van der Oost, J.; de Vos, W.M.; Young, M.J. Healthy human gut phageome. Proc. Natl. Acad. Sci. USA 2016, 113, 10400–10405. [Google Scholar] [CrossRef] [PubMed]
- Verbanic, S.; Kim, C.Y.; Deacon, J.M.; Chen, I.A. Improved single-swab sample preparation for re-covering bacterial and phage DNAfrom human skin and wound microbiomes. BMC Microbiol. 2019, 19, 214. [Google Scholar] [CrossRef] [PubMed]
- Andersen, B.M. Prevention and Control of Infections in Hospitals: Practice and Theory, 1st ed.; Springer Nature: Cham, Switzerland, 2018; pp. 337–437. [Google Scholar]
- Kerk, S.K.; Lai, H.Y.; Sze, S.K.; Ng, K.W.; Schmidtchen, A.; Adav, S.S. Bacteria Display Differential Growth and Adhesion Characteristics on Human Hair Shafts. Front. Microbiol. 2018, 9, 2145. [Google Scholar] [CrossRef]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; Green, E.D.; et al. Topographical and Temporal Diversity of the Human Skin Microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef]
- Murillo, N.; Raoult, D. Skin microbiota: Overview and role in the skin diseases acne vulgaris and rosacea. Future Microbiol. 2013, 8, 209–222. [Google Scholar] [CrossRef] [PubMed]
- Buerger, S. The Skin and Oral Microbiome: An Examination of Overlap and Potential Interactions between Microbiome Communities. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 45–58. [Google Scholar]
- Samaras, S.; Hoptroff, M. The Microbiome of Healthy Skin. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 3–32. [Google Scholar]
- Li, H.; Goh, B.N.; The, W.K.; Jiang, Z.; Goh, J.P.Z.; Goh, A.; Wu, G.; Hoon, S.S.; Raida, M.; Camattari, A.; et al. Skin Commensal Malassezia globosa Secreted Protease Attenuates Staphylococcus aureus Biofilm Formation. J. Investig. Dermatol. 2018, 138, 1137–1145. [Google Scholar] [CrossRef]
- Bay, L.; Barnes, C.J.; Fritz, B.G.; Thorsen, J.; Restrup, M.E.M.; Rasmussen, L.; Sørensen, J.K.; Hesselvig, A.B.; Odgaard, A.; Hansen, A.J.; et al. Universal dermal microbiome in human skin. mBio 2020, 11. [Google Scholar] [CrossRef]
- Gao, Z.; Tseng, C.; Pei, Z.; Blaser, M.L. Molecular analysis of human forearm superficial skin bacterial biota. Proc. Natl. Acad. Sci. USA 2007, 104, 2927–2932. [Google Scholar] [CrossRef] [PubMed]
- Romano-Bertrand, S.; Bourdier, A.; Aujoulat, F.; Michon, A.-L.; Masnou, A.; Parer, S.; Marchandin, H.; Jumas-Bilak, E. Skin microbiota is the main reservoir of Roseomonas mucosa, an emerging opportunistic pathogen so far assumed to be environmental. Clin. Microbiol. Infect. 2016, 22, 737.e1–737.e7. [Google Scholar] [CrossRef] [PubMed]
- Sherwani, M.A.; Tufail, S.; Muzaffar, A.F.; Yusuf, N. The skin microbiome and immune system: Potential target for chemoprevention? Photodermatol. Photoimmunol. Photomed. 2018, 34, 25–34. [Google Scholar] [CrossRef]
- Prescott, S.L.; Larcombe, D.L.; Logan, A.C.; West, C.; Burks, W.; Caraballo, L.; Levin, M.; Etten, E.V.; Horwitz, P.; Kozyrskyj, A.; et al. The skin microbiome: Impact of modern environments on skin ecology, barrier integrity, and systemic immune programming. World Allergy Organ. J. 2017, 10, 29. [Google Scholar] [CrossRef]
- Flowers, L.; Grice, E.A. The Skin Microbiota: Balancing Risk and Reward. Cell Host Microbe 2020, 28, 190–200. [Google Scholar] [CrossRef]
- Borkowski, A.W.; Gallo, R.L. The coordinated response of the physical and antimicrobial peptide barriers of the skin. J. Investig. Dermatol. 2011, 131, 285–287. [Google Scholar] [CrossRef] [PubMed]
- Bardan, A.; Nizet, V.; Gallo, R.L. Antimicrobial peptides and the skin. Exp. Opin. Biol. Ther. 2004, 4, 543–549. [Google Scholar] [CrossRef] [PubMed]
- Kiatsurayanon, C.; Ogawa, H.; Niyonsaba, F. The Role of Host Defense Peptide Human β-defensins in the Maintenance of Skin Barriers. Curr. Pharm. Des. 2018, 24, 1092–1099. [Google Scholar] [CrossRef]
- Farahmand, S. Microbiome of Compromised Skin. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 145–170. [Google Scholar]
- Bolla, B.S.; Erdei, L.; Urbán, E.; Burián, K.; Kemény, L.; Szabó, K. Cutibacterium acnes regulates the epidermal barrier properties of HPV-KER human immortalized keratinocyte cultures. Sci. Rep. 2020, 10, 12815. [Google Scholar] [CrossRef] [PubMed]
- Dimitriu, P.A.; Iker, B.; Malik, K.; Leung, H.; Mohn, W.W.; Hillebrand, G.G. New Insights into the Intrinsic and Extrinsic Factors That Shape the Human Skin Microbiome. mBio 2019, 10, e00839-19. [Google Scholar] [CrossRef]
- Brown, M.M.; Horswill, A.R. Staphylococcus epidermidis—Skin friend or foe? PLoS Pathog. 2020, 16, e1009026. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.R.; Gallo, R.L. Evidence that Human Skin Microbiome Dysbiosis Promotes Atopic Dermatitis. J. Investig. Dermatol. 2017, 137, 2460–2461. [Google Scholar] [CrossRef]
- Yamazaki, Y.; Nakamura, Y.; Núñez, G. Role of the microbiota in skin immunity and atopic dermatitis. Allergol. Int. 2017, 66, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Fischer, C.L.; Wertz, P.W. Effects of Endogenous Lipids on the Skin Microbiome. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 219–236. [Google Scholar]
- Prohic, A.; Sadikovic, T.J.; Krupalija-Fazlic, M.; Kuskunovic-Vlahovljak, S. Malassezia species in healthy skin and in dermatological conditions. Int. J. Dermatol. 2016, 55, 494–504. [Google Scholar] [CrossRef] [PubMed]
- Stehlikova, Z.; Kostovcik, M.; Kostovcikova, K.; Kverka, M.; Juzlova, K.; Rob, F.; Hercogova, J.; Bohac, P.; Pinto, Y.; Uzan, A.; et al. Dysbiosis of Skin Microbiota in Psoriatic Patients: Co-occurrence of Fungal and Bacterial Communities. Front. Microbiol. 2019, 10, 438. [Google Scholar] [CrossRef]
- James, A.G. The Axillary Microbiome and its Relationship with Underarm Odor. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 79–130. [Google Scholar]
- Chen, Y.E.; Tsao, H. The skin microbiome: Current perspectives and future challenges. J. Am. Acad. Dermatol. 2013, 69, 143–155. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Shi, L.; Ren, Y.; Tan, X.; Liu, W.; Liu, Z. Applications of Human Skin Microbiota in the Cutaneous Disorders for Ecology-Based Therapy. Front. Cell. Infect. Microbiol. 2020, 10, 570261. [Google Scholar] [CrossRef] [PubMed]
- Adamczyk, K.; Garncarczyk, A.; Antończak, P.; Wcisło-Dziadecka, D. The foot microbiome. J. Cosmet. Dermatol. 2020, 19, 1039–1043. [Google Scholar] [CrossRef]
- McCall, L.I.; Callewaert, C.; Zhu, Q.; Song, S.J.; Bouslimani, A.; Minich, J.J.; Ernst, M.; Ruiz-Calderon, J.F.; Cavallin, H.; Pereira, H.S.; et al. Home chemical and microbial transitions across urbanization. Nat. Microbiol. 2020, 5, 108–115. [Google Scholar] [CrossRef] [PubMed]
- Edmonds-Wilson, S.L.; Nurinova, N.I.; Zapka, C.A.; Fierer, N.; Wilson, M. Review of human hand microbiome research. J. Dermatol. Sci. 2015, 80, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Levy, G.; Solt, I. The Human Microbiome and Gender Medicine. Gender Genome 2018, 2, 123. [Google Scholar] [CrossRef]
- Dunn, A.B.; Jordan, S.; Baker, B.J.; Carlson, N.S. The Maternal Infant Microbiome: Considerations for Labor and Birth. MCN Am. J. Matern. Child. Nurs. 2017, 42, 318–325. [Google Scholar] [CrossRef]
- Baker, J.M.; Chase, D.M.; Herbst-Kralovetz, M.M. Uterine Microbiota: Residents, Tourists, or Invaders? Front. Immunol. 2018, 9, 208. [Google Scholar] [CrossRef]
- Aagaard, K.; Ma, J.; Antony, L.M.; Ganu, R.; Petrosino, J.; Versalovic, J. The Placenta Harbors a Unique Microbiome. Sci. Transl. Med. 2014, 6, 237–265. [Google Scholar] [CrossRef]
- Lauder, A.P.; Roche, A.M.; Sherrill-Mix, S.; Bailey, A.; Laughlin, A.L.; Bittinger, K.; Leite, R.; Elovitz, M.A.; Parry, S.; Bushman, F.D. Comparison of placenta samples with contamination controls does not provide evidence for a distinct placenta microbiota. Microbiome 2016, 4, 29. [Google Scholar] [CrossRef]
- Stamatas, G.N. Infant Skin Microbiome. In Skin Microbiome Handbook: From Basic Research to Product Development, 1st ed.; Dayan, N., Ed.; Scrivener Publishing LLC: Beverly, MA, USA, 2020; pp. 131–142. [Google Scholar]
- Zhu, T.; Liu, X.; Kong, F.Q.; Duan, Y.Y.; Yee, A.L.; Kim, M.; Galzote, C.; Gilbert, J.A.; Quan, Z.X. Age and Mothers: Potent Influences of Children’s Skin Microbiota. J. Investig. Dermatol. 2019, 139, 2497–2505. [Google Scholar] [CrossRef]
- Dominguez-Bello, M.G.; De Jesus-Laboy, K.M.; Shen, N.; Cox, L.M.; Amir, A.; Gonzalez, A.; Bokulich, N.A.; Song, S.J.; Hoashi, M.; Rivera-Vinas, J.I.; et al. Partial restoration of the microbiota of cesarean-born infants via vaginal microbial transfer. Nat. Med. 2016, 22, 250–253. [Google Scholar] [CrossRef] [PubMed]
- Stinson, L.F.; Payne, M.S.; Keelan, J.A. A Critical Review of the Bacterial Baptism Hypothesis and the Impact of Cesarean Delivery on the Infant Microbiome. Front. Med. (Lausanne) 2018, 5, 135. [Google Scholar] [CrossRef]
- Lehtimäki, J.; Karkman, A.; Laatikainen, T.; Paalanen, L.; Hertzen, L.; Haahtela, T.; Hanski, I.; Ruokolainen, L. Patterns in the skin microbiota differ in children and teenagers between rural and urban environments. Sci. Rep. 2017, 7, 45651. [Google Scholar] [CrossRef]
- Li, Z.; Bai, X.; Peng, T.; Yi, X.; Luo, L.; Yang, J.; Liu, J.; Wang, Y.; He, T.; Wang, X.; et al. New Insights Into the Skin Microbial Communities and Skin Aging. Front. Microbiol. 2020, 11, 565549. [Google Scholar] [CrossRef]
- Jo, J.-H.; Deming, C.; Kennedy, E.A.; Conlan, S.; Polley, E.C.; Ng, W.-L.; NISC Comparative Sequencing Program; Segre, J.A.; Kong, H.H. Diverse Human Skin Fungal Communities in Children Converge in Adulthood NISC Comparative Sequencing Program. J. Investig. Dermatol. 2016, 136, 2356–2363. [Google Scholar] [CrossRef]
- Shibagaki, N.; Suda, W.; Clavaud, C.; Bastien, P.; Takayasu, L.; Iioka, E.; Kurokawa, R.; Yamashita, N.; Hattori, Y.; Shindo, C.; et al. Aging-related changes in the diversity of women’s skin microbiomes associated with oral bacteria. Sci. Rep. 2017, 7, 10567. [Google Scholar] [CrossRef]
- Juge, R.; Rouaud-Tinguely, P.; Breugnot, J.; Servaes, K.; Grimaldi, C.; Roth, M.P.; Coppin, H.; Closs, B. Shift in skin microbiota of Western European women across aging. J. Appl. Microbiol. 2018, 125, 907–916. [Google Scholar] [CrossRef] [PubMed]
- Fournière, M.; Latire, T.; Souak, D.; Feuilloley, M.G.J.; Bedoux, G. Staphylococcus epidermidis and Cutibacterium acnes: Two Major Sentinels of Skin Microbiota and the Influence of Cosmetics. Microorganisms 2020, 8, 1752. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Budding, A.E.; van der Lugt-Degen, M.; Du-Thumm, L.; Vandeven, M.; Fan, A. The influence of age, gender and race/ethnicity on the composition of the human axillary microbiome. Int. J. Cosmet. Sci. 2019, 41, 371–377. [Google Scholar] [CrossRef]
- Ying, S.; Zeng, D.-N.; Chi, L.; Tan, Y.; Galzote, C.; Cardona, C.; Lax, S.; Gilbert, J.; Quan, Z.X. The Influence of Age and Gender on Skin-Associated Microbial Communities in Urban and Rural Human Populations. PLoS ONE 2015, 10, e0141842. [Google Scholar] [CrossRef]
- Ehlers, C.; Ivens, U.I.; Møller, M.L.; Senderovitz, T.; Serup, J. Females have lower skin surface pH than men. A study on the surface of gender, forearm site variation, right/left difference and time of the day on the skin surface pH. Skin Res. Technol. 2001, 7, 90–94. [Google Scholar] [CrossRef] [PubMed]
- Flak, M.B.; Neves, J.F.; Blumberg, R.S. Immunology. Welcome to the microgenderome. Science 2013, 339, 1044–1045. [Google Scholar] [CrossRef]
- Zapka, C.; Campbell, E.; Maxwell, S.; Gerba, C.; Dolan, M.; Arbogast, J.; Macinga, D. Bacterial hand contamination and transfer after use of contaminated bulk-soap-refillable dispensers. Appl. Environ. Microbiol. 2011, 77, 2898–2904. [Google Scholar] [CrossRef]
- Holland, K.T.; Bojar, R.A. Cosmetics: What is their influence on the skin microflora. Am. J. Clin. Dermatol. 2002, 3, 445–449. [Google Scholar] [CrossRef]
- Bouslimani, A.; da Silva, R.; Kosciolek, T.; Janssen, S.; Callewaert, C.; Amir, A.; Dorrestein, K.; Melnik, A.V.; Zaramela, L.S.; Kim, J.N.; et al. The impact of skin care products on skin chemistry and microbiome dynamics. BMC Biol. 2019, 17, 47. [Google Scholar] [CrossRef]
- Vandegrift, R.; Bateman, A.C.; Siemens, K.N.; Nguyen, M.; Wilson, H.E.; Green, J.L.; Van Den Wymelenberg, K.G.; Hickey, R.J. Cleanliness in context: Reconciling hygiene with a modern microbial perspective. Microbiome 2017, 5, 76. [Google Scholar] [CrossRef] [PubMed]
- Perez Perez, G.I.; Gao, Z.; Jourdain, R.; Ramirez, J.; Gany, F.; Clavaud, C.; Demaude, J.; Breton, L.; Blaser, M.J. Body Site Is a More Determinant Factor than Human Population Diversity in the Healthy Skin Microbiome. PLoS ONE 2016, 11, e0151990. [Google Scholar] [CrossRef] [PubMed]
- Leung, M.H.; Wilkins, D.; Lee, P.K. Insights into the pan-microbiome: Skin microbial communities of Chinese individuals differ from other racial groups. Sci. Rep. 2015, 5, 11845. [Google Scholar] [CrossRef] [PubMed]
- Hospodsky, D.; Pickering, A.J.; Julian, T.R.; Miller, D.; Gorthala, S.; Boehm, A.B.; Peccia, J. Hand bacterial communities vary across two different human populations. Microbiology 2014, 160, 1144–1152. [Google Scholar] [CrossRef] [PubMed]
- Trinh, P.; Zaneveld, J.R.; Safranek, S.; Rabinowitz, P.P. One Health Relationships between Human, Animal, and Environmental Microbiomes: A Mini-Review. Front Public Health 2018, 6, 235. [Google Scholar] [CrossRef] [PubMed]
- Capone, K.A.; Dowd, S.E.; Stamatas, G.N.; Nikolovski, J. Diversity of the human skinmicrobiome early in life. J. Investig. Dermatol. 2011, 131, 2026–2032. [Google Scholar] [CrossRef]
- Song, S.J.; Lauber, C.; Costello, E.K.; Lozupone, C.A.; Humphrey, G.; Berg-Lyons, D.; Caporaso, J.G.; Knights, D.; Clemente, J.C.; Nakielny, S.; et al. Cohabiting family members share microbiota with one another and with their dogs. eLife 2013, 2, e00458. [Google Scholar] [CrossRef] [PubMed]
- Kraemer, J.G.; Ramette, A.; Aebi, S.; Oppliger, A.; Hilty, M. Influence of pig farming on the human nasal microbiota: Key role of airborne microbial communities. Appl. Environ. Microbiol. 2018, 84. [Google Scholar] [CrossRef] [PubMed]
- Mosites, E.; Sammons, M.; Otiang, E.; Eng, A.; Noecker, C.; Manor, O.; Hilton, S.; Thumbi, S.M.; Onyango, C.; Garland-Lewis, G.; et al. Microbiome sharing between children, livestock and household surfaces in western Kenya. PLoS ONE 2017, 12, e0171017. [Google Scholar] [CrossRef]
- Torres, S.; Clayton, J.B.; Danzeisen, J.L.; Ward, T.; Huang, H.; Knights, D.; Johnson, T.J. Diverse bacterial communities exist on canine skin and are impacted by cohabitation and time. PeerJ 2017, 5, 3075. [Google Scholar] [CrossRef]
- Grönroos, M.; Parajuli, A.; Laitinen, O.H.; Roslund, M.I.; Vari, H.K.; Hyöty, H.; Puhakka, R.; Sinkkonen, A. Short-term direct contact with soil and plant materials leads to an immediate increase in diversity of skin microbiota. MicrobiologyOpen 2019, 8, e00645. [Google Scholar] [CrossRef] [PubMed]
- Callewaert, C.; Ravard Helffer, K.; Lebaron, P. Skin Microbiome and its Interplay with the Environment. Am. J. Clin. Dermatol. 2020, 21, 4–11. [Google Scholar] [CrossRef]
- Leung, M.H.Y.; Lee, P.K.H. The roles of the outdoors and occupants in contributing to a potential pan-microbiome of the built environment: A review. Microbiome 2016, 4, 21. [Google Scholar] [CrossRef]
- Peng, M.; Biswas, D. Environmental Influences of High-Density Agricultural Animal Operation on Human Forearm Skin Microflora. Microorganisms 2020, 8, 1481. [Google Scholar] [CrossRef]
- Burns, E.M.; Ahmed, H.; Isedeh, P.N.; Kohli, I.; Van Der Pol, W.; Shaheen, A.; Muzaffar, A.F.; Al-Sadek, C.; Foy, T.M.; Abdelgawwad, M.S.; et al. Ultraviolet radiation, both UVA and UVB, influences the composition of the skin microbiome. Exp. Dermatol. 2019, 28, 136–141. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Zhu, W.; Shu, M.; Jiang, Y.; Gallo, R.L.; Liu, Y.T.; Huang, C.M. The response of human skin commensal bacteria as a reflection of UV radiation: UV-B decreases porphyrin production. PLoS ONE 2012, 7, e47798. [Google Scholar] [CrossRef]
- Chien, A.L.; Tsai, J.; Leung, S.; Mongodin, E.F.; Nelson, A.M.; Kang, S.; Garza, L.A. Association of Systemic Antibiotic Treatment of Acne With Skin Microbiota Characteristics. JAMA Dermatol. 2019, 155, 425–434. [Google Scholar] [CrossRef]
- Park, S.Y.; Kim, H.S.; Lee, S.H.; Kim, S. Characterization and Analysis of the Skin Microbiota in Acne: Impact of Systemic Antibiotics. J. Clin. Med. 2020, 9, 168. [Google Scholar] [CrossRef]
- Xu, H.; Li, H. Acne, the Skin Microbiome, and Antibiotic Treatment. Am. J. Clin. Dermatol. 2019, 20, 335–344. [Google Scholar] [CrossRef] [PubMed]
- Sardana, K.; Gupta, T.; Kumar, B.; Gautam, H.K.; Garg, V.K. Cross-sectional Pilot Study of Antibiotic Resistance in Propionibacterium Acnes Strains in Indian Acne Patients Using 16S-RNA Polymerase Chain Reaction: A Comparison Among Treatment Modalities Including Antibiotics, Benzoyl Peroxide, and Isotretinoin. Indian J. Dermatol. 2016, 61, 45–52. [Google Scholar] [CrossRef]
- Lim, J.S.; Park, H.S.; Cho, S.; Yoon, H.S. Antibiotic Susceptibility and Treatment Response in Bacterial Skin Infection. Ann. Dermatol. 2018, 30, 186–191. [Google Scholar] [CrossRef]
- Fukunaga, B.T.; Sumida, W.K.; Taira, D.A.; Davis, J.W.; Seto, T.B. Hospital-Acquired Methicillin-resistant Staphylococcus aureus Bacteremia Related to Medicare Antibiotic Prescriptions: A State-Level Analysis. Hawaii J. Med. Public Health 2016, 75, 303–309. [Google Scholar] [PubMed]
- Kourtis, A.P.; Hatfield, K.; Baggs, J.; Mu, Y.; See, I.; Epson, E.; Nadle, J.; Kainer, M.A.; Dumyati, G.; Petit, S.; et al. Vital Signs: Epidemiology and Recent Trends in Methicillin-Resistant and in Methicillin-Susceptible Staphylococcus aureus Bloodstream Infections—United States. MMWR Morb. Mortal. Wkly. Rep. 2019, 68, 214–219. [Google Scholar] [CrossRef]
- Callewaert, C.; Knödlseder, N.; Karoglan, A.; Güell, M.; Paetzold, B. Skin microbiome transplantation and manipulation: Current state of the art. Comput. Struct. Biotechnol. J. 2021, 19, 624–631. [Google Scholar] [CrossRef]
- Costello, E.K.; Lauber, C.L.; Hamady, M.; Fierer, N.; Gordon, J.I.; Knight, R. Bacterial community variation in human body habitats across space and time. Science 2009, 326, 1694–1697. [Google Scholar] [CrossRef]
- Perin, B.; Addetia, A.; Qin, X. Transfer of skin microbiota between two dissimilar autologous microenvironments: A pilot study. PLoS ONE 2019, 14, e0226857. [Google Scholar] [CrossRef] [PubMed]
- Paetzold, B.; Willis, J.R.; Pereira De Lima, J.; Knödlseder, N.; Brüggemann, H.; Quist, S.R.; Gabaldón, T.; Güell, M. Skin microbiome modulation induced by probiotic solutions. Microbiome 2019, 7, 95. [Google Scholar] [CrossRef]
- Myles, I.A.; Earland, N.J.; Anderson, E.D.; Moore, I.N.; Kieh, M.D.; Williams, K.W.; Saleem, A.; Fontecilla, N.M.; Welch, P.A.; Darnell, D.A.; et al. First-in-human topical microbiome transplantation with Roseomonas mucosa for atopic dermatitis. JCI Insight 2018, 3, e120608. [Google Scholar] [CrossRef]
- O’Sullivan, J.N.; Rea, M.C.; O’Connor, P.M.; Hill, C.; Ross, R.P. Human skin microbiota is a rich source of bacteriocin-producing staphylococci that kill human pathogens. FEMS Microbiol. Ecol. 2019, 95, fiy241. [Google Scholar] [CrossRef]
- Nakatsuji, T.; Chen, T.H.; Narala, S.; Chun, K.A.; Two, A.M.; Tong, Y.; Shafiq, F.; Kotol, P.F.; Bouslimani, A.; Melnik, A.V.; et al. Antimicrobials from human skin commensal bacteria protect against Staphylococcus aureus and are deficient in atopic dermatitis. Sci. Transl. Med. 2017, 9, eaah4680. [Google Scholar] [CrossRef]
- Callewaert, C.; Hutapea, P.; Van de Wiele, T.; Boon, N. Deodorants and antiperspirants affect the axillary bacterial community. Arch. Dermatol. Res. 2014, 306, 701–710. [Google Scholar] [CrossRef]
- Moskovicz, V.; Gross, A.; Mizrahi, B. Extrinsic Factors Shaping the Skin Microbiome. Microorganisms 2020, 8, 1023. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.J.; Jeong, S.E.; Lee, S.; Kim, S.; Han, H.; Jeon, C.O. Effects of cosmetics on the skin microbiome of facial cheeks with different hydration levels. MicrobiologyOpen 2018, 7, e00557. [Google Scholar] [CrossRef] [PubMed]
- Butler, É.; Lundqvist, C.; Axelsson, J. Lactobacillus reuteri DSM 17938 as a Novel Topical Cosmetic Ingredient: A Proof of Concept Clinical Study in Adults with Atopic Dermatitis. Microorganisms 2020, 8, 1026. [Google Scholar] [CrossRef]
- Frerejacques, M.; Rousselle, C.; Gauthier, L.; Cottet-Emard, S.; Derobert, L.; Roynette, A.; Lerch, T.Z.; Changey, F. Human Skin Bacterial Community Response to Probiotic (Lactobacillus reuteri DSM 17938) Introduction. Microorganisms 2020, 8, 1223. [Google Scholar] [CrossRef]
- Van Herreweghen, F.; Amberg, C.; Marques, R.; Callewaert, C. Biological and Chemical Processes that Lead to Textile Malodour Development. Microorganisms 2020, 8, 1709. [Google Scholar] [CrossRef]
- Munk, S.; Johansen, C.; Stahnke, L.H.; Adler-Nissen, J. Microbial survival and odor in laundry. J. Surfactants Deterg. 2001, 4, 385–394. [Google Scholar] [CrossRef]
- McQueen, H.R.; Harynuk, J.J.; Wismer, V.W.; Keelan, M.; Xu, Y.; Paulina, A. Axillary odour build-up in knit fabrics following multiple use cycles. Int. J. Cloth. Sci. Technol. 2014, 26, 274–290. [Google Scholar] [CrossRef]
- Lin, Q.; Lim, J.Y.C.; Xue, K.; Pek, Y.; Cally, O.; Pei Lin, C.; Jun, X. Sanitizing agents for virus inactivation and disinfection. Viewpoints 2020, 1, e16. [Google Scholar] [CrossRef]
- Ramirez-Arcos, S.; Goldman, M. Skin disinfection methods: Prospective evaluation and postimplementation results. Transfusion 2010, 50, 59–64. [Google Scholar] [CrossRef]
- Kundrapu, S.; Sunkesula, V.; Jury, I.; Deshpande, A.; Donskey, C.J. A randomized trial of soap and water hand wash versus alcohol hand rub for removal of Clostridium difficile spores from hands of patients. Infect. Control Hosp. Epidemiol. 2014, 35, 204–206. [Google Scholar] [CrossRef]
- Foddai, A.C.G.; Grant, I.R.; Dean, M. Efficacy of instant hand sanitizers against foodborne pathogens compared with hand washing with soap and water in food preparation settings: A systematic review. J. Food Prot. 2016, 79, 1040–1054. [Google Scholar] [CrossRef] [PubMed]
- Two, A.M.; Nakatsuji, T.; Kotol, P.F.; Arvanitidou, E.; Du-Thumm, L.; Hata, T.R.; Gallo, R.L. The Cutaneous Microbiome and Aspects of Skin Antimicrobial Defense System Resist Acute Treatment with Topical Skin Cleansers. J. Investig. Dermatol. 2016, 136, 1950–1954. [Google Scholar] [CrossRef]
- Golin, A.P.; Choi, D.; Ghahary, A. Hand sanitizers: A review of ingredients, mechanisms of action, modes of delivery, and efficacy against coronaviruses. Am. J. Infect. Control 2020, 48, 1062–1067. [Google Scholar] [CrossRef] [PubMed]
- San Miguel, A.J.; Meisel, J.S.; Horwinski, J.; Zheng, Q.; Bradley, C.W.; Grice, E.A. Antiseptic Agents Elicit Short-Term, Personalized, and Body Site-Specific Shifts in Resident Skin Bacterial Communities. J. Investig. Dermatol. 2018, 138, 2234–2243. [Google Scholar] [CrossRef]
- Sandle, T. Standards and controls for skin disinfection. Clin. Svcs. J. 2016, 15, 25–28. [Google Scholar]
- Sauerbrei, A. Bactericidal and virucidal activity of ethanol and povidone-iodine. MicrobiologyOpen 2020, 9, e1097. [Google Scholar] [CrossRef]
- Tolcher, M.C.; Whitham, M.D.; El-Nashar, S.A.; Clark, S.L. Chlorhexidine-Alcohol Compared with Povidone-Iodine Preoperative Skin Antisepsis for Cesarean Delivery: A Systematic Review and Meta-Analysis. Am. J. Perinatol. 2019, 36, 118–123. [Google Scholar] [PubMed]
- Mimoz, O.; Lucet, J.C.; Kerforne, T.; Pascal, J.; Souweine, B.; Goudet, V.; Mercat, A.; Bouadma, L.; Lasocki, S.; Alfandari, S.; et al. Skin antisepsis with chlorhexidine-alcohol versus povidone iodine-alcohol, with and without skin scrubbing, for prevention of intravascular-catheter-related infection (CLEAN): An open-label, multicentre, randomised, controlled, two-by-two factorial trial. Lancet 2015, 386, 2069–2077. [Google Scholar] [CrossRef]
- Heckmann, N.; Sivasundaram, L.; Heidari, K.S.; Weber, A.E.; Mayer, E.N.; Omid, R.; Vangsness, C.T., Jr.; Hatch, G.F., III. Propionibacterium Acnes Persists Despite Various Skin Preparation Techniques. Arthroscopy 2018, 34, 1786–1789. [Google Scholar] [CrossRef]
- Van Diek, F.M.; Pruijn, N.; Spijkers, K.M.; Mulder, B.; Kosse, N.M.; Dorrestijn, O. The presence of Cutibacterium acnes on the skin of the shoulder after the use of benzoyl peroxide: A placebo-controlled, double-blinded, randomized trial. J. Shoulder Elbow Surg. 2020, 29, 768–774. [Google Scholar] [CrossRef]
- Hancock, D.S.; Rupasinghe, S.L.; Elkinson, I.; Bloomfield, M.G.; Larsen, P.D. Benzoyl peroxide + chlorhexidine versus chlorhexidine alone skin preparation to reduce Propionibacterium acnes: A randomized controlled trial. ANZ J. Surg. 2018, 88, 1182–1186. [Google Scholar] [CrossRef]
- Nakase, K.; Fukushima, H.; Yukawa, T.; Nakaminami, H.; Takeshi Fujii, T.; Noguchi, N. Propionibacterium acnes Has Low Susceptibility to Chlorhexidine Digluconate. Surg. Infect. 2018, 19, 298–302. [Google Scholar] [CrossRef]
- Steinmann, J.; Paulmann, D.; Becker, B.; Bischoff, B.; Steinmann, E.; Steinmann, J. Comparison of virucidal activity of alcohol-based hand sanitizers versus antimicrobial hand soaps in vitro and in vivo. J. Hosp. Infect. 2012, 82, 277–280. [Google Scholar] [CrossRef] [PubMed]
- Tuladhar, E.; Hazeleger, W.C.; Koopmans, M.; Zwietering, M.H.; Duizer, E.; Beumer, R.R. Reducing viral contamination from finger pads: Handwashing is more effective than alcohol-based hand disinfectants. J. Hosp. Infect. 2015, 90, 226–234. [Google Scholar] [CrossRef]
- Kampf, G.; Todt, D.; Pfaender, S.; Steinmann, E. Persistence of coronaviruses on inanimate surfaces and their inactivation with biocidal agents. J. Hosp. Infect. 2020, 104, 246–251. [Google Scholar] [CrossRef] [PubMed]
- De Pessemier, B.; Grine, L.; Debaere, M.; Maes, A.; Paetzold, B.; Callewaert, C. Gut–Skin Axis: Current Knowledge of the Interrelationship between Microbial Dysbiosis and Skin Conditions. Microorganisms 2021, 9, 353. [Google Scholar] [CrossRef] [PubMed]
- Yazdanbakhsh, M.; Kremsner, P.G.; Van Ree, R. Allergy, parasites, and hygiene hypothesis. Science 2002, 296, 490–494. [Google Scholar] [CrossRef] [PubMed]
- Humbert, P.; Pelletier, F.; Dreno, B.; Puzenat, E.; Aubin, F. Gluten intolerance and skin diseases. Eur. J. Dermatol. 2006, 16, 4–11. [Google Scholar] [PubMed]
- Grossi, E.; Cazzaniga, S.; Crotti, S.; Naldi, L.; Di Landro, A.; Ingordo, V.; Cusano, F.; Atzori, L.; Tripodi Cutrì, F.; Musumeci, M.; et al. The constellation of dietary factors in adolescent acne: A semantic connectivity map approach. J. Eur. Acad. Dermatol. Venereol. 2016, 30, 96–100. [Google Scholar] [CrossRef]
- Bosman, E.S.; Albert, A.Y.; Lui, H.; DUTZ, J.P.; Vallance, B.A. Skin exposure to Narrow Band Ultraviolet (UVB) light modulates the human intestinal microbiome. Front. Microbiol. 2019, 10, 2410. [Google Scholar] [CrossRef] [PubMed]
- Brough, H.A.; Liu, A.H.; Sicherer, S.; Makinson, K.; Douiri, A.; Brown, S.J.; Stephens, A.C.; McLean, W.I.; Turcanu, V.; Wood, R.A.; et al. Atopic dermatitis increases the effect of exposure to peanut antigen in dust on peanut sensitization and likely peanut allergy. J. Allergy Clin. Immunol. 2015, 135, 164–170. [Google Scholar] [CrossRef]
- Bartnikas, L.M.; Gurish, M.F.; Burton, O.T.; Leisten, S.; Janssen, E.; Oettgen, H.C.; Beaupré, J.; Lewis, C.N.; Austen, K.F.; Schulte, S.; et al. Epicutaneous sensitization results in IgE-dependent intestinal mast cell expansion and food-induced anaphylaxis. J. Allergy Clin. Immunol. 2013, 131, 451–460. [Google Scholar] [CrossRef] [PubMed]
| Skin Sites | |||||
|---|---|---|---|---|---|
| Moist | Sebaceous | Dry | Foot | ||
| Body sites location | groin, axilla, antecubital fossa, toe web | face, chest, back, | arm, leg, volar forearm | Moist—toe interdigital web space dry—plantar heel | |
| Fungi | Malassezia spp. | 36% CH–80%AD | 65% CH–99%AD | 35% CH–83%AD | 53–80% |
| Other fungi | Ascomycota: Aspergillus, Epicoccum, Phoma (levels >5%): 9.5%AD–40.2%CH Cladosporium, Cryptococcus | Cryptococcus, Aspergillus, Rhodotorula, Epicoccum, Saccharomyces, Candida, Epidermophyton Microsporum, Trichophyton | |||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Skowron, K.; Bauza-Kaszewska, J.; Kraszewska, Z.; Wiktorczyk-Kapischke, N.; Grudlewska-Buda, K.; Kwiecińska-Piróg, J.; Wałecka-Zacharska, E.; Radtke, L.; Gospodarek-Komkowska, E. Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota. Microorganisms 2021, 9, 543. https://doi.org/10.3390/microorganisms9030543
Skowron K, Bauza-Kaszewska J, Kraszewska Z, Wiktorczyk-Kapischke N, Grudlewska-Buda K, Kwiecińska-Piróg J, Wałecka-Zacharska E, Radtke L, Gospodarek-Komkowska E. Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota. Microorganisms. 2021; 9(3):543. https://doi.org/10.3390/microorganisms9030543
Chicago/Turabian StyleSkowron, Krzysztof, Justyna Bauza-Kaszewska, Zuzanna Kraszewska, Natalia Wiktorczyk-Kapischke, Katarzyna Grudlewska-Buda, Joanna Kwiecińska-Piróg, Ewa Wałecka-Zacharska, Laura Radtke, and Eugenia Gospodarek-Komkowska. 2021. "Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota" Microorganisms 9, no. 3: 543. https://doi.org/10.3390/microorganisms9030543
APA StyleSkowron, K., Bauza-Kaszewska, J., Kraszewska, Z., Wiktorczyk-Kapischke, N., Grudlewska-Buda, K., Kwiecińska-Piróg, J., Wałecka-Zacharska, E., Radtke, L., & Gospodarek-Komkowska, E. (2021). Human Skin Microbiome: Impact of Intrinsic and Extrinsic Factors on Skin Microbiota. Microorganisms, 9(3), 543. https://doi.org/10.3390/microorganisms9030543






