Microfluidics as a Novel Technique for Tuberculosis: From Diagnostics to Drug Discovery
Abstract
1. Introduction
1.1. Tuberculosis and Its Global Health Threat
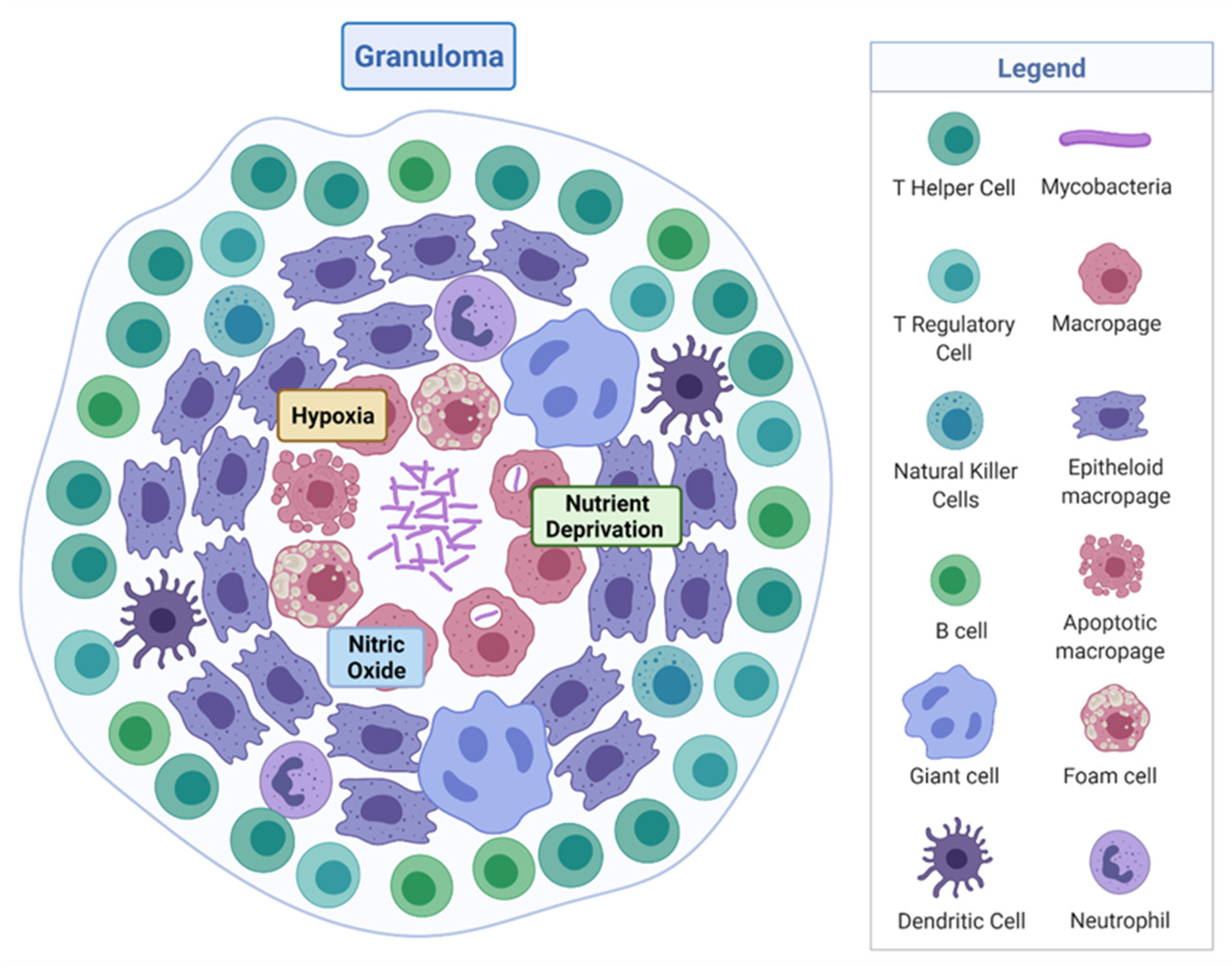
1.2. Mycobacterium Tuberculosis and the Pathogenesis of TB
1.3. Current Diagnostics, Research Methods, and Treatment
2. Microfluidics
2.1. Technological Advancement of Microfluidics
2.2. The Physics of Microfluidics
2.3. Droplet Microfluidics
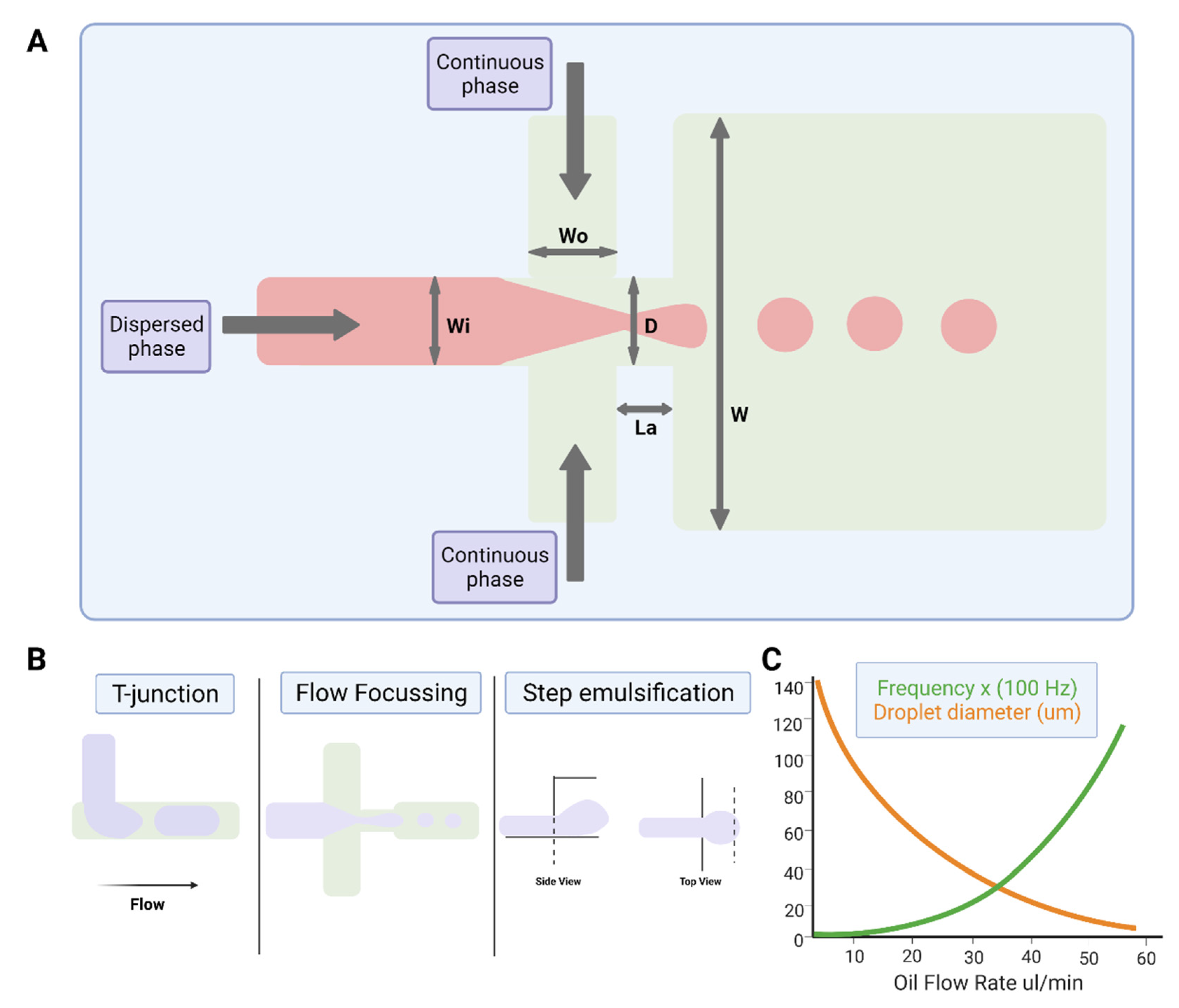
2.3.1. Physics of Droplet Microfluidics
2.3.2. Device Geometries for Droplet Production
2.3.3. Active Drop Formation
2.3.4. Droplet Sensing and Manipulation
2.4. Microfluidic Chip Materials and Microfabrication
3. Recent Application of Microfluidics for TB
3.1. Applications of Microfluidics for Diagnostics and Detecting Drug-Resistant Strains
3.2. Applications of Microfluidics for TB Drug Discovery
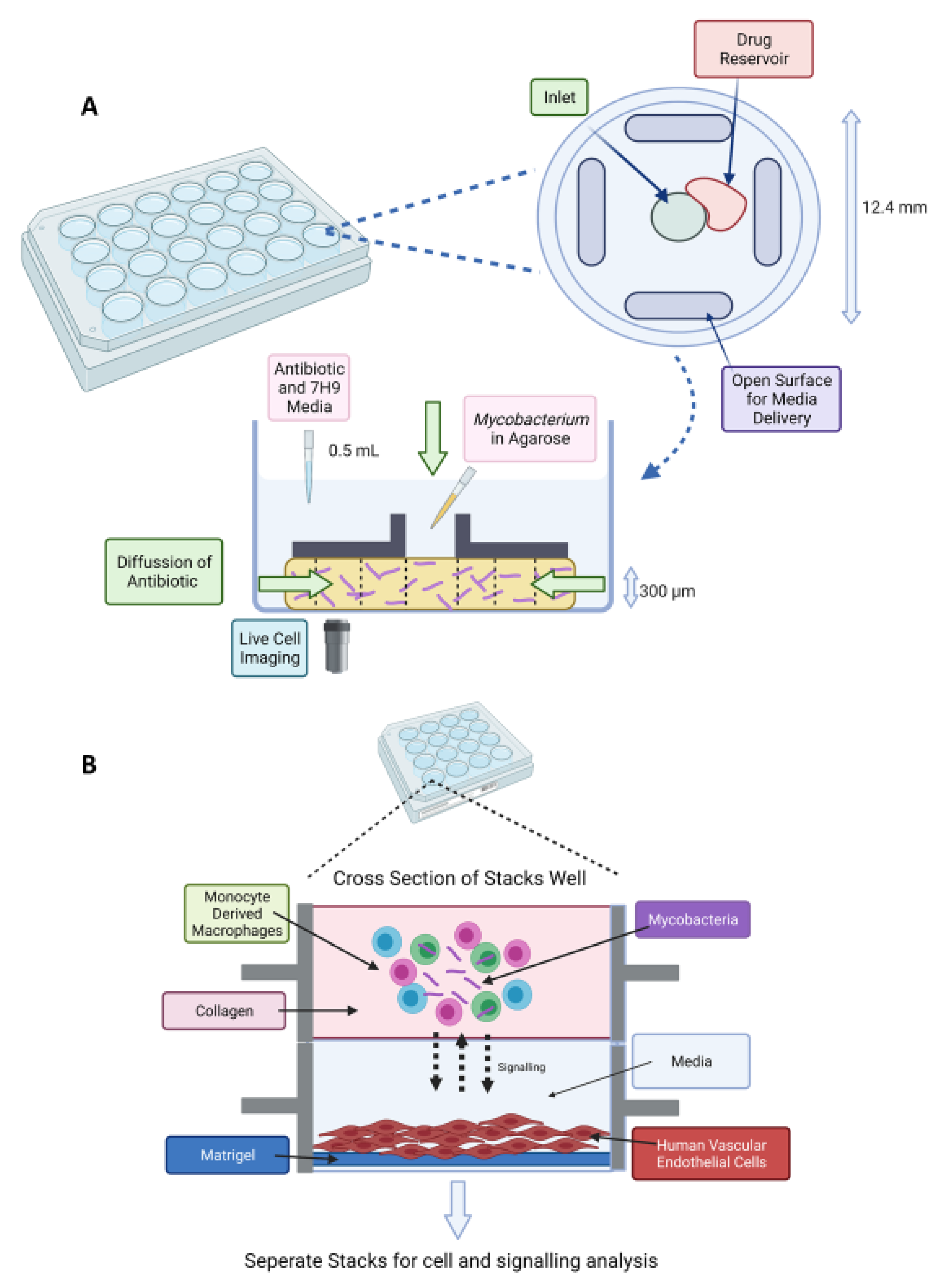
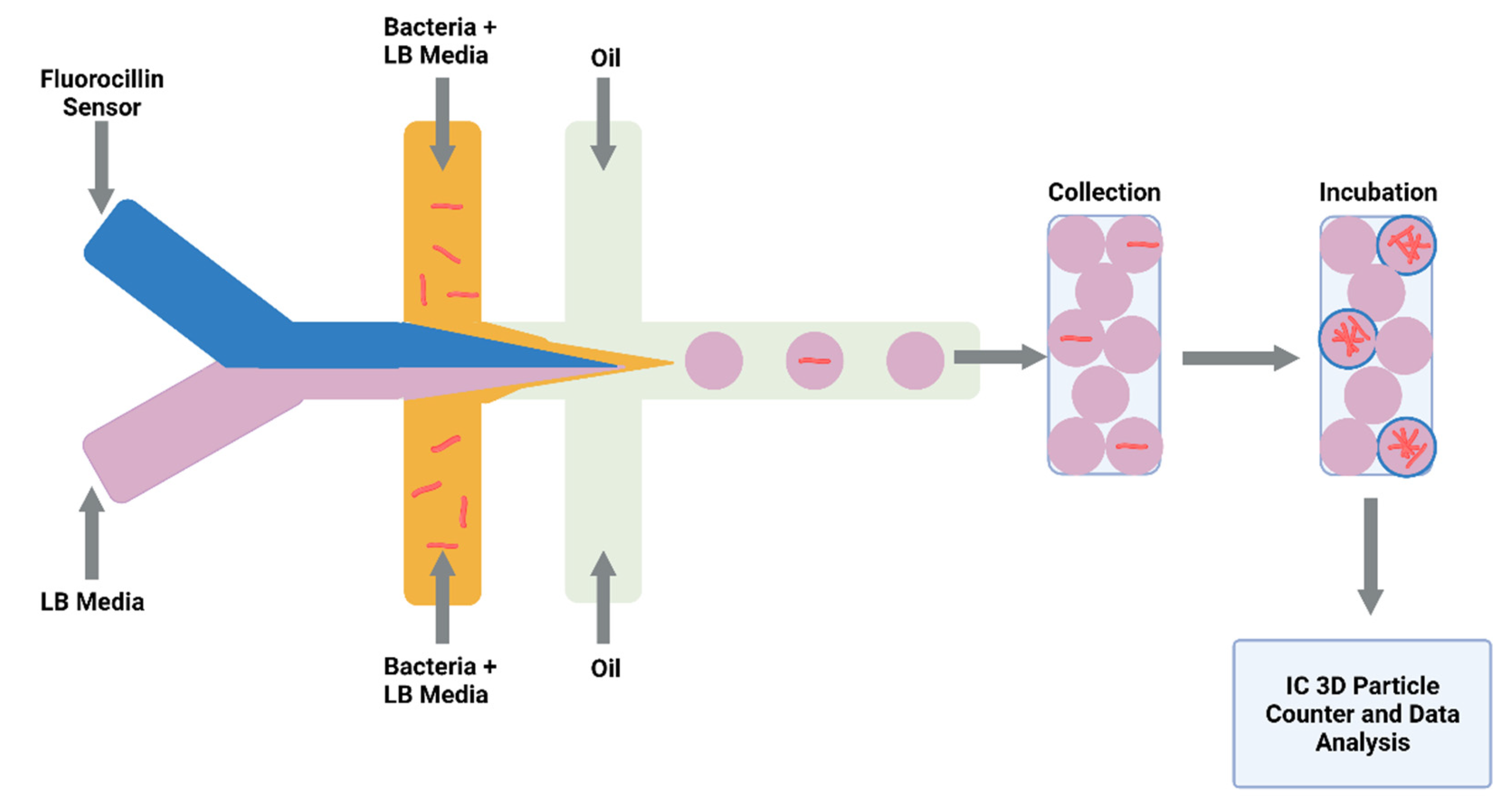
3.3. Future Application of Drug Screening in Microdroplets
4. Discussion and Future Perspectives
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- World Health Organization. Global Tuberculosis Report 2021; World Health Organization: Albany, NY, USA, 2021. [Google Scholar]
- Lee, S.H. Tuberculosis Infection and Latent Tuberculosis. Tuberc. Respir. Dis. 2016, 79, 201–206. [Google Scholar] [CrossRef]
- Dorman, S.E.; Nahid, P.; Kurbatova, E.V.; Phillips, P.P.J.; Bryant, K.; Dooley, K.E.; Engle, M.; Goldberg, S.V.; Phan, H.T.T.; Hakim, J.; et al. Four-Month Rifapentine Regimens with or without Moxifloxacin for Tuberculosis. N. Engl. J. Med. 2021, 384, 1705–1718. [Google Scholar] [CrossRef]
- END-TB. Expand New Drug Markets for TB; END-TB 2021; 2016; pp. 1–30. Available online: https://unitaid.org/assets/Swiss-TPH-endTB-MTE-final-report-rvd-final-13-Jun-18.pdf (accessed on 26 October 2021).
- World Health Organization. The END-TB Strategy; World Health Organization: Albany, NY, USA, 2012. [Google Scholar]
- Burki, T.K. The global cost of tuberculosis. Lancet Respir. Med. 2018, 6, 13. [Google Scholar] [CrossRef]
- Parish, T.; Stoker, N.G. Mycobacteria: Bugs and bugbears (Two steps forward and one step back). Mol. Biotechnol. 1999, 13, 191–200. [Google Scholar] [CrossRef]
- Batt, S.M.; Minnikin, D.E.; Besra, G.S. The thick waxy coat of mycobacteria, a protective layer against antibiotics and the host’s immune system. Biochem. J. 2020, 477, 1983–2006. [Google Scholar] [CrossRef]
- Minnikin, D.E.; Kremer, L.; Dover, L.G.; Besra, G.S. The methyl-branched fortifications of Mycobacterium tuberculosis. Chem. Biol. 2002, 9, 545–553. [Google Scholar] [CrossRef][Green Version]
- Jee, B. Understanding the early host immune response against Mycobacterium tuberculosis. Cent. Eur. J. Immunol. 2020, 45, 99–103. [Google Scholar] [CrossRef]
- Getahun, H.; Matteelli, A.; Chaisson, R.E.; Raviglione, M. Latent Mycobacterium tuberculosis infection. N. Engl. J. Med. 2015, 372, 2127–2135. [Google Scholar] [CrossRef]
- Ernst, J.D. The immunological life cycle of tuberculosis. Nat. Rev. Immunol. 2012, 12, 581–591. [Google Scholar] [CrossRef]
- Pieters, J. Entry and survival of pathogenic mycobacteria in macrophages. Microbes Infect. 2001, 3, 249–255. [Google Scholar] [CrossRef]
- Ramakrishnan, L. Revisiting the role of the granuloma in tuberculosis. Nat. Rev. Immunol. 2012, 12, 352–366. [Google Scholar] [CrossRef]
- Russell, D.G.; Cardona, P.-J.; Kim, M.-J.; Allain, S.; Altare, F. Foamy macrophages and the progression of the human tuberculosis granuloma. Nat. Immunol. 2009, 10, 943–948. [Google Scholar] [CrossRef]
- Martin, C.J.; Carey, A.F.; Fortune, S.M. A bug’s life in the granuloma. Semin. Immunopathol. 2016, 38, 213–220. [Google Scholar] [CrossRef]
- Houben, R.M.G.J.; Dodd, P.J. The Global Burden of Latent Tuberculosis Infection: A Re-estimation Using Mathematical Modelling. PLoS Med. 2016, 13, e1002152. [Google Scholar] [CrossRef]
- Singhal, R.; Myneedu, V.P. Microscopy as a diagnostic tool in pulmonary tuberculosis. Int. J. Mycobacteriol. 2015, 4, 1–6. [Google Scholar] [CrossRef]
- Walzl, G.; McNerney, R.; du Plessis, N.; Bates, M.; McHugh, T.D.; Chegou, N.N.; Zumla, A. Tuberculosis: Advances and challenges in development of new diagnostics and biomarkers. Lancet Infect. Dis 2018, 18, e199–e210. [Google Scholar] [CrossRef]
- Pai, M.; Nicol, M.P.; Boehme, C.C. Tuberculosis Diagnostics: State of the Art and Future Directions. Microbiol. Spectr. 2016, 4, 4–5. [Google Scholar] [CrossRef]
- Lee, A.; Xie, Y.L.; Barry, C.E.; Chen, R.Y. Current and future treatments for tuberculosis. BMJ 2020, 368, m216. [Google Scholar] [CrossRef]
- Boehme, C.C.; Nabeta, P.; Hillemann, D.; Nicol, M.P.; Shenai, S.; Krapp, F.; Allen, J.; Tahirli, R.; Blakemore, R.; Rustomjee, R.; et al. Rapid molecular detection of tuberculosis and rifampin resistance. N. Engl. J. Med. 2010, 363, 1005–1015. [Google Scholar] [CrossRef]
- Meehan, C.J.; Goig, G.A.; Kohl, T.A.; Verboven, L.; Dippenaar, A.; Ezewudo, M.; Farhat, M.R.; Guthrie, J.L.; Laukens, K.; Miotto, P.; et al. Whole genome sequencing of Mycobacterium tuberculosis: Current standards and open issues. Nat. Rev. Microbiol. 2019, 17, 533–545. [Google Scholar] [CrossRef]
- Toniolo, C.; Rutschmann, O.; McKinney, J.D. Do chance encounters between heterogeneous cells shape the outcome of tuberculosis infections? Curr. Opin. Microbiol. 2021, 59, 72–78. [Google Scholar] [CrossRef]
- Singh, A.K.; Gupta, U.D. Animal models of tuberculosis: Lesson learnt. Indian J. Med. Res. 2018, 147, 456–463. [Google Scholar] [CrossRef]
- Rhoades, E.R.; Frank, A.A.; Orme, I.M. Progression of chronic pulmonary tuberculosis in mice aerogenically infected with virulent Mycobacterium tuberculosis. Tuber. Lung Dis. 1997, 78, 57–66. [Google Scholar] [CrossRef]
- Gibson, S.E.R.; Harrison, J.; Cox, J.A.G. Modelling a Silent Epidemic: A Review of the In Vitro Models of Latent Tuberculosis. Pathogens 2018, 7, 88. [Google Scholar] [CrossRef]
- Parish, T. In vitro drug discovery models for Mycobacterium tuberculosis relevant for host infection. Expert Opin. Drug Discov. 2020, 15, 349–358. [Google Scholar] [CrossRef]
- Batyrshina, Y.R.; Schwartz, Y.S. Modeling of Mycobacterium tuberculosis dormancy in bacterial cultures. Tuberculosis 2019, 117, 7–17. [Google Scholar] [CrossRef]
- Schön, T.; Werngren, J.; Machado, D.; Borroni, E.; Wijkander, M.; Lina, G.; Mouton, J.; Matuschek, E.; Kahlmeter, G.; Giske, C.; et al. Antimicrobial susceptibility testing of Mycobacterium tuberculosis complex isolates—The EUCAST broth microdilution reference method for MIC determination. Clin. Microbiol. Infect. Off. Publ. Eur. Soc. Clin. Microbiol. Infect. Dis. 2020, 26, 1488–1492. [Google Scholar] [CrossRef]
- Kim, S.J. Drug-susceptibility testing in tuberculosis: Methods and reliability of results. Eur. Respir. J. 2005, 25, 564–569. [Google Scholar] [CrossRef]
- Centers for Disease Control and Prevention. Emergence of Mycobacterium tuberculosis with extensive resistance to second-line drugs—Worldwide, 2000–2004. MMWR. Morb. Mortal. Wkly. Rep. 2006, 55, 301–305. [Google Scholar]
- Whitesides, G.M. The origins and the future of microfluidics. Nature 2006, 442, 368–373. [Google Scholar] [CrossRef]
- Squires, T.M.; Quake, S.R. Microfluidics: Fluid physics at the nanoliter scale. Rev. Mod. Phys. 2005, 77, 977–1026. [Google Scholar] [CrossRef]
- Tabeling, P.; Chen, S. Introduction to Microfluidics; OUP Oxford: Oxford, UK, 2005. [Google Scholar]
- Novotný, J.; Foret, F. Fluid manipulation on the micro-scale: Basics of fluid behavior in microfluidics. J. Sep. Sci. 2017, 40, 383–394. [Google Scholar] [CrossRef]
- Beebe, D.J.; Mensing, G.A.; Walker, G.M. Physics and Applications of Microfluidics in Biology. Annu. Rev. Biomed. Eng. 2002, 4, 261–286. [Google Scholar] [CrossRef]
- Shim, J.U.; Ranasinghe, R.T.; Smith, C.A.; Ibrahim, S.M.; Hollfelder, F.; Huck, W.T.; Klenerman, D.; Abell, C. Ultrarapid generation of femtoliter microfluidic droplets for single-molecule-counting immunoassays. ACS Nano 2013, 7, 5955–5964. [Google Scholar] [CrossRef]
- Vyawahare, S.; Griffiths, A.D.; Merten, C.A. Miniaturization and parallelization of biological and chemical assays in microfluidic devices. Chem. Biol. 2010, 17, 1052–1065. [Google Scholar] [CrossRef]
- Theberge, A.B.; Courtois, F.; Schaerli, Y.; Fischlechner, M.; Abell, C.; Hollfelder, F.; Huck, W.T.S. Microdroplets in Microfluidics: An Evolving Platform for Discoveries in Chemistry and Biology. Angew. Chem. Int. Ed. 2010, 49, 5846–5868. [Google Scholar] [CrossRef]
- Abou-Hassan, A.; Sandre, O.; Cabuil, V. Microfluidics in Inorganic Chemistry. Angew. Chem. Int. Ed. 2010, 49, 6268–6286. [Google Scholar] [CrossRef]
- Zheng, B.; Gerdts, C.J.; Ismagilov, R.F. Using nanoliter plugs in microfluidics to facilitate and understand protein crystallization. Curr. Opin. Struct. Biol. 2005, 15, 548–555. [Google Scholar] [CrossRef]
- Shim, J.-u.; Cristobal, G.; Link, D.R.; Thorsen, T.; Fraden, S. Using Microfluidics to Decouple Nucleation and Growth of Protein Crystals. Cryst. Growth Des. 2007, 7, 2192–2194. [Google Scholar] [CrossRef]
- Guo, M.T.; Rotem, A.; Heyman, J.A.; Weitz, D.A. Droplet microfluidics for high-throughput biological assays. Lab Chip 2012, 12, 2146–2155. [Google Scholar] [CrossRef]
- Shembekar, N.; Chaipan, C.; Utharala, R.; Merten, C.A. Droplet-based microfluidics in drug discovery, transcriptomics and high-throughput molecular genetics. Lab Chip 2016, 16, 1314–1331. [Google Scholar] [CrossRef] [PubMed]
- Collins, D.J.; Neild, A.; deMello, A.; Liu, A.-Q.; Ai, Y. The Poisson distribution and beyond: Methods for microfluidic droplet production and single cell encapsulation. Lab Chip 2015, 15, 3439–3459. [Google Scholar] [CrossRef] [PubMed]
- Cabane, B.; Hénon, S. Liquides: Solutions, Dispersions, Émulsions, Gels; Belin Éducation: Paris, France, 2007. [Google Scholar]
- Dai, B.; Leal, L.G. The mechanism of surfactant effects on drop coalescence. Phys. Fluids 2008, 20, 040802. [Google Scholar] [CrossRef]
- Zhao, C.-X.; Middelberg, A.P.J. Two-phase microfluidic flows. Chem. Eng. Sci. 2011, 66, 1394–1411. [Google Scholar] [CrossRef]
- Yow, H.N.; Routh, A.F. Formation of liquid core–polymer shell microcapsules. Soft Matter 2006, 2, 940–949. [Google Scholar] [CrossRef]
- Müller, R.H.; Mäder, K.; Gohla, S. Solid lipid nanoparticles (SLN) for controlled drug delivery—A review of the state of the art. Eur. J. Pharm. Biopharm. 2000, 50, 161–177. [Google Scholar] [CrossRef]
- Kawaguchi, H. Functional polymer microspheres. Prog. Polym. Sci. 2000, 25, 1171–1210. [Google Scholar] [CrossRef]
- Stolnik, S.; Illum, L.; Davis, S.S. Long circulating microparticulate drug carriers. Adv. Drug Deliv. Rev. 1995, 16, 195–214. [Google Scholar] [CrossRef]
- Astete, C.E.; Sabliov, C.M. Synthesis and characterization of PLGA nanoparticles. J. Biomater. Sci. Polym. Ed. 2006, 17, 247–289. [Google Scholar] [CrossRef]
- Koh, W.-G.; Pishko, M.V. Fabrication of cell-containing hydrogel microstructures inside microfluidic devices that can be used as cell-based biosensors. Anal. Bioanal. Chem. 2006, 385, 1389–1397. [Google Scholar] [CrossRef]
- Steinbacher, J.L.; McQuade, D.T. Polymer chemistry in flow: New polymers, beads, capsules, and fibers. J. Polym. Sci. Part A Polym. Chem. 2006, 44, 6505–6533. [Google Scholar] [CrossRef]
- Seo, M.; Nie, Z.; Xu, S.; Mok, M.; Lewis, P.C.; Graham, R.; Kumacheva, E. Continuous Microfluidic Reactors for Polymer Particles. Langmuir 2005, 21, 11614–11622. [Google Scholar] [CrossRef] [PubMed]
- Hung, L.-H.; Lee, A. Microfluidic devices for the synthesis of nanoparticles and biomaterials. J. Med. Biol. Eng. 2006, 27, 1–6. [Google Scholar]
- Sesen, M.; Alan, T.; Neild, A. Droplet control technologies for microfluidic high throughput screening (μHTS). Lab Chip 2017, 17, 2372–2394. [Google Scholar] [CrossRef] [PubMed]
- Tran, T.M.; Lan, F.; Thompson, C.S.; Abate, A.R. From tubes to drops: Droplet-based microfluidics for ultrahigh-throughput biology. J. Phys. D Appl. Phys. 2013, 46, 114004. [Google Scholar] [CrossRef]
- Cramer, C.; Fischer, P.; Windhab, E.J. Drop formation in a co-flowing ambient fluid. Chem. Eng. Sci. 2004, 59, 3045–3058. [Google Scholar] [CrossRef]
- Umbanhowar, P.B.; Prasad, V.; Weitz, D.A. Monodisperse emulsion generation via drop break off in a coflowing stream. Langmuir 2000, 16, 347–351. [Google Scholar] [CrossRef]
- McDonald, J.C.; Duffy, D.C.; Anderson, J.R.; Chiu, D.T.; Wu, H.; Schueller, O.J.; Whitesides, G.M. Fabrication of microfluidic systems in poly(dimethylsiloxane). Electrophoresis 2000, 21, 27–40. [Google Scholar] [CrossRef]
- Utada, A.S.; Chu, L.Y.; Fernandez-Nieves, A.; Link, D.R.; Holtze, C.; Weitz, D.A. Dripping, Jetting, Drops, and Wetting: The Magic of Microfluidics. MRS Bull. 2007, 32, 702–708. [Google Scholar] [CrossRef]
- Sugiura, S.; Nakajima, M.; Tong, J.; Nabetani, H.; Seki, M. Preparation of Monodispersed Solid Lipid Microspheres Using a Microchannel Emulsification Technique. J. Colloid Interface Sci. 2000, 227, 95–103. [Google Scholar] [CrossRef]
- Stolovicki, E.; Ziblat, R.; Weitz, D.A. Throughput enhancement of parallel step emulsifier devices by shear-free and efficient nozzle clearance. Lab Chip 2018, 18, 132–138. [Google Scholar] [CrossRef] [PubMed]
- Link, D.R.; Grasland-Mongrain, E.; Duri, A.; Sarrazin, F.; Cheng, Z.D.; Cristobal, G.; Marquez, M.; Weitz, D.A. Electric control of droplets in microfluidic devices. Angew. Chem. Int. Ed. 2006, 45, 2556–2560. [Google Scholar] [CrossRef] [PubMed]
- Zhu, P.; Wang, L. Passive and active droplet generation with microfluidics: A review. Lab Chip 2017, 17, 34–75. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.-H.; Nguyen, N.-T.; Yobas, L.; Kang, T.G. Formation and manipulation of ferrofluid droplets at a microfluidicT-junction. J. Micromech. Microeng. 2010, 20, 045004. [Google Scholar] [CrossRef]
- Haeberle, S.; Zengerle, R.; Ducrée, J. Centrifugal generation and manipulation of droplet emulsions. Microfluid. Nanofluidics 2007, 3, 65–75. [Google Scholar] [CrossRef]
- Park, S.-Y.; Wu, T.-H.; Chen, Y.; Teitell, M.A.; Chiou, P.-Y. High-speed droplet generation on demand driven by pulse laser-induced cavitation. Lab Chip 2011, 11, 1010–1012. [Google Scholar] [CrossRef]
- Murshed, S.M.S.; Tan, S.H.; Nguyen, N.T.; Wong, T.N.; Yobas, L. Microdroplet formation of water and nanofluids in heat-induced microfluidic T-junction. Microfluid. Nanofluid. 2009, 6, 253–259. [Google Scholar] [CrossRef]
- Xu, J.; Attinger, D. Drop on demand in a microfluidic chip. J. Micromech. Microeng. 2008, 18, 065020. [Google Scholar] [CrossRef]
- Collins, D.J.; Alan, T.; Helmerson, K.; Neild, A. Surface acoustic waves for on-demand production of picoliter droplets and particle encapsulation. Lab Chip 2013, 13, 3225–3231. [Google Scholar] [CrossRef]
- Schmid, L.; Franke, T. SAW-controlled drop size for flow focusing. Lab Chip 2013, 13, 1691–1694. [Google Scholar] [CrossRef]
- Abate, A.R.; Romanowsky, M.B.; Agresti, J.J.; Weitz, D.A. Valve-based flow focusing for drop formation. Appl. Phys. Lett. 2009, 94, 023503. [Google Scholar] [CrossRef]
- Zeng, S.; Li, B.; Su, X.; Qin, J.; Lin, B. Microvalve-actuated precise control of individual droplets in microfluidic devices. Lab Chip 2009, 9, 1340–1343. [Google Scholar] [CrossRef]
- Jin, S.H.; Jeong, H.-H.; Lee, B.; Lee, S.S.; Lee, C.-S. A programmable microfluidic static droplet array for droplet generation, transportation, fusion, storage, and retrieval. Lab Chip 2015, 15, 3677–3686. [Google Scholar] [CrossRef] [PubMed]
- Josephides, D.; Davoli, S.; Whitley, W.; Ruis, R.; Salter, R.; Gokkaya, S.; Vallet, M.; Matthews, D.; Benazzi, G.; Shvets, E.; et al. Cyto-Mine: An Integrated, Picodroplet System for High-Throughput Single-Cell Analysis, Sorting, Dispensing, and Monoclonality Assurance. SLAS TECHNOLOGY Transl. Life Sci. Innov. 2020, 25, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Nguyen, N.-T.; Lassemono, S.; Chollet, F.A. Optical detection for droplet size control in microfluidic droplet-based analysis systems. Sens. Actuators B Chem. 2006, 117, 431–436. [Google Scholar] [CrossRef]
- Robert de Saint Vincent, M.; Cassagnère, S.; Plantard, J.; Delville, J.-P. Real-time droplet caliper for digital microfluidics. Microfluid. Nanofluid. 2012, 13, 261–271. [Google Scholar] [CrossRef]
- Baret, J.C.; Miller, O.J.; Taly, V.; Ryckelynck, M.; El-Harrak, A.; Frenz, L.; Rick, C.; Samuels, M.L.; Hutchison, J.B.; Agresti, J.J.; et al. Fluorescence-activated droplet sorting (FADS): Efficient microfluidic cell sorting based on enzymatic activity. Lab Chip 2009, 9, 1850–1858. [Google Scholar] [CrossRef]
- Cole, M.C.; Kenis, P.J.A. Multiplexed electrical sensor arrays in microfluidic networks. Sens. Actuators B Chem. 2009, 136, 350–358. [Google Scholar] [CrossRef]
- Moiseeva, E.V.; Fletcher, A.A.; Harnett, C.K. Thin-film electrode based droplet detection for microfluidic systems. Sens. Actuators B Chem. 2011, 155, 408–414. [Google Scholar] [CrossRef]
- Niu, X.; Zhang, M.; Peng, S.; Wen, W.; Sheng, P. Real-time detection, control, and sorting of microfluidic droplets. Biomicrofluidics 2007, 1, 44101. [Google Scholar] [CrossRef]
- Liu, W.-w.; Zhu, Y. “Development and application of analytical detection techniques for droplet-based microfluidics”—A review. Anal. Chim. Acta 2020, 1113, 66–84. [Google Scholar] [CrossRef]
- Zhu, Y.; Fang, Q. Analytical detection techniques for droplet microfluidics—A review. Anal. Chim. Acta 2013, 787, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Liu, B.F.; Li, J.; Liu, X. Advances in coupling microfluidic chips to mass spectrometry. Mass Spectrom. Rev. 2015, 34, 535–557. [Google Scholar] [CrossRef] [PubMed]
- Oedit, A.; Vulto, P.; Ramautar, R.; Lindenburg, P.W.; Hankemeier, T. Lab-on-a-Chip hyphenation with mass spectrometry: Strategies for bioanalytical applications. Curr. Opin. Biotechnol. 2015, 31, 79–85. [Google Scholar] [CrossRef] [PubMed]
- Jahn, I.J.; Žukovskaja, O.; Zheng, X.S.; Weber, K.; Bocklitz, T.W.; Cialla-May, D.; Popp, J. Surface-enhanced Raman spectroscopy and microfluidic platforms: Challenges, solutions and potential applications. Analyst 2017, 142, 1022–1047. [Google Scholar] [CrossRef] [PubMed]
- Basova, E.Y.; Foret, F. Droplet microfluidics in (bio)chemical analysis. Analyst 2015, 140, 22–38. [Google Scholar] [CrossRef] [PubMed]
- Tenje, M.; Fornell, A.; Ohlin, M.; Nilsson, J. Particle Manipulation Methods in Droplet Microfluidics. Anal. Chem. 2018, 90, 1434–1443. [Google Scholar] [CrossRef] [PubMed]
- Miltenyi, S.; Müller, W.; Weichel, W.; Radbruch, A. High gradient magnetic cell separation with MACS. Cytometry 1990, 11, 231–238. [Google Scholar] [CrossRef]
- Mirowski, E.; Moreland, J.; Zhang, A.; Russek, S.E.; Donahue, M.J. Manipulation and sorting of magnetic particles by a magnetic force microscope on a microfluidic magnetic trap platform. Appl. Phys. Lett. 2005, 86, 243901. [Google Scholar] [CrossRef]
- Yalcin, S.E.; Sharma, A.; Qian, S.; Joo, S.W.; Baysal, O. Manipulating particles in microfluidics by floating electrodes. Electrophoresis 2010, 31, 3711–3718. [Google Scholar] [CrossRef]
- Zhang, C.; Khoshmanesh, K.; Mitchell, A.; Kalantar-Zadeh, K. Dielectrophoresis for manipulation of micro/nano particles in microfluidic systems. Anal. Bioanal. Chem. 2010, 396, 401–420. [Google Scholar] [CrossRef] [PubMed]
- Yunus, N.A.M.; Nili, H.; Green, N.G. Continuous separation of colloidal particles using dielectrophoresis. Electrophoresis 2013, 34, 969–978. [Google Scholar] [CrossRef] [PubMed]
- Kim, U.; Qian, J.; Kenrick, S.A.; Daugherty, P.S.; Soh, H.T. Multitarget dielectrophoresis activated cell sorter. Anal. Chem. 2008, 80, 8656–8661. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.M.; Tu, E.; Raymond, D.E.; Yang, J.M.; Zhang, H.; Hagen, N.; Dees, B.; Mercer, E.M.; Forster, A.H.; Kariv, I.; et al. Microfluidic sorting of mammalian cells by optical force switching. Nat. Biotechnol. 2005, 23, 83–87. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.B.; Yoon, S.Y.; Sung, H.J.; Kim, S.S. Cross-Type Optical Particle Separation in a Microchannel. Anal. Chem. 2008, 80, 2628–2630. [Google Scholar] [CrossRef]
- Lee, C.-Y.; Lin, Y.-H.; Lee, G.-B. A droplet-based microfluidic system capable of droplet formation and manipulation. Microfluid. Nanofluidics 2009, 6, 599–610. [Google Scholar] [CrossRef]
- O’Rorke, R.D.; Wood, C.D.; Wälti, C.; Evans, S.D.; Davies, A.G.; Cunningham, J.E. Acousto-microfluidics: Transporting microbubble and microparticle arrays in acoustic traps using surface acoustic waves. J. Appl. Phys. 2012, 111, 094911. [Google Scholar] [CrossRef]
- Geislinger, T.M.; Franke, T. Sorting of circulating tumor cells (MV3-melanoma) and red blood cells using non-inertial lift. Biomicrofluidics 2013, 7, 044120. [Google Scholar] [CrossRef]
- Augustsson, P.; Magnusson, C.; Nordin, M.; Lilja, H.; Laurell, T. Microfluidic, label-free enrichment of prostate cancer cells in blood based on acoustophoresis. Anal. Chem. 2012, 84, 7954–7962. [Google Scholar] [CrossRef]
- Nielsen, J.B.; Hanson, R.L.; Almughamsi, H.M.; Pang, C.; Fish, T.R.; Woolley, A.T. Microfluidics: Innovations in Materials and Their Fabrication and Functionalization. Anal. Chem. 2020, 92, 150–168. [Google Scholar] [CrossRef]
- Becker, H.; Gärtner, C. Polymer microfabrication technologies for microfluidic systems. Anal. Bioanal. Chem. 2008, 390, 89–111. [Google Scholar] [CrossRef] [PubMed]
- Faustino, V.; Catarino, S.O.; Lima, R.; Minas, G. Biomedical microfluidic devices by using low-cost fabrication techniques: A review. J. Biomech. 2016, 49, 2280–2292. [Google Scholar] [CrossRef] [PubMed]
- Mata, A.; Fleischman, A.J.; Roy, S. Characterization of Polydimethylsiloxane (PDMS) Properties for Biomedical Micro/Nanosystems. Biomed. Microdevices 2005, 7, 281–293. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Duarte, R. Microfabrication technologies in dielectrophoresis applications—A review. Electrophoresis 2012, 33, 3110–3132. [Google Scholar] [CrossRef]
- Gencturk, E.; Mutlu, S.; Ulgen, K.O. Advances in microfluidic devices made from thermoplastics used in cell biology and analyses. Biomicrofluidics 2017, 11, 051502. [Google Scholar] [CrossRef]
- Gong, M.M.; Sinton, D. Turning the Page: Advancing Paper-Based Microfluidics for Broad Diagnostic Application. Chem. Rev. 2017, 117, 8447–8480. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Si, J.; Li, Z. Fabrication techniques for microfluidic paper-based analytical devices and their applications for biological testing: A review. Biosens. Bioelectron. 2016, 77, 774–789. [Google Scholar] [CrossRef]
- Yang, Y.; Noviana, E.; Nguyen, M.P.; Geiss, B.J.; Dandy, D.S.; Henry, C.S. Paper-Based Microfluidic Devices: Emerging Themes and Applications. Anal. Chem. 2017, 89, 71–91. [Google Scholar] [CrossRef]
- Li, H.; Steckl, A.J. Paper Microfluidics for Point-of-Care Blood-Based Analysis and Diagnostics. Anal. Chem. 2019, 91, 352–371. [Google Scholar] [CrossRef]
- Ziaie, B.; Baldi, A.; Lei, M.; Gu, Y.; Siegel, R.A. Hard and soft micromachining for BioMEMS: Review of techniques and examples of applications in microfluidics and drug delivery. Adv. Drug Deliv. Rev. 2004, 56, 145–172. [Google Scholar] [CrossRef]
- Nge, P.N.; Rogers, C.I.; Woolley, A.T. Advances in Microfluidic Materials, Functions, Integration, and Applications. Chem. Rev. 2013, 113, 2550–2583. [Google Scholar] [CrossRef] [PubMed]
- Italia, V.; Giakoumaki, A.N.; Bonfadini, S.; Bharadwaj, V.; Le Phu, T.; Eaton, S.M.; Ramponi, R.; Bergamini, G.; Lanzani, G.; Criante, L. Laser-Inscribed Glass Microfluidic Device for Non-Mixing Flow of Miscible Solvents. Micromachines 2019, 10, 23. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Whitesides, G.M. Soft Lithography. Angew. Chem. Int. Ed. Engl. 1998, 37, 550–575. [Google Scholar] [CrossRef]
- Weibel, D.B.; DiLuzio, W.R.; Whitesides, G.M. Microfabrication meets microbiology. Nat. Rev. Microbiol. 2007, 5, 209–218. [Google Scholar] [CrossRef] [PubMed]
- Lokensgard, E. Industrial Plastics: Theory and Applications; Cengage Learning: Boston, MA, USA, 2016. [Google Scholar]
- Wu, Z.; Chen, X.; Wu, Z.; Zhang, Q.; Gao, Q. Experimental study of fabricating a four-layers Cantor fractal microfluidic chip by CO2 laser system. Microsyst. Technol. 2019, 25, 1251–1256. [Google Scholar] [CrossRef]
- Piruska, A.; Nikcevic, I.; Lee, S.H.; Ahn, C.; Heineman, W.R.; Limbach, P.A.; Seliskar, C.J. The autofluorescence of plastic materials and chips measured under laser irradiation. Lab Chip 2005, 5, 1348–1354. [Google Scholar] [CrossRef] [PubMed]
- Wright, W.W. Plastics materials (5th edition) J. A. Brydson, Butterworths, London, 1989. pp. 864, price £57.50. ISBN 0-408-00721-4. Br. Polym. J. 1989, 21, 525. [Google Scholar] [CrossRef]
- Abdel-Wahab, A.A.; Ataya, S.; Silberschmidt, V.V. Temperature-dependent mechanical behaviour of PMMA: Experimental analysis and modelling. Polym. Test. 2017, 58, 86–95. [Google Scholar] [CrossRef]
- Becker, H.; Gärtner, C. Polymer microfabrication methods for microfluidic analytical applications. Electrophoresis 2000, 21, 12–26. [Google Scholar] [CrossRef]
- Sher, M.; Zhuang, R.; Demirci, U.; Asghar, W. Paper-based analytical devices for clinical diagnosis: Recent advances in the fabrication techniques and sensing mechanisms. Expert Rev. Mol. Diagn. 2017, 17, 351–366. [Google Scholar] [CrossRef]
- Institution of Engineering and Technology; Moorefi, M.; Ohta, A.; Shiroma, W. Paper-based capillary action. Electron. Lett. 2017, 53, 1339. [Google Scholar] [CrossRef]
- Songok, J.; Toivakka, M. Enhancing Capillary-Driven Flow for Paper-Based Microfluidic Channels. ACS Appl. Mater. Interfaces 2016, 8, 30523–30530. [Google Scholar] [CrossRef] [PubMed]
- Jing, W.; Jiang, X.; Zhao, W.; Liu, S.; Cheng, X.; Sui, G. Microfluidic Platform for Direct Capture and Analysis of Airborne Mycobacterium tuberculosis. Anal. Chem. 2014, 86, 5815–5821. [Google Scholar] [CrossRef]
- Das, D.; Dsouza, A.; Kaur, N.; Soni, S.; Toley, B.J. Paper Stacks for Uniform Rehydration of Dried Reagents in Paper Microfluidic Devices. Sci. Rep. 2019, 9, 15755. [Google Scholar] [CrossRef]
- Luo, J.; Fang, X.; Ye, D.; Li, H.; Chen, H.; Zhang, S.; Kong, J. A real-time microfluidic multiplex electrochemical loop-mediated isothermal amplification chip for differentiating bacteria. Biosens. Bioelectron. 2014, 60, 84–91. [Google Scholar] [CrossRef] [PubMed]
- Mühlig, A.; Bocklitz, T.; Labugger, I.; Dees, S.; Henk, S.; Richter, E.; Andres, S.; Merker, M.; Stöckel, S.; Weber, K.; et al. LOC-SERS: A Promising Closed System for the Identification of Mycobacteria. Anal. Chem. 2016, 88, 7998–8004. [Google Scholar] [CrossRef] [PubMed]
- Lyu, F.; Xu, M.; Cheng, Y.; Xie, J.; Rao, J.; Tang, S.K. Quantitative detection of cells expressing BlaC using droplet-based microfluidics for use in the diagnosis of tuberculosis. Biomicrofluidics 2015, 9, 044120. [Google Scholar] [CrossRef]
- Ip, K.; Chang, J.; Liu, T.; Dou, H.; Lee, G. An integrated microfluidic system for identification of live mycobacterium tuberculosis by real-time polymerase chain reaction. In Proceedings of the 2018 IEEE Micro Electro Mechanical Systems (MEMS), Belfast, UK, 21–25 January 2018; pp. 1124–1127. [Google Scholar]
- Domínguez, C.M.; Kosaka, P.M.; Sotillo, A.; Mingorance, J.; Tamayo, J.; Calleja, M. Label-Free DNA-Based Detection of Mycobacterium tuberculosis and Rifampicin Resistance through Hydration Induced Stress in Microcantilevers. Anal. Chem. 2015, 87, 1494–1498. [Google Scholar] [CrossRef]
- Zribi, B.; Roy, E.; Pallandre, A.; Chebil, S.; Koubaa, M.; Mejri, N.; Magdinier Gomez, H.; Sola, C.; Korri-Youssoufi, H.; Haghiri-Gosnet, A.M. A microfluidic electrochemical biosensor based on multiwall carbon nanotube/ferrocene for genomic DNA detection of Mycobacterium tuberculosis in clinical isolates. Biomicrofluidics 2016, 10, 014115. [Google Scholar] [CrossRef]
- Islamov, M.; Sypabekova, M.; Kanayeva, D.; Rojas-Solórzano, L. CFD Modeling of Chamber Filling in a Micro-Biosensor for Protein Detection. Biosensors 2017, 7, 45. [Google Scholar] [CrossRef]
- Mbano, I.M.; Mandizvo, T.; Rogich, J.; Kunota, T.T.R.; Mackenzie, J.S.; Pillay, M.; Balagaddé, F.K. Light Forge: A Microfluidic DNA Melting-based Tuberculosis Test. J. Appl. Lab. Med. 2020, 5, 440–453. [Google Scholar] [CrossRef] [PubMed]
- Minero, G.A.S.; Bagnasco, M.; Fock, J.; Tian, B.; Garbarino, F.; Hansen, M.F. Automated on-chip analysis of tuberculosis drug-resistance mutation with integrated DNA ligation and amplification. Anal. Bioanal. Chem. 2020, 412, 2705–2710. [Google Scholar] [CrossRef] [PubMed]
- Law, I.L.G.; Loo, J.F.C.; Kwok, H.C.; Yeung, H.Y.; Leung, C.C.H.; Hui, M.; Wu, S.Y.; Chan, H.S.; Kwan, Y.W.; Ho, H.P.; et al. Automated real-time detection of drug-resistant Mycobacterium tuberculosis on a lab-on-a-disc by Recombinase Polymerase Amplification. Anal. Biochem. 2018, 544, 98–107. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Cherukury, H.; Labanieh, L.; Zhao, W.; Kang, D.-K. Rapid Detection of β-Lactamase-Producing Bacteria Using the Integrated Comprehensive Droplet Digital Detection (IC 3D) System. Sensors 2020, 20, 4667. [Google Scholar] [CrossRef] [PubMed]
- Linger, Y.; Knickerbocker, C.; Sipes, D.; Golova, J.; Franke, M.; Calderon, R.; Lecca, L.; Thakore, N.; Holmberg, R.; Qu, P.; et al. Genotyping Multidrug-Resistant Mycobacterium tuberculosis from Primary Sputum and Decontaminated Sediment with an Integrated Microfluidic Amplification Microarray Test. J. Clin. Microbiol. 2018, 56, e01652-17. [Google Scholar] [CrossRef]
- Kukhtin, A.V.; Norville, R.; Bueno, A.; Qu, P.; Parrish, N.; Murray, M.; Chandler, D.P.; Holmberg, R.C.; Cooney, C.G. A Benchtop Automated Sputum-to-Genotype System Using a Lab-on-a-Film Assembly for Detection of Multidrug-Resistant Mycobacterium tuberculosis. Anal. Chem. 2020, 92, 5311–5318. [Google Scholar] [CrossRef]
- Evans, D.; Papadimitriou, K.I.; Greathead, L.; Vasilakis, N.; Pantelidis, P.; Kelleher, P.; Morgan, H.; Prodromakis, T. An Assay System for Point-of-Care Diagnosis of Tuberculosis using Commercially Manufactured PCB Technology. Sci. Rep. 2017, 7, 685. [Google Scholar] [CrossRef]
- Evans, D.; Papadimitriou, K.I.; Vasilakis, N.; Pantelidis, P.; Kelleher, P.; Morgan, H.; Prodromakis, T. A Novel Microfluidic Point-of-Care Biosensor System on Printed Circuit Board for Cytokine Detection. Sensors 2018, 18, 4011. [Google Scholar] [CrossRef]
- Cabibbe, A.M.; Miotto, P.; Moure, R.; Alcaide, F.; Feuerriegel, S.; Pozzi, G.; Nikolayevskyy, V.; Drobniewski, F.; Niemann, S.; Reither, K.; et al. Lab-on-Chip-Based Platform for Fast Molecular Diagnosis of Multidrug-Resistant Tuberculosis. J. Clin. Microbiol. 2015, 53, 3876–3880. [Google Scholar] [CrossRef]
- Lazzeri, E.; Santoro, F.; Oggioni, M.R.; Iannelli, F.; Pozzi, G. Novel primer-probe sets for detection and identification of mycobacteria by PCR-microarray assay. J. Clin. Microbiol. 2012, 50, 3777–3779. [Google Scholar] [CrossRef]
- Ou, X.; Li, Q.; Su, D.; Xia, H.; Wang, S.; Zhao, B.; Zhao, Y. A pilot study: VereMTB detection kit for rapid detection of multidrug-resistant mycobcterium tuberculosis in clinical sputum samples. PLoS ONE 2020, 15, e0228312. [Google Scholar] [CrossRef] [PubMed]
- Bielecka, M.K.; Tezera, L.B.; Zmijan, R.; Drobniewski, F.; Zhang, X.; Jayasinghe, S.; Elkington, P. A Bioengineered Three-Dimensional Cell Culture Platform Integrated with Microfluidics To Address Antimicrobial Resistance in Tuberculosis. mBio 2017, 8, e02073-02016. [Google Scholar] [CrossRef] [PubMed]
- Aldridge, B.B.; Fernandez-Suarez, M.; Heller, D.; Ambravaneswaran, V.; Irimia, D.; Toner, M.; Fortune, S.M. Asymmetry and aging of mycobacterial cells lead to variable growth and antibiotic susceptibility. Science 2012, 335, 100–104. [Google Scholar] [CrossRef]
- Richardson, K.; Bennion, O.T.; Tan, S.; Hoang, A.N.; Cokol, M.; Aldridge, B.B. Temporal and intrinsic factors of rifampicin tolerance in mycobacteria. Proc. Natl. Acad. Sci. USA 2016, 113, 8302–8307. [Google Scholar] [CrossRef] [PubMed]
- Rego, E.H.; Audette, R.E.; Rubin, E.J. Deletion of a mycobacterial divisome factor collapses single-cell phenotypic heterogeneity. Nature 2017, 546, 153–157. [Google Scholar] [CrossRef]
- Szafran, M.J.; Kołodziej, M.; Skut, P.; Medapi, B.; Domagała, A.; Trojanowski, D.; Zakrzewska-Czerwińska, J.; Sriram, D.; Jakimowicz, D. Amsacrine Derivatives Selectively Inhibit Mycobacterial Topoisomerase I (TopA), Impair M. smegmatis Growth and Disturb Chromosome Replication. Front. Microbiol. 2018, 9, 1592. [Google Scholar] [CrossRef]
- Trojanowski, D.; Ginda, K.; Pióro, M.; Hołówka, J.; Skut, P.; Jakimowicz, D.; Zakrzewska-Czerwińska, J. Choreography of the Mycobacterium replication machinery during the cell cycle. mBio 2015, 6, e02125. [Google Scholar] [CrossRef]
- Trojanowski, D.; Kołodziej, M.; Hołówka, J.; Müller, R.; Zakrzewska-Czerwińska, J. Watching DNA Replication Inhibitors in Action: Exploiting Time-Lapse Microfluidic Microscopy as a Tool for Target-Drug Interaction Studies in Mycobacterium. Antimicrob. Agents Chemother. 2019, 63, e00739-00719. [Google Scholar] [CrossRef]
- Golchin, S.A.; Stratford, J.; Curry, R.J.; McFadden, J. A microfluidic system for long-term time-lapse microscopy studies of mycobacteria. Tuberculosis 2012, 92, 489–496. [Google Scholar] [CrossRef]
- Baron, V.A.-O.; Chen, M.A.-O.; Hammarstrom, B.A.-O.; Hammond, R.J.H.; Glynne-Jones, P.A.-O.; Gillespie, S.H.; Dholakia, K. Real-time monitoring of live mycobacteria with a microfluidic acoustic-Raman platform. Commun. Biol. 2020, 3, 236. [Google Scholar] [CrossRef]
- Khara, J.S.; Mojsoska, B.; Mukherjee, D.; Langford, P.R.; Robertson, B.D.; Jenssen, H.; Ee, P.L.R.; Newton, S.M. Ultra-Short Antimicrobial Peptoids Show Propensity for Membrane Activity Against Multi-Drug Resistant Mycobacterium tuberculosis. Front. Microbiol. 2020, 11, 417. [Google Scholar] [CrossRef] [PubMed]
- Choi, J.; Yoo, J.; Kim, K.J.; Kim, E.G.; Park, K.O.; Kim, H.; Kim, H.; Jung, H.; Kim, T.; Choi, M.; et al. Rapid drug susceptibility test of Mycobacterium tuberculosis using microscopic time-lapse imaging in an agarose matrix. Appl. Microbiol. Biotechnol. 2016, 100, 2355–2365. [Google Scholar] [CrossRef]
- Jung, Y.G.; Kim, H.; Lee, S.; Kim, S.; Jo, E.; Kim, E.G.; Choi, J.; Kim, H.J.; Yoo, J.; Lee, H.J.; et al. A rapid culture system uninfluenced by an inoculum effect increases reliability and convenience for drug susceptibility testing of Mycobacterium tuberculosis. Sci. Rep. 2018, 8, 8651. [Google Scholar] [CrossRef] [PubMed]
- Elitas, M.; Dhar, N.; McKinney, J.D. Revealing Antibiotic Tolerance of the Mycobacterium smegmatis Xanthine/Uracil Permease Mutant Using Microfluidics and Single-Cell Analysis. Antibiotics 2021, 10, 794. [Google Scholar] [CrossRef] [PubMed]
- Thacker, V.V.; Dhar, N.; Sharma, K.; Barrile, R.; Karalis, K.; McKinney, J.D. A lung-on-chip model of early\textit Mycobacterium tuberculosis infection reveals an essential role for alveolar epithelial cells in controlling bacterial growth. eLife 2020, 9, e59961. [Google Scholar] [CrossRef] [PubMed]
- Yu, J.; Berthier, E.; Craig, A.; de Groot, T.E.; Sparks, S.; Ingram, P.N.; Jarrard, D.F.; Huang, W.; Beebe, D.J.; Theberge, A.B. Reconfigurable open microfluidics for studying the spatiotemporal dynamics of paracrine signalling. Nat. Biomed. Eng. 2019, 3, 830–841. [Google Scholar] [CrossRef] [PubMed]
- Berry, S.B.; Gower, M.S.; Su, X.; Seshadri, C.; Theberge, A.B. A Modular Microscale Granuloma Model for Immune-Microenvironment Signaling Studies in vitro. Front. Bioeng. Biotechnol. 2020, 8, 931. [Google Scholar] [CrossRef]
- Horka, M.; Sun, S.; Ruszczak, A.; Garstecki, P.; Mayr, T. Lifetime of Phosphorescence from Nanoparticles Yields Accurate Measurement of Concentration of Oxygen in Microdroplets, Allowing One To Monitor the Metabolism of Bacteria. Anal. Chem. 2016, 88, 12006–12012. [Google Scholar] [CrossRef]
- Weaver, J.C.; Williams, G.B.; Klibanov, A.; Demain, A.L. Gel Microdroplets: Rapid Detection and Enumeration of Individual Microorganisms by their Metabolic Activity. Bio/Technology 1988, 6, 1084–1089. [Google Scholar] [CrossRef]
- Ryan, C.; Nguyen, B.T.; Sullivan, S.J. Rapid assay for mycobacterial growth and antibiotic susceptibility using gel microdrop encapsulation. J. Clin. Microbiol. 1995, 33, 1720–1726. [Google Scholar] [CrossRef]
- Boedicker, J.Q.; Li, L.; Kline, T.R.; Ismagilov, R.F. Detecting bacteria and determining their susceptibility to antibiotics by stochastic confinement in nanoliter droplets using plug-based microfluidics. Lab Chip 2008, 8, 1265–1272. [Google Scholar] [CrossRef] [PubMed]
- Eun, Y.J.; Utada, A.S.; Copeland, M.F.; Takeuchi, S.; Weibel, D.B. Encapsulating bacteria in agarose microparticles using microfluidics for high-throughput cell analysis and isolation. ACS Chem. Biol. 2011, 6, 260–266. [Google Scholar] [CrossRef] [PubMed]
- Scanlon, T.C.; Dostal, S.M.; Griswold, K.E. A high-throughput screen for antibiotic drug discovery. Biotechnol. Bioeng. 2014, 111, 232–243. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Painter, R.E.; Enesa, K.; Holmes, D.; Whyte, G.; Garlisi, C.G.; Monsma, F.J.; Rehak, M.; Craig, F.F.; Smith, C.A. High-throughput screening of antibiotic-resistant bacteria in picodroplets. Lab Chip 2016, 16, 1636–1643. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, S.; Montalbán-López, M.; Peterhoff, D.; Deng, J.; Wagner, R.; Held, M.; Kuipers, O.P.; Panke, S. Analysis of modular bioengineered antimicrobial lanthipeptides at nanoliter scale. Nat. Chem. Biol. 2019, 15, 437–443. [Google Scholar] [CrossRef] [PubMed]
- Suea-Ngam, A.; Howes, P.D.; Srisa-Art, M.; deMello, A.J. Droplet microfluidics: From proof-of-concept to real-world utility? Chem. Commun. 2019, 55, 9895–9903. [Google Scholar] [CrossRef] [PubMed]
- Sathitratanacheewin, S.; Sunanta, P.; Pongpirul, K. Deep learning for automated classification of tuberculosis-related chest X-Ray: Dataset distribution shift limits diagnostic performance generalizability. Heliyon 2020, 6, e04614. [Google Scholar] [CrossRef]
- Galan, E.A.; Zhao, H.; Wang, X.; Dai, Q.; Huck, W.T.S.; Ma, S. Intelligent Microfluidics: The Convergence of Machine Learning and Microfluidics in Materials Science and Biomedicine. Matter 2020, 3, 1893–1922. [Google Scholar] [CrossRef]
- Feretzakis, G.; Sakagianni, A.; Loupelis, E.; Kalles, D.; Skarmoutsou, N.; Martsoukou, M.; Christopoulos, C.; Lada, M.; Petropoulou, S.; Velentza, A.; et al. Machine Learning for Antibiotic Resistance Prediction: A Prototype Using Off-the-Shelf Techniques and Entry-Level Data to Guide Empiric Antimicrobial Therapy. Healthc. Inform. Res. 2021, 27, 214–221. [Google Scholar] [CrossRef]
- Feretzakis, G.; Loupelis, E.; Sakagianni, A.; Kalles, D.; Lada, M.; Christopoulos, C.; Dimitrellos, E.; Martsoukou, M.; Skarmoutsou, N.; Petropoulou, S.; et al. Using Machine Learning Algorithms to Predict Antimicrobial Resistance and Assist Empirical Treatment. Stud. Health Technol. Inform. 2020, 272, 75–78. [Google Scholar] [CrossRef]
| Sample | Devices Applied | Applications |
|---|---|---|
| Direct bodily fluids Detection of clinical isolates, model organisms, attenuated strains Detection of drug resistant strains | Microfluidic open chips Lab-on-a-chip Droplet Fluidics | Polymerase Chain Reaction Loop-mediated isothermal amplification (LAMP) Recombinase polymerase amplification (RPA) Electrochemical Biosensing Fluorescence detection Surface-enhanced Raman spectroscopy Microscopy |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Molloy, A.; Harrison, J.; McGrath, J.S.; Owen, Z.; Smith, C.; Liu, X.; Li, X.; Cox, J.A.G. Microfluidics as a Novel Technique for Tuberculosis: From Diagnostics to Drug Discovery. Microorganisms 2021, 9, 2330. https://doi.org/10.3390/microorganisms9112330
Molloy A, Harrison J, McGrath JS, Owen Z, Smith C, Liu X, Li X, Cox JAG. Microfluidics as a Novel Technique for Tuberculosis: From Diagnostics to Drug Discovery. Microorganisms. 2021; 9(11):2330. https://doi.org/10.3390/microorganisms9112330
Chicago/Turabian StyleMolloy, Antonia, James Harrison, John S. McGrath, Zachary Owen, Clive Smith, Xin Liu, Xin Li, and Jonathan A. G. Cox. 2021. "Microfluidics as a Novel Technique for Tuberculosis: From Diagnostics to Drug Discovery" Microorganisms 9, no. 11: 2330. https://doi.org/10.3390/microorganisms9112330
APA StyleMolloy, A., Harrison, J., McGrath, J. S., Owen, Z., Smith, C., Liu, X., Li, X., & Cox, J. A. G. (2021). Microfluidics as a Novel Technique for Tuberculosis: From Diagnostics to Drug Discovery. Microorganisms, 9(11), 2330. https://doi.org/10.3390/microorganisms9112330








